Abstract
Here we report a heterozygous tandem duplication at the ASIP (agouti signaling protein) gene locus causing ubiquitous, ectopic ASIP expression in a female patient with extreme childhood obesity. The mutation places ASIP under control of the ubiquitously active itchy E3 ubiquitin protein ligase promoter, driving the generation of ASIP in patient-derived native and induced pluripotent stem cells for all germ layers and hypothalamic-like neurons. The patient’s phenotype of early-onset obesity, overgrowth, red hair and hyperinsulinemia is concordant with that of mutant mice ubiquitously expressing the homolog nonagouti. ASIP represses melanocyte-stimulating hormone-mediated activation as a melanocortin receptor antagonist, which might affect eating behavior, energy expenditure, adipocyte differentiation and pigmentation, as observed in the index patient. As the type of mutation escapes standard genetic screening algorithms, we rescreened the Leipzig Childhood Obesity cohort of 1,745 patients and identified four additional patients with the identical mutation, ectopic ASIP expression and a similar phenotype. Taken together, our data indicate that ubiquitous ectopic ASIP expression is likely a monogenic cause of human obesity.
Similar content being viewed by others
Main
Although there is doubtless a genetic component of common obesity, monogenic forms of human obesity are very rare1,2. Most of these single-gene defects affect the leptin–melanocortin 4 receptor (MC4R) pathway, which centrally regulates food intake and energy balance3,4. Detection of these5 and new monogenic obesity traits6 will not only improve our understanding of obesity mechanisms in humans but guide the development of new targeted pharmacotherapies increasingly available for affected patients2,7.
In a bottom-up approach, taking advantage of patient-derived adipose cells, we identified a potential new genetic cause of melanocortin obesity syndrome, phenocopying mouse and human proopiomelanocortin (POMC) deficiency and mouse ASIP mutants (so-called ‘agouti mice’).
Case report
A girl of non-consanguineous parents of European ancestry first presented at the age of 1.9 years with overgrowth and severe obesity. From infancy onwards, she was reported to have a constant desire for food and developed progressive extreme obesity and tall stature (Fig. 1a,b) accompanied by first signs of severe hyperinsulinemia and hepatic steatosis in early childhood (Fig. 1c–h).
a,b, Course of BMI (a) and height (b) of the patient (red dots) from birth to adolescence in relation to 3rd (P3), 50th (P50) and 97th (P97) sex-specific reference percentiles according to Kromeyer-Hauschild et al.40. Data from the patient were corrected for gestational age until the age of 2 years. c–g, Parameters of glucose metabolism with the respective reference ranges of either the local hospital laboratory depicted as gray areas or with prediabetes cut-offs according to the American Diabetes Association65 indicated by a dotted line. h, Serum levels of liver enzyme alanine aminotransferase (ALAT) in comparison to the reference values according to Bussler et al. 66 (shaded in gray). Blue arrows mark the time point of bariatric surgery at the age of 12.4 years. i, Pedigree of the family; females are indicated by circles, the father by a square. Individuals carrying the ASIP tandem duplication are indicated by black symbols, the open symbol indicates the mother not carrying the mutation. The index patient is indicated by a red arrow. Photographs present patient at age of 3.5 years and father at age 52 years. HbA1c, glycated hemoglobin; HOMA-IR, homeostatic model assessment for insulin resistance; 2 h glucose, glucose levels after 2 h of oral glucose tolerance testing.
Both parents also suffer from obesity (Fig. 1i). The father has type 2 diabetes, hypertension and gout. In distinct phenotypic features (red hair color, pale skin and freckles), the girl resembles her father. Clinically relevant variants in genes related to monogenic obesity and other inherited diseases were excluded by whole exome sequencing. At the age of 12.4 years with a body mass index (BMI) of 47.6 kg m−2 (BMI s.d. score (SDS) 3.8, weight 156 kg, height 181 cm and 3.34 SDS), she underwent bariatric surgery by sleeve gastrectomy leading to weight loss of 40 kg. She subsequently regained weight although metabolic improvement was retained (Fig. 1a–h).
During childhood, the patient reported being continuously hungry, except after eating very large amounts of food. She tried consciously to control food intake. At the age of 16 years, she scored high on the Eating Disorder Examination Questionnaire and the Dutch Eating Behavior Questionnaire, showing high cognitive restraint (>85th percentile) as well as extreme eating, weight and shape concerns (>95th percentile). She did not report binge eating, external (<5th percentile) or emotional eating (<25th percentile).
A clinical–psychological exploration noted remarkably low physical activity. Furthermore, indirect calorimetry revealed a reduced resting metabolic rate (1,660 kcal d−1, accounting for only 77.4% of the expected).
Hence, this phenotype of early-onset extreme obesity accompanied by tall stature, hypopigmentation and alterations in appetite and energy expenditure was suggestive for an underlying genetic or biological cause.
Results
Biological alterations and ectopic ASIP expression in adipose tissue
We obtained a sample of subcutaneous adipose tissue from the patient during bariatric surgery. Considering the severe overgrowth and obesity, we hypothesized that the adipose tissue-derived stromal vascular fraction (SVF) cells showed alterations in proliferative and/or adipogenic capacity. The patient’s SVF cells showed pronounced adipocyte differentiation that was sustained in long-term cultivation compared to SVF cells from four age-matched controls with obesity (Supplementary Appendix, Supplementary Table 1 and Fig. 2a–d), whereas proliferation of the patient’s SVF cells was not enhanced (Fig. 2e,f). We further found reduced mitochondrial maximum respiration, spare capacity and proton leak in the patient’s SVF cells as proxies for reduced energy expenditure (Fig. 2g).
a, At indicated time points of cultivation, subcutaneous SVF cells from the patient and control (Ctr) children, all presenting with overweight or obesity (n = 4; 3 females, 1 male), matched for age (mean age 13.8 years) were differentiated into adipocytes. Nuclei were stained with Hoechst (blue) and lipids with Nile red (green). b, Percentage of differentiated cells determined by cell counting. c, Using the same differentiated cells adipogenic capacity was quantified using Oil Red O. d, Relative expression of the adipogenic marker genes adipocyte protein 2 (AP2), peroxisome proliferator-activated receptor γ (PPARG) and adiponectin (ADIPOQ) in differentiated SVF cells from the patient and controls (n = 4). The differentiation was examined over n = 3 independent experiments (cells seeded for differentiation at days 65, 83 and 98 of cultivation). Data are presented as fold change in relation to the controls. e, Doubling time of SVF cells from the patient and control children (Ctr, n = 4) shown as mean with s.e. from n = 2 experiments at day 49 and 71 of cultivation. f, Cell viability was assessed by WST-1 assay in n = 3 experiments at day 49, 71 and 89 of cultivation. g, Mitochondrial function of SVF cells from the patient and controls (n = 4) measured in n = 3 independent experiments using the Seahorse Mito Stress Test kit. Data are presented as fold change in relation to the controls. The data points of each independent experiment are indicated for each individual (b–g). Bar plots represent the mean ± s.e.m. All bar plots from controls are shown in blue and from the patient in red. Statistical significance was assessed using two-sided Student’s t-test without adjustment of multiple comparisons.
To identify potential molecular causes for the enhanced adipogenesis, we searched for differentially expressed genes in the patient’s SVF cells compared to SVF cells from control children through transcriptome-wide analyses. Only one gene encoding ASIP was highly overexpressed in the index patient’s cells (Fig. 3a).
a, Differentially expressed genes between undifferentiated SVF cells from the patient and control children (Ctr; n = 4) presented as the log2 fold change of gene expression across −log10 false discovery rate (FDR). Similar results have been shown in a second independent experiment. b–d, ASIP gene expression levels are shown for adipocytes (b), SVF cells (c) and peripheral blood leukocytes (PBLs) (d). As PBL RNA samples from control children 1–4 were not available, measurements were performed in control children 5–8 (Supplementary Appendix and Supplementary Table 1). e, ASIP protein was detected by immunoblotting in cell lysates and conditioned medium from untreated and brefeldin A and monensin (B/M)-treated SVF cells from the patient and two control children, 1 and 3. Similar results were obtained by a second independent experiment. Gene expression data shown in b–d were normalized to the expression of β-actin and TATA-box-binding protein. Bar plots represent mean ± s.e.m. For b–d the fold changes in relation to mean of controls ± s.e.m. are indicated. All bar plots from controls are shown in blue and from the patient in red.
We confirmed ectopic ASIP expression in several cell types, including isolated adipocytes, SVF cells and peripheral blood leukocytes, indicating ubiquitous expression in the patient (Fig. 3b–d, Supplementary Appendix and Supplementary Table 2). Concordantly, we found ectopic ASIP protein in SVF cell lysates and conditioned medium in the patient’s cells. Inhibition of the secretory pathway lead to cellular retention of ASIP, confirming that the ectopically expressed ASIP is secreted8 (Fig. 3e).
