Abstract
Single-cell motility is spatially heterogeneous and driven by metabolic energy. Directly linking cell motility to cell metabolism is technically challenging but biologically important. Here, we use single-cell metabolic imaging to measure glycolysis in individual endothelial cells with genetically encoded biosensors capable of deciphering metabolic heterogeneity at subcellular resolution. We show that cellular glycolysis fuels endothelial activation, migration and contraction and that sites of high lactate production colocalize with active cytoskeletal remodelling within an endothelial cell. Mechanistically, RhoA induces endothelial glycolysis for the phosphorylation of cofilin and myosin light chain in order to reorganize the cytoskeleton and thus control cell motility; RhoA activation triggers a glycolytic burst through the translocation of the glucose transporter SLC2A3/GLUT3 to fuel the cellular contractile machinery, as demonstrated across multiple endothelial cell types. Our data indicate that Rho-GTPase signalling coordinates energy metabolism with cytoskeleton remodelling to regulate endothelial cell motility.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Gene Expression Omnibus datasets (GEOD) were downloaded from EndoDB (https://endotheliomics.shinyapps.io/endodb/): E-GEOD-15760, E-GEOD-20741, E-MTAB-5921, E-GEOD-40999, E-GEOD-16067, E-GEOD-11870, E-GEOD-1576, E-GEOD-47067. Original data that support our findings in this study are available from the corresponding author upon reasonable request or at https://doi.org/10.5281/zenodo.4638059. Source data are provided with this paper.
Code availability
The code generated during this study is available at https://github.com/wulab-code/laconic without restriction.
References
Kim, B., Li, J., Jang, C. & Arany, Z. Glutamine fuels proliferation but not migration of endothelial cells. EMBO J. 36, 2321–2333 (2017).
Wu, D. et al. HIF-1α is required for disturbed flow-induced metabolic reprogramming in human and porcine vascular endothelium. eLife 6, e25217 (2017).
Feng, S. et al. Mechanical activation of hypoxia-inducible factor 1α drives endothelial dysfunction at atheroprone sites. Arterioscler Thromb. Vasc. Biol. 37, 2087–2101 (2017).
Doddaballapur, A. et al. Laminar shear stress inhibits endothelial cell metabolism via KLF2-mediated repression of PFKFB3. Arterioscler Thromb. Vasc. Biol. 35, 137–145 (2015).
van Nieuw Amerongen, G. P., Koolwijk, P., Versteilen, A. & van Hinsbergh, V. W. Involvement of RhoA/Rho kinase signaling in VEGF-induced endothelial cell migration and angiogenesis in vitro. Arterioscler Thromb. Vasc. Biol. 23, 211–217 (2003).
Van Valen, D. A. et al. Deep learning automates the quantitative analysis of individual cells in live-cell imaging experiments. PLoS Comput. Biol. 12, e1005177 (2016).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. 234–241 (2015).
San Martin, A. et al. A genetically encoded FRET lactate sensor and its use to detect the Warburg effect in single cancer cells. PLoS ONE 8, e57712 (2013).
Csurka, G., Larlus, D. & Perronnin, F. What is a good evaluation measure for semantic segmentation? In Proc. British Machine Vision Conference (BMVA Press, 2013).
Halestrap, A. P. Monocarboxylic acid transport. Compr. Physiol. 3, 1611–1643 (2013).
Sonveaux, P. et al. Targeting the lactate transporter MCT1 in endothelial cells inhibits lactate-induced HIF-1 activation and tumor angiogenesis. PLoS ONE 7, e33418 (2012).
San Martin, A. et al. Imaging mitochondrial flux in single cells with a FRET sensor for pyruvate. PLoS ONE 9, e85780 (2014).
Takanaga, H., Chaudhuri, B. & Frommer, W. B. GLUT1 and GLUT9 as major contributors to glucose influx in HepG2 cells identified by a high sensitivity intramolecular FRET glucose sensor. Biochim. Biophys. Acta 1778, 1091–1099 (2008).
Imamura, H. et al. Visualization of ATP levels inside single living cells with fluorescence resonance energy transfer-based genetically encoded indicators. Proc. Natl Acad. Sci. USA 106, 15651–15656 (2009).
