Abstract
Cholesterol contributes to the structural basis of biological membranes and functions as a signaling molecule, whose dysregulation has been associated with various human diseases. Here, we report that the long non-coding RNA (lncRNA) SNHG6 increases progression from non-alcoholic fatty liver disease (NAFLD) to hepatocellular carcinoma (HCC) by modulating cholesterol-induced mTORC1 activation. Mechanistically, cholesterol binds ER-anchored FAF2 protein to promote the formation of a SNHG6–FAF2–mTOR complex. As a putative cholesterol effector, SNHG6 enhances cholesterol-dependent mTORC1 lysosomal recruitment and activation via enhancing FAF2–mTOR interaction at ER–lysosome contacts, thereby coordinating mTORC1 kinase cascade activation with cellular cholesterol biosynthesis in a self-amplified cycle to accelerate cholesterol-driven NAFLD–HCC development. Notably, loss of SNHG6 inhibits mTORC1 signaling and impairs growth of patient-derived xenograft liver cancer tumors, identifyifng SNHG6 as a potential target for liver cancer treatment. Together, our findings illustrate the crucial role of organelle-associated lncRNA in organelle communication, nutrient sensing, and kinase cascades.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq datasets have been deposited in the NCBI under accession number PRJNA769502. RNA pulldown–MS data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD034782. All data supporting the findings of this study are available from the corresponding authors on request. Source data are provided with this paper.
Change history
06 September 2022
A Correction to this paper has been published: https://doi.org/10.1038/s42255-022-00650-5
References
Chu, B. B. et al. Cholesterol transport through lysosome-peroxisome membrane contacts. Cell 161, 291–306 (2015).
Luo, J., Yang, H. & Song, B. L. Mechanisms and regulation of cholesterol homeostasis. Nat. Rev. Mol. Cell Biol. 21, 225–245 (2020).
Luo, J., Jiang, L., Yang, H. & Song, B. L. Routes and mechanisms of post-endosomal cholesterol trafficking: a story that never ends. Traffic 18, 209–217 (2017).
Luo, J., Yang, H. & Song, B. L. Mechanisms and regulation of cholesterol homeostasis. Nat. Rev. Mol. Cell Biol. 21, 225–245 (2020).
Attard, G., Cooper, C. S. & Bono, J. Steroid hormone receptors in prostate cancer: a hard habit to break? Cancer Cell 16, 458–462 (2009).
Degirolamo, C., Modica, S., Palasciano, G. & Moschetta, A. Bile acids and colon cancer: solving the puzzle with nuclear receptors. Trends Mol. Med. 17, 564–572 (2011).
Zhang, X., Coker, O.O., Chu, E.S., Fu, K. & Yu, J. Dietary cholesterol drives fatty liver-associated liver cancer by modulating gut microbiota and metabolites. Gut 70, 761–774 (2020).
Liu, G. Y. & Sabatini, D. M. mTOR at the nexus of nutrition, growth, ageing and disease. Nat. Rev. Mol. Cell Biol. 21, 183–203 (2020).
Anandapadamanaban, M., Masson, G. R., Perisic, O., Berndt, A. & Williams, R. L. Architecture of human Rag GTPase heterodimers and their complex with mTORC1. Science 366, 203–210 (2019).
Mossmann, D., Park, S. & Hall, M. N. mTOR signalling and cellular metabolism are mutual determinants in cancer. Nat. Rev. Cancer 18, 744–757 (2018).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9–Niemann-Pick C1 signaling complex. Science 355, 1306 (2017).
Lim, C. Y. et al. ER–lysosome contacts enable cholesterol sensing by mTORC1 and drive aberrant growth signalling in Niemann-Pick type C. Nat. Cell Biol. 21, 1206–1218 (2019).
Sang, L. et al. Mitochondrial long non-coding RNA GAS5 tunes TCA metabolism in response to nutrient stress. Nat. Metab. 3, 90–106 (2021).
Lin, A. et al. The LINK-A lncRNA interacts with PtdIns(3,4,5)P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Nat. Cell Biol. 19, 238–251 (2017).
