Abstract
Genome-wide association studies have identified 240 independent loci associated with type 2 diabetes (T2D) risk, but this knowledge has not advanced precision medicine. In contrast, the genetic diagnosis of monogenic forms of diabetes (including maturity-onset diabetes of the young (MODY)) are textbook cases of genomic medicine. Recent studies trying to bridge the gap between monogenic diabetes and T2D have been inconclusive. Here, we show a significant burden of pathogenic variants in genes linked with monogenic diabetes among people with common T2D, particularly in actionable MODY genes, thus implying that there should be a substantial change in care for carriers with T2D. We show that, among 74,629 individuals, this burden is probably driven by the pathogenic variants found in GCK, and to a lesser extent in HNF4A, KCNJ11, HNF1B and ABCC8. The carriers with T2D are leaner, which evidences a functional metabolic effect of these mutations. Pathogenic variants in actionable MODY genes are more frequent than was previously expected in common T2D. These results open avenues for future interventions assessing the clinical interest of these pathogenic mutations in precision medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
All relevant data have been included in the manuscript and/or in its supplementary tables and figures. Given the sensitivity and risk of re-identification, all clinical data linked with NGS data for this study is available only upon request from the corresponding authors. We used the following web links for publicly available datasets: (1) T2D Knowledge Portal. http://www.type2diabetesgenetics.org/gene/geneInfo/XXX, where XXX is the gene name; (2) Genome Aggregation Database (gnomAD). https://gnomad.broadinstitute.org/; (3) dbNSFP. https://sites.google.com/site/jpopgen/dbNSFP; (4) dbSNP. https://www.ncbi.nlm.nih.gov/snp/. Source data are provided with this paper.
Code availability
Code to perform analyses related to bioinformatics and biostatistics in this manuscript are available at https://github.com/umr1283/MODY_GENES (https://doi.org/10.5281/zenodo.4005715).
References
Mathers, C. D. & Loncar, D. Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med. 3, e442 (2006).
Abajobir, A. A. et al. Global, regional, and national under-5 mortality, adult mortality, age-specific mortality, and life expectancy, 1970–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet 390, 1084–1150 (2017).
Dieleman, J. L. et al. US spending on personal health care and public health, 1996–2013. JAMA 316, 2627–2646 (2016).
Willemsen, G. et al. The concordance and heritability of type 2 diabetes in 34,166 twin pairs from international twin registers: The discordant twin (DISCOTWIN) consortium. Twin Res. Hum. Genet. 18, 762–771 (2015).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Bonnefond, A. & Froguel, P. Rare and common genetic events in type 2 diabetes: what should biologists know? Cell Metab. 21, 357–368 (2015).
Vaxillaire, M. & Froguel, P. Monogenic diabetes: implementation of translational genomic research towards precision medicine. J. Diabetes 8, 782–795 (2016).
Babenko, A. P. et al. Activating mutations in the ABCC8 gene in neonatal diabetes mellitus. N. Engl. J. Med. 355, 456–466 (2006).
Pearson, E. R. et al. Switching from insulin to oral sulfonylureas in patients with diabetes due to Kir6.2 mutations. N. Engl. J. Med. 355, 467–477 (2006).
Shepherd, M., Shields, B., Ellard, S., Rubio-Cabezas, O. & Hattersley, A. T. A genetic diagnosis of HNF1A diabetes alters treatment and improves glycaemic control in the majority of insulin-treated patients. Diabet. Med. 26, 437–441 (2009).
Pearson, E. R. et al. Genetic cause of hyperglycaemia and response to treatment in diabetes. Lancet 362, 1275–1281 (2003).
Pearson, E. R. et al. Molecular genetics and phenotypic characteristics of MODY caused by hepatocyte nuclear factor 4α mutations in a large European collection. Diabetologia 48, 878–885 (2005).
