Abstract
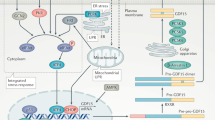
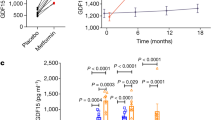
Metformin is the most commonly prescribed medication for type 2 diabetes, owing to its glucose-lowering effects, which are mediated through the suppression of hepatic glucose production (reviewed in refs. 1,2,3). However, in addition to its effects on the liver, metformin reduces appetite and in preclinical models exerts beneficial effects on ageing and a number of diverse diseases (for example, cognitive disorders, cancer, cardiovascular disease) through mechanisms that are not fully understood1,2,3. Given the high concentration of metformin in the liver and its many beneficial effects beyond glycemic control, we reasoned that metformin may increase the secretion of a hepatocyte-derived endocrine factor that communicates with the central nervous system4. Here we show, using unbiased transcriptomics of mouse hepatocytes and analysis of proteins in human serum, that metformin induces expression and secretion of growth differentiating factor 15 (GDF15). In primary mouse hepatocytes, metformin stimulates the secretion of GDF15 by increasing the expression of activating transcription factor 4 (ATF4) and C/EBP homologous protein (CHOP; also known as DDIT3). In wild-type mice fed a high-fat diet, oral administration of metformin increases serum GDF15 and reduces food intake, body mass, fasting insulin and glucose intolerance; these effects are eliminated in GDF15 null mice. An increase in serum GDF15 is also associated with weight loss in patients with type 2 diabetes who take metformin. Although further studies will be required to determine the tissue source(s) of GDF15 produced in response to metformin in vivo, our data indicate that the therapeutic benefits of metformin on appetite, body mass and serum insulin depend on GDF15.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rena, G., Pearson, E. R. & Sakamoto, K. Molecular mechanism of action of metformin: old or new insights? Diabetologia 56, 1898–1906 (2013).
Foretz, M., Guigas, B., Bertrand, L., Pollak, M. & Viollet, B. Metformin: from mechanisms of action to therapies. Cell Metab. 20, 953–966 (2014).
Rena, G., Hardie, D. G. & Pearson, E. R. The mechanisms of action of metformin. Diabetologia 60, 1577–1585 (2017).
Steinberg, G. R. Cellular energy sensing and metabolism—implications for treating diabetes: The 2017 outstanding scientific achievement award lecture. Diabetes 67, 169–179 (2018).
Chandel, N. S. et al. Are metformin doses used in murine cancer models clinically relevant? Cell Metab. 23, 569–570 (2016).
Madiraju, A. K. et al. Metformin inhibits gluconeogenesis via a redox-dependent mechanism in vivo. Nat. Med. 24, 1384–1394 (2018).
Jensen, J. B. et al. [11C]-labeled metformin distribution in the liver and small intestine using dynamic positron emission tomography in mice demonstrates tissue-specific transporter dependency. Diabetes 65, 1724–1730 (2016).
Gormsen, L. C. et al. In vivo imaging of human 11C-metformin in peripheral organs: dosimetry, biodistribution, and kinetic analyses. J. Nucl. Med. 57, 1920–1926 (2016).
Frid, A. et al. Novel assay of metformin levels in patients with type 2 diabetes and varying levels of renal function: clinical recommendations. Diabetes Care 33, 1291–1293 (2010).
Hunter, R. W. et al. Metformin reduces liver glucose production by inhibition of fructose-1-6-bisphosphatase. Nat. Med. 24, 1395–1406 (2018).
Fullerton, M. D. et al. Single phosphorylation sites in Acc1 and Acc2 regulate lipid homeostasis and the insulin-sensitizing effects of metformin. Nat. Med. 19, 1649–1654 (2013).
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015).
Wu, H. et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23, 850–858 (2017).
Duca, F. A. et al. Metformin activates a duodenal Ampk–dependent pathway to lower hepatic glucose production in rats. Nat. Med. 21, 506–511 (2015).
Maida, A., Lamont, B. J., Cao, X. & Drucker, D. J. Metformin regulates the incretin receptor axis via a pathway dependent on peroxisome proliferator-activated receptor-α in mice. Diabetologia 54, 339–349 (2011).