Physiologically, ASIP is expressed in the skin and regulates hair pigmentation through interaction with the melanocortin-1 receptor (MC1R). Ubiquitous expression of the murine ASIP homolog nonagouti in so-called agouti mice leads to yellow fur and obesity due to increased food intake, lipid storage in adipose tissue and reduced energy expenditure9,10,11,12. Hence, the concordant phenotype associated with ectopic expression of ASIP in our participants is likely a new monogenic obesity cause in humans.
Chromosomal rearrangement causes aberrant ectopic ASIP expression
Based on our finding of ectopic ASIP expression in the patient, we hypothesized an ASIP gene locus alteration. Genetic analyses including trio whole-genome sequencing (WGS), screen for copy-number variation (CNV) and cloning of the breakpoint sequence revealed a heterozygous 183-kbp tandem duplication on chromosome 20 encompassing the neighboring genes ASIP and itchy E3 ubiquitin protein ligase (ITCH) on the forward strand and adenosylhomocysteinase (AHCY) on the reverse strand (Fig. 4a). Additional clinically relevant variants or CNVs in genes related to obesity, such as MC4R, POMC and LEPR, were excluded by exome sequencing and a TruSight One Sequencing panel.
a, Schematic of the wild-type allele and the 183-kbp tandem duplication at the ASIP, AHCY and ITCH locus found in the patient. Locations of the breakpoints are presented according to the Dec. 2013 GRCh38/hg38 assembly. b, ASIP expression in PBLs in the patient, mother and father normalized to β-actin and TBP. c, Quantification of breakpoint-specific ITCH–ASIP fusion genomic DNA (gDNA) copy number in the patient, mother and father normalized to the copy number of β-actin gDNA. d, 5′-RACE–PCR from the patient and n = 2 controls (Ctr). The lower band contained the ITCH–ASIP fusion mRNA sequence depicted below. The upper bands are incompletely spliced mRNA variants. e,f, ASIP (e) and ITCH (f) expression across human tissues normalized to expression of β-actin and TBP. Tissue RNA pooled from several individuals was purchased TakaraBio (three technical replicates). g, Luciferase assays in HEK293 cells transfected with vectors with or without a 2.8-kbp ITCH promoter sequence upstream of the luciferase (Luc) reporter gene. Firefly Luc activity was normalized to Renilla Luc activity. Bar plots represent mean ± s.e.m. of n = 3 independent experiments. Statistical significance was assessed using two-sided Student’s t-test. h, Immunoblot of ASIP in cell lysates and supernatants from HEK293 cells transfected with the Taconic plasmid containing the ITCH–ASIP mRNA sequence under control of the 2.8-kbp ITCH promoter. i, Immunoblot of ASIP in cell lysates and supernatants from HEK293 cells transfected with plasmids containing the physiological ASIP coding sequence or the ITCH–ASIP fusion mRNA sequence downstream of no promoter or the ITCH promoter. Asterisk indicates a cloning artifact as an additional band due to introduction of an additional translation start codon upstream of the mRNA sequence from the restriction enzyme NcoI (C’CATGG). This generated a functional open reading frame, exclusively for the ITCH–ASIP mRNA sequence, leading to an ASIP protein with additional N-terminal amino acids. The results shown in h,i were each confirmed by a second independent experiment.
Both, AHCY and ITCH were not differentially regulated in transcriptome-wide analyses (Fig. 3a) and results from quantitative real-time PCR from SVF cells showed an approximately 1.5-fold increase in AHCY expression and no difference in ITCH expression (Extended Data Fig. 1a,b), whereas ASIP is overexpressed several hundredfold (Extended Data Fig. 1c). Similar results were obtained for peripheral blood leukocytes (Fig. 3d and Extended Data Fig. 1d,e). This is in line with the expectation from the genomic rearrangement resulting in three copies of the AHCY gene, no change in the number of ITCH coding exons and a switch of promoter usage driving ASIP gene expression by the ubiquitously active ITCH promoter.
As we saw a slight increase in AHCY expression, we investigated the effects of AHCY in more detail; however, experimental overexpression of AHCY in human Simpson–Golabi–Behmel syndrome (SGBS) preadipocytes13 did not affect adipocyte differentiation or the expression of peroxisome proliferator-activated receptor-γ (PPARG) as the master regulator of adipogenesis (Extended Data Fig. 1f). Hence, the main consequence of the chromosomal rearrangement mutation is ectopic ASIP expression.
Our index patient inherited the mutation from her father, as evidenced by ectopic ASIP expression in his peripheral blood leukocytes (Fig. 4b) and the presence of the duplication-specific ITCH–ASIP breakpoint sequence confirmed by WGS and breakpoint-specific quantitative polymerase chain reaction (qPCR) (Fig. 4c). The father harbors the same chromosomal rearrangement and according to self-reports had been taller and heavier than his peers from birth onwards. Starting at the age of 10 years, he engaged in competitive athletic sports and lost some weight. Nonetheless, he reported weighing 125 kg at the age of 18 years (BMI 36 kg m−2). He did not, however, score conspicuously on eating behavior questionnaires at the age of 52 years.
Both the patient and her father have red hair color and pale skin with freckles (Fig. 1i), which is another characteristic concordant with the yellow fur of the agouti mice ectopically expressing ASIP and is due to the action of ASIP as an antagonist at MC1R in the skin. We verified that neither the patient nor the father carries any genetic variant known to be associated with red hair, for example in MC1R14 (Supplementary Appendix and Supplementary Table 3).
By 5′ rapid amplification of complementary DNA ends and PCR (5′-RACE–PCR) and RNA-seq from peripheral blood and SVF cells, we demonstrated that the genetic rearrangement creates an ITCH–ASIP transcript consisting of the first two non-coding ITCH exons (5′ untranslated region (UTR)) fused to the ASIP coding exons (Fig. 4d). This indicates that ASIP expression is under the control of the ITCH promoter. Concordant with the GTEx catalog, we found high ASIP gene expression in skin and low expression in other healthy human tissues (Fig. 4e). As ITCH is ubiquitously expressed in human tissues (Fig. 4f and concordant with the GTEx catalog), it explains the ubiquitous expression of ASIP in our patient.
We verified functional ITCH–ASIP interaction in luciferase reporter assays showing that a 2.8-kbp genomic region located upstream of ITCH exon 1 is capable of driving gene expression (Fig. 4g). Furthermore, we generated an artificial fusion gene by cloning the ITCH–ASIP fusion cDNA sequence under control of the 2.8-kbp ITCH promoter and confirmed increased production and secretion of ASIP protein (Fig. 4h) independent of the presence or absence of the ITCH 5′-UTR (Fig. 4i).
Thus, the chromosomal rearrangement positioning ASIP under the control of the ITCH promoter is the cause of the ubiquitous expression of ASIP in our patient and may thus be responsible for the clinical phenotype of our patient with early-onset extreme obesity, altered eating behavior, reduced energy expenditure and hypopigmentation.
Retained ectopic ASIP expression in patient-derived iPSCs
To cause this agouti-like phenotype would require ASIP expression in central (hypothalamic) circuits. As these are naturally not directly accessible from patient samples and to provide further support that the genetic rearrangement in the patient causes ubiquitous ASIP expression in cells of all origins, we generated induced pluripotent stem cells (iPSCs) from patient-derived SVF cells and two control children. Ectopic ASIP expression persists in iPSCs from the patient compared to the controls (about 500-fold compared to controls; Fig. 5a), whereas AHCY is only slightly increased (about 1.25-fold) and ITCH is not differently expressed compared to controls (Extended Data Fig. 1g). We confirmed that ASIP protein is also secreted into the supernatant from patient iPSCs (Fig. 5b). We furthermore differentiated patient and control iPSCs into cells of the three germ layers mesoderm, ectoderm and endoderm and found ectopic ASIP expression in patient-derived iPSCs but not controls after differentiation in all germ layers (Fig. 5c). As the suspected mode of action is ectopic ASIP action in the hypothalamus, we went on and confirmed ASIP overexpression in patient-derived iPSCs differentiated into hypothalamic-like neurons via neuronal progenitor cells (Fig. 5d).