Stride, A. et al. Cross-sectional and longitudinal studies suggest pharmacological treatment used in patients with glucokinase mutations does not alter glycaemia. Diabetologia 57, 54–56 (2014).
Garg, V. et al. GATA4 mutations cause human congenital heart defects and reveal an interaction with TBX5. Nature 424, 443–447 (2003).
Bockenhauer, D. & Jaureguiberry, G. HNF1B-associated clinical phenotypes: the kidney and beyond. Pediatr. Nephrol. 31, 707–714 (2016).
Kodo, K. et al. GATA6 mutations cause human cardiac outflow tract defects by disrupting semaphorin-plexin signaling. Proc. Natl Acad. Sci. USA 106, 13933–13938 (2009).
Naylor, R. N. et al. Cost-effectiveness of MODY genetic testing: translating genomic advances into practical health applications. Diabetes Care 37, 202–209 (2014).
GoodSmith, M. S., Skandari, M. R., Huang, E. S. & Naylor, R. N. The impact of biomarker screening and cascade genetic testing on the cost-effectiveness of MODY genetic testing. Diabetes Care 42, 2247–2255 (2019).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Flannick, J. et al. Exome sequencing of 20,791 cases of type 2 diabetes and 24,440 controls. Nature 570, 71–76 (2019).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Sun, J., Zheng, Y. & Hsu, L. A unified mixed-effects model for rare-variant association in sequencing studies. Genet. Epidemiol. 37, 334–344 (2013).
Bansal, V. et al. Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6,888 individuals. BMC Med. 15, 213 (2017).
Froguel, P. et al. Close linkage of glucokinase locus on chromosome 7p to early-onset non-insulin-dependent diabetes mellitus. Nature 356, 162–164 (1992).
Flannick, J. et al. Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes. Nat. Genet. 45, 1380–1385 (2013).
Bonnefond, A. et al. Whole-exome sequencing and high throughput genotyping identified KCNJ11 as the thirteenth MODY gene. PLoS ONE 7, e37423 (2012).
Bonnefond, A. et al. GATA6 inactivating mutations are associated with heart defects and, inconsistently, with pancreatic agenesis and diabetes. Diabetologia 55, 2845–2847 (2012).
De Franco, E. et al. GATA6 mutations cause a broad phenotypic spectrum of diabetes from pancreatic agenesis to adult-onset diabetes without exocrine insufficiency. Diabetes 62, 993–997 (2013).
Meur, G. et al. Insulin gene mutations resulting in early-onset diabetes: marked differences in clinical presentation, metabolic status, and pathogenic effect through endoplasmic reticulum retention. Diabetes 59, 653–661 (2010).
Castel, S. E. et al. Modified penetrance of coding variants by cis-regulatory variation contributes to disease risk. Nat. Genet. 50, 1327–1334 (2018).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Baier, L. J. et al. ABCC8 R1420H loss-of-function variant in a southwest American Indian community: association with increased birth weight and doubled risk of type 2 diabetes. Diabetes 64, 4322–4332 (2015).
Balkau, B. [An epidemiologic survey from a network of French Health Examination Centres, (D.E.S.I.R.): epidemiologic data on the insulin resistance syndrome]. Rev. Epidemiol. Sante Publique 44, 373–375 (1996).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Meyre, D. et al. Genome-wide association study for early-onset and morbid adult obesity identifies three new risk loci in European populations. Nat. Genet. 41, 157–159 (2009).
Romon, M. et al. Relationships between physical activity and plasma leptin levels in healthy children: the Fleurbaix–Laventie Ville Santé II Study. Int. J. Obes. Relat. Metab. Disord. 28, 1227–1232 (2004).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Van Hout, C. V. et al. Whole exome sequencing and characterization of coding variation in 49,960 individuals in the UK Biobank. Preprint at bioRxiv https://doi.org/10.1101/572347 (2019).
