Abstract
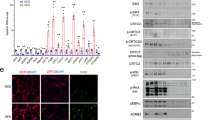
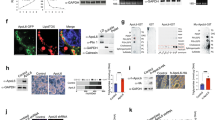
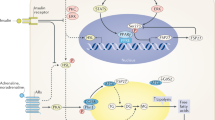
Impaired adipose tissue insulin signalling is a critical feature of insulin resistance. Here we identify a pathway linking the lipolytic enzyme hormone-sensitive lipase (HSL) to insulin action via the glucose-responsive transcription factor ChREBP and its target, the fatty acid elongase ELOVL6. Genetic inhibition of HSL in human adipocytes and mouse adipose tissue results in enhanced insulin sensitivity and induction of ELOVL6. ELOVL6 promotes an increase in phospholipid oleic acid, which modifies plasma membrane fluidity and enhances insulin signalling. HSL deficiency–mediated effects are suppressed by gene silencing of ChREBP and ELOVL6. Mechanistically, physical interaction between HSL, independent of lipase activity, and the isoform activated by glucose metabolism ChREBPα impairs ChREBPα translocation into the nucleus and induction of ChREBPβ, the isoform with high transcriptional activity that is strongly associated with whole-body insulin sensitivity. Targeting the HSL–ChREBP interaction may allow therapeutic strategies for the restoration of insulin sensitivity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding author upon reasonable request.
References
Abel, E. D. et al. Adipose-selective targeting of the GLUT4 gene impairs insulin action in muscle and liver. Nature 409, 729–733 (2001).
Morley, T. S., Xia, J. Y. & Scherer, P. E. Selective enhancement of insulin sensitivity in the mature adipocyte is sufficient for systemic metabolic improvements. Nat. Commun. 6, 7906 (2015).
Shearin, A. L., Monks, B. R., Seale, P. & Birnbaum, M. J. Lack of AKT in adipocytes causes severe lipodystrophy. Mol. Metab. 5, 472–479 (2016).
Softic, S. et al. Lipodystrophy due to adipose tissue specific insulin receptor knockout results in progressive NAFLD. Diabetes 65, 2187–2200 (2016).
Rondinone, C. M. et al. Insulin receptor substrate (IRS) 1 is reduced and IRS-2 is the main docking protein for phosphatidylinositol 3-kinase in adipocytes from subjects with non-insulin-dependent diabetes mellitus. Proc. Natl Acad. Sci. USA 94, 4171–4175 (1997).
Carvalho, E., Jansson, P. A., Nagaev, I., Wenthzel, A. M. & Smith, U. Insulin resistance with low cellular IRS-1 expression is also associated with low GLUT4 expression and impaired insulin-stimulated glucose transport. FASEB J. 15, 1101–1103 (2001).
Frojdo, S., Vidal, H. & Pirola, L. Alterations of insulin signaling in type 2 diabetes: a review of the current evidence from humans. Biochim. Biophys. Acta 1792, 83–92 (2009).
Nyman, E. et al. A single mechanism can explain network-wide insulin resistance in adipocytes from obese patients with type 2 diabetes. J. Biol. Chem. 289, 33215–33230 (2014).
Samuel, V. T. & Shulman, G. I. Mechanisms for insulin resistance: common threads and missing links. Cell 148, 852–871 (2012).
Karpe, F., Dickmann, J. R. & Frayn, K. N. Fatty acids, obesity, and insulin resistance: time for a reevaluation. Diabetes 60, 2441–2449 (2011).
Girousse, A. et al. Partial inhibition of adipose tissue lipolysis improves glucose metabolism and insulin sensitivity without alteration of fat mass. PLoS Biol. 11, e1001485 (2013).
Bezaire, V. et al. Contribution of adipose triglyceride lipase and hormone-sensitive lipase to lipolysis in hMADS adipocytes. J. Biol. Chem. 284, 18282–18291 (2009).
Barquissau, V. et al. White-to-brite conversion in human adipocytes promotes metabolic reprogramming towards fatty acid anabolic and catabolic pathways. Mol. Metab. 5, 352–365 (2016).