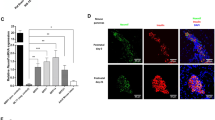
a, ASIP expression in SVF cell-derived iPSCs from n = 3 clones (as biological independent samples) of the patient and n = 2 clones from each control (Ctr) 1 and 10. b, Immunoblot of ASIP in cell lysates and supernatants from untreated iPSCs from the patient and the two control children. Similar results were obtained by a second independent experiment. c, Persisting ASIP expression in iPSCs of the patient (n = 3 clones as biologically independent samples) but not of control children (n = 2 clones each) differentiated into the three germ layers. Successful differentiation was confirmed by downregulation of the pluripotency markers OCT4, SOX2 and NANOG and upregulation of germ-layer-specific markers (MESP, T and MIXL for mesoderm; PAX6, SOX1 and Nestin for ectoderm; SOX17, FOXA2 and CXCR4 for endoderm). d, Persisting ASIP expression in patient-derived iPSCs (n = 2 clones; red) but not HMGU1 iPSC control cell line (blue) differentiated into hypothalamic-like neurons (HTNs) via neuronal progenitor cells (NPCs). Expression of hypothalamic marker genes LEPR, POMC, AGRP and MC4R confirmed successful differentiation. Gene expression levels were normalized to β-actin and TATA-box-binding protein. ASIP expression is presented as fold change in relation to the mean ± s.e.m. of the controls. Relative expression levels of germ-layer marker genes were calculated by delta-delta Ct method normalized to β-actin. e, Antagonistic activity of ASIP in absence and presence of α-MSH at human MC1R and MC4R. Basal cAMP levels concentration-dependently decreased in presence of increasing concentrations of ASIP (MC1R IC50 = 4.4 nM; MC4R IC50 = 16.9 nM). α-MSH stimulates both MC1R (EC50 = 2.3 nM) and MC4R (EC50 = 10.6 nM). Presence of 100 nM ASIP lowers the potency of α-MSH at MC1R (EC50 = 65 nM) and MC4R (EC50 = 149 nM). Data are mean ± s.e.m. of n = 3 independent experiments each performed in duplicate.
Finally, to experimentally confirm that ASIP indeed acts as an antagonist not only at MC1R but also at MC4R, we performed cAMP accumulation assays in CHO-K1 cells transiently transfected with human MC1R or MC4R. Increasing concentrations of ASIP cause a reduction of basal activity of both human MC1R and MC4R activity with half-maximum inhibitory concentration (IC50) values of 4.4 nM and 16.9 nM, respectively. The natural ligand α-melanocyte-stimulating hormone (α-MSH) exerted agonistic activity at both receptors (MC1R half-maximum effective concentration (EC50) = 2.3 nM; MC4R EC50 = 10.6 nM), but presence of 100 nM ASIP caused a reduction of α-MSH potency at both receptors, MC1R (EC50 = 65 nM) and MC4R (EC50 = 149 nM) (Fig. 5e).
Hence, ubiquitous expression of the nonagouti homolog ASIP is found in patient-derived cells of all three germ layers supporting the hypothesis that ectopic ASIP antagonizes MC4R signaling in the hypothalamus of our patient, which would be required to affect processes related to eating behavior and energy expenditure and hence the obesity phenotype.
Identification of additional patients with ASIP mutation
The type of genetic rearrangement is not readily detectable by standard genetic screening classically focusing on single-nucleotide variations. This may explain why there are no cases with detected ASIP mutations in the UK Biobank deposit or in the literature yet.
We screened a targeted 20 patients with a phenotype indicative of defects in the POMC pathway (extreme obesity and red hair color) for ASIP variants originally suspected but not diagnosed for POMC deficiency. However, we did not identify a patient with a similar ITCH–ASIP fusion (Fig. 6a).
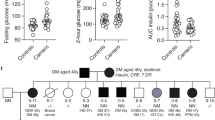
a, Cohort of patients originally suspected for POMC deficiency (n = 20) from the University of Berlin and the Leipzig Childhood Obesity cohort were analyzed for copy numbers of the genomic ITCH–ASIP fusion sequence normalized to the copy numbers of β-actin. Patients A–E were identified with ASIP mutations. Samples from the index patient (patient A, red) were used as a positive control. A patient not harboring the ASIP mutation served as control. Data are given as fold change compared to patient A. b, ASIP gene expression in PBLs of patients normalized to β-actin and TATA-box-binding protein. Data are given as fold change to the control patients (consisting of the twin of patient E, the mother of patient A and three patients of the Leipzig Childhood Obesity cohort). c, Comparison of BMI SDS and height SDS of patients with ASIP mutation (as indicated) with healthy population (n = 1,868; male n = 908; mean age 10.0 years; open black) and with Leipzig Obesity Childhood cohort (n = 1,734; male n = 861; mean age 11.6 years; open red) shown as violin plots with median and quartiles indicated as dashed and dotted lines, respectively. Patients with ASIP mutation have significantly higher height SDS (P = 0.0419 on two-sided Mann–Whitney U-test).
Targeted screening of our Leipzig Childhood Obesity cohort (n = 1,745) revealed four additional patients (three girls and one boy) carrying exactly the same mutation with the identical 5′ITCH–ASIP fusion as our index patient (Table 1). For three of them, RNA from peripheral blood was available and ectopic ASIP expression was confirmed (Fig. 6b). All four patients suffered from severe childhood obesity (Table 1). Compared to the obesity cohort, most patients carrying the ASIP mutation have a BMI SDS and height SDS above the median (Fig. 6c). Notably, one of these patients has a twin brother, who does not harbor the mutation. Even though obesity is present in both brothers, they are discordant in the extent of obesity (>25 kg difference in body weight, difference BMI SDS 0.67; height difference of 8.8 cm, difference height SDS 1.18; all at the age of 11 years; Table 1).
To further support the link between the identified tandem duplication and childhood obesity, we screened healthy populational childhood cohorts (n = 2,051). We performed power calculation assuming that the prevalence in the general pediatric population were identical to the one we observed in our obesity group (0.28%). According to this, with a power of 80% (significance level 0.05), screening of 2,154 probands should be sufficient to identify six carriers of the ASIP mutation (corresponding to 1 in 359). We did not find any in the healthy population cohort of 2,051 probands that we screened (P = 0.02 with both Fisher’s exact test and a chi-squared test with Monte Carlo simulation), further supporting that the tandem duplication is indeed associated with obesity.
Hence, identification of the additional, unrelated patients even in our local sample holds promise of detecting more patients with ubiquitous ASIP expression if genetic diagnostic algorithms are extended to chromosomal rearrangements.
Discussion
We report an agouti-like human monogenic obesity trait that presents with extreme early-onset obesity, accelerated linear growth beyond what would be expected for children with obesity15 and hypopigmentation. Based on experimental data from the analyses of patient-derived native and iPS cells, we identified ubiquitous ectopic ASIP expression due to chromosomal rearrangement as the likely cause of the phenotype (Fig. 7).
Starting out from a case of a patient with severe early-onset obesity, accelerated linear growth, red hair and hyperinsulinemia (gray shaded box), we found enhanced adipogenesis of her SVF adipose-derived cells. Transcriptomic profiling identified aberrant ASIP expression, which is caused by chromosomal rearrangement with tandem duplication placing ASIP under the control of the ITCH promoter found in genetic profiling. Functionally verified aberrant promoter usage subsequently leads to ectopic ASIP expression as verified in native and patient-derived iPSCs of the index patient. ASIP antagonizes MC4R signaling and thereby ubiquitous ectopic ASIP expression can induce a phenotype phenocopying the agouti mouse models. Cohort screening identified four additional patients carrying the same ASIP mutation, whereas none was identified in a populational cohort. Carriers of the ASIP mutation are shown as red-shaded symbols, controls with obesity as white-shaded symbols and healthy children as gray symbols.
The identified tandem duplication leads to the generation of an ectopically expressed ITCH–ASIP fusion messenger RNA under control of the naturally ubiquitously active ITCH promoter and results in increased levels of secreted ASIP protein. In line with this, we found ectopic ASIP expression in several cell types of the patient (for example adipocytes and peripheral blood leukocytes). Even though desirable, we are not able to show ectopic ASIP expression directly in functional hypothalamic/brain circuits as we naturally do not have access to these patient tissues; however, patient-derived iPSCs differentiated into cells of all germ layers and further on into hypothalamic-like neurons, maintaining ectopic ASIP expression. These findings together with the fact that ITCH expression was detected in the brain and hypothalamus and that ASIP acts as an antagonist at the MC1R and MC4R, support that ASIP is ectopically expressed in the hypothalamus of the patient.
The murine ASIP homolog nonagouti is well known to be linked to obesity in rodents. The coding region of the human ASIP gene is 85% homologous to the mouse gene16 and there is high functional conservation17. Concordant with our patient, several rodent models with ubiquitous ectopic expression of nonagouti develop increased obesity, body length, yellow fur and hyperinsulinemia9,11,12. Notably, similar to that in naturally occurring agouti mice9, ectopic ASIP expression in our patient is caused by a gene rearrangement resulting in a change in promoter usage and hence ubiquitous and ectopic ASIP expression. Of note, there is a transgenic agouti mouse model, in which expression of the murine ASIP homolog is driven by the ubiquitously active human β-actin promoter, hence a very similar genetic constellation as we identified in our patient. These mice show overexpression of ASIP in the brain, leading to an obese phenotype with hyperphagia and hyperinsulinemia10.
