Abstract
Osteoclasts are the exclusive bone-resorbing cells, playing a central role in bone metabolism, as well as the bone damage that occurs under pathological conditions1,2. In postnatal life, haematopoietic stem-cell-derived precursors give rise to osteoclasts in response to stimulation with macrophage colony-stimulating factor and receptor activator of nuclear factor-κB ligand, both of which are produced by osteoclastogenesis-supporting cells such as osteoblasts and osteocytes1,2,3. However, the precise mechanisms underlying cell fate specification during osteoclast differentiation remain unclear. Here, we report the transcriptional profiling of 7,228 murine cells undergoing in vitro osteoclastogenesis, describing the stepwise events that take place during the osteoclast fate decision process. Based on our single-cell transcriptomic dataset, we find that osteoclast precursor cells transiently express CD11c, and deletion of receptor activator of nuclear factor-κB specifically in CD11c-expressing cells inhibited osteoclast formation in vivo and in vitro. Furthermore, we identify Cbp/p300-interacting transactivator with Glu/Asp-rich carboxy-terminal domain 2 (Cited2) as the molecular switch triggering terminal differentiation of osteoclasts, and deletion of Cited2 in osteoclast precursors in vivo resulted in a failure to commit to osteoclast fate. Together, the results of this study provide a detailed molecular road map of the osteoclast differentiation process, refining and expanding our understanding of the molecular mechanisms underlying osteoclastogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding author on reasonable request. The raw sequencing data are deposited in the Gene Expression Omnibus under accession GSE147174. Other databases and resources used in this study include: RIKEN (http://www2.clst.riken.jp/arg/methods.html and http://www2.clst.riken.jp/arg/Cassette/CassetteMap_12.html) and the NCBI protein dataset (NCBInr RefSeq Release 90; https://www.ncbi.nlm.nih.gov/refseq/). Source data are provided with this paper.
References
Tsukasaki, M. & Takayanagi, H. Osteoimmunology: evolving concepts in bone-immune interactions in health and disease. Nat. Rev. Immunol. 19, 626–642 (2019).
Okamoto, K. et al. Osteoimmunology: the conceptual framework unifying the immune and skeletal systems. Physiol. Rev. 97, 1295–1349 (2017).
Jacome-Galarza, C. E. et al. Developmental origin, functional maintenance and genetic rescue of osteoclasts. Nature 568, 541–545 (2019).
Tsukasaki, M. et al. Host defense against oral microbiota by bone-damaging T cells. Nat. Commun. 9, 701 (2018).
Kong, Y. Y. et al. OPGL is a key regulator of osteoclastogenesis, lymphocyte development and lymph-node organogenesis. Nature 397, 315–323 (1999).
Danks, L. et al. RANKL expressed on synovial fibroblasts is primarily responsible for bone erosions during joint inflammation. Ann. Rheum. Dis. 75, 1187–1195 (2016).
Asano, T. et al. Soluble RANKL is physiologically dispensable but accelerates tumour metastasis to bone. Nat. Metab. 1, 868–875 (2019).
Udagawa, N. et al. Origin of osteoclasts: mature monocytes and macrophages are capable of differentiating into osteoclasts under a suitable microenvironment prepared by bone marrow-derived stromal cells. Proc. Natl Acad. Sci. USA 87, 7260–7264 (1990).
Takahashi, N. et al. Postmitotic osteoclast precursors are mononuclear cells which express macrophage-associated phenotypes. Dev. Biol. 163, 212–221 (1994).
Yasuda, H. et al. Osteoclast differentiation factor is a ligand for osteoprotegerin/osteoclastogenesis-inhibitory factor and is identical to TRANCE/RANKL. Proc. Natl Acad. Sci. USA 95, 3597–3602 (1998).
Tsukasaki, M. et al. LOX fails to substitute for RANKL in osteoclastogenesis. J. Bone Miner. Res. 32, 434–439 (2017).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Rivollier, A. et al. Immature dendritic cell transdifferentiation into osteoclasts: a novel pathway sustained by the rheumatoid arthritis microenvironment. Blood 104, 4029–4037 (2004).