Sang, L. J. et al. LncRNA CamK-A regulates Ca2+-signaling-mediated tumor microenvironment remodeling. Mol. Cell 72, 71–83(2018).
Li, R. H. et al. A phosphatidic acid-binding lncRNA SNHG9 facilitates LATS1 liquid–liquid phase separation to promote oncogenic YAP signaling. Cell Res 31, 1088–1105 (2021).
Xing, Z. et al. lncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell 159, 1110–1125 (2014).
Satriano, L., Lewinska, M., Rodrigues, P. M., Banales, J. M. & Andersen, J. B. Metabolic rearrangements in primary liver cancers: cause and consequences. Nat. Rev. Gastroenterol. Hepatol. 16, 748–766 (2019).
Fazal, F. M., Han, S., Kaewsapsak, P., Parker, K. R. & Ting, A. Y. Atlas of subcellular RNA localization revealed by APEX-seq. Cell 178, 473–490 (2019).
Julien, B. et al. Membrane-associated RNA-binding proteins orchestrate organelle-coupled translation. Trends Cell Biol. 29, 178–188 (2018).
Chu, B.-B. et al. Cholesterol transport through lysosome-peroxisome membrane contacts. Cell 161, 291–306 (2015).
Derler, I., Jardin, I., Stathopulos, P. B., Muik, M. & Romanin, C. Cholesterol modulates Orai1 channel function. Sci. Signal. 9, ra10 (2016).
Liu, D. et al. Squalene epoxidase drives NAFLD-induced hepatocellular carcinoma and is a pharmaceutical target. Sci. Transl. Med. 10, eaap9840 (2018).
Huang, D. Q., El-Serag, H. B. & Loomba, R. Global epidemiology of NAFLD-related HCC: trends, predictions, risk factors and prevention. Nat. Rev. Gastroenterol. Hepatol. 18, 223–238 (2021).
Bechmann, L. P. et al. The interaction of hepatic lipid and glucose metabolism in liver diseases. J. Hepatol. 56, 952–964 (2012).
Huang, B., Song, B. L. & Xu, C. Cholesterol metabolism in cancer: mechanisms and therapeutic opportunities. Nat. Metab. 2, 132–141 (2020).
Wang, Y. Q., Huang, G., Chen, J., Cao, H. & Xu, W. T. LncRNA SNHG6 promotes breast cancer progression and epithelial-mesenchymal transition via miR-543/LAMC1 axis. Breast Cancer Res. Treat. 188, 1–14 (2021).
Cao, C. et al. The long non-coding RNA, SNHG6-003, functions as a competing endogenous RNA to promote the progression of hepatocellular carcinoma. Oncogene 36, 1112–1122 (2017).
Wang, X. et al. LncRNA SNHG6 promotes proliferation, invasion and migration in colorectal cancer cells by activating TGF-beta/Smad signaling pathway via targeting UPF1 and inducing EMT via regulation of ZEB1. Int J. Med Sci. 16, 51–59 (2019).
Xu, M. et al. lncRNA SNHG6 regulates EZH2 expression by sponging miR-26a/b and miR-214 in colorectal cancer. J. Hematol. Oncol. 12, 3 (2019).
Zhang, Y., An, J. & Pei, Y. LncRNA SNHG6 promotes LMO3 expression by sponging miR-543 in glioma. Mol. Cell. Biochem. 472, 9–17 (2020).
Wang, H. et al. lncRNA SNHG6 promotes hepatocellular carcinoma progression by interacting with HNRNPL/PTBP1 to facilitate SETD7/LZTFL1 mRNA destabilization. Cancer Lett. 520, 121–131 (2021).
Lee, J. N., Zhang, X., Feramisco, J. D., Gong, Y. & Ye, J. Unsaturated fatty acids inhibit proteasomal degradation of Insig-1 at a postubiquitination step. J. Biol. Chem. 283, 33772–33783 (2008).
Kim, H. et al. UAS domain of Ubxd8 and FAF1 polymerizes upon interaction with long-chain unsaturated fatty acids. J. Lipid Res. 54, 2144–2152 (2013).
Lee, J. N. et al. Unsaturated fatty acids inhibit proteasomal degradation of Insig-1 at a postubiquitination step. J. Biol. Chem. 283, 33772–33783 (2008).
