Abstract
Following activation, conventional T (Tconv) cells undergo an mTOR-driven glycolytic switch. Regulatory T (Treg) cells reportedly repress the mTOR pathway and avoid glycolysis. However, here we demonstrate that human thymus-derived Treg (tTreg) cells can become glycolytic in response to tumour necrosis factor receptor 2 (TNFR2) costimulation. This costimulus increases proliferation and induces a glycolytic switch in CD3-activated tTreg cells, but not in Tconv cells. Glycolysis in CD3–TNFR2-activated tTreg cells is driven by PI3-kinase–mTOR signalling and supports tTreg cell identity and suppressive function. In contrast to glycolytic Tconv cells, glycolytic tTreg cells do not show net lactate secretion and shuttle glucose-derived carbon into the tricarboxylic acid cycle. Ex vivo characterization of blood-derived TNFR2hiCD4+CD25hiCD127lo effector T cells, which were FOXP3+IKZF2+, revealed an increase in glucose consumption and intracellular lactate levels, thus identifying them as glycolytic tTreg cells. Our study links TNFR2 costimulation in human tTreg cells to metabolic remodelling, providing an additional avenue for drug targeting.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All RNA-seq data are available in the GEO database under accession code GSE138603 and GSE138604. GSEAs were performed with GSEA software using the hallmark gene sets listed in the Molecular Signatures Database (MSigDB) available through https://www.gsea-msigdb.org/gsea/index.jsp. The proteomics data set used in this study is published by Cuadrado et al.14. All other data that support the findings of this study are available from the corresponding author on reasonable request.
Code availability
For RNA-sequencing, the script Itreecount was used to count the number of reads per gene. Itreecount is publicly available through https://github.com/NKI-GCF/itreecount.
References
Wing, J. B., Tanaka, A. & Sakaguchi, S. Human FOXP3+ regulatory T cell heterogeneity and function in autoimmunity and cancer. Immunity 50, 302–316 (2019).
Shevach, E. M. & Thornton, A. M. tTregs, pTregs, and iTregs: similarities and differences. Immunol. Rev. 259, 88–102 (2014).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Floess, S. et al. Epigenetic control of the foxp3 locus in regulatory T cells. PLoS Biol. 5, e38 (2007).
Blazar, B. R., MacDonald, K. P. A. & Hill, G. R. Immune regulatory cell infusion for graft-versus-host disease prevention and therapy. Blood 131, 2651–2660 (2018).
Riley, J. L., June, C. H. & Blazar, B. R. Human T regulatory cell therapy: take a billion or so and call me in the morning. Immunity 30, 656–665 (2009).
Koenen, H. J. et al. Human CD25highFoxp3pos regulatory T cells differentiate into IL-17-producing cells. Blood 112, 2340–2352 (2008).
Medler, J. & Wajant, H. Tumor necrosis factor receptor-2 (TNFR2): an overview of an emerging drug target. Expert Opin. Ther. Targets 23, 295–307 (2019).
Salomon, B. L. et al. Tumor necrosis factor alpha and regulatory T cells in oncoimmunology. Front. Immunol. 9, 444 (2018).
Huynh, A. et al. Control of PI(3) kinase in Treg cells maintains homeostasis and lineage stability. Nat. Immunol. 16, 188–196 (2015).
Zeiser, R. et al. Differential impact of mammalian target of rapamycin inhibition on CD4+CD25+Foxp3+ regulatory T cells compared with conventional CD4+ T cells. Blood 111, 453–462 (2008).
Sauer, S. et al. T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc. Natl Acad. Sci. USA 105, 7797–7802 (2008).
Cuadrado, E. et al. Proteomic analyses of human regulatory T cells reveal adaptations in signaling pathways that protect cellular identity. Immunity 48, 1046–1059 (2018).
Menk, A. V. et al. Early TCR signaling induces rapid aerobic glycolysis enabling distinct acute T cell effector functions. Cell Rep. 22, 1509–1521 (2018).
Frauwirth, K. A. et al. The CD28 signaling pathway regulates glucose metabolism. Immunity 16, 769–777 (2002).
Basu, S., Hubbard, B. & Shevach, E. M. Foxp3-mediated inhibition of Akt inhibits Glut1 (glucose transporter 1) expression in human T regulatory cells. J. Leukoc. Biol. 97, 279–283 (2015).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Shi, L. Z. et al. HIF1alpha-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
He, N. et al. Metabolic control of regulatory T cell (Treg) survival and function by Lkb1. Proc. Natl Acad. Sci. USA 114, 12542–12547 (2017).
