Abstract
Corynebacterium glutamicum is a promising host for production of valuable polyketides. Propionate addition, a strategy known to increase polyketide production by increasing intracellular methylmalonyl-CoA availability, causes growth inhibition in C. glutamicum. The mechanism of this inhibition was unclear before our work. Here we provide evidence that accumulation of propionyl-CoA and methylmalonyl-CoA induces growth inhibition in C. glutamicum. We then show that growth inhibition can be relieved by introducing methylmalonyl-CoA-dependent polyketide synthases. With germicidin as an example, we used adaptive laboratory evolution to leverage the fitness advantage of polyketide production in the presence of propionate to evolve improved germicidin production. Whole-genome sequencing revealed mutations in germicidin synthase, which improved germicidin titer, as well as mutations in citrate synthase, which effectively evolved the native glyoxylate pathway to a new methylcitrate pathway. Together, our results show that C. glutamicum is a capable host for polyketide production and we can take advantage of propionate growth inhibition to drive titers higher using laboratory evolution or to screen for production of polyketides.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The accession codes (SRR22028101, SRR22028102, SRR22028103, SRR22028104, SRR22028105, SRR22028106, SRR22028107, SRR22028108, SRR22028109, SRR22028110, SRR22028111, SRR22028112, SRR22028113, SRR22028114, SRR22028115, SRR22028116, SRR22028117, SRR22028118, SRR22028119, SRR22028120, SRR22028121, SRR22028122 and SRR22028123) for the genome sequence data of evolved strains reported in this paper are available at the Sequence Read Archive at PRJNA893829. All RNA-seq data are provided in the supplementary files. Source data are provided with this paper.
References
Moore, B. S. & Hopke, J. N. Discovery of a new bacterial polyketide biosynthetic pathway. ChemBioChem 2, 35–38 (2001).
Smith, C. J. et al. Novel polyketide metabolites from a species of marine fungi. J. Nat. Prod. 63, 142–145 (2000).
Mapari, S. A. et al. Exploring fungal biodiversity for the production of water-soluble pigments as potential natural food colorants. Curr. Opin. Biotechnol. 16, 231–238 (2005).
Flores-Sanchez, I. J. & Verpoorte, R. Plant polyketide synthases: a fascinating group of enzymes. Plant Physiol. Biochem. 47, 167–174 (2009).
Doull, J. L., Ayer, S. W., Singh, A. K. & Thibault, P. Production of a novel polyketide antibiotic, jadomycin B, by Streptomyces venezuelae following heat shock. J. Antibiot. 46, 869–871 (1993).
Bangera, M. G. & Thomashow, L. S. Identification and characterization of a gene cluster for synthesis of the polyketide antibiotic 2,4-diacetylphloroglucinol from Pseudomonas fluorescens Q2-87. J. Bacteriol. 181, 3155–3163 (1999).
Piao, S. J. et al. Hippolachnin A, a new antifungal polyketide from the South China Sea sponge Hippospongia lachne. Org. Lett. 15, 3526–3529 (2013).
Shi, Y. M. et al. Georatusin, a specific antiparasitic polyketide-peptide hybrid from the fungus Geomyces auratus. Org. Lett. 20, 1563–1567 (2018).
Ratnayake, R., Lacey, E., Tennant, S., Gill, J. H. & Capon, R. J. Kibdelones: novel anticancer polyketides from a rare Australian actinomycete. Chemistry 13, 1610–1619 (2007).
Hong, H., Demangel, C., Pidot, S. J., Leadlay, P. F. & Stinear, T. Mycolactones: immunosuppressive and cytotoxic polyketides produced by aquatic mycobacteria. Nat. Prod. Rep. 25, 447–454 (2008).
Chang, M. C. & Keasling, J. D. Production of isoprenoid pharmaceuticals by engineered microbes. Nat. Chem. Biol. 2, 674–681 (2006).
Yuzawa, S., Kim, W., Katz, L. & Keasling, J. D. Heterologous production of polyketides by modular type I polyketide synthases in Escherichia coli. Curr. Opin. Biotechnol. 23, 727–735 (2012).