Wakkach, A. et al. Bone marrow microenvironment controls the in vivo differentiation of murine dendritic cells into osteoclasts. Blood 112, 5074–5083 (2008).
Gallois, A. et al. Genome-wide expression analyses establish dendritic cells as a new osteoclast precursor able to generate bone-resorbing cells more efficiently than monocytes. J. Bone Miner. Res. 25, 661–672 (2010).
Alnaeeli, M., Penninger, J. M. & Teng, Y. T. Immune interactions with CD4+ T cells promote the development of functional osteoclasts from murine CD11c+ dendritic cells. J. Immunol. 177, 3314–3326 (2006).
Zhao, B. et al. Interferon regulatory factor-8 regulates bone metabolism by suppressing osteoclastogenesis. Nat. Med. 15, 1066–1071 (2009).
Kurotaki, D. et al. IRF8 inhibits C/EBPα activity to restrain mononuclear phagocyte progenitors from differentiating into neutrophils. Nat. Commun. 5, 4978 (2014).
Kurotaki, D. et al. Epigenetic control of early dendritic cell lineage specification by the transcription factor IRF8 in mice. Blood 133, 1803–1813 (2019).
Hanada, R. et al. Central control of fever and female body temperature by RANKL/RANK. Nature 462, 505–509 (2009).
Caton, M. L., Smith-Raska, M. R. & Reizis, B. Notch–RBP-J signaling controls the homeostasis of CD8− dendritic cells in the spleen. J. Exp. Med. 204, 1653–1664 (2007).
Shinohara, M. et al. Tyrosine kinases Btk and Tec regulate osteoclast differentiation by linking RANK and ITAM signals. Cell 132, 794–806 (2008).
Berlow, R. B., Dyson, H. J. & Wright, P. E. Hypersensitive termination of the hypoxic response by a disordered protein switch. Nature 543, 447–451 (2017).
Sakai, M. et al. CITED2 links hormonal signaling to PGC-1α acetylation in the regulation of gluconeogenesis. Nat. Med. 18, 612–617 (2012).
Sakai, M. et al. The GCN5-CITED2-PKA signalling module controls hepatic glucose metabolism through a cAMP-induced substrate switch. Nat. Commun. 7, 13147 (2016).
Kranc, K. R. et al. Cited2 is an essential regulator of adult hematopoietic stem cells. Cell Stem Cell 5, 659–665 (2009).
Du, J. et al. HIF-1α deletion partially rescues defects of hematopoietic stem cell quiescence caused by Cited2 deficiency. Blood 119, 2789–2798 (2012).
Shin, S. H. et al. Aberrant expression of CITED2 promotes prostate cancer metastasis by activating the nucleolin-AKT pathway. Nat. Commun. 9, 4113 (2018).
Bamforth, S. D. et al. Cardiac malformations, adrenal agenesis, neural crest defects and exencephaly in mice lacking Cited2, a new Tfap2 co-activator. Nat. Genet. 29, 469–474 (2001).
Barbera, J. P. et al. Folic acid prevents exencephaly in Cited2-deficient mice. Hum. Mol. Genet. 11, 283–293 (2002).
Mizoguchi, T. et al. Identification of cell cycle-arrested quiescent osteoclast precursors in vivo. J. Cell Biol. 184, 541–554 (2009).
Tanaka, S. et al. Macrophage colony-stimulating factor is indispensable for both proliferation and differentiation of osteoclast progenitors. J. Clin. Invest. 91, 257–263 (1993).
Maeda, K. et al. Wnt5a–Ror2 signaling between osteoblast-lineage cells and osteoclast precursors enhances osteoclastogenesis. Nat. Med. 18, 405–412 (2012).
Cox, T. R. et al. The hypoxic cancer secretome induces pre-metastatic bone lesions through lysyl oxidase. Nature 522, 106–110 (2015).
