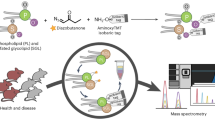
Abstract
Despite the crucial roles of lipids in metabolism, we are still at the early stages of comprehensively annotating lipid species and their genetic basis. Mass spectrometry–based discovery lipidomics offers the potential to globally survey lipids and their relative abundances in various biological samples. To discover the genetics of lipid features obtained through high-resolution liquid chromatography–tandem mass spectrometry, we analysed liver and plasma from 384 diversity outbred mice, and quantified 3,283 molecular features. These features were mapped to 5,622 lipid quantitative trait loci and compiled into a public web resource termed LipidGenie. The data are cross-referenced to the human genome and offer a bridge between genetic associations in humans and mice. Harnessing this resource, we used genome–lipid association data as an additional aid to identify a number of lipids, for example gangliosides through their association with B4galnt1, and found evidence for a group of sex-specific phosphatidylcholines through their shared locus. Finally, LipidGenie’s ability to query either mass or gene-centric terms suggests acyl-chain-specific functions for proteins of the ABHD family.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Genotypes and additional phenotype data associated with the DO mouse population have been deposited with Dryad (https://doi.org/10.5061/dryad.pj105; data files: Attie Islet eQTL data) (see Keller et al., ref. 32, for details). In addition, the data reported here are available for download and interactive web-based analysis at https://churchilllab.jax.org/qtlviewer/attie/islets. Genotyping used the Mouse Universal Genotyping Array (GigaMUGA; 143,259 markers).
MS data have been deposited in Chorus (http://chorusproject.org/) under ID 1610 (direct links to cell experiments https://chorusproject.org/anonymous/download/experiment/4984245205453479277, DO liver https://chorusproject.org/anonymous/download/experiment/a639bcc5602c441c9a1df94f4340d626, DO plasma https://chorusproject.org/anonymous/download/experiment/f8b273d222364f2a9d92cfdd0eb601b6, FS liver https://chorusproject.org/anonymous/download/experiment/c930cd419eb34dfebda7f53508c6969e, and FS plasma https://chorusproject.org/anonymous/download/experiment/9d4d025df0114687924d4075f3c927ca). Human Mouse homologues were obtained from the MGI homology database (available here: http://www.informatics.jax.org/downloads/reports/HOM_MouseHumanSequence.rpt). SNP associations were performed accessing variants from the database cc_variants.sqlite (available here: https://ndownloader.figshare.com/files/18533342) and genes from mouse_genes_mgi.sqlite (available here: https://ndownloader.figshare.com/files/17609252). Figures 1–5 and Extended Data Figs. 1–6 have associated raw data. Source data are provided with this paper.
Code availability
The data preparation and QTL mapping analysis are reproducibly documented in UNIX shell and R scripts posted on github (https://github.com/dmgatti/AttieMetabolomics). Code for data analysis and plotting is available at https://github.com/vanilink/DOLipids/ with input from Supplementary Tables 8 and 9. The genome–lipid associations are also accessible through an interactive web-based analysis tool that will allow users to replicate the analyses reported here (http://lipidgenie.com/). The source code for this resource can be found at https://github.com/coongroup/LipidGenie.
References
Han, X. Lipidomics for studying metabolism. Nat. Rev. Endocrinol. 12, 668–679 (2016).
Yang, L. et al. Recent advances in lipidomics for disease research. J. Sep. Sci. 39, 38–50 (2016).
Kind, T. et al. LipidBlast in silico tandem mass spectrometry database for lipid identification. Nat. Methods 10, 755–758 (2013).
Gross, R. W. & Han, X. Lipidomics at the interface of structure and function in systems biology. Chem. Biol. 18, 284–291 (2011).
Cajka, T. & Fiehn, O. Comprehensive analysis of lipids in biological systems by liquid chromatography-mass spectrometry. Trends Anal. Chem. 61, 192–206 (2014).
Tabassum, R. et al. Genetic architecture of human plasma lipidome and its link to cardiovascular disease. Nat. Commun. 10, 4329 (2019).
Kiyonami, R., Peake, D. A., Liu, X. & Huang, Y. Large-scale lipid profiling of a human serum lipidome using a high-resolution, accurate-mass LC/MS/MS approach Application Note No. 647 (Thermo Fisher Scientific, 2016).