Physiologically, the ASIP gene is expressed in hair follicles and antagonizes MC1R function on melanocytes and its ectopic expression, hence, causes yellow fur in mice. Although the physiological function of human ASIP does not seem to be entirely clear, we and others have shown it to be pharmacologically active and to antagonize the human melanocortin receptors17. Furthermore, genome-wide association studies revealed associations of variants at the ASIP locus with skin color, melanoma risk and red hair color, pointing toward a physiological function in hair and skin pigmentation similar to other mammals. Nevertheless, we did not find variants in ASIP or MC1R associated with red hair in our index patient, thus indicating that the pigmentation phenotype derives from the functional overexpression of ASIP. Apart from this, a study by Voisey et al. indicated that ASIP mRNA levels in adipocytes were sex-specifically correlated with BMI in adult human probands with opposite directions in men and women18. In contrast to this we found no or only spurious ASIP expression in subcutaneous adipose tissue samples of children from our Leipzig Adipose Tissue Childhood cohort19 (Extended Data Fig. 2).
The obesity phenotype is explained by murine nonagouti/ASIP and human ASIP sharing structural similarity with agouti-related peptide20. When ectopically expressed in the hypothalamus, ASIP is hypothesized to interfere with the leptin–melanocortin axis by competitively antagonizing the natural ligand α-melanocyte-stimulating hormone at the MC4R and thereby centrally affecting food intake and energy expenditure; however, ubiquitous expression of ASIP in mice has been reported to result in rather moderate hyperphagia compared to leptin- or leptin-receptor-deficient mice11. Similarly, our patient reported struggling with satiety before bariatric surgery and showed restrained eating, indicating a conscious counteracting of the hyperphagic drives that she sensed. Increased levels of incretins after bariatric surgery may have affected neuronal MC4R signaling in the patient, further facilitating restrained eating21. Besides hyperphagia, other mechanisms may affect the development of obesity11,22. Agouti-related peptide in the hypothalamus affects energy expenditure23, locomotor activity24 and sympathetic tone as well as shifting substrate utilization25 in peripheral tissues. Ectopic ASIP may have analogous effects. Indeed, we observed reduced spontaneous physical activity in our patient and observed a decreased resting metabolic rate at the whole body and cellular level, which might at least partially contribute to the obesity phenotype. The father suffers from obesity, diabetes and has red hair. In childhood and adolescence he engaged intensely in competitive sports, which affects energy balance beneficially, but did not entirely prevent obesity. Variable phenotype–genotype concordance is also known in other monogenic causes of obesity, for example MC4R mutations26. It has been described that MC4R mutations and variants exhibit partial penetrance and age-dependent expressivity27. Of note, our index patient reported that her hair changed with age, from deep red (as a toddler) to more reddish and brown (now as a young adult), which might indicate similar altered expressivity of the genotype–phenotype association. Although ectopic ASIP most likely promotes obesity primarily via central effects, ectopic ASIP may also contribute to obesity through peripheral effects as described for enhanced insulin secretion in the pancreas, leptin secretion from adipocytes, reduced lipolytic activity in adipocytes or regulation of adipogenic transcription factors22,28 and it may even affect activation of brown AT29. In line with this, the patient showed hyperinsulinemia and her SVF cells differentiated better into adipocytes than SVF cells from controls, indicating additional peripheral obesity-related effects of ubiquitous ASIP expression. Potentially, primary hyperinsulinemia may also drive the linear overgrowth, as recently shown for epigenetically driven hyperinsulinemia causing gigantism in mice30.
Notably, although the patient experienced prominent weight regain after bariatric surgery, she showed sustained metabolic improvement. Apart from the described restrictive eating behavior of the patient and/or a potential sustained effect on incretin balance21, another potential explanation for this might be the young age of the patient at surgery, as it has been shown previously that the benefit in remission of diabetes and hypertension from bariatric surgery was higher in adolescents compared to older patients31.
Likely there are more patients with undiagnosed ectopic ASIP expression, as the locus around ASIP is susceptible to genetic rearrangements32. In fact, patients with CNVs have been described in publicly available databases (for example DECIPHER). These are mainly larger deletions of >1 Mbps. The CNV gain per se (with retained endogenous promoter linkage) is, however, not sufficient for ubiquitous and ectopic ASIP expression. This is fully dependent on the position of the breakpoints and rejoining points. The crucial consequence of the specific tandem duplication and the breakpoint/reinsertion point in our patient(s) is that one copy of ASIP is placed under control of the ubiquitous ITCH promoter and this results in ectopic ASIP expression.
Such gain-of-function mutations have been described in other species9,33,34 and similar duplications resulting in an ITCH–ASIP fusion gene have been observed in sheep35 and in quail36. The chromosomal rearrangement is, however, not readily detectable by routine sequencing approaches but requires techniques identifying CNVs. Only by targeted re-evaluation we were able to find four additional patients in a cohort of 1,745 children with overweight and obesity carrying the same genomic rearrangement as our index patient. That this is also associated with ectopic ASIP expression in blood cells further supports the finding of the tandem duplication causing the ectopic ASIP expression and the obese phenotype. In fact, the frequency of ~0.28% ASIP mutations in this pediatric obesity cohort may even reach the range of MC4R mutation prevalence, so far the most frequent monogenic obesity trait37. The screening of more patients to confirm this and to specify the phenotype of this monogenic trait caused by ectopic ASIP expression is pertinent, particularly as MCR agonists, such as setmelanotide as recently approved for deficiencies in components of the leptin–melanocortin signaling pathway7, provide a promising treatment option for these patients; however, concomitant stimulation of MC1R with setmelanotide causes skin pigmentation and may change melanoma risk38. Another treatment option might be glucagon-like peptide-1 receptor (GLP-1R) agonists as they have been shown to mediate weight loss in patients with MC4R mutations to a similar degree as in patients with polygenic obesity39; however, they would not target the underlying pathomechanisms at the MC4R. Agonists that are selective for MC4R or specifically target hypothalamic neurons would constitute a promising treatment option2.
In summary, we report that a genomic rearrangement leads to ubiquitous expression of ASIP in patients with severe early-onset obesity and overgrowth. This phenotype is concordant with mouse agouti obesity models ectopically expressing the homolog nonagouti/ASIP. Both, murine and human ASIP interfere with the melanocortin receptor signaling, which likely explains the obese phenotypes.
Methods
Oversight
The index patient and children of the control group were participants in the Leipzig Adipose Tissue Childhood19 (NCT02208141, https://clinicaltrials.gov/ct2/show/NCT02208141, first registered August 2014) and the Leipzig Childhood Obesity (NCT04491344, https://clinicaltrials.gov/ct2/show/NCT04491344, first registered July 2020) cohorts15. Children from the age of 12 years and their guardians provided informed consent. The studies were approved by the Ethics Committee of the University of Leipzig (265/08, 007/04).
Clinical and eating behavior data, energy expenditure
Height and weight were measured using calibrated devices to the nearest 1 mm and 0.1 kg, respectively. BMI and height were referenced to sex and age according to current guidelines40 and were given as SDSs. Children were categorized into overweight (1.28 < BMI SDS ≤ 1.88) or obese (BMI SDS > 1.88) groups according to current national consensus guidelines41.
Blood parameters were measured by standard laboratory procedures in the Institute of Laboratory Medicine of the University of Leipzig.
Eating behavior was assessed by self-reports using validated questionnaires. The Eating Disorder Examination Questionnaire42,43 assesses eating disorder psychopathology with subscales restraint, eating, weight and shape concern and behavioral features (for example binge eating). Patient scores were compared to a reference population of 1,354 women <44 years of age44 and father’s scores to a reference population of 1,166 men aged 44–64 years (ref. 44). The Dutch Eating Behavior Questionnaire45 evaluates emotional eating, external eating and restraint using sum scores from Likert scales. Patient scores were compared to a reference population of 1,394 women between 14–94 years of age45.
Resting energy expenditure of the patient was examined at the age of 15 years following an overnight fast using an indirect calorimeter (Quark RMR, Cosmed Germany) according to the manufacturer’s protocol. Expected metabolic rate was calculated according to her age, sex, height and body mass.
Isolation and cultivation of SVF cells from adipose tissue
Subcutaneous AT samples were obtained from children undergoing bariatric or elective orthopedic surgery (Supplementary Table 1). SVF cells and adipocytes were isolated and preserved for later RNA isolation as previously described19,46. The remaining SVF cells were filtered through a nylon mesh with 30-µm pore size. Erythrocytes were removed with erythrocyte lysis buffer (0.154 M NH4Cl (Sigma-Aldrich), 0.01 M KHCO3 (Merck) and 0.1 mM EDTA (Sigma-Aldrich)). SVF cells were frozen in liquid nitrogen in Dulbecco’s modified Eagle medium/Ham F-12 culture medium (DMEM/F-12) (Life Technologies) containing 10% fetal bovine serum (FBS) (Biochrom) and 10% dimethyl sulfoxide (DMSO) (Sigma-Aldrich). Cells were thawed and 24 h after seeding cells were washed three times with phosphate-buffered saline to select for adipocyte precursor cells via plastic adherence. Isolated SVF cells were cultivated in DMEM/Ham F-12 culture medium containing 10% FBS and 100 U penicillin and 0.1 mg ml−1 streptomycin (Sigma-Aldrich) at 37 °C and 5% CO2 and passaged every 3–4 d.