Tanaka, S. RANKL-independent osteoclastogenesis: a long-standing controversy. J. Bone Miner. Res. 32, 431–433 (2017).
Lam, J. et al. TNF-α induces osteoclastogenesis by direct stimulation of macrophages exposed to permissive levels of RANK ligand. J. Clin. Invest. 106, 1481–1488 (2000).
Serra, D. et al. Self-organization and symmetry breaking in intestinal organoid development. Nature 569, 66–72 (2019).
Sasai, Y. Cytosystems dynamics in self-organization of tissue architecture. Nature 493, 318–326 (2013).
Goolam, M. et al. Heterogeneity in Oct4 and Sox2 targets biases cell fate in four-cell mouse embryos. Cell 165, 61–74 (2016).
Hanna, J. et al. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462, 595–601 (2009).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Novack, D. V. et al. The IκBB function of NF-κB2 p100 controls stimulated osteoclastogenesis. J. Exp. Med. 198, 771–781 (2003).
Yagi, M. et al. DC-STAMP is essential for cell–cell fusion in osteoclasts and foreign body giant cells. J. Exp. Med. 202, 345–351 (2005).
Nishikawa, K. et al. Blimp1-mediated repression of negative regulators is required for osteoclast differentiation. Proc. Natl Acad. Sci. USA 107, 3117–3122 (2010).
Sato, K. et al. Regulation of osteoclast differentiation and function by the CaMK–CREB pathway. Nat. Med. 12, 1410–1416 (2006).
Stauffer, D., Chang, B., Huang, J., Dunn, A. & Thayer, M. p300/CREB-binding protein interacts with ATR and is required for the DNA replication checkpoint. J. Biol. Chem. 282, 9678–9687 (2007).
Kim, K. et al. MMP-9 facilitates selective proteolysis of the histone H3 tail at genes necessary for proficient osteoclastogenesis. Genes Dev. 30, 208–219 (2016).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR–Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Nitta, T. et al. The thymic cortical epithelium determines the TCR repertoire of IL-17-producing γδT cells. EMBO Rep. 16, 638–653 (2015).
Kanki, H., Suzuki, H. & Itohara, S. High-efficiency CAG-FLPe deleter mice in C57BL/6J background. Exp. Anim. 55, 137–141 (2006).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Nestorowa, S. et al. A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. Blood 128, e20–e31 (2016).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, D377–D386 (2013).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Masuda, T., Tomita, M. & Ishihama, Y. Phase transfer surfactant-aided trypsin digestion for membrane proteome analysis. J. Proteome Res. 7, 731–740 (2008).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Boersema, P. J., Raijmakers, R., Lemeer, S., Mohammed, S. & Heck, A. J. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat. Protoc. 4, 484–494 (2009).
Acknowledgements
We thank N. Mizushima, K. Kusubata, Y. Morishita, T. Tsubokawa, K. Nagumo, S. Yin, M. Tsutsumi, A. Suematsu, T. Asano and K. Kubo for thoughtful discussion and valuable technical assistance. This work was supported in part by Grants-in-Aid for Specially Promoted Research (15H05703), Scientific Research B (18H02919 and 19H03485), Challenging Research (18K19438), Young Scientists (19K18943), Research Fellowship for Young Scientists (18J00744) and an International Research Fellow (18F18095) from the Japan Society for the Promotion of Science; AMED under grant number JP20ek0410073; and AMED-CREST under grant number JP20gm1210008.