Golay, A. Metformin and body weight. Int. J. Obes. 32, 61–72 (2008).
Diabetes Prevention Program Research Group. Long-term safety, tolerability, and weight loss associated with metformin in the Diabetes Prevention Program Outcomes Study. Diabetes Care 35, 731–737 (2012).
Lee, A. & Morley, J. E. Metformin decreases food consumption and induces weight loss in subjects with obesity with type II non-insulin-dependent diabetes. Obes. Res. 6, 47–53 (1998).
Wang, Q. et al. Metformin suppresses diabetes-accelerated atherosclerosis via the inhibition of Drp1-mediated mitochondrial fission. Diabetes 66, 193–205 (2017).
Ning, H.-H. et al. The effects of metformin on simple obesity: a meta-analysis. Endocrine 62, 528–534 (2018).
Solymár, M. et al. Metformin induces significant reduction of body weight, total cholesterol and LDL levels in the elderly - a meta-analysis. PLoS ONE 13, e0207947 (2018).
Stumvoll, M., Nurjhan, N., Perriello, G., Dailey, G. & Gerich, J. E. Metabolic effects of metformin in non-insulin-dependent diabetes mellitus. N. Engl. J. Med. 333, 550–554 (1995).
Kim, H.-J. et al. Metformin decreases meal size and number and increases c-Fos expression in the nucleus tractus solitarius of obese mice. Physiol. Behav. 110–111, 213–220 (2013).
Meinken, J., Walker, G., Cooper, C. R. & Min, X. J. MetazSecKB: the human and animal secretome and subcellular proteome knowledgebase. Database 2015, bav077 (2015).
McInnes, N. et al. Piloting a remission strategy in type 2 diabetes: results of a randomized controlled trial. J. Clin. Endocrinol. Metab. 102, 1596–1605 (2017).
Patel, S. et al. GDF15 provides an endocrine signal of nutritional stress in mice and humans. Cell Metab. 29, 707–718 (2019).
Hsu, J.-Y. et al. Non-homeostatic body weight regulation through a brainstem-restricted receptor for GDF15. Nature 550, 255–259 (2017).
Mullican, S. E. et al. GFRAL is the receptor for GDF15 and the ligand promotes weight loss in mice and nonhuman primates. Nat. Med. 23, 1150–1157 (2017).
Emmerson, P. J. et al. The metabolic effects of GDF15 are mediated by the orphan receptor GFRAL. Nat. Med. 23, 1215–1219 (2017).
Yang, L. et al. GFRAL is the receptor for GDF15 and is required for the anti-obesity effects of the ligand. Nat. Med. 23, 1158–1166 (2017).
Quentin, T., Steinmetz, M., Poppe, A. & Thoms, S. Metformin differentially activates ER stress signaling pathways without inducing apoptosis. Dis. Model. Mech. 5, 259–269 (2012).
Kim, K. H. et al. Metformin-induced inhibition of the mitochondrial respiratory chain increases FGF21 expression via ATF4 activation. Biochem. Biophys. Res. Commun. 440, 76–81 (2013).
Gerstein, H. C. et al. Growth differentiation factor 15 as a novel biomarker for metformin. Diabetes Care 40, 280–283 (2017).
Natali, A. et al. Metformin is the key factor in elevated plasma growth differentiation factor-15 levels in type 2 diabetes: a nested, case-control study. Diabetes Obes. Metab. 21, 412–416 (2019).
Chrysovergis, K. et al. NAG-1/GDF-15 prevents obesity by increasing thermogenesis, lipolysis and oxidative metabolism. Int. J. Obes. (Lond.) 38, 1555–1564 (2014).
Johnen, H. et al. Tumor-induced anorexia and weight loss are mediated by the TGF-β superfamily cytokine MIC-1. Nat. Med. 13, 1333–1340 (2007).
Kim, J. M. et al. NAG-1/GDF15 transgenic mouse has less white adipose tissue and a reduced inflammatory response. Mediators Inflamm. 2013, 641851 (2013).