Functional effects of AHCY overexpression
Human SGBS cells were kindly supplied by M. Wabitsch (University of Ulm) and cultured as previously described13.
pCMV plasmid transfections were performed with the Neon Transfection System 100 µl kit (Invitrogen). The electroporation protocol was optimized to pulse voltage 1,300 V, pulse width 20 ms, pulse number 2 and a cell density of 6 × 106 cells ml−1. SGBS cells were transfected with 5 µg of pCMV-AHCY plasmid, for AHCY overexpression or the pCMV-Entry (both oriGene) as a control, per 1 × 106 cells. After electroporation, 500,000 cells per well were seeded in a 12-well format for differentiation assay.
Proliferation assay
For assessing proliferation, 500 SVF cells were seeded at day 49, 71 and 89 of cultivation per well in 96-well plates in five replicates and cultivated for 8 d. After 1 and 8 d, cell viability was assessed by WST-1 assay (Roche, Applied Bioscience) according to the manufacturer’s protocol. Afterwards, cells were fixed, stained with Hoechst 33342 (Sigma-Aldrich) and counted by microscope. Doubling time (days) between d1 and d8 was calculated as (duration (8 d) × log10(2)) / (log10(cell number per image d1) – log10(cell number per image d8))47.
Differentiation assay
For assessing adipogenic capacity, in technical triplicates 30,000 SVF cells at day 34, 49, 71 and 89 of cultivation were seeded per well in a 48-well dish or transfected SGBS cells in a 12-well format were used. Cells were grown to confluence for 24 to 48 h and differentiated according to the Poietics human adipose-derived stem cell-adipogenesis protocol (Lonza) for 8 d (SVF cells) or 10 d (transfected SGBS cells).
Differentiated cells were fixed with Roti-Histofix 4% (Carl Roth), double-stained with 10 µg ml−1 Nile red (Sigma-Aldrich) and 40 µg ml−1 Hoechst 33342 (Sigma-Aldrich). The percentage of differentiated cells is given as mean from n = 3 technical replicates calculated as (number of counted differentiated cell per microscopic image) / (total number of cells per image). A cell was defined to be differentiated when containing at least two lipid droplets. Additionally, the cells were stained with Oil Red O (Sigma-Aldrich) solution (0.3% in 60% isopropanol) for 15 min and washed with H2O. Oil Red O was extracted with isopropanol and absorption was measured at 540 nm.
Bioenergetic profiling
Mitochondrial function was assessed using the Seahorse XFe24 Analyzer (Agilent Technologies) and the XF Cell Mito Stress Test kit (Agilent Technologies) using concentrations of 2 µM oligomycin, 3 µM carbonyl cyanide-4 (trifluoromethoxy) phenylhydrazone and 1 µM rotenone/antimycin A. A total of 15,000 SVF cells from the patient or control children were seeded into gelatin-coated wells of XFe24-plates in 3–4 technical replicates. Measurements were performed 48 h post-transfection using XF Base Medium (Agilent Technologies) containing 2 mM pyruvate (Sigma-Aldrich), 10 mM glucose (Sigma-Aldrich) and 2 mM glutamine (Sigma-Aldrich). Results were normalized to total protein per well (µg) by lysing the cells in 30 µl 50 mM NaCl solution (Carl Roth) with subsequent protein quantification using the Pierce BCA Protein Assay kit (Thermo Fisher Scientific).
Isolation of nucleic acids
RNA was isolated from SVF cells and reverse-transcribed as previously described48. From blood, RNA was extracted using the PAXgene Blood RNA Tubes protocol (PreAnalytics). RNA from different human tissues pooled from several individuals were obtained from Clontech-TakaraBio. Genomic DNA was extracted from EDTA blood samples using the QIAamp DNA Blood Mini kit (QIAGEN).
Transcriptome-wide analyses
Gene expression was measured using a HumanHT-12 v4 BeadChip array (Illumina). The R package limma was used to perform preprocessing (that is, background correction and quantile normalization) and differential gene expression analyses49 (https://doi.org/10.5281/zenodo.7223530). Fold changes were log2-transformed and P values were adjusted for multiple testing using the Benjamini–Hochberg procedure (FDR).
qRT–PCR
Quantitative real-time PCR (qRT–PCR) was performed as previously described48 using the QuantStudio3 Real-Time PCR System (Applied Biosystems) with the qPCR Master Mix Plus - Low ROX (Eurogentec) or the Takyon Low ROX Probe Master Mix dTTP Blue (Eurogentec). Gene expression was quantified using a standard curve of serial dilutions from a linearized plasmid containing the target sequence or from a reference sample. Gene expressions were normalized to the expression of β-actin and TATA-box-binding protein. For measurement of ASIP expression in subcutaneous adipose tissue samples from the Leipzig Adipose Tissue Childhood cohort hypoxanthine-guanine-phosphoribosyltransferase was used as a third reference gene. Supplementary Appendix and Supplementary Table 4 lists all sequences of primers and probes and TaqMan Gene Expression Assays (Thermo Fisher Scientific) used for qRT–PCR.
5′-RACE–PCR
5′-RACE–PCR was performed from peripheral blood RNA using the SMARTer RACE cDNA Amplification kit (Clontech Laboratories) using primers in Supplementary Appendix and Supplementary Table 5.
RNA-seq and transcript analyses
RNA was isolated from SVF cells of the patient and control children 1–4 as well as from peripheral blood mononuclear cells of the patient and control 9 using TRI REAGENT (Sigma-Aldrich). RNA quantity was measured with a spectrometer (Nanodrop ND 1000). We included only RNA samples with RIN values >8 analyzed on the Agilent 2100 bioanalyzer using the RNA 6000 Nano Chip (Agilent Technologies). Indexed cDNA libraries were generated using TruSeq RNA Sample Preparation kits v2 (Illumina). The average library size was 300 bp as determined on the Agilent 2100 Bioanalyzer with DNA 1000 Chips. The libraries were sequenced on the Illumina HiScanSQ Sequencing System (Macrogen).
Reads were mapped to the reference human genome (GRCh38.p13, INSDC Assembly GCA_000001405.28, Dec 2013) using TopHat (v.2.1.1)50. After indexing with samtools (v.1.9)51 the mapped reads were assembled to transcripts and quantified by StringTie (v.2.1.3b)52,53. For TopHat, we used the ‘default’ parameters. StringTie parameters ‘read coverage’ (–c), ‘transcript length’ (–m) and ‘bases on both sides of a junction a spliced read has to cover’ (–a) were set to minimal values. The parameter ‘fraction of most abundant transcript at one locus’ (–f) was lowered from default (0.01) to 0 as correction for artifacts and incompletely processed mRNA with a 1% cutoff was performed after the comparative analysis. For all other StringTie parameters default values were used. Assembled transcripts were inspected with the Integrated Genome Viewer (Broad Institute)54,55 and samples showing a visible 3′ bias due to oligonucleotide-dT/poly-A primer selection were not included.
Due to small sizes of the exons of ITCH and ASIP coverage, analyses of single exons were not possible. Instead, a de novo assembly of ITCH and ASIP transcripts was performed using StringTie. Three new transcript variants occurred containing only the first two annotated exons of ITCH. The fragments per kilo base per million mapped reads for these variants and also of the ASIP and AHCY transcripts were compared between the patient and the controls.
Genomic and genetic analyses
Trio WGS and bioinformatic analyses were performed at the DRESDEN-concept Genome Center. After ultrasonic shearing of 800 ng gDNA (LE220, Covaris) the DNA library preparation was performed using the Kapa HyperPlus kit (Roche) according the manufacturer’s instructions. After ligation with uniquely dual indexed adaptors (IDT), non-ligated adaptors were removed by adding XP beads (Beckmann Coulter) in a ratio of 1:0.9. The DNA libraries were then size selected with XP beads to an average insert size of 300 bp and quantified by qPCR (LightCycler 480, Roche) and the Fragment Analyzer (Agilent). Libraries were sequenced paired end 2 × 150 bp to a coverage >38× on a NovaSeq 6000 (Illumina). Bioinformatic analysis was performed using the Illumina DRAGEN pipeline (07.021.595.3.7.5).
The TruSight One Sequencing panel was performed after enrichment with Nextera DNA Flex Pre-Enrichment LibraryPrep and Enrichment, IDT for Illumina Nextera UD Indexes and the NextSeq 500/550 High Output v2 kit (300 cycles) using the Illumina NextSeq 500/550. Data were analyzed using the software Varvis and Varfeed (Limbus). All analyses were performed according to American College of Medical Genetics and Genomics criteria.
Molecular karyotyping of genomic DNA was performed by array comparative genomic hybridization using the Infinium CytoSNP-850K v.1.2 BeadChip (Illumina) and analyzed using BlueFuse Multi (v.4.4).
