Abstract
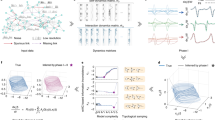
Epidemics, voting behaviour and cascading failures in power grids are examples of natural, social and technological phenomena that can be modelled as dynamical processes on networks. The study of such important complex systems requires approximation, but the assumptions that underpin the standard mean-field approaches are routinely violated by dynamics on real-world networks, leading to uncontrolled errors and even controversial results. Consequently, determining the approximation precision has been recognised as a key challenge. We present a micro-scale foundation for mean-field approximation of a wide range of dynamics on networks that facilitates quantification of approximation error, elucidating its connection to network structure and model dynamics. We show that our coarse-graining approach minimises approximation error and we obtain an upper bound on this uncertainty. We illustrate our approach using epidemic dynamics on real-world networks.
Similar content being viewed by others
Introduction
The study of natural, social and technological phenomena in complex systems invariably requires approximations that coarse-grain and simplify, so that insights can be obtained about the causal mechanisms at work. A case in point, and our focus, is the study of dynamical processes on complex networks1, such as models of epidemics2,3, opinion dynamics4,5,6, the diffusion of innovations7,8,9,10, the evolution of languages11,12,13 and cultural polarisation14,15. The standard approach to analyse dynamics on networks is via mean-field approximations, which range in accuracy and complexity2,16,17,18,19. While such methods have provided important insights, the assumptions that underpin mean-field approximations—the absence of clustering (‘a friend of a friend is my friend’), modularity (community structure) and dynamical correlations (‘I’m similar to my neighbours’)—are routinely violated by dynamical processes on real-world complex networks and it is generally difficult to quantify how well a particular approximation will do a priori, given the network or dynamical process20. Mean-field approximation has also resulted in controversy concerning the critical epidemic threshold in scale-free networks17,21,22,23. Because of these issues, the quantification of approximation error has been recognised as one of the key challenges for network epidemic modellers24.
In this article we address these critical issues by presenting a foundation for mean-field approximations of dynamics on networks, which builds from the micro-scale description of the dynamics and facilitates the quantification of approximation error. We use approximate lumping to derive low-dimensional mean-field equations for a broad class of Markov chain dynamics on networks which includes models of epidemics and opinion dynamics. The coarse-grained states are based on the number of each type of ‘vertex-state’, such as the number of susceptible and infected vertices in the susceptible–infected–susceptible (SIS) model of epidemics. In contrast to standard mean-field approximations, the transition rates between these coarse-grained states are derived directly from the exact evolution of the probability distribution over states—known as the master equation or forward Kolmogorov equation—and are shown to minimise approximation error, in the sense that they are closest to an exact lumping. This provides a theoretical underpinning that simplifies and standardises the process of deriving mean-field approximations for practitioners: the microscopic formulation of a model can be easily translated into a mean-field approximation using the formulae we have obtained. Furthermore, this approach enables us to derive a bound on the approximate lumping error and compare this to errors computed from stochastic simulation of epidemic dynamics on several benchmark real-world networks.
Results and discussion
We consider Markov chain dynamics on finite, connected networks with undirected, unweighted edges and no self-loops, where each vertex in the network can be in one of a finite number of “vertex-states”. For example, in models of epidemics the vertex-states correspond to individuals’ disease status, which could be susceptible to infection, infected, recovered, etc. In models of voting behaviour, the vertex-states correspond to the party that each person plans to vote for. If M is the number of vertex-states and N is the number of vertices, then there are MN possible states, i.e. configurations of vertex-states on the network. Thus the size of the full state-space for Markov chain dynamics on networks is extremely large, even for moderate N, and consequently, unless the network contains significant symmetry25,26, approximation is essential. Despite this, the state-space is finite so we denote the probability distribution at time t over state-space by \(X(t)={({X}_{1}(t),{X}_{2}(t),\ldots ,{X}_{{M}^{N}}(t))}^{{{{{{{{\rm{T}}}}}}}}}\), where Xk(t) is the probability of being in the kth state. Variables related to the full state-space will be upper-case Latin letters and the indices k and l indicate that the index is over the full state-space. In continuous time t, the evolution of X(t) is described by the forward Kolmogorov or master equation27,
where Q is the infinitesimal generator, an MN by MN matrix in which each off-diagonal component Qkl gives the transition rate from state S[k] to state S[l], and the diagonal components ensure that rows sum to zero. Bold variables indicate matrices. Our approach can also be adapted to discrete time models.