Slatter, D. A. et al. Mapping the human platelet lipidome reveals cytosolic phospholipase A2 as a regulator of mitochondrial bioenergetics during activation. Cell Metab. 23, 930–944 (2016).
Contrepois, K. et al. Cross-platform comparison of untargeted and targeted lipidomics approaches on aging mouse plasma. Sci. Rep. 8, 17747 (2018).
Blaženović, I. et al. Increasing compound identification rates in untargeted lipidomics research with liquid chromatography drift time-ion mobility mass spectrometry. Anal. Chem. 90, 10758–10764 (2018).
Mahieu, N. G. & Patti, G. J. Systems-Level annotation of a metabolomics data set reduces 25 000 features to fewer than 1000 unique metabolites. Anal. Chem. 89, 10397–10406 (2017).
Blaženović, I., Kind, T., Ji, J. & Fiehn, O. Software tools and approaches for compound identification of LC–MS/MS data in metabolomics. Metabolites https://doi.org/10.3390/metabo8020031 (2018).
Gross, R. W. The evolution of lipidomics through space and time. Biochim. Biophys. Acta–Mol. Cell Biol. Lipids 1862, 731–739 (2017).
Koelmel, J. P. et al. LipidMatch: an automated workflow for rule-based lipid identification using untargeted high-resolution tandem mass spectrometry data. BMC Bioinf. 18, 331 (2017).
Hartler, J. et al. Deciphering lipid structures based on platform-independent decision rules. Nat. Methods 14, 1171–1174 (2017).
Hutchins, P. D., Russell, J. D. & Coon, J. J. LipiDex: an integrated software package for high-confidence lipid identification. Cell Syst. 6, 621–625.e5 (2018).
Hutchins, P. D., Russell, J. D. & Coon, J. J. Mapping lipid fragmentation for tailored mass spectral libraries. J. Am. Soc. Mass. Spectrom. 30, 659–668 (2019).
Kostyukevich, Y. et al. Hydrogen/deuterium exchange aiding compound identification for LC–MS and MALDI imaging lipidomics. Anal. Chem. 91, 13465–13474 (2019).
Stefely, J. A. et al. Mitochondrial protein functions elucidated by multi-omic mass spectrometry profiling. Nat. Biotechnol. 34, 1191–1197 (2016).
Dumas, M.-E. et al. Topological analysis of metabolic networks integrating co-segregating transcriptomes and metabolomes in type 2 diabetic rat congenic series. Genome Med. 8, 101 (2016).
Cazier, J.-B. et al. Untargeted metabolome quantitative trait locus mapping associates variation in urine glycerate to mutant glycerate kinase. J. Proteome Res. 11, 631–642 (2012).
Krumsiek, J. et al. Mining the unknown: a systems approach to metabolite identification combining genetic and metabolic information. PLoS Genet. 8, e1003005 (2012).
Shin, S.-Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 46, 543–550 (2014).
Rueedi, R. et al. Metabomatching: using genetic association to identify metabolites in proton NMR spectroscopy. PLoS Comput. Biol. 13, e1005839 (2017).
Raffler, J. et al. Identification and MS-assisted interpretation of genetically influenced NMR signals in human plasma. Genome Med. 5, 13 (2013).
Gatti, D. M. et al. Quantitative trait locus mapping methods for diversity outbred mice. G3 4, 1623–1633 (2014).
Broman, K. W. et al. R/qtl2: Software for mapping quantitative trait loci with high-dimensional data and multiparent populations. Genetics 211, 495–502 (2019).
Svenson, K. L. et al. High-resolution genetic mapping using the mouse diversity outbred population. Genetics 190, 437–447 (2012).
Chesler, E. J. et al. Diversity outbred mice at 21: maintaining allelic variation in the face of selection. G3 6, 3893–3902 (2016).
Mayer, R. et al. Common themes and cell type specific variations of higher order chromatin arrangements in the mouse. BMC Cell Biol. 6, 44 (2005).
Aylor, D. L. et al. Genetic analysis of complex traits in the emerging collaborative cross. Genome Res. 21, 1213–1222 (2011).
Keller, M. P. et al. Genetic drivers of pancreatic islet function. Genetics 209, 335–356 (2018).
Keller, M. P. et al. Gene loci associated with insulin secretion in islets from nondiabetic mice. J. Clin. Invest. 129, 4419–4432 (2019).
Chick, J. M. et al. Defining the consequences of genetic variation on a proteome-wide scale. Nature 534, 500–505 (2016).