Thelen, A. M. & Zoncu, R. Emerging roles for the lysosome in lipid metabolism. Trends Cell Biol. 27, 833–850 (2017).
Hao, F. et al. Rheb localized on the Golgi membrane activates lysosome-localized mTORC1 at the Golgi-lysosome contact site. J. Cell Sci. 131, jcs208017 (2018).
Korolchuk, V. I. et al. Lysosomal positioning coordinates cellular nutrient responses. Nat. Cell Biol. 13, 453–460 (2011).
Lu, X. Y. et al. Feeding induces cholesterol biosynthesis via the mTORC1–USP20–HMGCR axis. Nature 588, 479–484 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wilhelm, L. P. et al. STARD3 mediates endoplasmic reticulum-to-endosome cholesterol transport at membrane contact sites. EMBO J. 36, 1412–1433 (2017).
Zhan, Y. Y. et al. The orphan nuclear receptor Nur77 regulates LKB1 localization and activates AMPK. Nat. Chem. Biol. 8, 897–904 (2012).
Chen, J. et al. SAR1B senses leucine levels to regulate mTORC1 signalling. Nature 596, 281–284 (2021).
Jung, J. W. et al. Transmembrane 4 L six family member 5 senses arginine for mTORC1 signaling. Cell Metab. 29, 1306–1319 e1307 (2019).
Acknowledgements
We thank X. Li and S.-F. Xie (Westlake University) for assistance for protein MS analysis. We thank H. Ju (Sun Yat-sen University) for the support in the liver cancer PDX tumor growth analysis and for the help with tissue collection. We thank Sagene eBioart for the help with making pattern diagrams in Fig. 8q. Schematic illustrations of NAFLD–HCC and PDX mice models were created using BioRender.com (license numbers IV2433S552 and RJ2433PEHR). This work was supported in part by the National Key Research and Development Program of China (no. 2021YFC2700903), the National Natural Science Foundation of China (no. 81672791, no. 81872300), and the Zhejiang Provincial Natural Science Fund for Distinguished Young Scholars of China (no. LR18C060002).
Author information
Authors and Affiliations
Contributions
A. L. and T. Z. conceived and designed the research. F. L., T. T., and Z. Z. performed most of the biochemical and molecular experiments and bioinformatics analysis, with assistance from S. X., L. Z., J. Y., C. S., L. S., Z. Y., and L. Q. T. T. and H.-Q. J. performed liver cancer PDX tumor-growth analysis. H.-L. P. and W. W. performed cellular cholesterol measurements and MS analyses. K. G. performed homology modeling and molecular docking. Z. Y. and L. Q. conducted bioinformatics analysis. J. Y. and F. L. performed in vivo experiments. T. Z., J. S., W. W., H.-L. P., J. L., Q. Y., H.-Q. J., and X. L. contributed to discussion and data interpretation. A. L., T. Z., and W. W. edited the manuscript. A. L. and F. L wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Jose Carlos Fernandez-Checa, Jan-Wilhelm Kornfeld, Hande Topel Batarlar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary handling editor: Alfredo Gimenez-Cassina in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cholesterol promotes HCC development by SNHG6.