Davies, P. F. Flow-mediated endothelial mechanotransduction. Physiol. Rev. 75, 519–560 (1995).
Mayor, R. & Etienne-Manneville, S. The front and rear of collective cell migration. Nat. Rev. Mol. Cell Biol. 17, 97–109 (2016).
Diebold, L. P. et al. Mitochondrial complex III is necessary for endothelial cell proliferation during angiogenesis. Nat. Metab. 1, 158–171 (2019).
Mole, D. R. et al. 2-Oxoglutarate analogue inhibitors of hif prolyl hydroxylase. Bioorg. Medicinal Chem. Lett. 13, 2677–2680 (2003).
Fazal, F. et al. Essential role of cofilin-1 in regulating thrombin-induced RelA/p65 nuclear translocation and intercellular adhesion molecule 1 (ICAM-1) expression in endothelial cells. J. Biol. Chem. 284, 21047–21056 (2009).
van Nieuw Amerongen, G. P., Vermeer, M. A. & van Hinsbergh, V. W. Role of RhoA and Rho kinase in lysophosphatidic acid-induced endothelial barrier dysfunction. Arterioscler Thromb. Vasc. Biol. 20, E127–E133 (2000).
van Nieuw Amerongen, G. P., van Delft, S., Vermeer, M. A., Collard, J. G. & van Hinsbergh, V. W. Activation of RhoA by thrombin in endothelial hyperpermeability: role of Rho kinase and protein tyrosine kinases. Circ. Res. 87, 335–340 (2000).
Blanchoin, L., Boujemaa-Paterski, R., Sykes, C. & Plastino, J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 94, 235–263 (2014).
Goddard, L. M. & Iruela-Arispe, M. L. Cellular and molecular regulation of vascular permeability. Thromb. Haemost. 109, 407–415 (2013).
Mehta, D. & Malik, A. B. Signaling mechanisms regulating endothelial permeability. Physiol. Rev. 86, 279–367 (2006).
Vandenbroucke, E., Mehta, D., Minshall, R. & Malik, A. B. Regulation of endothelial junctional permeability. Ann. N. Y. Acad. Sci. 1123, 134–145 (2008).
Tang, M. et al. Brain microvasculature defects and Glut1 deficiency syndrome averted by early repletion of the glucose transporter-1 protein. Nat. Commun. 8, 14152 (2017).
Fessel, J. P. et al. Metabolomic analysis of bone morphogenetic protein receptor type 2 mutations in human pulmonary endothelium reveals widespread metabolic reprogramming. Pulm. Circ. 2, 201–213 (2012).
Sasmal, D. K. et al. TCR–pMHC bond conformation controls TCR ligand discrimination. Cell Mol. Immunol. 17, 203–217 (2020).
Sasmal, D. K., Pulido, L. E., Kasal, S. & Huang, J. Single-molecule fluorescence resonance energy transfer in molecular biology. Nanoscale 8, 19928–19944 (2016).
Fraccaroli, A. et al. Endothelial α-parvin controls integrity of developing vasculature and is required for maintenance of cell-cell junctions. Circ. Res. 117, 29–40 (2015).
Khan, S. et al. EndoDB: a database of endothelial cell transcriptomics data. Nucleic Acids Res. 47, D736–D744 (2019).
Wu, C. et al. Mechanosensitive PPAP2B regulates endothelial responses to atherorelevant hemodynamic forces. Circ. Res. 117, e41–e53 (2015).
Yao, L., Xue, X., Yu, P., Ni, Y. & Chen, F. Evans blue dye: a revisit of its applications in biomedicine. Contrast Media Mol. Imaging 2018, 7628037 (2018).
Riedl, J. et al. Lifeact: a versatile marker to visualize F-actin. Nat. Methods 5, 605–607 (2008).
Pertz, O., Hodgson, L., Klemke, R. L. & Hahn, K. M. Spatiotemporal dynamics of RhoA activity in migrating cells. Nature 440, 1069–1072 (2006).
De Bock, K. et al. Role of PFKFB3-driven glycolysis in vessel sprouting. Cell 154, 651–663 (2013).