Kemis, J. H. et al. Genetic determinants of gut microbiota composition and bile acid profiles in mice. PLoS Genet. https://doi.org/10.1371/journal.pgen.1008073 (2019).
Gallego, S. F., Højlund, K. & Ejsing, C. S. Easy, fast, and reproducible quantification of cholesterol and other lipids in human plasma by combined high resolution MSX and FTMS analysis. J. Am. Soc. Mass. Spectrom. 29, 34–41 (2018).
Ogiso, H., Suzuki, T. & Taguchi, R. Development of a reverse-phase liquid chromatography electrospray ionization mass spectrometry method for lipidomics, improving detection of phosphatidic acid and phosphatidylserine. Anal. Biochem. 375, 124–131 (2008).
Fahy, E. et al. Update of the LIPID MAPS comprehensive classification system for lipids. J. Lipid Res. 50, S9–S14 (2009).
Liebisch, G. et al. Shorthand notation for lipid structures derived from mass spectrometry. J. Lipid Res. 54, 1523–1530 (2013).
Su, Z. et al. Genetic basis of HDL variation in 129/SvImJ and C57BL/6J mice: importance of testing candidate genes in targeted mutant mice. J. Lipid Res. 50, 116–125 (2009).
Kettunen, J. et al. Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nat. Commun. 7, 11122 (2016).
Zhang, W. et al. Genome-wide association mapping of quantitative traits in outbred mice. G3 2, 167–174 (2012).
Pamir, N. et al. Genetic control of the mouse HDL proteome defines HDL traits, function, and heterogeneity. J. Lipid Res. 60, 594–608 (2019).
Wang, X., Korstanje, R., Higgins, D. & Paigen, B. Haplotype analysis in multiple crosses to identify a QTL gene. Genome Res. 14, 1767–1772 (2004).
Blanco-Vaca, F., Escolà-Gil, J. C., Martín-Campos, J. M. & Julve, J. Role of apoA-II in lipid metabolism and atherosclerosis: advances in the study of an enigmatic protein. J. Lipid Res. 42, 1727–1739 (2001).
Kontush, A., Lhomme, M. & Chapman, M. J. Unraveling the complexities of the HDL lipidome. J. Lipid Res. 54, 2950–2963 (2013).
Murphy, R. C., Leiker, T. J. & Barkley, R. M. Glycerolipid and cholesterol ester analyses in biological samples by mass spectrometry. Biochim. Biophys. Acta 1811, 776 (2011).
Lerno, L. A. Jr, German, J. B. & Lebrilla, C. B. Method for the identification of lipid classes based on referenced Kendrick mass analysis. Anal. Chem. 82, 4236–4245 (2010).
Eilbeck, K. et al. The sequence ontology: a tool for the unification of genome annotations. Genome Biol. 6, R44 (2005).
Nagata, Y. et al. Expression cloning of beta 1,4 N-acetylgalactosaminyltransferase cDNAs that determine the expression of GM2 and GD2 gangliosides. J. Biol. Chem. 267, 12082–12089 (1992).
Dotta, F. et al. Pancreatic islet ganglioside expression in nonobese diabetic mice: comparison with C57BL/10 mice and changes after autoimmune beta-cell destruction. Endocrinology 130, 37–42 (1992).
Li, Z. et al. Impact of sphingomyelin synthase 1 deficiency on sphingolipid metabolism and atherosclerosis in mice. Arterioscler. Thromb. Vasc. Biol. 32, 1577–1584 (2012).
Bergfeld, A. K. et al. N-glycolyl groups of nonhuman chondroitin sulfates survive in ancient fossils. Proc. Natl Acad. Sci. USA 114, E8155–E8164 (2017).
Strømme, P. et al. X-linked Angelman-like syndrome caused by Slc9a6 knockout in mice exhibits evidence of endosomal–lysosomal dysfunction. Brain 134, 3369–3383 (2011).
Spessott, W., Uliana, A. & Maccioni, H. J. F. Defective GM3 synthesis in Cog2 null mutant CHO cells associates to mislocalization of lactosylceramide sialyltransferase in the golgi complex. Neurochem. Res. 35, 2161–2167 (2010).
Ledeen, R. W. & Wu, G. The multi-tasked life of GM1 ganglioside, a true factotum of nature. Trends Biochem. Sci. 40, 407–418 (2015).