Cirulli, E. T. et al. Genome-wide rare variant analysis for thousands of phenotypes in over 70,000 exomes from two cohorts. Nat. Commun. 11, 542 (2020).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Liu, X., Wu, C., Li, C. & Boerwinkle, E. dbNSFP v3.0: a one-stop database of functional predictions and annotations for human nonsynonymous and splice-site SNVs. Hum. Mutat. 37, 235–241 (2016).
Regier, A. A. et al. Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects. Nat. Commun. 9, 4038 (2018).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Kendig, K. I. et al. Sentieon DNASeq variant calling workflow demonstrates strong computational performance and accuracy. Front. Genet. 10, 736 (2019).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Vaser, R., Adusumalli, S., Leng, S. N., Sikic, M. & Ng, P. C. SIFT missense predictions for genomes. Nat. Protoc. 11, 1–9 (2016).
Schwarz, J. M., Cooper, D. N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat. Methods 11, 361–362 (2014).
Ellard, S., Colclough, K., Patel, K. A. & Hattersley, A. T. Prediction algorithms: pitfalls in interpreting genetic variants of autosomal dominant monogenic diabetes. J. Clin. Invest. 130, 14–16 (2020).
Abraham, G. & Inouye, M. Fast principal component analysis of large-scale genome-wide data. PLoS ONE 9, e93766 (2014).
Siva, N. 1000 Genomes project. Nat. Biotechnol. 26, 256 (2008).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Chen, H. et al. Efficient variant set mixed model association tests for continuous and binary traits in large-scale whole-genome sequencing studies. Am. J. Hum. Genet. 104, 260–274 (2019).
Moutsianas, L. et al. The power of gene-based rare variant methods to detect disease-associated variation and test hypotheses about complex disease. PLoS Genet. 11, e1005165 (2015).
Lee, S., Abecasis, G. R., Boehnke, M. & Lin, X. Rare-variant association analysis: study designs and statistical tests. Am. J. Hum. Genet. 95, 5–23 (2014).
Balduzzi, S., Rücker, G. & Schwarzer, G. How to perform a meta-analysis with R: a practical tutorial. Evid. Based Ment. Health 22, 153–160 (2019).
Acknowledgements
We are grateful to all individuals included in the different cohort studies. We thank O. Sand and I. Rabearivelo for their contribution to the first computer analyses. This research has been conducted using the UK Biobank Application Number 40436. We thank the Genome Aggregation Database (gnomAD) and the groups that provided exome and genome variant data to this resource. A full list of contributing groups can be found at https://gnomad.broadinstitute.org/about. We would like to thank the Type 2 Diabetes Knowledge Portal and the groups that provided data to this resource. This work was supported by grants from the French National Research Agency (ANR-10-LABX-46 (European Genomics Institute for Diabetes) and ANR-10-EQPX-07-01 (LIGAN-PM)), from the European Research Council (ERC GEPIDIAB – 294785, to P.F.; ERC Reg-Seq – 715575, to A. Bonnefond), and from the National Center for Precision Diabetic Medicine – PreciDIAB, which is jointly supported by the French National Agency for Research (ANR-18-IBHU-0001), by the European Union (FEDER), by the Hauts-de-France Regional Council and by the European Metropolis of Lille (MEL). Funding was also provided to the Renown Institute for Health Innovation by Renown Health and the Renown Health Foundation.
Author information
Authors and Affiliations
Contributions
A. Bonnefond and P.F. conceived the idea of the study, supervised the analyses and performed data interpretation; E.D., B.T. and E.V. made libraries and performed NGS, with help from A.D. and V.D.; S.G., J.-M.B., J.T.L., E.T.C., G.E., R.R., B.B., M.M., S.F., G.C., M.V., N.L.W., J.J.G and P.F. managed the collection of samples with clinical data; F.A. prepared samples; F.D.G. and D.L.G. performed computer analyses; A. Bonnefond, M.B. and A. Bolze performed data curation and statistical analysis, with help from L.Y. and M.C.; A. Bonnefond wrote the first draft of the paper, with help from P.F.; all authors critically reviewed the paper and approved the report for submission.