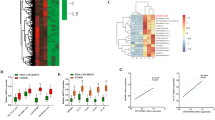
(A) Analyzing SNHG6 expression in liver cancer datasets. The horizontal line inside the box is the median; the top is the upper quartile; the bottom is the lower quartile; the upper whiskers extend from the upper quartile to the highest actual value; the lower whiskers extend from the lower quartile The number is extended to the lowest actual value. Two-sided Student’s t test. (B) Overall Survival analysis of SNHG6 in the liver cancer database. Kaplan-Meier analysis. (C) MTT assay of Vector-/SNHG6-Huh7 cells. Data are mean values ± S.D. Two-way ANOVA. (D-E) Colony formation of sgCtrl /sgSNHG6 HepG2 cells (D) and of shScr/shSNHG6 Huh7 cells (E). Data are mean values ± S.D. One-way ANOVA with Tukey’s test. (F-G) qRT-PCR detection of 12 candidates’ expression upon fatty acid stimulation (0.4 mM palmitic acid, 24 h) (F); glucose stimulation (24 h) (G). Data are presented as mean values ± S.D. Two-sided Student’s t-test. (H) qRT-PCR detection of SNHG6 expression at different cholesterol-incubated times (50 μM). Data are presented as mean values ± S.D. One-way ANOVA with Tukey’s test. (I) MTT assay of sgCtrl/sgSNHG6 HepG2 cells at different cholesterol-incubated times (50 μM). Data are presented as mean values ± S.D. Two-way ANOVA. (J) IB detection for Lysosomes/ mitochondria/ ER quality purified from Huh7 cells. (K-M) qRT-PCR detection of SNHG6 enrichment on ER (K)/ lysosomes (L)/ mitochondria (M) in Huh7 cells. Data are presented as mean values ± S.D. (N,O) Detecting cholesterol levels in sgCtrl/ sgSNHG6 HepG2 cells (N) and shScr/ shSNHG6 Huh7 cells (O). Data are presented as mean values ± S.D. One-way ANOVA with Tukey’s test. (P) Detecting cholesterol levels in vector-/ SNHG6-Huh7 cells. Data are mean values ± S.D. Two-sided Student’s t-test. (Q,R) qRT-PCR analysis of indicated genes in shScr/shSNHG6 Huh7 cells. Data are presented as mean values ± S.D. One-way ANOVA with Tukey’s test (Q); and in Vector-/SNHG6-Huh7 cells. Data are mean values ± S.D. Two-sided Student’s t-test (R).
Extended Data Fig. 2 The subcellular distribution of SNHG6 in hepatoma.
(A) IB detection the quality of lysosomal fraction. Lysosomes were purified from Huh7 cells and then were subjected to RNase treatment in vitro for 15 mins. (B) qRT-PCR assay of the lysosomal distribution of SNHG6 upon RNase treatment in Huh7 cells. Data are presented as mean values ± S.D. (C) IB detection the quality of ER fraction. ER were purified from Huh7 cells and then were subjected to RNase treatment in vitro for 15 mins. (D) qRT-PCR assay of the ER distribution of SNHG6 upon RNase treatment in Huh7 cells. Data are presented as mean values ± S.D. (E-G) The relative enrichment of SNHG6 at lysosome fractions was detected by qRT-PCR assay upon indicated nutrition stimulation. Huh7 cells were incubated with arginine (E), insulin (F) and glucose (G) stimulation for 24 h, followed qRT-PCR assay to detect indicated makers and SNHG6 expression. Data are presented as mean values ± S.D. Two-sided Student’s t-test. (H) In vitro-transcribed SNHG6-sense (FL), SNHG6-antisense (AS), SNHG6-D1, SNHG6-D2, and SNHG6-D3 transcripts.
Extended Data Fig. 3 SNHG6 directly binds to FAF2 and mTOR.
(A) Immunoblot detection of proteins retrieved by in vitro-transcribed SNHG6-sense (Sen.) or antisense (AS.) transcripts from Huh7 cell lysates. (B) Coomassie blue staining of recombinant protein. (C-F) In vitro RNA protein binding assay examining the interaction of biotinylated SNHG6 with bacterially purified MBP-His-tagged RHEB (C), RagA (D), P18 (E) and SFB-tagged Raptor purified from HEK293T cells (F). (G-I) RIP detection of SNHG6-mTOR interaction upon indicated stimulation. HepG2 cells were re-cultured in amino acid (G), insulin (H) or glucose (I) medium for 12 h. Data are presented as mean values ± S.D. Two-way ANOVA. (J-L) RIP detection of SNHG6-FAF2 interaction upon indicated stimulation. Amino acid (J), insulin (K) and glucose (L) treatment method for HepG2 cells were same as above described in G-I. Data are presented as mean values ± S.D. Two-way ANOVA. (M) IB detection of mTOR expression in shScr/shmTOR HepG2 cells.
Extended Data Fig. 4 SNHG6 regulates FAF2-mTOR interaction in hepatoma.