Yore, M. M. et al. Discovery of a class of endogenous mammalian lipids with anti-diabetic and anti-inflammatory effects. Cell 159, 318–332 (2014).
McLean, S., Davies, N. W., Nichols, D. S. & Mcleod, B. J. Triacylglycerol estolides, a new class of mammalian lipids, in the paracloacal gland of the brushtail possum (Trichosurus vulpecula). Lipids 50, 591–604 (2015).
Parker, B. L. et al. An integrative systems genetic analysis of mammalian lipid metabolism. Nature 567, 187–193 (2019).
Lord, C. C., Thomas, G. & Brown, J. M. Mammalian alpha beta hydrolase domain (ABHD) proteins: lipid metabolizing enzymes at the interface of cell signaling and energy metabolism. Biochim. Biophys. Acta 1831, 792–802 (2013).
Long, J. Z. et al. Metabolomics annotates ABHD3 as a physiologic regulator of medium-chain phospholipids. Nat. Chem. Biol. 7, 763–765 (2011).
Draisma, H. H. M. et al. Genome-wide association study identifies novel genetic variants contributing to variation in blood metabolite levels. Nat. Commun. 6, 7208 (2015).
Ha, C. Y. et al. The association of specific metabolites of lipid metabolism with markers of oxidative stress, inflammation and arterial stiffness in men with newly diagnosed type 2 diabetes. Clin. Endocrinol. 76, 674–682 (2012).
Demirkan, A. et al. Genome-wide association study identifies novel loci associated with circulating phospho- and sphingolipid concentrations. PLoS Genet. 8, e1002490 (2012).
Miller, M. R. et al. Unconventional endocannabinoid signaling governs sperm activation via the sex hormone progesterone. Science 352, 555–559 (2016).
Baggelaar, M. P., Maccarrone, M. & van der Stelt, M. 2-Arachidonoylglycerol: a signaling lipid with manifold actions in the brain. Prog. Lipid Res. 71, 1–17 (2018).
Jha, P. et al. Systems analyses reveal physiological roles and genetic regulators of liver lipid species. Cell Syst. 6, 722–733.e6 (2018).
Jha, P. et al. Genetic regulation of plasma lipid species and their association with metabolic phenotypes. Cell Syst. 6, 709–721.e6 (2018).
Stacey, D. et al. ProGeM: a framework for the prioritization of candidate causal genes at molecular quantitative trait loci. Nucleic Acids Res. 47, e3 (2019).
Kastenmüller, G., Raffler, J., Gieger, C. & Suhre, K. Genetics of human metabolism: an update. Hum. Mol. Genet. 24, R93–R101 (2015).
Mitok, K. A. et al. Islet proteomics reveals genetic variation in dopamine production resulting in altered insulin secretion. J. Biol. Chem. 293, 5860–5877 (2018).
Broman, K. W., Gatti, D. M., Svenson, K. L., Sen, Ś. & Churchill, G. A. Cleaning genotype data from diversity outbred mice. G3 9, 1571–1579 (2019).
Choi, K. B. churchill-lab/gbrs v.0.1.5 (Zenodo, 2017); https://doi.org/10.5281/zenodo.291787
Adusumilli, R. & Mallick, P. Data conversion with ProteoWizard msConvert. Methods Mol. Biol. 1550, 339–368 (2017).
Johnson, W. E., Evan Johnson, W., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Churchill, G. A. & Doerge, R. W. Empirical threshold values for quantitative trait mapping. Genetics 138, 963–971 (1994).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019).
RStudio Team. RStudio: Integrated Development Environment for R (RStudio, 2016).
Wickham, H., François, R., Henry, L. & Müller, K. dplyr: A grammar of data manipulation (2019).
Wickham, H. & Henry, L. tidyr: Tidy messy data (2019).
Wickham, H. et al. Reshaping data with the reshape package. J. Stat. Softw. 21, 1–20 (2007).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Neuwirth, E. RColorBrewer: ColorBrewer palettes (2014).
Sievert, C. plotly for R (2018).
Kolde, R. pheatmap: Pretty heatmaps (2019).
Holtz, Y. Manhattan plot in R: a review. R graph gallery https://www.r-graph-gallery.com/101_Manhattan_plot.html (2019).
Chong, J. et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 46, W486–W494 (2018).
Müller, K., Wickham, H., James, D. A. & Falcon, S. RSQLite: ‘SQLite’ interface for R (2019).