Wilhelm, K. et al. FOXO1 couples metabolic activity and growth state in the vascular endothelium. Nature 529, 216–220 (2016).
Karmaus, P. W. F. et al. Metabolic heterogeneity underlies reciprocal fates of TH17 cell stemness and plasticity. Nature 565, 101–105 (2019).
Bantug, G. R., Galluzzi, L., Kroemer, G. & Hess, C. The spectrum of T cell metabolism in health and disease. Nat. Rev. Immunol. 18, 19–34 (2018).
Hensley, ChristopherT. et al. Metabolic heterogeneity in human lung tumors. Cell 164, 681–694 (2016).
van Heerden, J. H. et al. Lost in transition: start-up of glycolysis yields subpopulations of nongrowing cells. Science 343, 1245114 (2014).
Huang, J. et al. A single peptide-major histocompatibility complex ligand triggers digital cytokine secretion in CD4+ T cells. Immunity 39, 846–857 (2013).
Masters, C. Interactions between glycolytic enzymes and components of the cytomatrix. J. Cell Biol. 99, 222s–225s (1984).
Huang, Y. et al. Normal glucose uptake in the brain and heart requires an endothelial cell-specific HIF-1α-dependent function. Proc. Natl Acad. Sci. USA 109, 17478–17483 (2012).
Kuo, A., Lee, M. Y. & Sessa, W. C. Lipid droplet biogenesis and function in the endothelium. Circ. Res. 120, 1289–1297 (2017).
Yu, P. et al. FGF-dependent metabolic control of vascular development. Nature 545, 224–228 (2017).
Sorbara, L. R. et al. Thrombin-induced translocation of GLUT3 glucose transporters in human platelets. Biochem. J. 328, 511–516 (1997).
Huang, R. T. et al. Experimental lung injury reduces Kruppel-like factor 2 to increase endothelial permeability via regulation of RAPGEF3–Rac1 signaling. Am. J. Respir. Crit. Care Med 195, 639–651 (2017).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Li, L. et al. Artificial virus delivers CRISPR–Cas9 system for genome editing of cells in mice. ACS Nano 11, 95–111 (2017).
Barbee, K. A., Davies, P. F. & Lal, R. Shear stress-induced reorganization of the surface topography of living endothelial cells imaged by atomic force microscopy. Circ. Res. 74, 163–171 (1994).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Acknowledgements
This work was supported by NIH grants R00AI106941 (J.H.), R21AI120010 (J.H.), R01HL138223 (Y.F.) and R01HL136765 (Y.F.), NIH New Innovator Award DP2AI144245 (J.H.), T32EB009412 (D.L.H.), T32HL007381 (D.L.H.) and F32HL134288 (D.W.) and NIH Pathway to Independence Award K99HL145113 (D.W.) and R00HL145113 (D.W.). NVIDIA GPU Grant (D.W.), CSCTR Early Career Development Award (D.W.) and NSF Career Award 1653782 (J.H.) also supported this work.
Author information
Authors and Affiliations
Contributions
D.W. and D.L.H. planned and executed experiments, analysed data and interpreted results. T.S. analysed data. C.-F.Y., A.M., T.-P.S., R.-T.H. and Z.Z. performed experiments. J.H. and Y.F. planned experiments and interpreted results. G.M.M. helped to design experiments. D.W., D.L.H., J.H. and Y.H. wrote and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Ali Ertürk, Marc Tramier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-cell metabolic assay utilizes deep learning-enabled segmentation.