(A) Co-IP detection of the FAF2-mTOR interaction at cholesterol starvation. HepG2 cells expressing SFB-FAF2 and GFP-mTOR plasmids (48 h) were subjected to MCD (0.5% w/v) for 3 h; (B) Cholesterol mediated FAF2-mTOR interaction in a dose-dependent manner. HepG2 cells expressing SFB-FAF2 and GFP-mTOR plasmids (48 h) were replenished indicated cholesterol concentrations for 3 h; (C) Co-IP detection of the FAF2-mTOR interaction at arginine treatment. HepG2 cells expressing indicated plasmids were pre-cultured in amino acid deficiency medium for 3 h and were changed to arginine sufficiency medium for 1 h; (D) Co-IP detection of the FAF2-mTOR interaction at insulin treatment. HepG2 cells expressing indicated plasmids were pre-cultured in serum deficiency medium for 3 h and then were subjected to insulin stimulation for 1 h. (E) Co-IP detection of the FAF2-mTOR interaction upon glucose treatment. HepG2 cells expressing indicated plasmids, followed by glucose starvation for 8 h and then glucose replenishment for 8 h. (F,G) The regulatory function of SNHG6 on FAF2-mTOR interaction was detected by Co-IP assay. GFP-mTOR, SFB-FAF2 and SNHG6 plasmids (F) or SNHG6 interfering RNA (G) were transfected to HepG2 cells for 48 h as described conditions. (H) Co-IP detection of the FAF2-mTOR interaction upon RNase treatment. Indicated plasmids were transfected into HepG2 cells for 48 h, whose cell lysates were incubated with RNase treatment for 15 mins in vitro, followed S-protein beads immunoprecipitation and IB detection using indicated antibodies. (I) Co-IP detection FAF2-mTOR interaction in Huh7 cells. shScr/shSNHG6 Huh7 cells were transfected with SFB-FAF2 plasmids for 48 h, followed by cholesterol replenishment (50 μM, 2 h). (J) Coomassie blue staining of GST-tagged FAF2 truncation.
Extended Data Fig. 5 SNHG6 promotes cholesterol biosynthesis and proliferation via mTORC1 signaling in liver cancer cells.
(A) IF staining of mTOR-LAMP2 in Huh7 cells. shScr/shSNHG6 Huh7 cells were subjected to cholesterol starvation and cholesterol replenishment (50 μM, 2 h). Scale bar, 10 μm. (B) Quantification of the colocalization of mTOR-LAMP2 in (A). Data are mean values ± S.D. n = 21 cells per condition. Two-way ANOVA. (C) IB detection of indicated proteins. Vector/ SNHG6 were transfected into HepG2 cells for 48 h, and lysosomes were purified from HepG2 cells, followed by IB detection using indicated antibodies. (D) Co-IP detection of RagA-Raptor interaction. SFB-Raptor, His-RagA plasmid, siCtrl and siRNA-SNHG6 were transfected into HepG2 cells as described conditions for 48 h. (E) Co-IP detection Raptor-RagA interaction in Huh7 cells. shScr/shSNHG6 Huh7 cells were transfected with SFB-Raptor plasmids for 48 h, followed by and cholesterol stimulation (50 μM, 2 h). (F) IB detection of indicated proteins in shScr/shSNHG6 Huh7 cells that were subjected to cholesterol stimulation (50 μM, 2 h). (G,H) IB detection of indicated proteins. HepG2 cells (G)/ Huh7 cells (H) overexpressing vector or SNHG6 were treated with cholesterol stimulation (50 μM, 2 h), followed IB detection for indicated proteins. (I-N) HepG2 cells overexpressing vector or SNHG6 were subjected to rapamycin treatment (100 nM) for 24 h. (I) IB detection of indicated proteins. (J-L) The mRNA levels of SREBF2, SQLE, HMGCS1 were detected by qRT-PCR. (M) Relative cholesterol content in HepG2 cells. (N) MTT assay of HepG2 cells with or without rapamycin. Data are mean values ± S.D. Two-way ANOVA. (O-Q) Vector/SNHG6 was transfected to PMH for 48 h. (O) IB detection of indicated protein expression. (P) qPCR analysis of Srebf2, Hmgcs1, Sqle and SNHG6 expression. (Q) Relative cholesterol concentration in PMH. Data are presented as mean values ± S.D. Two-sided Student’s t-test.