Beier, U. H. et al. Essential role of mitochondrial energy metabolism in Foxp3+ T-regulatory cell function and allograft survival. FASEB J. 29, 2315–2326 (2015).
Angelin, A. et al. Foxp3 reprograms T cell metabolism to function in low-glucose, high-lactate environments. Cell Metab. 25, 1282–1293 e1287 (2017).
Gerriets, V. A. et al. Foxp3 and Toll-like receptor signaling balance Treg cell anabolic metabolism for suppression. Nat. Immunol. 17, 1459–1466 (2016).
Howie, D. et al. Foxp3 drives oxidative phosphorylation and protection from lipotoxicity. JCI Insight 2, e89160 (2017).
Delgoffe, G. M. et al. The kinase mTOR regulates the differentiation of helper T cells through the selective activation of signaling by mTORC1 and mTORC2. Nat. Immunol. 12, 295–303 (2011).
Wei, J. et al. Autophagy enforces functional integrity of regulatory T cells by coupling environmental cues and metabolic homeostasis. Nat. Immunol. 17, 277–285 (2016).
Chapman, N. M. et al. mTOR coordinates transcriptional programs and mitochondrial metabolism of activated Treg subsets to protect tissue homeostasis. Nat. Commun. 9, 2095 (2018).
Procaccini, C. et al. The proteomic landscape of human ex vivo regulatory and conventional T cells reveals specific metabolic requirements. Immunity 44, 712 (2016).
De Rosa, V. et al. Glycolysis controls the induction of human regulatory T cells by modulating the expression of FOXP3 exon 2 splicing variants. Nat. Immunol. 16, 1174–1184 (2015).
Yang, S., Wang, J., Brand, D. D. & Zheng, S. G. Role of TNF–TNF receptor 2 Signal in regulatory T cells and Its therapeutic implications. Front. Immunol. 9, 784 (2018).
Atretkhany, K. N. et al. Intrinsic TNFR2 signaling in T regulatory cells provides protection in CNS autoimmunity. Proc. Natl Acad. Sci. USA 115, 13051–13056 (2018).
Marin Morales, J. M. et al. Automated clinical grade expansion of regulatory T cells in a fully closed system. Front. Immunol. 10, 38 (2019).
Opstelten, R. et al. GPA33: A marker to identify stable human regulatory T cells. J. Immunol. 204, 3139–3148 (2020).
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007).
Golovina, T. N. et al. CD28 costimulation is essential for human T regulatory expansion and function. J. Immunol. 181, 2855–2868 (2008).
Tanner, L. B. et al. Four key steps control glycolytic flux in mammalian cells. Cell Syst. 7, 49–62 e48 (2018).
Gerriets, V. A. et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J. Clin. Invest. 125, 194–207 (2015).
Shi, J. H. & Sun, S. C. Tumor necrosis factor receptor-associated factor regulation of nuclear factor κB and mitogen-activated protein kinase pathways. Front. Immunol. 9, 1849 (2018).
He, X. et al. A TNFR2-agonist facilitates high purity expansion of human low purity treg cells. PLoS ONE 11, e0156311 (2016).
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72 (2014).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Haas, R. et al. Intermediates of metabolism: from bystanders to signalling molecules. Trends Biochem. Sci. 41, 460–471 (2016).
Acuto, O. & Michel, F. CD28-mediated co-stimulation: a quantitative support for TCR signalling. Nat. Rev. Immunol. 3, 939–951 (2003).
Walker, L. S. & Sansom, D. M. Confusing signals: recent progress in CTLA-4 biology. Trends Immunol. 36, 63–70 (2015).
Grell, M. et al. The transmembrane form of tumor necrosis factor is the prime activating ligand of the 80 kDa tumor necrosis factor receptor. Cell 83, 793–802 (1995).
Urbano, P. C. M., Koenen, H., Joosten, I. & He, X. An autocrine TNFα-tumor necrosis factor receptor 2 loop promotes epigenetic effects inducing human treg stability in vitro. Front. Immunol. 9, 573 (2018).
Kleijwegt, F. S. et al. Critical role for tnf in the induction of human antigen-specific regulatory T cells by tolerogenic dendritic cells. J. Immunol. 185, 1412 (2010).
Chopra, M. et al. Exogenous TNFR2 activation protects from acute GvHD via host T reg cell expansion. J. Exp. Med. 213, 1881–1900 (2016).
