Abstract
Benign hepatosteatosis, affected by lipid uptake, de novo lipogenesis and fatty acid (FA) oxidation, progresses to non-alcoholic steatohepatitis (NASH) on stress and inflammation. A key macronutrient proposed to increase hepatosteatosis and NASH risk is fructose. Excessive intake of fructose causes intestinal-barrier deterioration and endotoxaemia. However, how fructose triggers these alterations and their roles in hepatosteatosis and NASH pathogenesis remain unknown. Here we show, using mice, that microbiota-derived Toll-like receptor (TLR) agonists promote hepatosteatosis without affecting fructose-1-phosphate (F1P) and cytosolic acetyl-CoA. Activation of mucosal-regenerative gp130 signalling, administration of the YAP-induced matricellular protein CCN1 or expression of the antimicrobial peptide Reg3b (beta) peptide counteract fructose-induced barrier deterioration, which depends on endoplasmic-reticulum stress and subsequent endotoxaemia. Endotoxin engages TLR4 to trigger TNF production by liver macrophages, thereby inducing lipogenic enzymes that convert F1P and acetyl-CoA to FA in both mouse and human hepatocytes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA-seq data have been deposited in NCBI Gene Expression Omnibus (GEO) database under accession number GSE119080. Microbiome sequencing data have been deposited in EMBL-EBI under accession number ERP110352. Source data are provided with this paper.
References
Spengler, E. K. & Loomba, R. Recommendations for diagnosis, referral for liver biopsy, and treatment of nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Mayo Clin. Proc. 90, 1233–1246 (2015).
Stickel, F. & Hellerbrand, C. Non-alcoholic fatty liver disease as a risk factor for hepatocellular carcinoma: mechanisms and implications. Gut 59, 1303–1307 (2010).
Tilg, H. & Moschen, A. R. Evolution of inflammation in nonalcoholic fatty liver disease: the multiple parallel hits hypothesis. Hepatology 52, 1836–1846 (2010).
Fukui, H. Increased intestinal permeability and decreased barrier function: does it really influence the risk of inflammation? Inflamm. Intest. Dis. 1, 135–145 (2016).
Lebeaupin, C. et al. Endoplasmic reticulum stress signalling and the pathogenesis of non-alcoholic fatty liver disease. J. Hepatol. 69, 927–947 (2018).
Puri, P. et al. Activation and dysregulation of the unfolded protein response in nonalcoholic fatty liver disease. Gastroenterology 134, 568–576 (2008).
Rahman, K. et al. Loss of junctional adhesion molecule a promotes severe steatohepatitis in mice on a diet high in saturated fat, fructose, and cholesterol. Gastroenterology 151, 733–746.e12 (2016).
Nakagawa, H. et al. ER stress cooperates with hypernutrition to trigger TNF-dependent spontaneous HCC development. Cancer Cell 26, 331–343 (2014).
Kim, J. Y. et al. ER stress drives lipogenesis and steatohepatitis via caspase-2 activation of S1P. Cell 175, 133–145.e15 (2018).
Vos, M. B. & Lavine, J. E. Dietary fructose in nonalcoholic fatty liver disease. Hepatology 57, 2525–2531 (2013).
Jin, R. et al. Fructose induced endotoxemia in pediatric nonalcoholic fatty liver disease. Int. J. Hepatol. 2014, 560620 (2014).
Kavanagh, K. et al. Dietary fructose induces endotoxemia and hepatic injury in calorically controlled primates. Am. J. Clin. Nutr. 98, 349–357 (2013).
Spruss, A., Kanuri, G., Stahl, C., Bischoff, S. C. & Bergheim, I. Metformin protects against the development of fructose-induced steatosis in mice: role of the intestinal barrier function. Lab. Investig. 92, 1020–1032 (2012).
Lambertz, J., Weiskirchen, S., Landert, S. & Weiskirchen, R. Fructose: a dietary sugar in crosstalk with microbiota contributing to the development and progression of non-alcoholic liver disease. Front. Immunol. 8, 1159 (2017).
Oh, J.-H. et al. Dietary fructose and microbiota-derived short-chain fatty acids promote bacteriophage production in the gut symbiont Lactobacillus reuteri. Cell Host Microbe 25, 273–284.e6 (2019).
Chang, P. V., Hao, L., Offermanns, S. & Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl Acad. Sci. USA 111, 2247–2252 (2014).