Extended Data Fig. 6 FAF2 promotes cholesterol-dependent mTORC1 signaling in liver cancer cells.
(A) IB detection of indicated proteins. Vector/ SFB-FAF2 were transfected into HepG2 cells for 48 h, followed by cholesterol stimulation (50 μM, 2 h). (B) IB detection of 70S6K and 4EBP activation. HepG2 cells stably expressing shScr/shFAF2 plasmids were subjected to cholesterol stimulation (50 μM, 2 h). (C) lysosomes were purified from shCtrl/shFAF2 HepG2 cells upon indicated cholesterol stimulation (50 μM, 2 h), followed IB detection. (D) MTT assay of HepG2 cells that were transfected with siFAF2-1/siFAF2-2 for 48 h. Data are mean values ± S.D. Two-way ANOVA. (E) Coomassie blue staining of recombinant GFT-FAF2 proteins. (F-G) Fluorescence intensity profile of GST (F)/GST-FAF2 (G) protein with increasing cholesterol concentration. (H) Coomassie blue staining of recombinant MBP-His-FAF2 mutations. (I) siRNA-SLC38A9 and SFB-FAF2 plasmids were transfected to HepG2 cells for 48 h, and then preformed cholesterol stimulation (50 μM, 2 h), followed by IB detection. (J) Transfecting SFB-vector and SFB-FAF2 plasmids to sgCtrl / sgSNHG6 HepG2 cells for 48 h and performing cholesterol treatment as indicated (50 μM, 2 h), followed IB detection.
Extended Data Fig. 7 Hepatic Snhg6 knockdown impaired NAFLD-HCC progression.
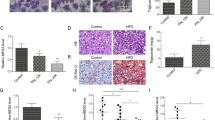
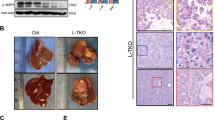
(A) Alignment of human and mouse Snhg6 sequences. (B) Detection of Snhg6 expression in Hepa1-6 cells. Data are presented as mean values ± S.D. One-way ANOVA with Tukey’s test. (C) IB detection of indicated proteins in shScr/shSnhg6 Hepa1-6 cells that were subjected to cholesterol stimulation (50 μM, 2 h). (D) qRT-PCR detection of Snhg6 expression in liver of mice fed with HFHC. n = 5 mice/normal group, n = 10 mice/NASH group, Data are presented as mean values ± S.D. Two-sided Mann-Whitney U. (E-H) AAV-Vector mice and AAV-shSnhg6 mice were fed with DEN and HFHC diet for 10 weeks. H&E staining (Scale bar, 100 μm) (E), liver weight and liver/body weight ratio (F), relative indicated RNA levels (G), and immunoblot of indicated proteins in liver sections from indicated groups (H). n = 5 mice/ group. Data are presented as mean values ± S.D. Two-sided Mann-Whitney U. (I-O) AAV-shScr mice and AAV-shSnhg6 mice were fed with DEN and HFHC diet for 18 weeks. H&E and sirius red staining (Scale bar, 100 μm) (I), quantification of sirius red staining (n = 7 fields of view per condition) (J), liver weight and liver/body weight ratio (K), serum ALT concentration (L), serum cholesterol concentration (M), relative RNA levels (N), IB detection of indicated protein expression (O) in liver of mice from indicated groups. n = 5 mice/ group. Data are presented as mean values ± S.D. Two-sided Mann-Whitney U. (P-U) AAV-Vector mice and AAV-shSnhg6 mice were fed with DEN and HFHC diet for 30 weeks. Tumor number (P), H&E, sirius red and Ki67 staining (Scale bar, 100 μm) (Q, left), quantification of sirius red and Ki67 (n = 7 fields of view per condition) (Q, right), liver weight and liver/body weight ratio (R), serum / total cholesterol concentration (S), relative RNA levels (T), and immunoblots of indicated proteins in liver of mice from indicated groups (U). Data are presented as mean values ± S.D. n = 5 mice/group. Two-sided Mann-Whitney U test.