In the “Methods” section we describe how the components of the infinitesimal generator relate to the microscopic dynamics, i.e. the transition rates of individual vertices between vertex-states. We assume that the positive entries of the infinitesimal generator are affine (i.e. constant plus linear) functions of the number of neighbouring vertices in each vertex-state. For example, in epidemic models, a susceptible vertex typically becomes infected at a rate proportional to the number of infected neighbours. We also focus on ‘homogeneous’ models where the micro-scale transition rates are identical for all vertices with the same number of neighbours in each vertex-state. These features define a class of network dynamics that we call ‘homogeneous single-vertex transition models’ (homogeneous SVTs) with ‘affine vertex-state transition matrices’ (affine VSTMs). Specifically, if a model has an affine VSTM then a vertex in vertex-state \({{{{{{{\mathcal{A}}}}}}}}\), with nm neighbours in the mth vertex-state, transitions to vertex-state \({{{{{{{\mathcal{B}}}}}}}}\) with rate
where the \({\delta }_{m}^{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}\) are arbitrary non-negative constants. This covers a broad range of dynamical processes on networks28, but in Supplementary Note 5 we also consider generalisations to heterogeneous and nonlinear network dynamics with quadratic VSTMs.
We coarse-grain such Markov chain network dynamics using a method called approximate lumping, in which states are grouped together (lumped) according to a pre-defined partition of state-space29. We consider approximate lumping partitions based on sets of states that have the same total number of vertices in each vertex-state, i.e. the number of susceptible and infected vertices in the SIS model. We refer to this type of approximate lumping as a population model approximation30. To make this precise, let \(s\in {{\mathbb{Z}}}_{\ge 0}^{M}\) be a lumped state, which is a vector of length M whose mth component, sm, denotes the number of vertices in the mth vertex-state. Lumped variables will be lower-case Latin letters and m will index vertex-states. It follows that there are \(r=\left(\begin{array}{c}{N+M-1}\\ {N}\end{array}\right)\) possible lumped states, since a lumped state is a combination of N vertex-states drawn from M possibilities with repetition. Thus we number the lumped states in the lumped state-space s[1], s[2], …, s[r] and we use Π = {Π1, Π2, …, Πr} to denote the corresponding lumping partition. Let \(x(t)={({x}_{1}(t),\ldots ,{x}_{r}(t))}^{{\mathrm {T}}}\) denote the time-dependent Markov chain probability distribution over Π, where xi(t) is the probability of being in the lumped state s[i]. We use indices i and j to indicate that the index is over the lumped state-space. The evolution of x(t) is then the solution to
where q is the approximate lumping generator, which needs to be determined.
The idea here is to use the coarse-grained generator q = DQC, where \({{{{{{{\bf{C}}}}}}}}\in {\{0,1\}}^{{M}^{N}\times r}\) is the collector matrix29, whose kjth component is one if S[k] ∈ Πj and zero otherwise, and \({{{{{{{\bf{D}}}}}}}}\in {{\mathbb{R}}}^{r\times {M}^{N}}\) is the distributor matrix, whose ilth component is 1/∣Πi∣ if S[l] ∈ Πi and zero otherwise. The effect of using q = DQC is to average the sum of rates out of states in one lumping partition cell and into another. This approach has the following advantages. Firstly it minimises error, in the sense that it is closest to an exact lumping where QC = Cq (details in the Methods section), which is made precise in the following theorem.
Theorem 2.1
The lumped infinitesimal generator q = DQC minimises ∣∣QC − Cq∣∣F (Frobenius norm).
Secondly, the matrix q can be explicitly derived for affine network dynamics, leading to the following theorem.
Theorem 2.2
Let Ω be the state-space of a homogeneous SVT with affine VSTM on a network with mean degree z, and let q = DQC be the lumped infinitesimal generator corresponding to the population model approximation Π = {Π1, Π2, …, Πr} with lumped states s[1], s[2], …, s[r]. If s[i] and s[j] correspond to a single vertex changing from vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) to \({{{{{{{\mathcal{B}}}}}}}}\) and \({s}_{1}^{[i]}\) is the number of vertices in vertex-state \({{{{{{{\mathcal{A}}}}}}}}\), then
These are the main theoretical results of the paper. Outlines of the proofs are given in the “Methods” section and further details are provided in the Supplementary Methods.