Kelly, C. J. et al. Crosstalk between microbiota-derived short-chain fatty acids and intestinal epithelial HIF augments tissue barrier function. Cell Host Microbe 17, 662–671 (2015).
Geidl-Flueck, B. & Gerber, P. A. Insights into the hexose liver metabolism—glucose versus fructose. Nutrients 9, 1026 (2017).
Softic, S., Cohen, D. E. & Kahn, C. R. Role of dietary fructose and hepatic de novo lipogenesis in fatty liver disease. Dig. Dis. Sci. 61, 1282–1293 (2016).
Softic, S. et al. Divergent effects of glucose and fructose on hepatic lipogenesis and insulin signaling. J. Clin. Invest. 127, 4059–4074 (2017).
Grivennikov, S. I. et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 491, 254–258 (2012).
Karin, M. & Clevers, H. Reparative inflammation takes charge of tissue regeneration. Nature 529, 307–315 (2016).
Barnett, M. P. G. et al. Changes in colon gene expression associated with increased colon inflammation in interleukin-10 gene-deficient mice inoculated with Enterococcus species. BMC Immunol. 11, 39 (2010).
Siddiqui, R. A. et al. Comparative study of the modulation of fructose/sucrose-induced hepatic steatosis by mixed lipid formulations varying in unsaturated fatty acid content. Nutr. Metab. (Lond.) 12, 41 (2015).
Landy, J. et al. Tight junctions in inflammatory bowel diseases and inflammatory bowel disease associated colorectal cancer. World J. Gastroenterol. 22, 3117–3126 (2016).
Jaeken, J., Pirard, M., Adamowicz, M., Pronicka, E. & van Schaftingen, E. Inhibition of phosphomannose isomerase by fructose 1-phosphate: an explanation for defective N-glycosylation in hereditary fructose intolerance. Pediatr. Res. 40, 764–766 (1996).
Kaser, A. & Blumberg, R. S. Endoplasmic reticulum stress and intestinal inflammation. Mucosal Immunol. 3, 11–16 (2010).
Balda, M. S. & Matter, K. Tight junctions. J. Cell Sci. 111(Pt 5), 541–547 (1998).
Shalapour, S. et al. Inflammation-induced IgA+ cells dismantle anti-liver cancer immunity. Nature 551, 340–345 (2017).
Vos, M. B., Kimmons, J. E., Gillespie, C., Welsh, J. & Blanck, H. M. Dietary fructose consumption among US children and adults: the Third National Health and Nutrition Examination Survey. Medscape J. Med. 10, 160 (2008).
Low, B. C. et al. YAP/TAZ as mechanosensors and mechanotransducers in regulating organ size and tumor growth. FEBS Lett. 588, 2663–2670 (2014).
Choi, J. S., Kim, K.-H. & Lau, L. F. The matricellular protein CCN1 promotes mucosal healing in murine colitis through IL-6. Mucosal Immunol. 8, 1285–1296 (2015).
Taniguchi, K. et al. A gp130–Src–YAP module links inflammation to epithelial regeneration. Nature 519, 57–62 (2015).
Miki, T., Holst, O. & Hardt, W.-D. The bactericidal activity of the C-type lectin RegIIIβ against Gram-negative bacteria involves binding to lipid A. J. Biol. Chem. 287, 34844–34855 (2012).
Schröder, M. & Kaufman, R. J. ER stress and the unfolded protein response. Mutat. Res. 569, 29–63 (2005).
Metidji, A. et al. The environmental sensor AHR protects from inflammatory damage by maintaining intestinal stem cell homeostasis and barrier integrity. Immunity 49, 353–362.e5 (2018).
Wiest, R., Albillos, A., Trauner, M., Bajaj, J. S. & Jalan, R. Targeting the gut–liver axis in liver disease. J. Hepatol. 67, 1084–1103 (2017).
Ammirante, M., Luo, J.-L., Grivennikov, S., Nedospasov, S. & Karin, M. B-cell-derived lymphotoxin promotes castration-resistant prostate cancer. Nature 464, 302–305 (2010).
Fu, S. et al. Polysome profiling in liver identifies dynamic regulation of endoplasmic reticulum translatome by obesity and fasting. PLoS Genet. 8, e1002902 (2012).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinf. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinf. 10, 161 (2009).
Luo, W. & Brouwer, C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Marotz, C. et al. DNA extraction for streamlined metagenomics of diverse environmental samples. Biotechniques 62, 290–293 (2017).