Author information
Authors and Affiliations
Contributions
M.T. designed and performed most of the experiments, interpreted the results and wrote the manuscript. N.C.-N.H. performed most of the computational analyses and contributed to data interpretation and manuscript preparation. K.O., R.M., N.K. and T.N. provided advice on project planning and contributed to data interpretation and manuscript preparation. Y. Kurikawa performed proteome analysis and contributed to data interpretation and manuscript preparation. A.T. and W.P. conducted bone histomorphometric analyses and contributed to data interpretation. T.A., H.K., T.O., T.M., M.S., M.M., Y. Kobayashi and J.M.P. provided genetically modified mice and contributed to data interpretation and manuscript preparation. H.T. directed the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The Department of Osteoimmunology is an endowment department, supported with an unrestricted grant from AYUMI Pharmaceutical, Chugai Pharmaceutical, MIKI HOUSE and Noevir.
Additional information
Peer review information Primary Handling Editors: Pooja Jha; George Caputa. Nature Metabolism thanks Luca Biasco and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The expression of osteoclastic genes and marker genes for dendritic cells during osteoclast culture.
a, The mRNA expression levels of Acp5, Ctsk, Mmp9, Dcstamp, Ocstamp, Atp6v0d2, Irf8, Bcl6 and Mafb during osteoclastogenesis analyzed by bulk-RNA seq (n=3). Data are represented as mean ± SEM. b, The expression pattern of Itgax, Cd74, H2-Aa, H2-Eb1, H2-Ab1, Cd80 and CD86 in the UMAP visualization.
Extended Data Fig. 2 Clustering and trajectory analyses by Monocle 3 and FateID.
a, Expression of marker genes for monocytic precursors (Tnfrsf11a and Csf1r), DC-like precursor (Itgax) and mature osteoclast (Acp5, Ctsk and Mmp9) analyzed by Monocle 3. b, Pseudotime analysis by Monocle3. c, d, Clustering (c) and pseudotime analysis (d) by FateID. The cluster number is the same as that in the manuscript.
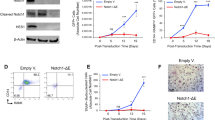
Extended Data Fig. 3 Effect of Cd11c-Cre mediated deletion of RANK on bone.
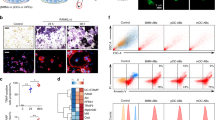
a, Bone formation parameters analyzed by bone histomorphometry using the tibiae of 11-week-old female Tnfrsf11aflox/Δ (n=4) and Tnfrsf11aflox/Δ CD11c-Cre (n=5) mice. Data are represented as mean ± SEM. P values were calculated using one-sided Student’s t-test. b, Representative data of the μCT analysis of the femur of 11-week-old male mice. Representative pictures of more than three independent experiments are shown. c, d, The μCT analysis of the femur of 4-week-old female Tnfrsf11aflox/flox (n=5) and Tnfrsf11aflox/flox CD11c-Cre (n=4) mice. Representative data (c) and quantification (d) are shown. Scale bars, 1mm. Data are represented as mean ± SEM. P values were calculated using one-sided Student’s t-test.
Extended Data Fig. 4 Marker genes for each cluster.
Heatmap of top 4 marker genes that were differentially expressed in each cluster.
Extended Data Fig. 5 Gating strategy for FACS analysis.
Representative flow cytometry profiles of gatig strategy used in Fig. 2j and Fig. 4e. Among single cells, B220−Ly6G−Vcam1− cells are subdivided into three fractions by using cell surface markers CSF1R and C5aR1.
Extended Data Fig. 6 Identification of Cited2 as a novel regulator of osteoclastogenesis.
a, The expression patterns of Cited2 during the trajectory of osteoclast differentiation. b, The expression pattern of Cited2 in the UMAP visualization. c, The expression pattern of Cited2 during osteoclastogenesis analyzed by bulk RNA-seq. Data are represented as mean ± SEM. d, The pictures of Cited2+/+ and Cited2–/ mice on embryonic day 16.5. The arrowhead indicates exencephaly. e, Osteoclast differentiation in Cited2+/+ and Cited2–/– cells. Representative data and quantification (n=6) are shown (right). Data are represented as mean ± SEM. P values were calculated using one-sided Student’s t-test. f, The expression pattern of genes enriched in Cited2+/+ (left) and Cited2−/− – cells (right) in the UMAP visualization. g, The differentiation trajectory of Cited2−/− – cells estimated by the cluster-based minimum spanning tree on a UMAP visualization. h, MA plot of significant genes that were differentially expressed in Cited2+/+ cells and Cited2−/− – cells (light blue dots). Cell cycle genes (Ccnd1, Ccna2, Nek6, E2f2, E2f1, Cdc6 and Dhfr) are denoted in red. i, Cell cycle analysis of G1, S and G2/M phases on the UMAP visualization.