Kallscheuer, N., Kage, H., Milke, L., Nett, M. & Marienhagen, J. Microbial synthesis of the type I polyketide 6-methylsalicylate with Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 103, 9619–9631 (2019).
Keatinge-Clay, A. T. The uncommon enzymology of cis-acyltransferase assembly lines. Chem. Rev. 117, 5334–5366 (2017).
Zha, W., Rubin-Pitel, S. B., Shao, Z. & Zhao, H. Improving cellular malonyl-CoA level in Escherichia coli via metabolic engineering. Metab. Eng. 11, 192–198 (2009).
Enjalbert, B., Millard, P., Dinclaux, M., Portais, J. C. & Letisse, F. Acetate fluxes in Escherichia coli are determined by the thermodynamic control of the Pta-AckA pathway. Sci. Rep. 7, 42135 (2017).
Takamura, Y. & Nomura, G. Changes in the intracellular concentration of acetyl-CoA and malonyl-CoA in relation to the carbon and energy metabolism of Escherichia coli K12. J. Gen. Microbiol. 134, 2249–2253 (1988).
Milke, L. & Marienhagen, J. Engineering intracellular malonyl-CoA availability in microbial hosts and its impact on polyketide and fatty acid synthesis. Appl. Microbiol. Biotechnol. 104, 6057–6065 (2020).
Mattozzi, M., Ziesack, M., Voges, M. J., Silver, P. A. & Way, J. C. Expression of the sub-pathways of the Chloroflexus aurantiacus 3-hydroxypropionate carbon fixation bicycle in E. coli: toward horizontal transfer of autotrophic growth. Metab. Eng. 16, 130–139 (2013).
Yuzawa, S. et al. Short-chain ketone production by engineered polyketide synthases in Streptomyces albus. Nat. Commun. 9, 4569 (2018).
Risdian, C., Mozef, T. & Wink, J. Biosynthesis of polyketides in Streptomyces. Microorganisms 7, 124 (2019).
Cruz-Morales, P. et al. Biosynthesis of polycyclopropanated high energy biofuels. Joule 6, 1590–1605 (2022).
Murli, S., Kennedy, J., Dayem, L. C., Carney, J. R. & Kealey, J. T. Metabolic engineering of Escherichia coli for improved 6-deoxyerythronolide B production. J. Ind. Microbiol. Biotechnol. 30, 500–509 (2003).
Weber, T., Welzel, K., Pelzer, S., Vente, A. & Wohlleben, W. Exploiting the genetic potential of polyketide producing streptomycetes. J. Biotechnol. 106, 221–232 (2003).
Lee, N. et al. Synthetic biology tools for novel secondary metabolite discovery in Streptomyces. J. Microbiol. Biotechnol. 29, 667–686 (2019).
Sano, C. History of glutamate production. Am. J. Clin. Nutr. 90, 728S–732S (2009).
Claes, W. A., Puhler, A. & Kalinowski, J. Identification of two prpDBC gene clusters in Corynebacterium glutamicum and their involvement in propionate degradation via the 2-methylcitrate cycle. J. Bacteriol. 184, 2728–2739 (2002).
Huser, A. T. et al. Development of a Corynebacterium glutamicum DNA microarray and validation by genome-wide expression profiling during growth with propionate as carbon source. J. Biotechnol. 106, 269–286 (2003).
Kirchner, O. & Tauch, A. Tools for genetic engineering in the amino acid-producing bacterium Corynebacterium glutamicum. J. Biotechnol. 104, 287–299 (2003).
Nesvera, J. & Patek, M. Tools for genetic manipulations in Corynebacterium glutamicum and their applications. Appl. Microbiol. Biotechnol. 90, 1641–1654 (2011).
Portevin, D. et al. A polyketide synthase catalyzes the last condensation step of mycolic acid biosynthesis in mycobacteria and related organisms. Proc. Natl Acad. Sci. USA 101, 314–319 (2004).