Amir, A. et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems 2, e00191–16 (2017).
Kuczynski, J. et al. Using QIIME to analyze 16S rRNA gene sequences from microbial communities. Curr. Protoc. Bioinformatics 36, 10.7 (2011).
Lee, W. N. et al. In vivo measurement of fatty acids and cholesterol synthesis using D2O and mass isotopomer analysis. Am. J. Physiol. Metab. 266, E699–E708 (1994).
Wallace, M. et al. Enzyme promiscuity drives branched-chain fatty acid synthesis in adipose tissues. Nat. Chem. Biol. 14, 1021–1031 (2018).
Kireeva, M. L., MO, F. E., Yang, G. P. & Lau, L. F. Cyr61, a product of a growth factor-inducible immediate-early gene, promotes cell proliferation, migration, and adhesion. Mol. Cell. Biol. 16, 1326–1334 (1996).
Mandal, S. et al. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb. Ecol. Health Dis. 26, 27663 (2015).
Acknowledgements
We thank M. Raffatellu for advice and discussion and V. Sheen, W. Gong, J. Yung and K. Lam for technical support. Research was supported by grants from the NIH (P42ES010337, R01DK120714, R01CA198103, R01AI043477, R01CA211794 and R01CA234128 to M.K.; R03CA223717 to J.T.; T32AI007469 and K22AI139444 to R.MN..; R01CA192642, R01CA218254 to M.T.D.-M.; R01DK108743, R01CA207177 and R01CA211794 to J.M.; U01AA027681 to S.S. and M.K.; and R01CA188652 to C.M.M.), JSPS KAKENHI (JP15K21775) and ‘Kibou Projects’ Startup Support for Young Researchers in Immunology (to K.T.), and the Australian NHMRC (APP112227) to M.A.F. and M.K.; work in F.R.G.’s laboratory was supported by institutional funds from the Georg-Speyer-Haus and by the LOEWE Center Frankfurt Cancer Institute (FCI), funded by the Hessen State Ministry for Higher Education, Research and the Arts (III L 5 - 519/03/03.001 - (0015)), NIH K01DK116917 and P30DK063491 pilot award to J.D.W.; and S10OD020025, R01ES027595, and P42ES010337 to M.J. M.A.F. is a senior principal research fellow of the NHMRC Australia (APP 1116936).
Author information
Authors and Affiliations
Contributions
J.T. designed the project, performed most experiments, analysed data and wrote the paper with M.K., who conceived and supervised the project; G.D.C. performed major experiments; S.R. performed and analysed RNA-seq data; D.C.H. and P.J.M. performed lipidomic studies; C.R.G. and C.M.M. designed and conducted DNL measurements; A.V. and R.K. conducted microbiome analysis and contributed to writing; K.T. generated gp130Act mice; F.C., C.C. and F.R.G. generated and analysed Reg3bIEC mice; L.F.L. produced recombinant CCN1; S.S. performed DSS treatment and tissue and data analysis; R.M.N. performed further IB experiments; J.M. and M.T.D.-M. supervised metabolomic studies; M.A.F. designed and supervised RNA-seq and lipidomic studies. X.L. and T.K. provided human hepatocytes, J.D.W., R.M., M.N. and M.J. planned, optimized and performed fructose metabolite measures. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.K. holds a US patent on the use of MUP-uPA mice to study NASH and HCC (10,034,462 B2) and had received research support from Jansen Pharmaceuticals. All other authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Elena Bellafante.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 HFrD stimulates HCC development without affecting body weight or white adipose tissue.