Herbertson, R. A. et al. Targeted chemoradiation in metastatic colorectal cancer: a phase I trial of 131I-huA33 with concurrent capecitabine. J. Nucl. Med. 55, 534–539 (2014).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Williams, N. C. & O’Neill, L. A. J. A role for the Krebs cycle intermediate citrate in metabolic reprogramming in innate immunity and inflammation. Front. Immunol. 9, 141 (2018).
Thurnher, M. & Gruenbacher, G. T lymphocyte regulation by mevalonate metabolism. Sci. Signal 8, re4 (2015).
Timilshina, M. et al. Activation of mevalonate pathway via LKB1 is essential for stability of treg cells. Cell Rep. 27, 2948–2961 e2947 (2019).
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013).
Brightbill, H. D. et al. NF-κB inducing kinase is a therapeutic target for systemic lupus erythematosus. Nat. Commun. 9, 179 (2018).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Chen, Y., Lun, A. & Smyth, G. From reads to genes to pathways: differential expression analysis of RNA-seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res 5, 1438 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Ross, K. L. & Dalluge, J. J. Liquid chromatography/tandem mass spectrometry of glycolytic intermediates: deconvolution of coeluting structural isomers based on unique product ion ratios. Anal. Chem. 81, 4021–4026 (2009).
Smith, C. A. et al. METLIN: a metabolite mass spectral database. Ther. Drug Monit. 27, 747–751 (2005).
Guijas, C. et al. METLIN: a technology platform for identifying knowns and unknowns. Anal. Chem. 90, 3156–3164 (2018).
van der Windt, G. J., Chang, C. H. & Pearce, E. L. Measuring bioenergetics in T cells using a seahorse extracellular flux analyzer. Curr. Protoc. Immunol. 113, 3.16B.11–13.16B.14 (2016).
Zhuo, C. et al. Higher FOXP3–TSDR demethylation rates in adjacent normal tissues in patients with colon cancer were associated with worse survival. Mol. Cancer 13, 153 (2014).
Acknowledgements
We thank the employees of the Flow Cytometry Facility and Genomics Facility of the Netherlands Cancer Institute and the Flow Cytometry Core Facility at Leiden University Medical Center for their technical assistance, M. Giera and S. Kostidis of the Center for Proteomics and Metabolomics Facility at Leiden University Medical Center for valuable discussions regarding the metabolomics experiments on freshly sorted cells and E. Schrama for assistance in flow-cytometric assays. M. A. Aslam was supported by the Institute of Molecular Biology and Biotechnology, Bahauddin Zakariya University, Multan, Pakistan. This work was supported by Oncode, ZonMW-TOP grant 40-00812-98-13071 and grants ICI-00014 and ICI-00025 from the Institute for Chemical Immunology, funded by ZonMW Gravitation.
Author information
Authors and Affiliations
Contributions
S.d.K., M.M., J.B. and C.R.B. designed the study; S.d.K., M.M. and A.T.H. designed and performed the experiments and analysed the data; I.B. performed confocal microscopy experiments, analysed the data and contributed to the manuscript; R.J.E.D. performed targeted metabolomics analysis on freshly sorted cells and contributed to the manuscript; D.B. performed the short-term metabolic assays; M.A.A. provided assistance in analyses of the transcriptome data; S.d.K., M.M., J.B. and C.R.B. wrote the manuscript; D.A. provided conceptual advice in the study design and contributed to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors declare to have no financial or non-financial competing interests.
Additional information
Peer review information Primary Handling Editors: George Caputa; Elena Bellafante.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cell sorting strategy and characterization of naïve CD4+ Tconv and tTreg cells from human peripheral blood.
a, Total human CD4+ T cells were isolated from human PBMC using MACS and subjected to fluorescence-activated cell sorting. Within the live (PI-) and single cell gates (FSC-A/FSC-H), CD4+ T cells were selected for further discrimination between Tconv and tTreg cells using CD127 and CD25 expression levels (G1 and G2). Subsequently, naïve Tconv and tTreg cells were gated based on CD45RA expression and the novel marker GPA33 to obtain tTreg cells with high purity, resulting in CD25lowCD127highCD45RA+GPA33int Tconv cells and CD25highCD127lowCD45RA+GPA33high tTreg cells. b, Flow cytometric analysis of FOXP3, IKZF2 and total CTLA4 protein expression in the naïve Tconv and tTreg cells (representative of n=6). Mean fluorescence intensity (MFI) is shown in the plots. Sample size (n) represents cells from individual donors, analyzed in independent experiments.