Corresponding authors
Ethics declarations
Competing interests
R.R. is an advisory panel member for AstraZeneca, AbbVie, Sanofi, Merck Sharp & Dohme, Eli Lilly, Janssen, Novo Nordisk, Diabnext, Vaiomer and Physiogenex; is a speaker for Bayer and Servier; and has received research funding and provided research support to Danone Research, Amgen, Sanofi and Novo Nordisk. A. Bolze, E.T.C., J.L. and N.L.W. are employees and shareholders of Helix. No other conflicts were reported.
Additional information
Peer review information Primary Handling Editor: Pooja Jha.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mean rate of rare coding variants of potential interest and samples which were filtered out by each QC across the 33 genes linked with monogenic diabetes, in the RaDiO study.
The figure shows the rate of excluded samples (on the left) or variants (on the right) per QC.
Extended Data Fig. 2 Association between P/LP variants in actionable MODY genes and age of T2D diagnosis, in the 2,178 participants with T2D from the RaDiO study.
Association analyses were performed using a linear regression adjusted for age, sex, BMI and genetic ancestry. β, mean effect; SD, standard deviation; SE, standard error.
Extended Data Fig. 3 Association between P/LP variants in actionable MODY genes and insulin intake, in the 2,178 participants with T2D from the RaDiO study.
Association analyses were performed using a logistic regression adjusted for age, sex, BMI and genetic ancestry. CI, confidence interval; NA, not available; OR, odds ratio.
Extended Data Fig. 4 Association between P/LP variants in actionable MODY genes and metformin intake, in the 2,178 participants with T2D from the RaDiO study.
Association analyses were performed using a logistic regression adjusted for age, sex, BMI and genetic ancestry. CI, confidence interval; NA, not available; OR, odds ratio.
Extended Data Fig. 5 Association between P/LP variants in actionable MODY genes and family history of T2D, in the 2,178 participants with T2D from the RaDiO study.
Association analyses were performed using a logistic regression adjusted for age, sex, BMI and genetic ancestry. CI, confidence interval; OR, odds ratio.
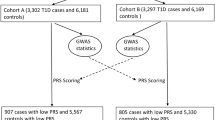
Extended Data Fig. 6 Study design reporting entry criteria for cases and controls in the RaDiO study, the UK Biobank, the HNP study and the AMP T2D knowledge portal.
The figure shows the study design for each population study. Cases are highlighted in orange, controls are highlighted in green and exclusions are highlighted in yellow.
Extended Data Fig. 7
Bootstrapping with varying sample sizes and (a.) controls with fasting glucose < 6.1 mmol/l or (b.) controls with fasting glucose < 7.0 mmol/l, in the RaDiO study. Average: 90 sets of bootstrap.
Supplementary information
Supplementary Information
Supplementary Figures 1–3 and References for Supplementary Tables
Source data
Source Data Fig. 1
Statistical Source Data
Rights and permissions
About this article
Cite this article
Bonnefond, A., Boissel, M., Bolze, A. et al. Pathogenic variants in actionable MODY genes are associated with type 2 diabetes. Nat Metab 2, 1126–1134 (2020). https://doi.org/10.1038/s42255-020-00294-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00294-3
This article is cited by
-
Pathogenic monoallelic variants in GLIS3 increase type 2 diabetes risk and identify a subgroup of patients sensitive to sulfonylureas
Diabetologia (2024)
-
Monogenic diabetes
Diabetology International (2024)
-
Whole-genome sequencing of multiple related individuals with type 2 diabetes reveals an atypical likely pathogenic mutation in the PAX6 gene
European Journal of Human Genetics (2023)
-
Monogenic diabetes
Nature Reviews Disease Primers (2023)
-
Human GLP1R variants affecting GLP1R cell surface expression are associated with impaired glucose control and increased adiposity
Nature Metabolism (2023)