Kavaler, S. et al. Pancreatic beta-cell failure in obese mice with human-like CMP-Neu5Ac hydroxylase deficiency. FASEB J. 25, 1887–1893 (2011).
Salama, A. et al. Neu5Gc and α1-3 GAL xenoantigen knockout does not affect glycemia homeostasis and insulin secretion in pigs. Diabetes 66, 987–993 (2017).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008).
Acknowledgements
This work was supported by National Institutes of Health grant nos. P41 GM108538 and 2R01DK101573. This research was performed using the compute resources and assistance of the UW–Madison Center For High Throughput Computing (CHTC) in the Department of Computer Sciences. The CHTC is supported by UW–Madison, the Advanced Computing Initiative, the Wisconsin Alumni Research Foundation, the Wisconsin Institutes for Discovery, and the National Science Foundation, and is an active member of the Open Science Grid, which is supported by the National Science Foundation and the US Department of Energy’s Office of Science. We thank B. Paulson for help with sample preparation and D. Hwang for help with figures. We thank J. Simcox (Department of Biochemistry, University of Wisconsin–Madison, Madison) for providing the His-tagged CMV6-GFP plasmid and Hepa1-6 cells.
Author information
Authors and Affiliations
Contributions
J.D.R., M.P.K., G.A.C., A.D.A. and J.J.C. designed the experiment. K.L.S., D.S.S., M.E.R. and M.P.K. assisted with sample collection. V.L., P.D.H., E.A.T. and T.R.R. performed the MS analysis. E.M.C. performed cell experiments. V.L., I.J.M., D.R.B., P.D.H., M.P.K., D.M.G., G.R.K., D.P. and G.A.C. analysed data. V.L., K.A.O., I.J.M., M.P.K., K.W.B., G.A.C., A.D.A. and J.J.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: Pooja Jha.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identified lipids and unidentified features occupy characteristic regions in the m/z vs. RT space.
a, In plasma, we quantified 1,721 lipidomic features, 621 of which were identified, and b, In liver, we quantified 1,562 lipidomic features, 615 of which were identified. Abbreviations: m/z (mass-to-charge), RT (retention time).
Extended Data Fig. 2 Lipid profiling and subsequent QTL mapping reveals clusters of associated lipids.
a, Lipid class distribution of all 1,721 plasma and b, 1,562 liver lipidomic features. c, 1,405 plasma and d, 1,190 lipid features showed at least one QTL with an LOD > 6 as displayed in a Manhattan plot (n = 3,353 and 2,269 total QTL, respectively). Hierarchical clustering of these features against the 69,005 markers on the mouse genome, resulted in clustering of lipid class based on hotspots at the genetic level. Abbreviations: Chr (chromosome), DO (diversity outbred), QTL (quantitative trait loci), LOD (logarithm of odds).
Extended Data Fig. 3 Apoa2 as the candidate gene at the largest lipid hotspot.
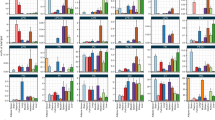
a, 255 plasma (black) features mapping to the apoa2 locus on chromosome 1 share an allele effect pattern with upregulation in the 129 allele, while 2 mapping liver features (white) do not share the pattern (based on hierarchical clustering on allele effects, with a Euclidean distance cutoff of h = 1.5). b, The allele effect is exemplary replicated in an independent experiment of founder strain plasma CE(18:2) levels (n = 4 for each sex and strain, boxplots are defined with the first and third quartiles (25th and 75th percentile) for lower and upper hinges, 1.5x interquartile range for the length of the whiskers, centre line at median (50% quantile)). c, The same pattern was not visible in previously reported34 Apoa2 liver protein and RNA allele effects. Abbreviations: CE (cholesteryl ester), FS (founder strain).
Extended Data Fig. 4 B4galnt1 as the candidate gene at the hotspot with the largest LOD.
a, The selection of B4galnt1 as the candidate gene for the chromosome 10:127 Mbp locus was corroborated by NOD-specific allele effects in previously reported liver eQTL and b, pQTL34. c, The allele effect patterns of the later as gangliosides identified features mapping to the B4galnt1 locus could further be validated in an independent experiment of founder strain mice (exemplar GM3 pattern, n = 4 for each sex and strain, boxplots are defined with the first and third quartiles (25th and 75th percentile) for lower and upper hinges, 1.5x interquartile range for the length of the whiskers, center line at median (50% quantile)). Abbreviations: FS (founder strain), Mbp (megabase pair).