Extended Data Fig. 2 Phenotypical and functional characterization of in vitro expanded Tconv and tTreg cells.
a, Schematic overview of Tconv and tTreg cell expansion cultures. The nature of “restimulation” on day 18 is indicated in the legends of the specific experiments. b, Flow cytometric analysis of FOXP3, IKZF2, cell surface CD25 and total CTLA4 protein levels in expanded Tconv and tTreg cells at 24 h after restimulation with anti-CD3/CD28 mAbs or control (-) (representative of n=6). c, Quantification of flow cytometric results obtained as indicated in b, showing the MFI for each protein (n=6), ****p=5.02x10-5, 3.34x10-6 and 5.58x10-5 for FOXP3, IKZF2 and CTLA4, respectively. d, Assessment of the suppressive capacity of tTreg cells on day 18 of the expansion cultures. Cells in the suppression assay were stimulated with agonistic mAb to CD3 and the assay was read out after 4 days. The percentage of dividing CD4+ and CD8+ Tconv cells is displayed (n=5), ****p=5.1x10-13, 4.99x10-13, 5.57x10-13, 8.67x10-13 and 3.32x10-11 for 1:1, 1:2, 1:4, 1:8 and 1:16 ratio versus 0:1 ratio, respectively, within CD3+CD4+ responder cells and ****p=5.81x10-13, 7.83x10-13, 2.67x10-12, 3.4x10-11 and 1.78x10-7 for 1:1, 1:2, 1:4, 1:8 and 1:16 ratio versus 0:1 ratio, respectively, within CD3+CD8+ responder cells. (c, d) Two-way ANOVA with Tukey’s post hoc test was used for statistical analysis. Data are presented as mean ± SEM. Sample size (n) represents cells from individual donors, analyzed in independent experiments (a–d).
Extended Data Fig. 3 PFK mRNA and protein expression by Tconv and tTreg cells upon CD3/CD28-mediated activation.
a, Venn diagrams depicting the number of overlapping and unique genes (left) or metabolites (right) that showed significant changes in abundance upon CD3/CD28-mediated activation for 24 h in Tconv and tTreg cells, as determined by transcriptomics and metabolomics (n=4). Statistical evaluation was performed using an unpaired two-sided Student’s t-test with the Benjamini-Hochberg method with FDR<0.05 (transcriptomics) or by unpaired two-sided Student’s t-test with p<0.05 (metabolomics). b, mRNA (top) and protein levels (bottom) of PFK isoforms in Tconv and tTreg cells at 24 h after stimulation as indicated. mRNA levels are expressed in TPM, as detected in our transcriptome data set (n=4), *p=0.0408. (c) Protein levels are expressed in LFQ intensity, as detected in an earlier proteomics data set14 (n=3). (b, c) Two-way ANOVA with Bonferroni’s post hoc test was used for statistical analysis. Data are presented as mean ± SEM. Sample size (n) represents independent expansion cultures of distinct donors (a–c).
Extended Data Fig. 4 Significant changes in glycolysis/TCA cycle intermediates in Tconv and tTreg cells activated via CD3/CD28.
Unpaired two-sided Student’s t-test was used for statistical analysis (*). Data is derived from 4 independent expansion cultures of distinct donors.
Extended Data Fig. 5 TNFR2 acts as a costimulatory receptor on tTreg cells.
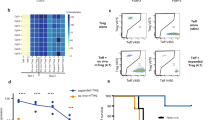
a, Flow cytometric assessment of tTreg cell division by CTV dilution upon restimulation for 96 h as indicated. The percentage of dividing cells is depicted. FOXP3, IKZF2 and total CTLA4 protein expression is shown for each cell division (representative of n=3). b, Left panel: Flow cytometric assessment of 6-NBDG uptake activity in Tconv and tTreg cells that were restimulated for 24 h as indicated. Dashed lines indicate the modal 6-NBDG uptake for unstimulated cells. Right panel: quantification of 6-NBDG uptake data, based on the geometric MFI, normalized to unstimulated Tconv cells (n=3), ****p=5.88x10-5. Two-way ANOVA with Tukey’s post hoc test was used for statistical analysis. Data are presented as mean ± SEM. Sample size (n) represents cells from individual donors, analyzed in independent experiments.
Extended Data Fig. 6 Transcriptomic analysis of Tconv and tTreg cells upon TNFR2 costimulation.
a, PCA plot of the transcriptomes of Tconv and tTreg cells that were activated via either CD3 or CD3/TNFR2 for 24 h (n=3 independent expansion cultures from different donors). b, Heat map showing hierarchical clustering of the 2664 genes that were differentially expressed between the comparative conditions. Z-scores showing relative gene expression values are color-coded. Grey-scale boxes (right) indicate clusters. Statistical evaluation was performed according to ANOVA with the Benjamini-Hochberg method with FDR<0.005.