Lee, J. Y., Park, J. S., Kim, H. J., Kim, Y. & Lee, H. S. Corynebacterium glutamicum whcB, a stationary phase-specific regulatory gene. FEMS Microbiol. Lett. 327, 103–109 (2012).
Gonzalez-Garcia, R. A., Nielsen, L. K. & Marcellin, E. Heterologous production of 6-deoxyerythronolide B in Escherichia coli through the Wood Werkman cycle. Metabolites 10, 228 (2020).
Haller, T., Buckel, T., Retey, J. & Gerlt, J. A. Discovering new enzymes and metabolic pathways: conversion of succinate to propionate by Escherichia coli. Biochemistry 39, 4622–4629 (2000).
Boghigian, B. A., Zhang, H. & Pfeifer, B. A. Multi-factorial engineering of heterologous polyketide production in Escherichia coli reveals complex pathway interactions. Biotechnol. Bioeng. 108, 1360–1371 (2011).
Plassmeier, J., Barsch, A., Persicke, M., Niehaus, K. & Kalinowski, J. Investigation of central carbon metabolism and the 2-methylcitrate cycle in Corynebacterium glutamicum by metabolic profiling using gas chromatography-mass spectrometry. J. Biotechnol. 130, 354–363 (2007).
Botella, L., Lindley, N. D. & Eggeling, L. Formation and metabolism of methylmalonyl coenzyme A in Corynebacterium glutamicum. J. Bacteriol. 191, 2899–2901 (2009).
Veit, A. et al. Pathway identification combining metabolic flux and functional genomics analyses: acetate and propionate activation by Corynebacterium glutamicum. J. Biotechnol. 140, 75–83 (2009).
Park, Y. K., Ledesma-Amaro, R. & Nicaud, J. M. De novo biosynthesis of odd-chain fatty acids in Yarrowia lipolytica enabled by modular pathway engineering. Front. Bioeng. Biotechnol. 7, 484 (2019).
Pang, B. et al. Investigation of indigoidine synthetase reveals a conserved active-site base residue of nonribosomal peptide synthetase oxidases. J. Am. Chem. Soc. 142, 10931–10935 (2020).
Chemler, J. A. et al. Biochemical and structural characterization of germicidin synthase: analysis of a type III polyketide synthase that employs acyl-ACP as a starter unit donor. J. Am. Chem. Soc. 134, 7359–7366 (2012).
Bai, Y. et al. Structure-based molecular networking for the target discovery of novel germicidin derivatives from the sponge-associated streptomyces sp. 18A01. J. Antibiot. 74, 799–806 (2021).
Parvez, A. et al. Novel type III polyketide synthases biosynthesize methylated polyketides in Mycobacterium marinum. Sci. Rep. 8, 6529 (2018).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47, D309–D314 (2019).
Metz, J. G. et al. Production of polyunsaturated fatty acids by polyketide synthases in both prokaryotes and eukaryotes. Science 293, 290–293 (2001).
Nivina, A., Yuet, K. P., Hsu, J. & Khosla, C. Evolution and diversity of assembly-line polyketide synthases. Chem. Rev. 119, 12524–12547 (2019).
Ridley, C. P., Lee, H. Y. & Khosla, C. Evolution of polyketide synthases in bacteria. Proc. Natl Acad. Sci. USA 105, 4595–4600 (2008).
Helfrich, E. J. N. et al. Evolution of combinatorial diversity in trans-acyltransferase polyketide synthase assembly lines across bacteria. Nat. Commun. 12, 1422 (2021).
Klaus, M. & Grininger, M. Correction: engineering strategies for rational polyketide synthase design. Nat. Prod. Rep. 38, 1409 (2021).
Chen, D. et al. Improvement of FK506 production in Streptomyces tsukubaensis by genetic enhancement of the supply of unusual polyketide extender units via utilization of two distinct site-specific recombination systems. Appl. Environ. Microbiol. 78, 5093–5103 (2012).