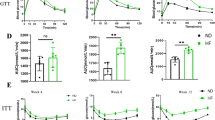
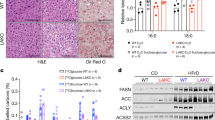
a, Food consumption by MUP- uPA and BL6 mice given CSD (for MUP-uPA n = 4 cages/9 male mice; for BL6 n = 5 cages/ 11 male mice) or HFrD (for MUP-uPA n = 4 cages/12 male mice; for BL6 n = 5 cages/ 12 male mice). b, Formalin-fixed, paraffin-embedded (FFPE) and frozen liver tissue sections of DEN- challenged male BL6 mice were stained with H&E or ORO (n = 6 per group). Representative images are shown. Scale bars, 100 μm. c, Total triglycerides in livers of DEN- challenged male BL6 mice (n = 6 per group) are presented as fold-change relative to CSD-fed mice. d, e, Body and WAT weight of male MUP-uPA (n = 9 per group) (d) and DEN-challenged male BL6 (n = 11 for CSD group at 6 months and n = 9 for CSD group at 9 months; n=12 for HFrD group at 6 months and n=9 for HFrD group at 12 months) (e) mice given CSD or HFrD were measured at the indicated time points. f, Colon length in indicated mice (MUP-uPA n = 9, BL6 n=12, per group). g, h, MUP-uPA mice (n = 15 per group) (g) or DEN- challenged BL6 mice (n = 14 per group) (h) fed HFrD (▲) or CSD (■) for 5 months were subjected to glucose tolerance tests (GTT). i, j, CSD- or HFrD- fed MUP-uPA mice (n = 4 male mice per group) (i) or DEN-challenged BL6 mice (n = 6 male mice per group) (j) were placed in metabolic cages for two-and-a-half 24-hour cycles composed of two 12-hour light periods and three dark periods. VO2, VCO2, heat production and body weights were measured. k, Non-fasting serum insulin was measured by ELISA in indicated male mice (n=8 per group). l, Liver tumor histology in MUP-uPA and DEN- challenged BL6 mice given either CSD, HFrD, or HFrD + Abx cocktail (MUP-uPA n = 7, BL6 n=12 per group). FFPE tumor sections were stained with hematoxylin and eosin (H&E). Representative images are shown. Scale bars, 100 μm. Unpaired two-sided Student’s t test was used in panels c to f and k, whereas two way ANOVA and Sidak’s multiple comparison test was used in panels g-j. Mean ± SEM, *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 2 Fructose effects on DNL and lipogenic enzymes.
a, Cytosolic acetyl-CoA concentrations in 6-month-old CSD and HFrD fed MUP - uPA male mice (n = 9 per CSD group, n=7 per HFrD group). b, Expression of genes encoding FA-oxidizing enzymes in CSD- or HFrD- fed (5 months) MUP-uPA (n=7 per group) and DEN-challenged BL6 male mice (n = 9 per group). c, A metabolic chart comparing pathways through which glucose and fructose are converted to acetyl-CoA in the cytoplasm. GK-glucokinase; KHK-ketohexokinase; GI-glucose isomerase; PFK-phosphofructokinase. Cytoplasmic acetyl-CoA is converted to malonyl-CoA by acetyl-CoA carboxylase (ACC) and malonyl- CoA serves as the building block for synthesis of C16:0 (palmitate) and C18:0 (stearate) by fatty acid synthase (FAS). Expression of ACC1 and FAS is strongly upregulated by prolonged HFrD feeding. d, e, Measurement of newly synthesized FA in livers (d) and jejunum (e) of MUP-uPA male mice using D2O as a tracer after a short-term feeding period (48 hours; n = 7 per group). f, Amounts of F1P in liver and jejunum expressed in arbitrary units (48 hours feeding; n = 7 per group). g, h, Expression of inflammatory and lipogenic genes and ACC1 protein in livers (g) and fecal albumin and serum FITC-dextran concentrations (h) in CSD- or HFrD-fed (48 hours) male mice (n = 7 per group). Unpaired two-sided Student’s t test was used in panels a, b and d-h. Mean ± SEM, *P < 0.05, **P < 0.01. Benjamini-Hochberg FDR adjustment for P-values has been performed on data presented in panels b and g.
Extended Data Fig. 3 HFrD feeding downregulates TJP mRNAs and induces ER stress, colonic inflammation and barrier deterioration.