Extended Data Fig. 5 Allele effects characterize genome-lipid hotspots.
a, Hierarchical clustering of allele effects at Chr 6:91 Mbp resulted in 21 features with matching A/J down effect (main cluster featuring the six B6 male specific features (red) after row-scaling and Ward clustering, cutoff at h=5). b, Consistently, the pattern of male >> female was observed for each of the FS except for A/J as visible in the example for m/z 1130 (n = 4 for each sex and strain, boxplots are defined with the first and third quartiles (25th and 75th percentile) for lower and upper hinges, 1.5x interquartile range for the length of the whiskers, center line at median (50% quantile).) c, Hierarchical clustering of allele effects at Chr 5:31 Mbp locus resulted in 10 features with matching B6 and NZO up effect (main cluster featuring LysoPC 14:0 (turquoise) after row-scaling and Ward clustering, cutoff at h=8). d, This pattern could be replicated in the FS (n = 4 for each sex and strain, boxplots are defined with the first and third quartiles (25th and 75th percentile) for lower and upper hinges, 1.5x interquartile range for the length of the whiskers, center line at median (50% quantile)), as shown for LysoPC 14:0, as well as e, in opposite directionality in a liver eQTL34. f, Hierarchical clustering of allele effects at Chr 7:79 Mbp locus resulted in 8 features with matching WSB down effect (main cluster featuring PUFA-containing phospholipids (turquoise) after row-scaling and Ward clustering, cutoff at h=2.5). g, The mapping phospholipids contained polyunsaturated fatty acids such as 20:4 and 22:6. h-i, Abhd2 liver RNA and protein allele effects34 matched with an opposite WSB high effect. Abbreviations: DO (diversity outbred), FS (founder strain), Chr (chromosome), Mbp (megabase pair), PC (phosphatidylcholine), PUFA (polyunsaturated fatty acid).
Extended Data Fig. 6 Overexpressing ABHD1 and ABHD3 results in distinct phospholipid signature.
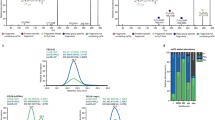
a, Experimental design of the validation experiment featuring three technical and four biological replicates of Hepa1-6 cells either untransfected (CTL), transfected with a His-tag GFP control (GFP), or transfected with MYC-tagged ABHD1 or ABHD3. b, Western blot of Hepa1-6 overexpression of ABHD1 and ABHD3. Shown is an overlay of membrane and ECL blot for MYC-tag. c, Heatmap of top 49 features from discovery lipidomics experiment with p < 0.05 (ANOVA, Fisher’s LSD post-hoc). Features were sum-normalized and log2-transformed. Hierarchical clustering (Ward clustering, Euclidean distance) shows two clusters with opposite fold changes distinguishing between ABHD1 and ABHD3 and the GFP control.
Extended Data Fig. 7 Lipid class abbreviations and identifications with respective adduct types.
As searched for in LipiDex databases (see Methods).
Supplementary information
Supplementary Tables
Supplementary Tables 1–12
Source data
Source Data Fig. 5
Statistical source data for Fig. 5k–m.
Source Data Extended Data Fig. 6
Unprocessed western blot from Extended Data Fig. 6b.
Rights and permissions
About this article
Cite this article
Linke, V., Overmyer, K.A., Miller, I.J. et al. A large-scale genome–lipid association map guides lipid identification. Nat Metab 2, 1149–1162 (2020). https://doi.org/10.1038/s42255-020-00278-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00278-3
This article is cited by
-
Plasma metabolomics supports non-fasted sampling for metabolic profiling across a spectrum of glucose tolerance in the Nile rat model for type 2 diabetes
Lab Animal (2023)
-
Genetic mapping of microbial and host traits reveals production of immunomodulatory lipids by Akkermansia muciniphila in the murine gut
Nature Microbiology (2023)
-
Sample multiplexing-based targeted pathway proteomics with real-time analytics reveals the impact of genetic variation on protein expression
Nature Communications (2023)
-
Spatial snapshots of amyloid precursor protein intramembrane processing via early endosome proteomics
Nature Communications (2022)
-
Measurement of lipid flux to advance translational research: evolution of classic methods to the future of precision health
Experimental & Molecular Medicine (2022)