Tsai, V. W.-W. et al. TGF-b superfamily cytokine MIC-1/GDF15 is a physiological appetite and body weight regulator. PLoS ONE 8, e55174 (2013).
Ding, Q. et al. Identification of macrophage inhibitory cytokine-1 in adipose tissue and its secretion as an adipokine by human adipocytes. Endocrinology 150, 1688–1696 (2009).
Macia, L. et al. Macrophage inhibitory cytokine 1 (MIC-1/GDF15) decreases food intake, body weight and improves glucose tolerance in mice on normal & obesogenic diets. PLoS ONE 7, e34868 (2012).
Grant, P. J. Beneficial effects of metformin on haemostasis and vascular function in man. Diabetes Metab. 29, 6S44–6S52 (2003).
Barzilai, N., Crandall, J. P., Kritchevsky, S. B. & Espeland, M. A. Metformin as a tool to target aging. Cell Metab. 23, 1060–1065 (2016).
Dzamko, N. et al. AMPK 1 deletion reduces appetite, preventing obesity and hepatic insulin resistance. J. Biol. Chem. 285, 115–122 (2010).
Zinszner, H. et al. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev. 12, 982–995 (1998).
Leonhard, W. N. et al. Salsalate, but not metformin or canagliflozin, slows kidney cyst growth in an adult-onset mouse model of polycystic kidney disease. EBioMedicine 47, 436–445 (2019).
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Breitling, R., Armengaud, P., Amtmann, A. & Herzyk, P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 573, 83–92 (2004).
Acknowledgements
The authors would like to thank A. Božović and V. Kulasingam for measuring serum metformin levels. E.A.D. was a recipient of an Ontario Graduate Scholarship (Queen Elizabeth II Graduate Scholarship in Science and Technology) and a Douglas C. Russell Memorial Scholarship. G.R.S. is a Canada Research Chair and the J. Bruce Duncan Chair in Metabolic Diseases. This study was supported by research grants from the Canadian Institutes of Health Research (201709FDN-CEBA-116200 to G.R.S.) and Diabetes Canada (DI-5-17-5302-GS). We thank Sanofi for providing heterozygous breeding pairs of GDF15-null mice.
Author information
Authors and Affiliations
Contributions
E.A.D., R.J.F., B.K.S., N.M., S.H., G.P., H.C.G. and G.R.S. designed the experiments. E.A.D., R.J.F., B.K.S., P.M.S., M.R.M., R.L. and R.M.G. performed the experiments and/or analysed the data. A.R.R. and A.G.M. provided bioinformatics analysis and support. M.K. generated GDF15-KO mice. E.A.D., R.J.F. and G.R.S. wrote the manuscript. All authors edited the manuscript and provided comments.
Corresponding author
Ethics declarations
Competing interests
S.H., G.P., H.C.G. and G.R.S. hold a patent entitled ‘Growth differentiation factor 15 as biomarker for metformin’ (WO/2017/108941). S.H. and M.K. are employees of Sanofi. All other authors have no competing interests.
Additional information
Peer review information Primary Handling Editor: Christoph Schmitt.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Metformin, phenformin and buformin increase GDF15 release independent of complex 1 inhibition or AMPK.
a, Volcano plot showing differentially regulated genes after 24 h of metformin treatment (metformin versus control, n = 4 per group). b,c, GDF15 release is stimulated in a dose-dependent manner by two biguanides that are structurally similar to metformin: (b) phenformin (0 μM n = 4, 10 μM n = 3, 30 μM n = 3 and 50 μM n = 4) and (c) buformin (0 μM n = 3, 10 μM n = 2, 30 μM n = 2 and 50 μM n = 3). d, The complex I inhibitor rotenone (0 μM, 0.1 μM, 1 μM, 5 μM n = 3) does not increase GDF15 release. e, Metformin increases GDF15 release in primary hepatocytes from WT (control n = 7, metformin n = 7), AMPK β1KO (control n = 4, metformin n = 3) and ACC DKI (control n = 3, metformin n = 3) mice. Data are presented as mean ± s.e.m. For b–d, * indicates P < 0.05 and *** indicates P < 0.001 for one-way ANOVA with Sidak multiple comparison test. For e, * indicates P < 0.05 for two-way ANOVA with Sidak multiple comparison test.