a, b, Expression of TJP genes in colon (a) and jejunum (b) of MUP- uPA male mice fed HFrD with (a, n = 6 or 7; b, n = 6) or without (a, n = 9; b, n = 7) concomitant Abx treatment. c, IB analysis of indicated TJP in colon tissue of above mice (n = 9 no Abx, n=7 plus Abx). Representative blots are shown. d, Expression of Il22 mRNA and IL-22 regulated genes in colon tissue of CSD-, HFrD-, or HFrD + Abx-fed MUP- uPA male mice (n = 9 no Abx, n = 7 plus Abx). e, f, Male BL6 mice (e, n = 8 per group; f = 10 per group) were given 30% fructose in drinking water and fed NCD for 5 (e) or 3 (f) months and mRNA amounts of TJP genes in colon and jejunum (e), colon length and FITC-dextran serum levels were measured (f). g, h, Male BL6 mice were given regular water with or without 30% sucrose and fed NCD for 3 months. Colon length and FITC-dextran serum levels (n = 10 per group) (g), colon TJP mRNAs (h) and ER stress marker mRNAs (i) (h and i, n = 9 per regular water group and n = 8 per 30% sucrose group) were measured. j, Chop and sXbp1 mRNA amounts in fructose and glucose treated organoids (isolated from 3 different male mice). j-m, Expression of indicated mRNAs in BL6 enteroids (isolated from 3 different male mice) maintained in media containing the indicated glucose and/or fructose concentrations (mM) and treated with KHK inhibitor (k and l), TUDCA (m) or vehicle. Unpaired two-sided Student’s t test was used in all panels, other than c, to determine Mean ± SEM, *P < 0.05, **P < 0.01, ***P < 0.001. Benjamini- Hochberg FDR adjustment for P-values has been performed on data presented in panels a, b, d, e and h-m. Experiments on enteroids have been repeated twice with similar results.
Extended Data Fig. 4 Fructose-induced alterations in the liver transcriptome.
a, KEGG pathway map of innate immune signaling with HFrD-induced genes indicated in red. b, Heatmap depicting the entire transcriptome of non-tumor liver tissue from DEN-treated BL6 male mice kept on CSD (n = 3) or HFrD (n = 2) for 5 months and analyzed immediately thereafter. c, Heatmap depicting the entire transcriptome of tumor-bearing liver tissue from DEN-treated BL6 male mice kept on CSD (n = 2) or HFrD (n = 3) until 9 months old. d, Heatmap depicting expression of HCC-related genes in non- tumor and tumor liver tissues from mice described in (b). The color code represents relative mRNA abundance where red shows overexpression and blue shows under expression. e, Most significant Gene Ontology enrichment terms for Biological Pathways for the genes shown in heatmap (b) comparing DEN-treated BL6 mice fed a CSD or HFrD at 6 months of age. The bigger the dot, the more genes fall in the respective category. The color code represents the adjusted P- value (all are significant and below a cut-off level of 0.01) determined using Benjamini-Hochberg FDR adjustment.
Extended Data Fig. 5 Fructose-induced endotoxemia, hepatosteatosis, HCC, liver fibrosis, glucose intolerance and colonic inflammation are inhibited by antibiotics.
a, CSD and HFrD-fed MUP-uPA (n=8 CSD, n=9 HfrD, n=7 HFrD+Abx) and DEN- challenged BL6 (n = 9 per group) male mice were treated or not with a broad-spectrum Abx cocktail for 5 months, after which serum endotoxin concentrations were measured. b, HCC burden in MUP-uPA (n=9 HFrD, n=7 HFrD+Abx) and DEN- challenged BL6 (n=11 HFrD, n=9 HFrD+Abx) male mice given HFrD alone or HFrD plus Abx cocktail. c, Hepatosteatosis in MUP-uPA and DEN-challenged BL6 male mice given HFrD ± Abx for 5 months determined by ORO staining of liver sections (n = 6). Scale bars, 100 μm. d, Liver triglyceride content in above mice (n = 6 per group). e–g, Expression of lipogenic (e) and inflammatory (f) genes (n = 9 HFrD, n = 7 HFrD+Abx) and IB analysis of liver ACC1 and FAS (g). h, i, Liver F1P abundance (MUP-uPA; n = 7 samples from different mice per group) (h) and cytosolic acetyl-CoA (n = 8 HFrD, n = 7 HFrD+Abx samples from different mice per group; the HFrD bar is same as in Extended Data Fig. 2a). j, k, glucose tolerance (MUP-uPA, n = 10 serum samples from different mice per group, BL6, n = 7 serum samples from different mice per group) (j) and non- fasting serum insulin (n = 8 HFrD, n = 7 HFrD+Abx serum samples from different mice per group) (k). l, Total body and white adipose tissue weights in indicated male mice (MUP- UPA, n = 9 HFrD, n = 7 HFrD+Abx; BL6, n = 12 HFrD, n = 10 HFrD+Abx) (m) Liver fibrosis in HFrD-fed MUP-uPA mice was examined by Sirius Red staining (n = 7), representative images. n, Colon length in indicated mouse strains (MUP-uPA, n = 9 HFrD, n = 7 HFrD+Abx; BL6 n = 12 HFrD, n = 10 HFrD+Abx ; the HFrD bars are same as in Extended Data Fig. 1f). Scale bars, 100 μm. Unpaired two-sided Student’s t test was used in panels b, d-f, h, I, k, l and n to determine mean ± SEM, whereas panel a was analyzed by one way ANOVA and Tukey’s multiple comparison test and panel j was analyzed using two way ANOVA and Sidak’s multiple comparison test. *P < 0.05, **P < 0.01, ***P < 0.001. Benjamini-Hochberg FDR adjustment for P-values has been performed on data presented in panels e and f.