For concreteness, we illustrate the approximate lumping approach in Fig. 1 using the SIS model of epidemic dynamics, which has M = 2 and is an example of “binary-state dynamics”31. The vertex-states of the SIS model are referred to as susceptible (\({{{{{{{\mathcal{S}}}}}}}}\)) and infected (\({{{{{{{\mathcal{I}}}}}}}}\)). A susceptible vertex with n1 infected neighbours becomes infected with rate βn1 and an infected vertex recovers with rate γ, where β, γ > 0 are model parameters. In relation to our notation for affine VSTMs introduced in Eq. (1), we have \({\delta }_{1}^{{{{{{{{\mathcal{S}}}}}}}},{{{{{{{\mathcal{I}}}}}}}}}=\beta\), \({\delta }_{0}^{{{{{{{{\mathcal{I}}}}}}}},{{{{{{{\mathcal{S}}}}}}}}}=\gamma\) and all other \({\delta }_{m}^{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}\) are zero. Our approach partitions state-space into “levels”, so that the ith level, Πi, contains all states that have i infected vertices, and this reduces the size of state-space from 2N to N + 1. For SIS dynamics, we obtain a mean-field birth-death process with infection rates given by
and recovery rates
These rates will be unsurprising to those familiar with mean-field approximations of network dynamics, but note that here we have derived these directly from the full Markov chain description rather than via moment closures based on non-rigorous probabilistic arguments, as is typical2. For the SIS model and other binary-state dynamics, this approach gives rise to a birth–death process; for network dynamics with M > 2, it yields a Markov population model30.
a Illustrates the matrix multiplication DQC = q that lumps the infinitesimal generator Q of the full Markov chain using the collector and distributor matrices, C and D, respectively, to produce the tridiagonal approximate lumping infinitesimal generator q. Colour indicates the value of the corresponding matrix entry for the infection rate β = 4 and recovery rate γ = 1; zero entries are white. The horizontal and vertical lines indicate the different groupings of states by level; level 0 is in the left/top and level 4 is on the right/bottom. b Illustrates transitions from a state with two infected vertices that are accounted for by the full Markov chain. Blue vertices indicate susceptible and red vertices indicate infected. The transition rates are given next to the corresponding arrows. The vertical dots indicate that there are more states with two infected vertices. c Illustrates the corresponding transition rates for the approximate lumping from level two, i.e. two infected vertices. In general the lumped recovery rate is γi and the lumped infection rate is βzi(N−i)/(N−1), where i is the level (number of infected individuals); for the case illustrated N = 4, z = 2 and i = 2. d Illustrates the average number of infected vertices from solutions to the master equation for the full Markov chain (exact) and the approximate lumping (approximate). Note the log scale on the horizontal time axis.
In the lumped state-space, the error of our approximation is y(t) = CTX(t)−x(t) and so
This is an inhomogeneous linear system of ODEs, thus applying the variation of constants formula yields
where we have assumed that y(0) = 0, i.e. the lumped initial state CTX(0) is known. To simplify the error computation we assume that the initial distribution of the full Markov chain is stationary so that X(t) = X*. Quasi-stationary distributions can also be handled in an analogous way and are discussed in Supplementary Note 4. In the “Methods” section, we derive a bound on the stationary absolute mean error
for binary-state dynamics. However, this involves terms that depend on the full Markov chain, so we must resort to approximations to make further progress.