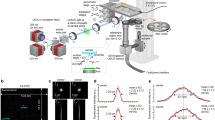
a, mTFP/Venus (or 1/FRET) change is linearly correlated over 6 orders of magnitude of log lactate (n = 287 cells; black line is semilog fit with R2 = 0.96, gray dotted lines are 95% confidence interval); error bars are SEM. Standard deviation of regression coefficients for R2 for (a) is 0.049. b, Deep learning architecture for semantic segmentation, modified from Ronneberger et al7. The U-Net consisted of a contracting path (encoder) and an expansive path (decoder). The contracting path consists of layers composed of two 3 × 3 convolutions, each followed by a rectified linear unit (ReLU) activation function. Each layer is followed by a 2 × 2 max pooling operation with a stride of two- which will double the number of feature channels being used. The convolutions start with 64 feature channels and 128 × 128 images and continue until they reach 512 channels and 16 × 16 images. In the expansive path, 2 × 2 up-convolutions (up sampling) of stride 2 to decrease by half the feature channels and increase the size of the image. The layers composed of 3 × 3 convolutions and ReLU are concatenated with the pair of layers in the contracting side, in order to reinclude the localization information. In the final layer, a 1 × 1 convolution layer is used to map the resulting 64-component feature channels vector to the 4 segmentation channels for background, boundary, cytoplasm, and nuclei. c, Ground Truth Fluorescence vs. Predicted Fluorescence, as evaluated with deep-learning-enabled semantic segmentation. (n ~ 20,000 cells) Ground truth fluorescence was defined as signal inside the cell boundary. C.I., confidence interval. d, Frequency distribution of fluorescence slopes. Data from (c) were randomly sampled 100 times and fitted to a line. This process was repeated 10,000 times and the distribution of fitted slopes was plotted (open black circles). The red line is a Gaussian fit with mean 0.9 and standard deviation of 0.05. Thus, the error in estimating slopes of fluorescence is approximately 0.05/0.9, or approximately 5%.
Extended Data Fig. 2 Single-cell laconic accurately mimics bioenergetics assays.
a, Extracellular flux assay of HAECs treated with glucose then pCMBA (n = 5) using Seahorse; biological replicates, error bars are s.e.m., P < 0.0001 for both comparisons. b, A single-cell glycolytic stress test using lactate FRET sensor. Cells were first starved in zero glucose buffer for 1 hour. Glucose addition increased intracellular lactate, which was further increased by adding oligomycin and glucose. Addition of 2-deoxyglucose (2-DG) blocked all glycolysis, causing intracellular lactate to drop. c, Overnight treatment of endothelial cells with 500 μM DMOG increases ECAR measured by extracellular flux relative to DMSO-treated cells using Seahorse. (n = 11 combined biological and technical replicates for each condition; error bars are s.e.m., P < 0.0001). Statistical significance determined by one-way ANOVA followed by Bonferroni test (a) or by two-sided Welch’s t-test (c). ***P <= 0.0005.
Extended Data Fig. 3 RhoA activity induces contraction using glycolysis.
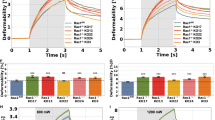
a, LPR of HAECs transfected with either RhoA-T19N-eBFP2 (dominant negative (DN) form, cyan) or RhoA-Q63L-mScarlet-i (constitutively active (CA) form, red), mixed, then deconvoluted by colour (n = 174 “cyan” and 52 “red”, respectively). Error bars are s.e.m., P = 0.0193. b, TEER measurement of HAECs after thrombin treatment, either pre-treated with Y27632 or 2DG compared to control (no Y27632 or 2DG). Statistical significance determined by two-sided Welch’s t-test. *P <= 0.05.
Extended Data Fig. 4 SLC2A3 but not SLC2A1 knockdown mitigates thrombin-induced phosphorylation of CFL and MLC and glycolysis.