Extended Data Fig. 8 SNHG6 promotes proliferation in hepatoma.
(A) In vivo generated tumors are shown. (B) IHC analyses of tumors were performed with the Ki67 and CD31 antibodies. Scale bar, 100 μm. (C,D) qRT-PCR analysis of SNHG6 expression in PDX tumors. Data are mean values ± S.D. n = 5 mice / group. Two-way ANOVA. (E) IHC analyses of PDX tumors were performed with the Ki67 and CyclinD1 antibodies. Scale bar, 100 μm. (F) IB detection of tumors was performed with indicated antibodies (n = 5 mice / group). (G) The amount of SNHG6 was associated with expression of Ki67 or CCNB1 in the dataset (TCGA-LIHC). Data are presented as individual data and mean, statistics by Kruskal-Wallis test. Bounds of box show the 25th and 75th percentiles, and the horizontal lines inside the box represent the median. The whiskers show 2.5th to 97.5th percentiles. (H) the expressional correlation between SQLE and SNHG6 in the dataset. The r values and P values from Spearman’s correlation analysis. (I-K) The expression of FAF2 in carcinoma and adjacent tissues in several datasets. The horizontal line inside the box is the median; the top is the upper quartile; the bottom is the lower quartile; the upper whiskers extend from the upper quartile to the highest actual value; the lower whiskers extend from the lower quartile The number is extended to the lowest actual value. Two-sided Student’s t test. (L) qRT-PCR detection of FAF2 expression in carcinoma and adjacent tissue. Data are presented as mean values ± S.D. SYSUCC, n = 48 biologically independent samples. Data are presented as mean values ± S.D. Two-sided Wilcoxon test. (M) Overall Survival was compared between samples with low and high amount of FAF2 in the liver cancer dataset (TCGA-LIHC). Kaplan-Meier analysis. (N) Overall survival analysis of FAF2 status in hepatic carcinoma detected by qRT-PCR (SYSUCC, n = 80 biologically independent samples; Kaplan-Meier analysis). (O) Correlation between FAF2 levels and MKi67 in the datasets (TCGA-LIHC). The r values and P values from Spearman’s correlation analysis.
Supplementary information
Supplementary Information
Supplementary Tables 1–7
Source data
Statistical Source Data Fig. 1
Statistical Source Data
Statistical Source Data Fig. 2
Statistical Source Data
Statistical Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3
Unprocessed Western Blots
Statistical Source Data Fig. 4
Statistical Source Data
Source Data Fig. 4
Unprocessed Western Blots
Statistical Source Data Fig. 5
Statistical Source Data
Source Data Fig. 5
Unprocessed Western Blots
Statistical Source Data Fig. 6
Statistical Source Data
Source Data Fig. 6
Unprocessed Western Blots
Statistical Source Data Fig. 7
Statistical Source Data
Source Data Fig. 7
Unprocessed Western Blots
Statistical Source Data Fig. 8
Statistical Source Data
Source Data Fig. 8
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 1
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 3
Unprocessed Western Blots
Source Data Extended Data Fig. 4
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 5
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 6
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 7
Unprocessed Western Blots
Statistical Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 8
Unprocessed Western Blots
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, F., Tian, T., Zhang, Z. et al. Long non-coding RNA SNHG6 couples cholesterol sensing with mTORC1 activation in hepatocellular carcinoma. Nat Metab 4, 1022–1040 (2022). https://doi.org/10.1038/s42255-022-00616-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-022-00616-7
This article is cited by
-
Integration of transcription regulation and functional genomic data reveals lncRNA SNHG6’s role in hematopoietic differentiation and leukemia
Journal of Biomedical Science (2024)
-
The roles and molecular mechanisms of non-coding RNA in cancer metabolic reprogramming
Cancer Cell International (2024)
-
Mobilization of cholesterol induces the transition from quiescence to growth in Caenorhabditis elegans through steroid hormone and mTOR signaling
Communications Biology (2024)
-
SOAT1 regulates cholesterol metabolism to induce EMT in hepatocellular carcinoma
Cell Death & Disease (2024)
-
LncFASA promotes cancer ferroptosis via modulating PRDX1 phase separation
Science China Life Sciences (2024)