Luciferase reporter and expression assays
DNA sequences were cloned according to the TOPO TA Cloning kit, Dual Promoter protocol and confirmed by sequencing by the Core Unit for DNA technologies of the University of Leipzig. Primers used for cloning are indicated in Supplementary Appendix and Supplementary Table 5.
For luciferase assays the ITCH promoter was inserted upstream the luciferase reporter gene into the pGL3-Basic vector using the restriction enzymes SacI and XhoI.
For generation of modified pGL3-Basic expression vectors containing the ASIP or the ITCH–ASIP coding sequence under control of ITCH or no promoter, the luciferase reporter gene was cut out of the pGL3-Basic vector using NcoI and XbaI and the cloned mRNA sequences were inserted. Subsequently, the promoter sequence of ITCH was inserted into those vectors using the restriction enzymes SacI and XhoI.
A second independent expression vector containing the coding sequence of ASIP fused to the 5′ UTR of ITCH under control of the ITCH promoter (2.8 kbp upstream of transcription start) was generated by Taconic Bioscience, as shown in Supplementary Appendix and Supplementary Fig. 1.
For luciferase reporter assays, 500,000 human embryo kidney cell line 293 (HEK293) (Sigma-Aldrich, ECACC 85120602) cells were seeded per well (six-well dish plate) and were co-transfected 24 h post-seeding with 1 µg of pGL3-Basic plasmids containing the firefly luciferase reporter under control of a 2,868 bp ITCH promoter sequence or no promoter and 50 ng of pRL-CMV control plasmid containing the luciferase gene from Renilla reniformis using 3 µl Fugene HD transfection reagent (Promega). Luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega) according to the manufacturer’s protocol 48 h post-transfection using a CLARIOstar plate reader (BMG Labtech). Experiments were performed in technical duplicates. For normalization, the sum of firefly luciferase signal was divided by the sum of Renilla luciferase signal.
Generation and differentiation of patient-derived iPSCs
The generation of iPSCs from SVF cells isolated from adipose tissue of the patient and two control patients, 1 and 10, was performed by the iPSC Core Facility, Institute of Stem Cell Research, Helmholtz Zentrum Munchen. Pluripotency was confirmed by staining of OCT4, SOX2, LIN28 and NANOG56. The iPSCs were cultivated on Geltrex (Thermo Fisher Scientific)-coated cell culture plates in mTeSR 1 medium (Stem Cell Technologies) at 37 °C and 5% CO2.
Characterization of the pluripotency of patient-derived iPSCs was performed as monolayers by directed differentiation into the ectodermal57, mesodermal58 and endodermal59 germ layers for a period of 5 d and subsequent qRT–PCR analyses of marker genes for each of the germ layers, as previously described56. Relative expression levels were calculated using the delta-delta Ct method normalized to β-actin.
Differentiation of patient-obtained iPSCs and the HMGU1 control iPSC line (kindly supplied by iPSC Core Facility, Institute of Stem Cell Research, Helmholtz Zentrum Munchen) into HTNs followed the previously described protocol of Wang et al.60. For generating hypothalamic NPCs, 500,000 iPSCs per well were cultured on Geltrex-coated 12-well-plates as described in the protocol and collected at day 12. For further differentiation, 300,000 single cells per well were plated on poly-l-ornithine- (0.01%, Sigma-Aldrich) and laminin-coated (4 µg ml−1; Thermo Fisher Scientific) 24-well plates. After cells had attached to the plate, the medium was replaced with N2 medium with B27 and 10 μM DAPT. Four days later, the medium was changed to N2 medium with B27 and 20 ng ml−1 BDNF (R&D) and cells were cultured until day 26.
Immunoblotting
For analysis of ASIP protein secretion 800,000 SVF cells were seeded on 15-cm plates, cells and supernatants were collected after 24 h of incubation with the secretory pathways inhibitors 5 µg ml−1 brefeldin A (in DMSO; BioLegend) and 2 µM monensin (in 70% ethanol; BioLegend) or the control medium containing the same volume of DMSO and ethanol.
For analysis of ASIP protein generation from the fusion construct, 600,000 HEK cells were seeded per six-well dish and transfected 24 h later with 2 µg plasmid using 8 µl using Fugene HD transfection reagent (Promega) per well. Medium was changed 24 h after transfection. Cells and supernatants were collected 48 h after medium change for immunoblotting.
For the analysis of ASIP protein generation and secretion in the iPSC lines, cells were grown in six-well-plates to confluence before the medium was changed to 1 ml knockout DMEM with 1× MEM NEAA, 1× β-mercaptoethanol, 0.1% BSA (Serva), 100 U penicillin, 0.1 mg ml−1 streptomycin and 2 mM glutamine. After 24 h, cells and supernatants were collected for immunoblotting.
Conditioned medium was centrifuged for 5 min at 800g and the supernatant lyophilized and re-suspended in H2O for receiving 10x concentrated medium. Cells were lysed in RIPA lysis buffer (50 mM Tris pH 7.5, 150 mM NaCl, 1% Triton X-100 and 0.1% SDS, with cOmplete Mini Protease Inhibitor Cocktail Tablets; Roche, Sigma-Aldrich) with additional break-up via QiaShredder Homogenizer Columns (QIAGEN). The protein concentration of cell lysates was measured using the Pierce BCA Protein Assay kit (Thermo Fisher Scientific). Equal amounts of protein or medium were resolved by 12% or 15% SDS–PAGE and ASIP was detected using an ASIP antibody (PA5-77052, Invitrogen, 1:1,000 dilution). Equal loading was confirmed by detection of β-actin using a β-actin antibody (ab8227, Abcam, 1:1,000 dilution).
Melanocortin receptor signaling by cAMP assay
A Chinese hamster ovary cell line CHO-K1 (ATCC CCL-61) was grown in DMEM/F-12 supplemented with 10% FBS, 100 U ml−1 penicillin and 100 µg ml−1 streptomycin. Cells were at seeded at 0.9 × 106 cells per 25-cm2 flask and transfected with plasmid (3 µg total amount of DNA) and Lipofectamine 2000 (Thermo Fisher Scientific). Plasmids encoding human MC1R61, human MC4R62 and empty vector pcDps have previously been described.
One day after transfection, cells were split into 96-well plates (2 × 104 cells per well) and serum-starved for 16 h before the experiment. cAMP content of cell extracts was determined by a non-radioactive assay with ALPHAScreen (PerkinElmer) technology as previously described63. In brief, stimulation with various α-MSH (Sigma-Aldrich) concentrations in the absence or presence of 100 nM ASIP (R&D systems) or with various concentrations of ASIP alone were performed 48 h after transfection. Reactions were stopped by aspiration of medium and cells were lysed in 20 μl of lysis buffer containing 1 mM 3-isobutyl-1-methylxanthine. From each well, 5 μl of lysate were transferred to a 384-well plate. Acceptor beads and donor beads were added according to the manufacturers’ protocol. Cyclic AMP accumulation data were analyzed using GraphPad Prism v.8.4.3.
Screening for patients with ASIP mutations
In unselected cohorts of n = 1,745 children of the Leipzig Childhood Obesity cohort (mean age, 11.6 years; female, n = 879; male, n = 867) as well as 2,051 children of healthy population childhood cohorts (Leipzig childhood cohort, n = 1,447; mean age, 11.7 years; female, n = 755; male, n = 692; Leipzig Lifestyle and environmental factors and their Influence on Newborns Allergy risk (LINA) cohort64, n = 604; mean age, 4 years; female, n = 291; male, n = 313) we screened for carriers of the genomic ITCH–ASIP fusion. The studies were approved by the ethics committee of the University of Leipzig (Leipzig childhood cohort, reg. nos. 007-04/027-04, 782-1998, and 029-2006; LINA cohort, reg. nos. 046-2006 and 144-10-31052010, registered in https://www.birthcohorts.net since 6 May 2019). Children from the age of 12 years on and their guardians provided informed consent. Twenty patients with obesity and red hair color, who were previously suspected for mutations in the POMC gene were additionally screened for genomic ITCH–ASIP fusion. These patients were recruited at the Freie Universitat Berlin, Humboldt-Universitat zu Berlin, Institute for Experimental Pediatric Endocrinology (mean age, 9.4 years; female, n = 13; male, n = 7). This analysis was approved by the local ethical committee (EA2/131/11).
gDNA copy numbers of the ITCH–ASIP fusion sequence were quantified using SYBR green qRT–PCR using the Maxima SYBR Green/ROX qPCR Master Mix (2×) (Thermo Fisher Scientific) and the ITCH–ASIP fusion primers (Supplementary Appendix and Supplementary Table 5) with an input of 2–10 ng gDNA per reaction. Copy numbers were normalized to copy numbers of β-actin. On each plate, samples from the patient were carried along as a positive control.