Extended Data Fig. 7 Fetal liver transfer experiments using Cited2−/ – mice.
a, Schematic of the experimental setting for the fetal liver transfer into wild-type mice using Cited2+/+ or Cited2−/− − fetal liver cells. b, The μCT parameters of the femur of Cited2+/+ (n=3) and Cited2–/− – (n=6) bone marrow chimeric mice. Data are represented as mean ± SEM. P values were calculated using one-sided Student’s t-test. c, d, Bone histomorphometric analysis of the tibiae of Cited2+/+ (n=3) and Cited2−/− − (n=6) bone marrow chimeric mice. Scale bars, 200 μm. Quantification of osteoclastic bone resorption (c) and osteoblastic bone formation (d) were shown. Data are represented as mean ± SEM. P values were calculated using one-sided Student’s t-test.
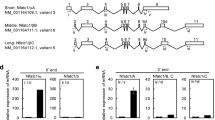
Extended Data Fig. 8 Conditional gene-targeting of Cited2.
a, Scheme of the targeting strategy. Primer 1 (P1): 5′—CACACTTTCCCGGATCCTAGG—3′, Primer 2 (P2): 5′—GAGCACGTCTCAAGCTGCAGAC—3′. b, Genotyping PCR of tail genomic DNA using the primer pairs shown in (a). Representative pictures of more than three independent experiments are shown. c, The Cited2 mRNA expression levels during in vitro osteoclastogenesis in Cited2flox/flox and Cited2flox/flox Tnfrsf11a-Cre mice (n=4). Data are represented as mean ± SEM.
Extended Data Fig. 9 The expression of negative regulators of osteoclastogenesis.
The expression pattern of Irf8, Bcl6, Mafb, Rbpj, Stat5a, Zbtb7a, Cyld, Tank, Foxo3, Eif2ak2, Commd1, Gna13, Nfkb2, and Traf3 in the UMAP visualization.
Extended Data Fig. 10 The effect of osteoclastogenic cytokines on the pre-osteoclasts (cluster 2, 3 and 4) and failed osteoclasts (cluster 6 and 8).
a, Scheme of the in vitro culture system. b, TRAP staining on the cells induced by indicated conditions. Representative pictures of triplicated experiments are shown. Scale bars; 200 μm.
Supplementary information
Supplementary Table
A list of the marker genes for each cluster
Source data
Source Data Extended Data Fig. 8
Unprocessed gel image.
Rights and permissions
About this article
Cite this article
Tsukasaki, M., Huynh, N.CN., Okamoto, K. et al. Stepwise cell fate decision pathways during osteoclastogenesis at single-cell resolution. Nat Metab 2, 1382–1390 (2020). https://doi.org/10.1038/s42255-020-00318-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00318-y
This article is cited by
-
Long noncoding RNA Malat1 protects against osteoporosis and bone metastasis
Nature Communications (2024)
-
Bone-targeting engineered small extracellular vesicles carrying anti-miR-6359-CGGGAGC prevent valproic acid-induced bone loss
Signal Transduction and Targeted Therapy (2024)
-
Profiling joint tissues at single-cell resolution: advances and insights
Nature Reviews Rheumatology (2024)
-
Osteoclast differentiation and dynamic mRNA expression during mice embryonic palatal bone development
Scientific Reports (2023)
-
Mechanical stimulation controls osteoclast function through the regulation of Ca2+-activated Cl− channel Anoctamin 1
Communications Biology (2023)