Eissing, L. et al. De novo lipogenesis in human fat and liver is linked to ChREBP-beta and metabolic health. Nat. Commun. 4, 1528 (2013).
Roberts, R. et al. Markers of de novo lipogenesis in adipose tissue: associations with small adipocytes and insulin sensitivity in humans. Diabetologia 52, 882–890 (2009).
Collins, J. M., Neville, M. J., Hoppa, M. B. & Frayn, K. N. De novo lipogenesis and stearoyl-CoA desaturase are coordinately regulated in the human adipocyte and protect against palmitate-induced cell injury. J. Biol. Chem. 285, 6044–6052 (2010).
Guillou, H., Zadravec, D., Martin, P. G. & Jacobsson, A. The key roles of elongases and desaturases in mammalian fatty acid metabolism: insights from transgenic mice. Prog. Lipid Res. 49, 186–199 (2010).
Hodson, L. & Fielding, B. A. Stearoyl-CoA desaturase: rogue or innocent bystander? Prog. Lipid Res. 52, 15–42 (2013).
Ohno, Y. et al. ELOVL1 production of C24 acyl-CoAs is linked to C24 sphingolipid synthesis. Proc. Natl Acad. Sci. USA 107, 18439–18444 (2010).
Nagase, T. et al. Synthesis and biological evaluation of a novel 3-sulfonyl-8-azabicyclo[3.2.1]octane class of long chain fatty acid elongase 6 (ELOVL6) inhibitors. J. Med. Chem. 52, 4111–4114 (2009).
Xin, Z. et al. Discovery of piperidine-aryl urea-based stearoyl-CoA desaturase 1 inhibitors. Bioorg. Med. Chem. Lett. 18, 4298–4302 (2008).
Antonny, B., Vanni, S., Shindou, H. & Ferreira, T. From zero to six double bonds: phospholipid unsaturation and organelle function. Trends Cell Biol. 25, 427–436 (2015).
Holzer, R. G. et al. Saturated fatty acids induce c-Src clustering within membrane subdomains, leading to JNK activation. Cell 147, 173–184 (2011).
Filhoulaud, G., Guilmeau, S., Dentin, R., Girard, J. & Postic, C. Novel insights into ChREBP regulation and function. Trends Endocrinol. Metab. 24, 257–268 (2013).
Herman, M. A. et al. A novel ChREBP isoform in adipose tissue regulates systemic glucose metabolism. Nature 484, 333–338 (2012).
Bae, J. S., Oh, A. R., Lee, H. J., Ahn, Y. H. & Cha, J. Y. Hepatic Elovl6 gene expression is regulated by the synergistic action of ChREBP and SREBP-1c. Biochem. Biophys. Res. Commun. 478, 1060–1066 (2016).
Langin, D., Laurell, H., Holst, L. S., Belfrage, P. & Holm, C. Gene organization and primary structure of human hormone-sensitive lipase: possible significance of a sequence homology with a lipase of Moraxella TA144, an antarctic bacterium. Proc. Natl Acad. Sci. USA 90, 4897–4901 (1993).
Stoeckman, A. K., Ma, L. & Towle, H. C. Mlx is the functional heteromeric partner of the carbohydrate response element-binding protein in glucose regulation of lipogenic enzyme genes. J. Biol. Chem. 279, 15662–15669 (2004).
Soderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Lou, D. Q. et al. Chicken ovalbumin upstream promoter-transcription factor II, a new partner of the glucose response element of the L-type pyruvate kinase gene, acts as an inhibitor of the glucose response. J. Biol. Chem. 274, 28385–28394 (1999).
Laurell, H. et al. Species-specific alternative splicing generates a catalytically inactive form of human hormone-sensitive lipase. Biochem. J. 328, 137–143 (1997).
Ray, H. et al. The presence of a catalytically inactive form of hormone-sensitive lipase is associated with decreased lipolysis in abdominal subcutaneous adipose tissue of obese subjects. Diabetes 52, 1417–1422 (2003).