a, Western blot of SLC2A1, SLC2A3, phospho-cofilin and phospho-MLC in the presence or absence of siRNA targeted towards SLC2A1 and thrombin treatment (n = 3 biological replicates). Loading control for each protein is ACTB unless specifically denoted in parentheticals. Phospho-proteins are additionally normalized to total. Original blot in Source Data c. b, Heatmap of glucose transporter expression relative to SLC2A1 in HAECs captured by qRT-PCR (n = 3 biological replicates) c-d, Western blot of SLC2A1, SLC2A3, phospho-cofilin and phospho-MLC in the presence or absence of two distinct siRNA (separate from the one used in Fig. 5a) (#1 in c and #2 in d) targeted towards SLC2A3 and thrombin treatment (n = 3 biological replicates. For c, P = 0.0190 for pCFL and 0.0490 for ppMLC. For d, P = 0.0272 for ppMLC.). Loading control for each protein is ACTB unless specifically denoted in parentheticals. Phospho-proteins are additionally normalized to total. Original blot in Source Data d-e. e, Single-cell LPR of HAECs treated with thrombin in the presence of siSLC2A1 or siSLC2A3 (n = 277 for thrombin vs. control, n = 281 for thrombin vs. thrombin + siSLC2A1, and n = 278 for thrombin vs. thrombin + siSLC2A3, P < 0.0001, P = 0.0384, P < 0.0001, respectively.). f-g, Glucose (f) and ATP levels (g) in single-cells in the presence or absence of SLC2A3 knockdown and thrombin treatment (f: n = 248, 248, 108, 190 from left to right, P < 0.0001 for all comparisons; g: 190, 598, 150, 287 from left to right, P < 0.0001 for all comparisons). h, Gap index of cells treated with or without thrombin in the presence or absence of SLC2A3 knockdown (n= 12, 16, 8, 32 from left to right). i-j, Representative images of HAECs treated with siSLC2A3 or control with varying concentrations of glucose (i) or with oligomycin (j) in the presence of thrombin. Red color is CDH5. Gap size index and gap index quantification of images (i, n = 10, 10, 7, 8 from left to right, p = 0.022 for gap size index and P = 0.0019; j, n = 6 for gap size index, P = 0.0013 for siSLC2A3 vs control and P = 0.040 for siSLC2A3 vs. siSLC2A3 + oligomycin, and n = 10, 11, 7 from left to right for gap index, P < 0.0001). White stars highlight gaps in the monolayer. Scale bars = 250 μm. Statistical significance determined by multiple unpaired two-tailed t-tests (a, c, d), one-way ANOVA followed by Bonferroni test (e-g, i) or by two-sided Welch’s t-test (h, j). All error bars are s.e.m. *P <= 0.05; **P <= 0.005; ***P <= 0.0005.
Extended Data Fig. 5 SLC2A3 inhibition abrogates thrombin-induced phosphorylation of CFL and MLC and contraction across multiple endothelial cell types.
a-b, Western blot analysis of glucose transporter expression, phosphorylated CFL, and phosphorylated MLC, in human umbilical vein endothelial cells (HUVEC, a, n = 3; biological replicates, P = 0.0257 for pCFL and 0.0414 or ppMLC) and human microvascular endothelial cells (HMVEC, b, n = 3 for SLC2A1, SLC2A3, n = 4 for CFL and n = 5 for MLC; biological replicates, P < 0.001 for pCFL and P= 0.0014 for ppMLC). Loading control for each protein is ACTB unless specifically denoted in parentheticals. Original blot in Source Data f-g. c, Representative images and cell size of contractile HUVECs subjected to thrombin stimulation in the absence or presence of SLC2A3 knockdown. (n = 1375, 1279, 760, 1162 from left to right, P < 0.0001) d, Representative images, gap size index, and gap index of contractile HMVECs subjected to thrombin stimulation in the absence or presence of SLC2A3 knockdown. Representative images are phalloidin. (n = 4 biological replicates, P = 0.0171 for gap size index and P = 0.0262 for gap index) Statistical significance determined by two-sided Welch’s t-test. All error bars are s.e.m. n.s., not significant. *P <= 0.05; **P <= 0.005; ***P <= 0.0005.
Extended Data Fig. 6 In vivo overexpression of SLC2A3 increased vascular leak.