Extended Data Fig. 6 Antibiotics attenuate fructose- induced hepatosteatosis, barrier deterioration and other alterations in BL6 mice.
a–g, Male BL6 mice were fed CSD or HFrD ± Abx. a, After five months, liver tissue sections were stained with ORO (n = 6 samples from different mice per group). Representative images are shown. Scale bars, 100 μm. b, c, Inflammatory and DNL-related mRNAs in liver were determined by Q-RT- PCR (n = 9 per group). d, e, Inflammatory and lipogenic proteins in liver lysates were analyzed by IB analysis (n = 7 protein lysates per group). Representative blots are shown. f, g, Q-RT-PCR analysis of indicated TJP genes in jejunum (f) and colon (g) (n = 9 per group). Two-sided Student’s t test was used to determine mean ± SEM, *P < 0.05, **P < 0.01, ***P < 0.001. Benjamini- Hochberg FDR adjustment for P-values has been performed on data presented in panels b, c, f, g.
Extended Data Fig. 7 Fructose drink enhances hepatosteatosis, expression of liver mRNAs encoding inflammatory cytokines and DNL-related enzymes and barrier deterioration.
a-c, BL6 male mice were placed on 30% fructose in drinking water or regular water and NCD for 5 months (n = 8 per group). a, Liver sections were ORO stained and representative images are shown (n = 6 sections per group). b, mRNA amounts of liver inflammatory and DNL- related genes were PCR analyzed (n = 8 samples per group). c, Total body, liver and white adipose tissue weights were measured (n = 8). d–h, Male BL6 mice placed on 30% fructose drink or regular water and HFD for 3 months. Body, liver and WAT weight and liver/body weight ratio were determined (n = 8 per group) (d). Liver sections were analyzed by ORO staining (n = 6 sections per group). Representative images are shown (e). Liver mRNAs encoding cytokines and DNL-related proteins were measured using Q-RT-PCR (n = 7 per group) (f). Expression of TJP mRNAs in colon (n = 7 per group) (g) and FITC-dextran in serum (n = 6 per group) (h) were also measured. Scale bars, 100 μm. Two-sided Student’s t test was used to determine mean ± SEM, *P < 0.05, **P < 0.01, ***P < 0.001. Benjamini-Hochberg FDR adjustment for P-values has been performed on data presented in panels b, f, g.
Extended Data Fig. 8 Fructose-induced upregulation of YAP target genes and attenuation of TJP mRNA downregulation, barrier deterioration, hepatosteatosis and induction of DNL-related proteins in MUP-uPA/gp130Act and CCN1-treated mice.