Hoffstedt, J., Forster, D. & Lofgren, P. Impaired subcutaneous adipocyte lipogenesis is associated with systemic insulin resistance and increased apolipoprotein B/AI ratio in men and women. J. Intern. Med. 262, 131–139 (2007).
Kursawe, R. et al. Decreased transcription of ChREBP-alpha/beta isoforms in abdominal subcutaneous adipose tissue of obese adolescents with prediabetes or early type 2 diabetes: associations with insulin resistance and hyperglycemia. Diabetes 62, 837–844 (2013).
Nilsson, E. et al. Altered DNA methylation and differential expression of genes influencing metabolism and inflammation in adipose tissue from subjects with type 2 diabetes. Diabetes 63, 2962–2976 (2014).
Soronen, J. et al. Adipose tissue gene expression analysis reveals changes in inflammatory, mitochondrial respiratory and lipid metabolic pathways in obese insulin-resistant subjects. BMC Med. Genom. 5, 9 (2012).
Matsuzaka, T. et al. Crucial role of a long-chain fatty acid elongase, Elovl6, in obesity-induced insulin resistance. Nat. Med. 13, 1193–1202 (2007).
Ryan, M. et al. Diabetes and the Mediterranean diet: a beneficial effect of oleic acid on insulin sensitivity, adipocyte glucose transport and endothelium-dependent vasoreactivity. QJM 93, 85–91 (2000).
Salas-Salvado, J. et al. Reduction in the incidence of type 2 diabetes with the Mediterranean diet: results of the PREDIMED-Reus nutrition intervention randomized trial. Diabetes Care 34, 14–19 (2011).
Ibarguren, M. et al. Partitioning of liquid-ordered/liquid-disordered membrane microdomains induced by the fluidifying effect of 2-hydroxylated fatty acid derivatives. Biochim. Biophys. Acta 1828, 2553–2563 (2013).
Pietilainen, K. H. et al. Association of lipidome remodeling in the adipocyte membrane with acquired obesity in humans. PLoS Biol. 9, e1000623 (2011).
Moon, Y. A., Ochoa, C. R., Mitsche, M. A., Hammer, R. E. & Horton, J. D. Deletion of ELOVL6 blocks the synthesis of oleic acid but does not prevent the development of fatty liver or insulin resistance. J. Lipid Res. 55, 2597–2605 (2014).
Kraemer, F. B. & Shen, W. J. Hormone-sensitive lipase: control of intracellular tri-(di-)acylglycerol and cholesteryl ester hydrolysis. J. Lipid Res. 43, 1585–1594 (2002).
Lafontan, M. & Langin, D. Lipolysis and lipid mobilization in human adipose tissue. Prog. Lipid Res. 48, 275–297 (2009).
Czech, M. P. Cellular basis of insulin insensitivity in large rat adipocytes. J. Clin. Invest. 57, 1523–1532 (1976).
Solinas, G., Boren, J. & Dulloo, A. G. De novo lipogenesis in metabolic homeostasis: more friend than foe?. Mol. Metab. 4, 367–377 (2015).
Skurk, T., Ecklebe, S. & Hauner, H. A novel technique to propagate primary human preadipocytes without loss of differentiation capacity. Obes. (Silver Spring) 15, 2925–2931 (2007).
Rossmeislova, L. et al. Weight loss improves the adipogenic capacity of human preadipocytes and modulates their secretory profile. Diabetes 62, 1990–1995 (2013).
Moon, Y. A., Shah, N. A., Mohapatra, S., Warrington, J. A. & Horton, J. D. Identification of a mammalian long chain fatty acyl elongase regulated by sterol regulatory element-binding proteins. J. Biol. Chem. 276, 45358–45366 (2001).
Bonneau, L. et al. Plasma membrane sterol complexation, generated by filipin, triggers signaling responses in tobacco cells. Biochim. Biophys. Acta 1798, 2150–2159 (2010).
Grober, J. et al. Characterization of the promoter of human adipocyte hormone-sensitive lipase. Biochem. J. 328, 453–461 (1997).