Li, S., Li, Z., Pang, S., Xiang, W. & Wang, W. Coordinating precursor supply for pharmaceutical polyketide production in Streptomyces. Curr. Opin. Biotechnol. 69, 26–34 (2021).
Ladner, C. C. New Strategies for Engineering the Substrate Specificity of Polyketide Synthases (North Carolina State Univ. Press, 2016).
Bergmann, S. et al. Genomics-driven discovery of PKS-NRPS hybrid metabolites from Aspergillus nidulans. Nat. Chem. Biol. 3, 213–217 (2007).
Yi, X., Khey, J., Kazlauskas, R. J. & Travisano, M. Plasmid hypermutation using a targeted artificial DNA replisome. Sci. Adv. 7, eabg8712 (2021).
Schafer, A. et al. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145, 69–73 (1994).
Kim, J. et al. Engineering Saccharomyces cerevisiae for isoprenol production. Metab. Eng. 64, 154–166 (2021).
Baidoo, E. E. K., Wang, G., Joshua, C. J., Benites, V. T. & Keasling, J. D. Liquid chromatography and mass spectrometry analysis of isoprenoid intermediates in Escherichia coli. Methods Mol. Biol. 1859, 209–224 (2019).
Acknowledgements
This work was part of the US Department of Energy (DoE) Joint BioEnergy Institute (jbei.org) supported by the US DoE Office of Science, Office of Biological and Environmental Research, through contract DE-AC02-05CH11231 between Lawrence Berkeley National Laboratory and the US DoE and by a DoE Office of Science Distinguished Scientist Award to J.D.K. The views and opinions of the authors expressed herein do not necessarily state or reflect those of the US Government or any agency thereof. Neither the US Government nor any agency thereof, nor any of their employees, makes any warranty, expressed or implied or assumes any legal liability or responsibility or the accuracy, completeness, or usefulness of any information, apparatus, product, or process disclosed or represents that its use would not infringe privately owned rights. The DoE will provide public access to the results of federally sponsored research in accordance with the DoE Public Access Plan (http://energy.gov/downloads/doe-public-access-plan). Funding for the open access charge is provided by the US DoE.
Author information
Authors and Affiliations
Contributions
The study was conceived by C.Z. and R.W.H. C.Z. designed the study, conducted experiments, processed and analyzed the data. N.L. analyzed transcription data, genome sequencing data and modifed figures, while G.L. conducted the ALE experiments. Q.D. performed protein simulation and E.E.K.B. and R.K. analyzed the data from the 13C labeled experiments. B.L. and Z.W. contributed to the construction of engineered strains. R.C.K. and J.M. performed genome sequencing and Y.L. reviewed and edited the manuscript. L.V. conducted fatty acid measurement. J.D.K. supervised the project, reviewed and edited the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
J.D.K. has financial interests in Amyris, Ansa Biotechnologies, Apertor Pharma, Berkeley Yeast, Cyklos Materials, Demetrix, Lygos, Napigen, ResVita Bio and Zero Acre Farms. All other authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Yongjin Zhou and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Giménez-Cassina in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genealogy of strains created in this manuscript and the changes that were made to create those strains.
This paper can be divided into two parts. In the first part, we demonstrated that the propionyl-CoA/methylmalonyl-CoA accumulation is the reason to cause growth inhibition in C. glutamicum (propionate uptake and propionate metabolism). Based on that, we demonstrated that introducing methylmalonyl-CoA dependent polyketide synthase can relieve this inhibition (PKS-based detoxification). The second part is focused on evolving polyketide synthase based on propionate inhibition (ALE and reverse engineering). To strengthen the selection pressure, we knocked out the 2-methylcitrate pathway and mutase AB (based on our [13C] labeling results showing that more than 99% propionate will goes into 2-methylcitrate pathway rather than into the methylmalonyl-CoA, which will significantly weaken the selection stress.
Extended Data Fig. 2 Carbon utilization efficiency comparison among glucose, branched-chain amino acids and propionate.