a, GEO datasets of primary endothelial mouse cells comparing Slc2a1 and Slc2a3 from different organs. Fold change indicates that data are normalized to Slc2a1 except for E-GEOD-47067 which was normalized to average of Slc2a1 in all tissues. Only controls of experiments in the GEO datasets were used in these analyses. (n = 3 independent biological replicates for each condition for E-GEOD-15760; n = 6 independent biological replicates for each condition for E-GEOD-20741; n = 3 independent biological replicates for each condition for E-MTAB-5921; n = 3 independent biological replicates for each condition for E-GEOD-40999; n = 3 independent biological replicates for each condition for E-GEOD-11870; n = 3 independent biological replicates for each condition for E-GEOD-1576; n = 3 independent biological replicates for condition in each organ for E-GEOD-47067) b, Immunofluorescence of an arterial section demonstrating enhanced mScarlet-i fluorescence (red) colocalized with ve-cadh (green) which marks the endothelium next to the lumen (yellow, merge) (Scale bar, 500 μm, representative of n = 4 mice). c, SLC2A3 expressing plasmid uses an endothelial specific CDH5 promoter. SLC2A3 is detected by qRT-PCR of the intima of the mouse aorta, and far less than in the media and adventitia. (n = 4 independent biological replicates, P = 0.0030) d, Evans blue (OD 620 nm) quantification of mouse aortas, normalized by weight. (n = 8 independent biological replicates for each condition, P = 0.0050). Scale bar is 1 mm. Statistical significance was determined by two-sided Welch’s t-test. All error bars are s.e.m. **P <= 0.005.
Extended Data Fig. 7 Subcellular RhoA activity, actin turnover, and LPR distribution in single motile endothelial cells.
a, Montage of actin timelapse in a migratory endothelial cell; maximum projection of change in actin structure (Lifeact difference image, obtained by sequential subtraction of preceding 10 images, taken 1 minute apart, followed by maximum intensity projection of the absolute value of the differences), per pixel LPR map (LPR), Lifeact difference and LPR overlap (“difference+LPR”, merge), scale bar = 20 µm. White arrows in montage indicate cell spreading. Green outline shows enhanced LPR near the regions of cell spreading. Representative of n = 14 cells, 3 separate experiments. b, Pearson’s correlation coefficient (PCC) and Manders’ overlap coefficient (MOC) of Lifeact difference and LPR, compared to random. The random PCC/MOC is calculated by randomizing the LPR image and computing with the Lifeact difference image. Fold change of experimental PCC/MOC over randomized is shown. (n = 14). Error bars are s.e.m, P = 0.014 for PCC and P = 0.0489 for MOC. c-l, RhoA activity measured by RhoA-FLARE in contractile endothelial cells. Representative data of total n = 9 cells in 3 replicates. (Scale bar = 16 μm in c, g, k. Scale bar = 7 μm in d, f, h, l). d,f, regions of contractile cell from (c) showing RhoA membrane activity. Circle in (f) notes moving RhoA activity. e, kymograph showing contractile edge from (d) and increasing RhoA activity over time. h, region of contractile cell from (g) showing increased RhoA membrane activity. i-j, kymograph showing non-contractile edge (i) without RhoA increase and contractile edge (j) from (g) and increasing RhoA activity over time. l, region of contractile cell from (k) showing RhoA membrane activity. Dotted ellipses notes highlight increasing RhoA activity along with active contraction. Statistical significance determined by two-sided Welch’s t-test. *P <= 0.05.
Supplementary information
Supplementary Information
Supplementary Methods
Supplementary Tables
Supplementary Tables 1–3
Source data
Source Data Fig. 4
Unmodified western blots with markers for Fig. 4f
Source Data Fig. 5
Unmodified western blots with markers for Fig. 5a
Source Data Extended Data Fig. 4
Unmodified western blots with markers for Extended Data Fig. 4a–c
Source Data Extended Data Fig. 5
Unmodified western blots with markers for Extended Data Fig. 5a,b
Rights and permissions
About this article
Cite this article
Wu, D., Harrison, D.L., Szasz, T. et al. Single-cell metabolic imaging reveals a SLC2A3-dependent glycolytic burst in motile endothelial cells. Nat Metab 3, 714–727 (2021). https://doi.org/10.1038/s42255-021-00390-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00390-y
This article is cited by
-
Lactate biosensors for spectrally and spatially multiplexed fluorescence imaging
Nature Communications (2023)
-
Monitoring NAD(H) and NADP(H) dynamics during organismal development with genetically encoded fluorescent biosensors
Cell Regeneration (2022)
-
The Vasculature in Pulmonary Fibrosis
Current Tissue Microenvironment Reports (2022)
-
Endothelial metabolism going single
Nature Metabolism (2021)