For comparison of BMI SDS and height SDS of identified carriers with ASIP mutation with the obesity cohort, we excluded eight patients with underlying syndromic or endocrine diseases (Prader–Willi syndrome, growth hormone deficiency, syndromic short stature or autoimmune polyendocrinopathy) and a further three patients had missing data for either height or BMI. Of the LINA birth cohort, from n = 183 probands, data and samples were only available at birth and were, therefore, excluded from comparative anthropometric analyses.
Statistical analyses
If not otherwise stated, statistical analyses were performed using a Student’s t-test (two-sided) in GraphPad Prism 6 (GraphPad Software). P values <0.05 were considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated during and/or analyzed during the current study are deposited as source files except for patient individualized datasets that underly patient confidentiality restrictions but are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Bouchard, C. Genetics of obesity: what we have learned over decades of research. Obesity 29, 802–820. (2021).
Hinney, A., Körner, A. & Fischer-Posovszky, P. The promise of new anti-obesity therapies arising from knowledge of genetic obesity traits. Nat. Rev. Endocrinol. https://doi.org/10.1038/s41574-022-00716-0 (2022).
Lotta, L. A. et al. Human gain-of-function MC4R variants show signaling bias and protect against obesity. Cell 177, 597–607 (2019).
van der Klaauw, A. A. & Farooqi, I. S. The hunger genes: pathways to obesity. Cell 161, 119–132. (2015).
Wabitsch, M. et al. Measurement of immunofunctional leptin to detect and monitor patients with functional leptin deficiency. Eur. J. Endocrinol. 176, 315–322. (2017).
Marenne, G. et al. Exome sequencing identifies genes and gene sets contributing to severe childhood obesity, linking PHIP variants to repressed POMC transcription. Cell Metab. 31, 1107–1119 (2020).
Clément, K. et al. Efficacy and safety of setmelanotide, an MC4R agonist, in individuals with severe obesity due to LEPR or POMC deficiency: single-arm, open-label, multicentre, phase 3 trials. Lancet Diabetes Endocrinol. 8, 960–970. (2020).
Ericson, M. D., Freeman, K. T., Schnell, S. M., Fleming, K. A. & Haskell-Luevano, C. Structure-activity relationship studies on a macrocyclic agouti-related protein (AGRP) scaffold reveal agouti signaling protein (ASP) residue substitutions maintain melanocortin-4 receptor antagonist potency and result in inverse agonist pharmacology at the melanocortin-5 receptor. J. Med. Chem. 60, 8103–8114. (2017).
Duhl, D. M., Vrieling, H., Miller, K. A., Wolff, G. L. & Barsh, G. S. Neomorphic agouti mutations in obese yellow mice. Nat. Genet. 8, 59–65 (1994).
Klebig, M. L., Wilkinson, J. E., Geisler, J. G. & Woychik, R. P. Ectopic expression of the agouti gene in transgenic mice causes obesity, features of type II diabetes, and yellow fur. Proc. Natl Acad. Sci. USA 92, 4728–4732 (1995).
Miltenberger, R. J., Mynatt, R. L., Wilkinson, J. E. & Woychik, R. P. The role of the agouti gene in the yellow obese syndrome. J. Nutr. 127, 1902S–1907SS (1997).
Moussa, N. M. & Claycombe, K. J. The yellow mouse obesity syndrome and mechanisms of agouti-induced obesity. Obes. Res. 7, 506–514 (1999).
Wabitsch, M. et al. Characterization of a human preadipocyte cell strain with high capacity for adipose differentiation. Int. J. Obes. Relat. Metab. Disord. 25, 8–15 (2001).
Morgan, M. D. et al. Genome-wide study of hair colour in UK Biobank explains most of the SNP heritability. Nat. Commun. 9, 5271 (2018).
Kempf, E. et al. Dynamic alterations in linear growth and endocrine parameters in children with obesity and height reference values. eClinicalMedicine 37, 100977 (2021).
Kwon, H. Y. et al. Molecular structure and chromosomal mapping of the human homolog of the agouti gene. Proc. Natl Acad. Sci. USA 91, 9760–9764 (1994).
Yang, Y.-K. et al. Effects of recombinant agouti-signaling protein on melanocortin action. Mol. Endocrinol. 11, 274–280 (1997).
Voisey, J., Imbeault, P., Hutley, L., Prins, J. B. & van Daal, A. Body mass index-related human adipocyte agouti expression is sex-specific but not depot-specific. Obes. Res. 10, 447–452 (2002).
Landgraf, K. et al. Evidence of early alterations in adipose tissue biology and function and its association with obesity-related inflammation and insulin resistance in children. Diabetes 64, 1249–1261 (2015).
Patel, M. P. et al. Loop-swapped chimeras of the agouti-related protein and the agouti signaling protein identify contacts required for melanocortin 1 receptor selectivity and antagonism. J. Mol. Biol. 404, 45–55 (2010).
He, Z. et al. Direct and indirect effects of liraglutide on hypothalamic POMC and NPY/AGRP neurons - Implications for energy balance and glucose control. Mol. Metab. 28, 120–134. (2019).
Voisey, J. & Van Daal, A. Agouti: from mouse to man, from skin to fat. Pigment Cell Res. 15, 10–18 (2002).
Small, C. J. et al. Chronic CNS administration of agouti-related protein (AGRP) reduces energy expenditure. Int. J. Obes. Relat. Metab. Disord. 27, 530–533 (2003).
Tang-Christensen, M. et al. Central administration of ghrelin and agouti-related protein (83-132) increases food intake and decreases spontaneous locomotor activity in rats. Endocrinology 145, 4645–4652 (2004).
Cavalcanti-de-Albuquerque, J. P., Bober, J., Zimmer, M. R. & Dietrich, M. O. Regulation of substrate utilization and adiposity by AGRP neurons. Nat. Commun. 10, 311 (2019).
Lubrano-Berthelier, C. et al. Melanocortin 4 receptor mutations in a large cohort of severely obese adults: prevalence, functional classification, genotype-phenotype relationship, and lack of association with binge eating. J. Clin. Endocrinol. Metab. 91, 1811–1818 (2006).
Winkler, T. W. et al. The influence of age and sex on genetic associations with adult body size and shape: a large-scale genome-wide interaction study. PLoS Genet. 11, e1005378 (2015).
Mynatt, R. L. & Stephens, J. M. Regulation of PPARγ and obesity by agouti/melanocortin signaling in adipocytes. Ann. NY Acad. Sci. 994, 141–146 (2003).
Schnabl, K., Westermeier, J., Li, Y. & Klingenspor, M. Opposing actions of adrenocorticotropic hormone and glucocorticoids on UCP1-mediated respiration in brown adipocytes. Front. Physiol. 9, 1931 (2018).
Yang, C. H. et al. Independent phenotypic plasticity axes define distinct obesity sub-types. Nat. Metab. 4, 1150–1165. (2022).
Inge, T. H. et al. Five-year outcomes of gastric bypass in adolescents as compared with adults. The. N. Engl. J. Med. 380, 2136–2145. (2019).
Nakayama, K. & Ishida, T. Alu-mediated 100-kb deletion in the primate genome: the loss of the agouti signaling protein gene in the lesser apes. Genome Res 16, 485–490 (2006).
Chandramohan, B., Renieri, C., La Manna, V., La & Terza, A. The alpaca agouti gene: genomic locus, transcripts and causative mutations of eumelanic and pheomelanic coat color. Gene 521, 303–310 (2013).
Girardot, M. et al. The insertion of a full-length Bos taurus LINE element is responsible for a transcriptional deregulation of the Normande Agouti gene. Pigment Cell Res. 19, 346–355 (2006).
Norris, B. J. & Whan, V. A. A gene duplication affecting expression of the ovine ASIP gene is responsible for white and black sheep. Genome Res. 18, 1282–1293 (2008).
Robic, A. et al. Two new structural mutations in the 5′ region of the ASIP gene cause diluted feather color phenotypes in Japanese quail. Genet Sel. Evol. 51, 12 (2019).
Wade, K. H. et al. Loss-of-function mutations in the melanocortin 4 receptor in a UK birth cohort. Nat. Med. 27, 1088–1096. (2021).
Taylor, N. J. et al. Inherited variation at MC1R and ASIP and association with melanoma-specific survival. Int. J. Cancer 136, 2659–2667 (2015).
Iepsen, E. W. et al. Patients with obesity caused by melanocortin-4 receptor mutations can be treated with a glucagon-like peptide-1 receptor agonist. Cell Metab. 28, 23–32 (2018).
Kromeyer-Hauschild, K. et al. Percentiles of body mass index in children and adolescents evaluated from different regional German studies. Monatsschr. Kinder 149, 807–18. (2001).
AWMF. Evidence-based (S3) Guideline of the Working Group on Childhood and Adolescent Obesity (AGA) of the German Obesity Society (DAG) and the German Society of Pediatrics and Adolescent Medicine (DGKJ). https://awmf.org/leitlinien/detail/ll/050-002.html (2019).