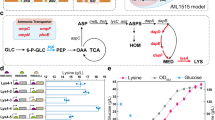
Glucose is one of the most common carbon sources. 2 mol CO2 are generated (during TCA cycle) when 1 mol glucose is converted to succinyl-CoA. Then, succinyl-CoA is converted into methylmalonyl-CoA, which is subsequently decarboxylated when it is loaded onto the PKS by AT domains. Hence, 3 mol CO2 will be lost when 1 mol glucose is used for polyketide production. The carbon utilization efficiency is approximately 50%. For branched chain amino acids like valine, approximately 40% carbon will be lost when it is utilized by PKSs. However, compared with the above two carbon sources, there is no carbon lost when propionate is utilized by PKSs because 1 CO2 will be fixed at the beginning, even though this carbon will be lost when methylmalonyl-CoA is loaded by PKSs. The above information indicates that compared with other carbon sources, propionate is better because the carbon utilization efficiency is much higher. Glu, Glucose; G6P, Glucose 6-phosphate; F6P, Fructose 6-phosphate; Pyr, Pyruvate; Suc-CoA, Succinyl-CoA; MM-CoA, Methylmalonyl-CoA.
Extended Data Fig. 3 CgI2569 is involved in propionate utilization.
a: SDS-PAGE of CgI2569, ?marked bands represent purified CgI2569 protein. 1-4 represents four different tubes during elution. b: LC-MS analysis of the reaction between propionate and succinyl-CoA, indicating that purified CgI2569 catalyzed the reaction between propionate and succinyl-CoA. c: LC-MS analysis of the reaction between propionate and free CoA, indicating that purified CgI2569 did not catalyze the reaction between propionate and free CoA. d: LC-MS analysis of the reaction between propionate and acetyl-CoA, showing that purified CgI2569 catalyzed the reaction between propionate and acetyl-CoA. e: LC-MS analysis of the reaction between propionate and methylmalonyl-CoA, showing that purified CgI2569 did not catalyze the reaction between propionate and methylmalonyl-CoA. f: Growth curves of wild-type in GXII minimal medium with different concentrations of propionate (0, 0.2, 0.6 and 1 g/L), showing that cell growth is inhibited when propionate concentration is increased. g: Growth curves of different strains in CGXII minimal medium, which show that knockout pta-ack or cgl2569 did not affect cell growth in glucose medium. Cz01 (?pta-ack), Cz02 (?cgl2569), Cz03 (?cgl2569?pta-ack). All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples).
Extended Data Fig. 4 Knocking out cgI2537 (acdh) further inhibits cell growth in propionate medium.
a: Alternate metabolic pathways for propionate consumption in wild type cells with percentage of propionyl-CoA funneled through the 2-methylcitrate and methylmalonyl-CoA pathways indicated. b: Acyl-CoA dehydrogenase-based propionyl-CoA utilization pathway. Transcriptomic data indicate that acdh was significantly upregulated in propionate medium (log2 (Fold change) = 4.5), indicating that acdh may be involved in propionate inhibition. c: Growth curves of different strains (WT, Cz05, Cz06, Cz23, Cz24, Cz25) in CGXII minimal medium. d: Growth curves of strains encoding acdh mutations in CGXII minimal medium with 0.5 g/L propionate. Cz05 (?prpDBC1/2). Cz06 (?prpDBC1/2, ?mcmAB). Cz23 (?acdh). Cz24 (?acdh, ?prpDBC1/2). Cz25 (?acdh, ?prpDBC1/2, ?mcmAB). Growth results indicate that deleting acdh in Cz05 or Cz06 further inhibits cell growth. All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples). Based on the results shown in (a), we surmise that most of the propionate is funneled through the 2-methylcitrate pathway with little flowing through the methylmalonyl-CoA pathway, indicating that the 2-methylcitrate pathway is the main pathway to utilize propionate, consistent with Fig. 3a.