Langin, D. et al. Adipocyte lipases and defect of lipolysis in human obesity. Diabetes 54, 3190–3197 (2005).
Iizuka, K., Bruick, R. K., Liang, G., Horton, J. D. & Uyeda, K. Deficiency of carbohydrate response element-binding protein (ChREBP) reduces lipogenesis as well as glycolysis. Proc. Natl Acad. Sci. USA 101, 7281–7286 (2004).
Tan, C. Y. et al. Brown adipose tissue thermogenic capacity is regulated by Elovl6. Cell Rep. 13, 2039–2047 (2015).
Klimcakova, E. et al. Worsening of obesity and metabolic status yields similar molecular adaptations in human subcutaneous and visceral adipose tissue: decreased metabolism and increased immune response. J. Clin. Endocrinol. Metab. 96, E73–E82 (2011).
Del Prato, S. et al. Characterization of cellular defects of insulin action in type 2 (non-insulin-dependent) diabetes mellitus. J. Clin. Invest. 91, 484–494 (1993).
DeFronzo, R. A., Tobin, J. D. & Andres, R. Glucose clamp technique: a method for quantifying insulin secretion and resistance. Am. J. Physiol. 237, E214–E223 (1979).
Dahlman, I. et al. The fat cell epigenetic signature in post-obese women is characterized by global hypomethylation and differential DNA methylation of adipogenesis genes. Int. J. Obes. (Lond.). 39, 910–919 (2015).
Acknowledgements
The authors acknowledge N. Venteclef (Centre de Recherche des Cordeliers, Paris) and J. Boucher (AstraZeneca, Göteborg, Sweden) for critical reading and comments on the manuscript. E. Courty and J. Personnaz participated in mouse studies during internship at I2MC. The GenoToul Animal Care, Anexplo, Imaging-TRI (especially F. Gaits-Iacovoni for helpful discussion) and Quantitative Transcriptomics facilities contributed to the work. This work was supported by Inserm, Paul Sabatier University, Fondation pour la Recherche Médicale (DEQ20170336720 to D.L.), Agence Nationale de la Recherche (ANR-12-BSV1-0025Obelip and ANR-17-CE14-0015Hepadialogue to D.L.), Région Midi-Pyrénées (OBELIP and ILIP projects to D.L.), FORCE/F-CRIN for clinical research on obesity, EU/EFPIA Innovative Medicines Initiative Joint Undertaking (EMIF grant 115372 to P.A., A.V.P. and D.L.) and AstraZeneca France (TALIP project to D.L.). D.L. is a member of Institut Universitaire de France.
Author information
Authors and Affiliations
Contributions
P.M. and M. Houssier performed the majority of in vitro experiments and analyzed data with the contribution of A. Mairal, C.G., F.B., B.M., E.R., P.D.D., V. Sramkova, V.B., D.B., M.M., C.L., L.L., F.L. and M. Harms. P.M., M. Houssier, E. Mouisel, G.T., S.V., L.M., S.G., B.M.-R., T.S., H.G., C.H., A.V.P. and C.P. performed and analyzed in vivo data from mouse models. P.M., S.B., M.M., B.F., A.A., E. Meugnier, C.L., R.R.L., W.S., V. Stich, P.A., M.R., N.V. and H.V. performed and analyzed in vivo data in human clinical studies. S.C.-B., S.V. and J.B.-M. analyzed lipidomics data. A. Mazars and M.Z. performed and analyzed FRAP experiments. B.P., C.M., N.V., S.H. and H.V. interpreted the data. P.M., M. Houssier and D.L. conceived the study, interpreted the data and wrote the manuscript. D.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
T.S. is an employee of Physiogenex. M. Harms and S.H. are employees of AstraZeneca. The other authors declare no competing financial and non-financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Methods
Rights and permissions
About this article
Cite this article
Morigny, P., Houssier, M., Mairal, A. et al. Interaction between hormone-sensitive lipase and ChREBP in fat cells controls insulin sensitivity. Nat Metab 1, 133–146 (2019). https://doi.org/10.1038/s42255-018-0007-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-018-0007-6