Extended Data Fig. 7 Significant changes in glycolysis/TCA cycle intermediates in Tconv and tTreg cells activated via CD3/TNFR2.
Unpaired two-sided Student’s t-test was used for statistical analysis (*). Data is derived from 4 independent expansion cultures of distinct donors.
Extended Data Fig. 8 Complete mass isotopologue analysis of glycolysis and TCA cycle intermediates upon [13C6]-glucose tracing in Tconv and tTreg cells.
Levels of all detected mass isotopologues of the indicated metabolites in Tconv and tTreg cells that were unstimulated or activated via CD3 alone or CD3/TNFR2 for 24 h in the presence of [13C6]-glucose, as measured by LC-MS (n=4 independent expansion cultures, as in Fig. 5), ***p=0.0002 for HexP, *p=0.0381 for F-1,6-BP, **p=0.0029 for DHAP, *p=0.0397 for pyruvate, *p=0.0137 for α-ketoglutarate and **p=0.0029 for malate. Two-way ANOVA with Bonferroni’s post hoc test was used for statistical analysis. Data are presented as mean ± SEM.
Extended Data Fig. 9 Phosphorylation of mTOR and S6 in Tconv cells.
a, Left panel: flow cytometric analysis of phosphorylated mTOR (Ser2448) levels in Tconv cells following stimulation for 24 h as indicated. Right panel: quantification based on MFI (n=4), *p=0.0131 and 0.0197 for αCD3/28 and αCD3/TNFR2 versus unstimulated, respectively. b, Left panel: flow cytometric analysis of phosphorylated S6 (Ser235/236) levels in Tconv cells following stimulation for 24 h as indicated. Right panel: quantification based on the MFI (n=4), **p=0.0026 and ***p=0.0003 for αCD3/28 and αCD3/TNFR2 versus unstimulated, respectively. (a, b) One-way repeated measures ANOVA with Bonferroni’s post hoc test was used for statistical analysis. c, Quantification of phosphorylated mTOR (Ser2448) levels based on MFI in ex vivo naïve Tconv and tTreg cells stimulated for 24 h as indicated (n=3), **p=0.0039. d, Quantification of phosphorylated S6 (Ser235/236) levels based on MFI in ex vivo naïve Tconv and tTreg cells stimulated for 24 h as indicated (n=4), *p=0.0175. e, Quantification of 6-NBDG uptake activity in ex vivo naïve Tconv and tTreg cells stimulated for 24 h as indicated (n=3), **p=0.0015. f, g, Quantification of flow cytometric analysis of phosphorylated mTOR (Ser2448) (f) or S6 (Ser235/Ser236) (g) levels in Tconv cells following stimulation for 24 h as indicated in presence or absence of selective inhibitors of PI3K (LY294002) or NIK (NIK-SMI1) in the culture medium (n=5), ****p=4.54x10-6, 2.13x10-6 and 5.2x10-10 for phosphorylated mTOR comparing medium versus LY294002 for αCD3, αCD3/28 and αCD3/TNFR2, respectively, and ****p=1.91x10-5, 8.55x10-7 and 4.64x10-9 for phosphorylated S6 comparing medium versus LY294002 for αCD3, αCD3/28 and αCD3/TNFR2, respectively. (c–g) Two-way ANOVA with Tukey’s post hoc test was used for statistical analysis. Data are presented as mean ± SEM. Sample size (n) represents cells from individual donors, analyzed in independent experiments.
Supplementary information
Supplementary Information
Supplementary Figure 1
Rights and permissions
About this article
Cite this article
de Kivit, S., Mensink, M., Hoekstra, A.T. et al. Stable human regulatory T cells switch to glycolysis following TNF receptor 2 costimulation. Nat Metab 2, 1046–1061 (2020). https://doi.org/10.1038/s42255-020-00271-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00271-w
This article is cited by
-
Cellular metabolism regulates the differentiation and function of T-cell subsets
Cellular & Molecular Immunology (2024)
-
TNF and TNF receptors as therapeutic targets for rheumatic diseases and beyond
Nature Reviews Rheumatology (2023)
-
The TNFα/TNFR2 axis mediates natural killer cell proliferation by promoting aerobic glycolysis
Cellular & Molecular Immunology (2023)
-
TNFR2 antagonistic antibody induces the death of tumor infiltrating CD4+Foxp3+ regulatory T cells
Cellular Oncology (2023)
-
Proteomics reveals unique identities of human TGF-β-induced and thymus-derived CD4+ regulatory T cells
Scientific Reports (2022)