Extended Data Fig. 5 Free CoA and acetyl-CoA in WT, Cz05 and, Cz06 strains.
a: Free CoA and acetyl-CoA measurement in wild type, Cz05 and Cz06 cultures grown in CGXII minimal medium with 1 g/L propionate. b: Free CoA measurement in Cz05 and Cz06 cultures grown in CGXII minimal medium with 1 g/L propionate and 10 μM vitamin B12. Cz05 (?prpDBC1/2). Cz06 (?prpDBC1/2, ?mcmAB). Compared with wild-type, free CoA and acetyl-CoA were both depleted in Cz05 and Cz06 strains when cells were inoculated into propionate-containing medium. Moreover, compared to Cz06, the free CoA concentration increased in Cz05 when cells were inoculated into propionate-containing medium with 10 μM vitamin B12. This is consistent with our hypothesis that accumulation of methylmalonyl-CoA is causing slower growth by depleting acetyl-CoA and free CoA-SH. This result is consistent with previous results2,3, in which propionate or its derivatives were shown to inhibit acetyl-CoA production. All cells were inoculated in BHI medium (30 °C, 200 rpm for 16 h), then transferred into CGXII minimal medium (30 °C, 200 rpm for 16 h) and finally inoculated into CGXII minimal medium with 1 g/L propionate. Samples were collected during early exponential phase (OD600 is around 0.5). All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples). Statistical analysis was performed using two-tailed Student’s t test.
Extended Data Fig. 6 Free CoA and vitamin B12 rescue cell growth in propionate.
a: Growth curves of different strains in CGXII minimal medium with 1 g/L propionate and 10 µM sodium pantothenate. b: Growth curves of Cz05 in CGXII minimal medium with 0.6 g/L propionate with 10 µM vitamin B12 or 10/100 µM sodium pantothenate. c: Growth curve of Cz05 in CGXII minimal medium with 1 g/L propionate with 10 µM vitamin B12 or 10/100 µM sodium pantothenate. d: Growth curve of Cz05 in CGXII minimal medium with 1.6 g/L propionate with 10 µM vitamin B12 or 10/100 µM sodium pantothenate. e: Transcriptomic analysis of genes in the CoA biosynthesis pathway in wild type strain. Samples 1 and 2 were inoculated in CGXII minimal medium with 1 g/L propionate and control samples were inoculated into CGXII minimal medium. Samples were collected during the early exponential phase at an OD600 of approximately 0.5. Cz04 (?mcmAB). Cz05 (?prpDBC1/2). Cz06 (?prpDBC1/2, ?mcmAB). All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples).
Extended Data Fig. 7 Propionate tolerance assay and ALE strategy to evolve germicidin initial production strain (Cz12).
a: Growth curve of Cz34 (?prpDBC1, prpDBC2::sfp, ?mcmAB) in CGXII minimal medium with various concentrations of propionate. b: Lag phase of Cz34 in CGXII minimal medium with various concentrations of propionate. All strains were pre-cultivated in BHI overnight (16 h), inoculated into CGII minimal medium and grown for 16 h. Then cells were inoculated into CGXII minimal medium with various concentrations of propionate. All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples). c: ALE workflow, cells were cultivated in BHI medium overnight (30 °C, 16 h), then inoculated into CGXII minimal medium and finally transferred into fresh CGXII medium with increasing concentration of propionate. Cells were passaged 10 times. d: OD600 measurement during ALE process. Three colonies (colony-1, colony-2 and colony-3) were chosen to perform ALE. The duration and propionate concentration of each passage is indicated above.
Extended Data Fig. 8 Evaluate cell growth after ALE.
a: Biomass production of different strains after 7 h. b: Biomass production of different strains after 12 h. c: Biomass production of different strains after 17 h. d: Biomass production of different strains after 21 h. e: Growth curves of Cz34, initial strain (Cz12) and various evolved strains. f: Growth rate of initial strain and various evolved strains. All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples). Statistical analysis was performed using two-tailed Student’s t test (**P < 0.01, ***P < 0.001).