Hilbert, A. & Tuschen-Caffier, B. Eating Disorder Examination-Questionnaire: Deutschsprachige Übersetzung (Verlag für Psychotherapie, 2006).
Fairburn, C. G. Cognitive Behavior Therapy and Eating Disorders (Guilford Publications, 2008).
Hilbert, A., de Zwaan, M. & Braehler, E. How frequent are eating disturbances in the population? Norms of the eating disorder examination-questionnaire. PLoS ONE 7, e29125 (2012).
Nagl, M., Hilbert, A., de Zwaan, M., Braehler, E. & Kersting, A. The German version of the Dutch Eating Behavior Questionnaire: psychometric properties, measurement invariance, and population-based norms. PLoS ONE 11, e0162510 (2016).
Landgraf, K. et al. The obesity-susceptibility gene TMEM18 promotes adipogenesis through activation of PPARG. Cell Rep. https://doi.org/10.1016/j.celrep.2020.108295 (2020).
Doubling Time Computing. http://www.doubling-time.com/compute.php (2006).
Bernhard, F. et al. Functional relevance of genes implicated by obesity genome-wide association study signals for human adipocyte biology. Diabetologia 56, 311–322 (2013).
Smyth, G., Thorne, N. & Wettenhall, J. limma: Linear models for microarray data user’s guide. Bioinformatics and computational biology solutions using R and bioconductor2011.
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Eberherr, A. C. et al. Rescue of STAT3 function in hyper-IgE syndrome using adenine base editing. CRISPR J. 4, 178–190. (2021).
Shi, Y., Kirwan, P. & Livesey, F. J. Directed differentiation of human pluripotent stem cells to cerebral cortex neurons and neural networks. Nat. Protoc. 7, 1836–1846 (2012).
Borchin, B., Chen, J. & Barberi, T. Derivation and FACS-mediated purification of PAX3+/PAX7+ skeletal muscle precursors from human pluripotent stem cells. Stem Cell Rep. 1, 620–631 (2013).
Ori, C. et al. Single cell trajectory analysis of human pluripotent stem cells differentiating towards lung and hepatocyte progenitors. Preprint at bioRxiv https://doi.org/10.1101/2021.02.23.432413 (2021).
Wang, L., Egli, D. & Leibel, R. L. Efficient generation of hypothalamic neurons from human pluripotent stem cells. Curr. Protoc. Hum. Genet. 90, 21.5.1–21.5.14 (2016).
Lalueza-Fox, C. et al. A melanocortin 1 receptor allele suggests varying pigmentation among Neanderthals. Science 318, 1453–1455 (2007).
Stäubert, C. et al. Evolutionary aspects in evaluating mutations in the melanocortin 4 receptor. Endocrinology 148, 4642–4648 (2007).
Peters, A. et al. Natural biased signaling of hydroxycarboxylic acid receptor 3 and G protein-coupled receptor 84. Cell Commun. Signal. 18, 31 (2020).
Hinz, D. et al. Cord blood Tregs with stable FOXP3 expression are influenced by prenatal environment and associated with atopic dermatitis at the age of one year. Allergy 67, 380–389 (2012).
American Diabetes Association. 2. Classification and diagnosis of diabetes: standards of medical care in diabetes-2020. Diabetes Care 43, S14–S31 (2020).
Bussler, S. et al. New pediatric percentiles of liver enzyme serum levels (alanine aminotransferase, aspartate aminotransferase, γ-glutamyltransferase): effects of age, sex, body mass index, and pubertal stage. Hepatology 68, 1319–1330 (2018).
Acknowledgements
We thank the children and parents who participated in the study, the physicians and technical and clinical assistants who helped with the study and U. Stenzel for the bioinformatic transcript analyses. We also thank E. Rusha and A. Pertek from the iPSC Core Facility, Institute of Stem Cell Research, Helmholtz Zentrum München for generating iPSCs. We thank A.-D. Liebing from the Rudolf Schönheimer Institute of Biochemistry for technical support. We thank H. Schiffmann for his support with the collection of patient data. This work was supported by the German Research Foundation for the Clinical Research Center ‘Obesity Mechanisms’ SFB1052/CRC1052 (no. 209933838) projects C05 to A. Körner, C09 to A. Körner and K.L., and KO3512/3-1 to A. Körner and E.Z., and by the German Diabetes Association to A. Körner. R.S. was funded by the joint Clinician Scientist Program of the Medical Faculty and the Helmholtz Institute for Metabolic, Obesity and Vascular Research of the Helmholtz Zentrum München at the University of Leipzig. The funders of the study had no role in the study design, in the collection, analysis and interpretation of data or in the writing of the report.
Author information
Authors and Affiliations
Contributions
A. Körner was the project principal investigator and led the design of the study, established the project and funding and supervised experiments, analyses and data interpretation. A. Körner assumes responsibility for the completeness and integrity of the data and the fidelity of the report. W.K. contributed to discussion of manuscript. E.K., K.L., R.S., M.H., A.H., R.A.J., P.B., A. Kühnapfel, Y.H.T., C.S., T.S., W.R., E.Z. and A. Körner performed and supervised experiments, performed clinical assessments, data analyses and interpretation and drafted the manuscript. G.H. and P.K. provided cohort samples for mutation screening. M.B. contributed to project management, data discussions and edited the manuscript. All authors contributed to the interpretation of study findings and had the opportunity to review and revise the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 AHCY overexpression in different cell types.
Gene expression levels of (A) adenosylhomocysteinase (AHCY) and (B) itchy E3 ubiquitin protein ligase (ITCH) in stromal vascular fraction (SVF) cells before and after adipocyte differentiation and compared to that of (C) agouti signaling protein (ASIP) in differentiated SVF cells are shown from the patient and controls (Ctr) patients 1-4 (n=4; 3 females, 1 male) matched for age (mean age 13.8 years) and body weight state (all presenting with overweight or obesity). Data points of n=3 independent experiments are indicated for each subject. In peripheral blood leukocytes (PBL) AHCY (D) and ITCH (E) expression are shown from the patient and controls. Data points are indicated for each subject. (F) Relative AHCY expression in Simpson–Golabi-Behmel syndrome (SGBS) cells after transfection with pCMV-Entry control vector (Ctr) or pCMV-AHCY (AHCY) in n=3 independent experiments. Transfected SGBS cells were differentiated into adipocytes and the percentage of differentiated cells was determined by cell counting after Hoechst and Nile red staining. Using the same cells adipogenic capacity was compared by Oil red O staining (representative microscopic pictures from one of three independent experiments; scale bar = 500µM) and gene expression of the adipogenic marker gene peroxisome proliferator-activated receptor gamma (PPARG) after differentiation in n=2 independent experiments was determined. Bar plots show the mean ± SEM. (G) AHCY and ITCH expression levels are shown for SVF cell-derived induced pluripotent stem cells (iPSCs) from three clones of the patient and two clones from each of the control (Ctr) children, 1 and 10. Data points for each clone from each subject are indicated. The bar plots A-E and G present the mean values ± SEM as a fold change of the controls. Gene expression data were normalized to the expression of β-actin and TATA-box-binding protein. All bars from controls are shown in blue and from the patient in red. P values derived using two-sided Student’s t-test are given.
Extended Data Fig. 2 ASIP expression in adipose tissue samples of children.
Gene expression levels of agouti signaling protein (ASIP) in subcutaneous adipose tissue (AT) of 328 children included in the Leipzig AT childhood cohort are shown in comparison to patient A. Data points are indicated for each subject. Gene expression data were normalized to the expression of β-actin, TATA-box-binding protein and hypoxanthine-guanine-phosphoribosyltransferase. Data points from controls are shown in blue and from the patient in red.
Supplementary information
Supplementary Information
Supplementary Tables 1–5, Supplementary Fig. 1 and Supplementary References.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 3
Unprocessed and uncropped images of western blots; RAW images Fig. 3.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 4
Unprocessed and uncropped images of agarose gel and western blots; RAW images Fig. 4.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 5
Unprocessed and uncropped images of western blots; RAW images Fig. 5.
Source Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kempf, E., Landgraf, K., Stein, R. et al. Aberrant expression of agouti signaling protein (ASIP) as a cause of monogenic severe childhood obesity. Nat Metab 4, 1697–1712 (2022). https://doi.org/10.1038/s42255-022-00703-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-022-00703-9
This article is cited by
-
Obesity: exploring its connection to brain function through genetic and genomic perspectives
Molecular Psychiatry (2024)
-
A sequence of SVA retrotransposon insertions in ASIP shaped human pigmentation
Nature Genetics (2024)
-
New variant for monogenic obesity
Nature Reviews Endocrinology (2023)
-
Rare genetic forms of obesity in childhood and adolescence, a comprehensive review of their molecular mechanisms and diagnostic approach
European Journal of Pediatrics (2023)
-
The discovery of human agouti-induced obesity and its implication for genetic diagnosis
Nature Metabolism (2022)