Extended Data Fig. 2 Acute metformin treatment did not alter energy expenditure but reduced RER.
a, Serum metformin 1 h after acute saline (n = 4) or metformin gavage (n = 7). *P < 0.05, for unpaired two-sided t-test. b, Serum GLP-1 10 min after acute metformin gavage in wild-type (n = 8) and GDF15-KO (n = 6) mice fed a 45% HFD. c–f, Wild-type (control n = 9, metformin n = 9) and GDF15-KO (control n = 6, metformin n = 6) mice were fed a 45% HFD, placed in metabolic cages, and allowed to acclimatize for approximately 24 h before a single oral gavage of metformin (250 mg kg–1) or appropriate volume of saline 2 h before the onset of the dark period. RER, beam breaks and energy expenditure were measured for 24 h after gavage, and data are presented as mean ± s.e.m. *P < 0.05 between control and metformin, two-way ANOVA with Sidak multiple comparison test.
Extended Data Fig. 3 Chronic metformin treatment does not alter lean mass, RER or energy expenditure.
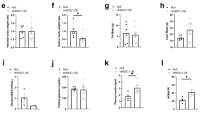
Wild-type (control n = 8, metformin n = 9) and GDF15-KO (control n = 8, metformin n = 8) mice were fed a 45% HFD for 4 weeks prior to being switched to control (tap water) or metformin water (3 g l−1) for 10 weeks. a,b, Body composition was assessed at week 4 of treatment (wild-type control n = 8, wild-type metformin n = 9, GDF15-KO control n = 8 and GDF15-KO metformin n = 8). c–i, Wild-type (control n = 7, metformin n = 8) and GDF15-KO (control n = 8, metformin n = 8) mice were placed in metabolic cages and allowed to acclimatize for approximately 24 h. Food intake, activity, RER and energy expenditure were measured over 48 h. Energy expenditure is shown (e) uncorrected, (f,g) corrected for body mass, and (h,i) corrected for lean mass, and data are presented as mean ± s.e.m.
Extended Data Fig. 4 Metformin in drinking water elicits clinically relevant serum metformin levels.
Wild-type and GDF15-KO mice were fed a 45% HFD for 4 weeks prior to being switched to control (tap water) or metformin water (3 g l−1) for 10 weeks. Wild-type (control n = 6, metformin n = 8) and GDF15-KO (control n = 6, metformin n = 7) mice were placed in metabolic cages and allowed to acclimatize for approximately 24 h. a, Water intake was measured over 48 h. b, Metformin dose was calculated based on water intake and body mass. c, Serum metformin was measured after mice were killed at the onset of the light period. d, Food intake of 45% HFD was monitored every 3–4 days in mice fed ad libitum (n = 9), mice fed ad libitum treated with metformin (n = 10), and mice pair-fed with metformin-treated animals (n = 10). e, Serum GDF15 was assessed at 3 weeks. Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.005, ***P < 0.001 between control and metformin, two-way ANOVA with Sidak multiple comparison test (a–c), one-way ANOVA with Sidak multiple comparison test (d,e).
Supplementary information
Supplementary Information
Supplementary Tables 1–3
Source data
Source Data Fig. 2
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Day, E.A., Ford, R.J., Smith, B.K. et al. Metformin-induced increases in GDF15 are important for suppressing appetite and promoting weight loss. Nat Metab 1, 1202–1208 (2019). https://doi.org/10.1038/s42255-019-0146-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-019-0146-4
This article is cited by
-
Obesity and the kidney: mechanistic links and therapeutic advances
Nature Reviews Endocrinology (2024)
-
mTORC1 in energy expenditure: consequences for obesity
Nature Reviews Endocrinology (2024)
-
Metformin and feeding increase levels of the appetite-suppressing metabolite Lac-Phe in humans
Nature Metabolism (2024)
-
The gastrointestinal tract is a major source of the acute metformin-stimulated rise in GDF15
Scientific Reports (2024)
-
Artesunate treats obesity in male mice and non-human primates through GDF15/GFRAL signalling axis
Nature Communications (2024)