Extended Data Fig. 9 Reverse Engineering to evaluate beneficial mutation.
a: Modeled surface structure of GltA2 (Citrate Synthase) based on module 4TVM (PDB)4. Residues that were mutated in evolved strains are labeled. Green represents mutations E239G, R310C and S60F. Active site catalytic triad residues are shown in pink. b: SDS-PAGE gel results of purified GltA2. Lanes 1-3 are three fractions of purified GltA2; we moved forward with the fraction in lane 2 for subsequent experiments. GltA2 molecular weight is approximately 47.98 KD. c: Growth of different strains in CGXII minimal medium. d: In vitro citrate synthase assay using purified GltA2 proteins. Acetyl-CoA and/or propionyl-CoA are consumed in the reaction, releasing free CoA-SH which is measured colorimetrically. CzEv208, single colony from evolved strain Cz13 (population). Cz34 (?prpDBC2, ?mcmAB, prpDBC1::sfp). Cz12 (entire germicidin production pathway integrated into Cz34). Cz16 (GltA2 Glu239Gly in Cz12), Cz17 (GltA2 Arg310Cyc in Cz12), Cz18 (GltA2 Ser60Phe in Cz12). All data represent the mean ± SD and error bars indicate the standard error (n = 3, biologically independent samples).
Extended Data Fig. 10 LC-MS results to detect 13C labeled intermediates for germicidin product.
a: Measure the level of 13C-labeled different intermediates among Cz12 and Cz15. b: Measure the level of 13C-labeled 2-methylcitrate and germicidin in CzEv208. To further confirm that evolved GltA2 can catalyze the reaction between propionyl-CoA and oxaloacetate and the generated pyruvate can be used to produce acetyl-CoA and malonyl-CoA, Cz12 and Cz15 (native GltA2 in Cz12 replaced with evolved GltA2) were pre-cultivated in BHI liquid medium (30 °C, 200 rpm for 16 h). Then, the cells were transferred into CGXII minimal medium (30 °C, 200 rpm for 16 h). Finally, Cz12 and Cz15 were inoculated into CGXII minimal medium with 1 g/L [13C3] propionate. Samples were collected during exponential phase (OD600 around 0.5). Compared to Cz12, the concentrations of [13C] 2-methylcitrate, [13C] acetyl-CoA and [13C] malonyl-CoA were significantly increased, indicating that evolved GltA2 can catalyze the reaction between propionyl-CoA and oxalacetate to produce 2-methylcitrate. All data represent the mean ± SD, and error bars indicate the standard error (n = 3, biologically independent samples). Statistical analysis was performed using two-tailed Student’s t test (**P < 0.05, ***P < 0.001).
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Supplementary Tables 1 and 2.
Supplementary Data 1
Source data for supplementary figures.
Source data
Source Data Fig. 2
Statistical Source Data for Fig. 2.
Source Data Fig. 3
Statistical Source Data for Fig. 3.
Source Data Fig. 4
Statistical Source Data for Fig. 4.
Source Data Fig. 5
Statistical Source Data for Fig. 5.
Source Data Fig. 6
Statistical Source Data for Fig. 6.
Source Data Extended Data Fig. 3
Statistical Source Data for Extended Fig. 3.
Source Data Extended Data Fig. 4
Statistical Source Data for Extended Fig. 4.
Source Data Extended Data Fig. 5
Statistical Source Data for Extended Fig. 5.
Source Data Extended Data Fig. 6
Statistical Source Data for Extended Fig. 6.
Source Data Extended Data Fig. 7
Statistical Source Data for Extended Fig. 7.
Source Data Extended Data Fig. 8
Statistical Source Data for Extended Fig. 8.
Source Data Extended Data Fig. 9
Statistical Source Data for Extended Fig. 9.
Source Data Extended Data Fig. 10
Statistical Source Data for Extended Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhan, C., Lee, N., Lan, G. et al. Improved polyketide production in C. glutamicum by preventing propionate-induced growth inhibition. Nat Metab 5, 1127–1140 (2023). https://doi.org/10.1038/s42255-023-00830-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00830-x