a, b, Expression of indicated mRNAs in BL6 enteroids maintained in media containing the indicated glucose and fructose concentrations (mM) (n = 3, enteroids isolated from three separate male mice; the experiment was repeated twice with similar results). a, and in colonic mucosa of MUP-uPA mice (n=8 per group) (b) was measured by Q-RT- PCR. c, Enteroids from MUP-uPA and MUP- uPA/gp130Act mice (n = 4, enteroids isolated from four different mice per group) were incubated with 20 mM fructose. Expression of indicated mRNAs was measured by Q-RT-PCR. This experiment was repeated twice with similar results. d, e, Expression of antimicrobial (d) and TJP (e) mRNAs in colon tissues of MUP- uPA/gp130Act (MUP-uPA/Tg) mice fed CSD (n = 9 per group) or HFrD (n = 8 per group) compared to MUP-uPA mice fed HFrD for five months (n = 9 per group). Asterisks indicate significant changes between MUP-uPA and MUP-uPA/Tg mice fed HFrD. f, IB analysis of claudin-1 in colons of MUP-uPA/gp130Act and MUP-uPA mice kept on CSD or HFrD as indicated. A representative blot is shown. g, FITC-dextran serum concentrations in indicated mice (n = 6 serum samples per group). h, Serum endotoxin concentrations in indicated mice measured by ELISA (n = 8 serum samples per group). i, F1P abundance in liver and jejunum of indicated mice (arbitrary units; n = 7 samples per group). j–p, 6-week-old MUP-uPA male mice were fed HFrD for 3 months and i.p. injected 2 μg CCN1 or vehicle (PBS) every other day for the last 4 weeks of HFrD feeding. Length (j) and TJP mRNA expression (k) were determined on excised colons (n = 7 per group). Fecal albumin was measured by ELISA (n = 8 serum samples per group) (l). ORO staining (n = 6 tissue samples) and triglyceride concentrations (n = 8 samples per group) (m), inflammatory and lipogenic gene expression (n = 7 per group) (n) and FAS protein amounts (n = 7 per group), representative blots (o) were determined in liver tissue. p, Body weight was measured at the end of the CCN1 treatment course on 7 male mice per group (n = 7). q, Endotoxin levels in serum of indicated mice (n = 3 per group). Scale bars, 100 μm. In panels a, d, e, g, h and q one-way ANOVA with Tukey’s multiple comparison test was used to determine mean ± SEM. Two-sided Student’s t test was used to determine mean ± SEM in panels b, c, i-n, and p. *P < 0.05, **P < 0.01, ***P < 0.001. Benjamini-Hochberg FDR adjustment for P-values has been performed on data presented in panels c, k and n.
Extended Data Fig. 9 Low-dose LPS treatment stimulates hepatosteatosis and increases lipogenic and inflammatory gene expression and effect of fructose on microbiome diversity.
a–c, Six-week-old MUP- uPA male mice were fed CSD and received daily i.p. injections of LPS (0.25 mg/kg) or vehicle (PBS). a, ORO staining of liver sections (n = 7 per group; scale bars-100 μm), representative images are shown. b, Liver triglyceride concentrations in above mice (n = 7 per group). c, Q-RT-PCR analysis of lipogenic and inflammatory mRNAs (n =8 per group). d–f, Six-week-old MUP-uPA/Tg mice were fed HFrD and received daily i.p. injections of LPS or PBS. d, ORO staining of liver sections (n = 7 per group; scale bars-100 μm), representative images are shown. e, Liver triglyceride concentrations (n = 7 per group). f, Relative mRNA amounts of lipogenic and inflammatory genes (n = 8 per group). g–i, Relative abundance of significantly differing taxa. Significance was determined by ANCOM. Box- plots of observed operational taxonomic units (OTUs) α- diversity indices comparing stool samples of MUP-uPA (g) and MUP-uPA/Tg (i) mice treated as indicated. h–j, Principal coordinate analysis of microbiome data using unweighted UniFrac distances; difference between CSD and HFrD was significant in both MUP-uPA (h) (pseudo-F statistic = 2.88, P < 0.001, n = 10 and 24) and MUP-uPA/Tg (j) (pseudo-F statistic = 9.19, P < 0.001, n = 11 and 12) mice as determined by PERMANOVA. Two-sided Student’s t test was used to determine mean ± SEM in panels b, c, e and f. *P < 0.05, **P < 0.01, ***P < 0.001. Benjamini-Hochberg FDR adjustment for P-values has been performed on data presented in panels c and f.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2
Source data
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Unprocessed western blots
Source Data Fig. 4
Unprocessed western blots
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 5
Unprocessed western blots
Source Data Extended Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 8
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Todoric, J., Di Caro, G., Reibe, S. et al. Fructose stimulated de novo lipogenesis is promoted by inflammation. Nat Metab 2, 1034–1045 (2020). https://doi.org/10.1038/s42255-020-0261-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-0261-2
This article is cited by
-
D-mannose alleviates intervertebral disc degeneration through glutamine metabolism
Military Medical Research (2024)
-
Fructose regulates the pentose phosphate pathway and induces an inflammatory and resolution phenotype in Kupffer cells
Scientific Reports (2024)
-
Dectin-1 plays a deleterious role in high fat diet-induced NAFLD of mice through enhancing macrophage activation
Acta Pharmacologica Sinica (2023)
-
Acetyl-CoA metabolism in cancer
Nature Reviews Cancer (2023)
-
Physiological and pathological roles of lipogenesis
Nature Metabolism (2023)