We focus on the SISa model32, which is similar to the SIS model but has an additional ‘ambient’ infection rate α, so a susceptible vertex with n1 infected neighbours becomes infected with rate α + βn1. Recovery is the same as in the SIS model. Unlike the SIS model, where the state with all susceptible vertices is absorbing, the SISa model has a stationary distribution. In the “Methods” section we obtain a bound on the stationary absolute mean error of the SISa model that depends on \({a}_{i}^{+}\), which is a constant related to the state that has the largest or smallest number of edges between susceptible and infected vertices in the ith level. Unfortunately, computing \({a}_{i}^{+}\) is computationally difficult (an algorithm that did so would provide a solution to the Max-Cut problem, which is NP-complete33). Thus we settle for an estimate, \({\widetilde{a}}_{i}^{+} > \,0\), obtained from a tractable greedy algorithm, described in detail in the “Methods” section, that sequentially picks susceptible vertices to become infected which introduce the largest or smallest number of edges between susceptible and infected vertices. Our numerically tractable bound depends on an assumption about \({\widetilde{a}}_{i}^{+}{x}_{i}^{* }\) and the full system, which is made precise in the “Methods” section. In Supplementary Note 3 we show that while this assumption does not always hold, we typically obtain an informative bound regardless. We also propose an approximation \({a}_{i}^{* }{x}_{i}^{* }\) based on averaging the minimum and maximum number of edges between susceptible and infected vertices at each level, although this approximation does not have a rigorous foundation.
Application to real-world networks
To illustrate the application of our results on a topical example, we use the SIR model of epidemics on a real-world contact network derived from GPS data. There are three vertex-states in the SIR model, namely susceptible, infected and recovered, which we denote by \({{{{{{{\mathcal{S}}}}}}}}\), \({{{{{{{\mathcal{I}}}}}}}}\) and \({{{{{{{\mathcal{R}}}}}}}}\) respectively. A susceptible vertex with n1 infected neighbours becomes infected at a rate βn1, and an infected vertex recovers at a rate γ. There are 3N states in the full Markov chain and (N + 2)(N + 1)/2 lumped states, corresponding to distinct numbers of vertices in each of the vertex-states. The lumped transition rate qij from the ith lumped state with \({s}_{{{{{{{{\mathcal{S}}}}}}}}}^{[i]}\) susceptible vertices and \({s}_{{{{{{{{\mathcal{I}}}}}}}}}^{[i]}\) infected vertices, to the jth lumped state in which a susceptible vertex has become infected is
(Note that here it is convenient to use the vertex-states \({{{{{{{\mathcal{I}}}}}}}}\) and \({{{{{{{\mathcal{S}}}}}}}}\) rather than an integer to index the lumped state s[i]). Similarly, if an infected vertex recovers then the lumped transition rate is \({{{{{{{{\bf{q}}}}}}}}}_{ij}=\gamma {s}_{{{{{{{{\mathcal{I}}}}}}}}}^{[i]}.\) There are N + 1 lumped absorbing states in which there are no infected vertices and the number of recovered vertices ranges from zero to N.
We used a real-world contact network derived from data collected as part of the BBC documentary ‘Contagion! The BBC Four Pandemic’34,35. This study collected GPS traces of people who downloaded the ‘BBC Pandemic’ smart phone application. Data made publicly available from this study consists of timestamped anonymised pairwise distances within 50 m between 469 participants around the town of Haselmere, UK. We aggregated these data to create a static network between participants that came within 1 m of each other. We used the largest connected component of this network, which consists of N = 369 people and has mean degree z = 5.53. We refer to this as the ‘Haselmere 1m’ network. We used parameters γ = 1 and β = γR0(N−1)/(zN), where R0 = 3, since this would give a reproduction number of R0 in the corresponding compartmental model equations. Initially five vertices were selected uniformly at random to be infected.
In Fig. 2a, b we compare stochastic simulations (red) of the SIR model on the Haselmere 1m network with the corresponding approximate lumping (blue). Figure 2a illustrates the mean number of infected vertices over time (thick solid lines) and the corresponding 90-percentile of the simulated and approximate lumping distributions (shading). We also include, for comparison, results from homogeneous, heterogeneous and individual-based mean-field approximations (dashed, dot, and dash-dot lines respectively—see Supplementary Note 1 and Kiss et al. 2 for details), illustrating that the accuracy of our approach is comparable. However, our approach also produces a full probability distribution over the lumped states, which we use to compute the percentiles in Fig. 2a. This distribution could also be used for Bayesian parameter estimation and even data assimilation. Furthermore, with our approach we are able to compute absorption statistics and in Fig. 2b we compare the absorption probability into each absorbing state (i.e. the total number of infected individuals) of stochastic simulations (grey) and our approximate lumping (blue).
a Illustrates the evolution of the mean number of infected vertices from 3000 stochastic simulations (thick red line) and the approximate lumping (thick blue line) for the SIR model on the Haselmere 1m network. The red and blue shading illustrate the 90-percentile of the corresponding distributions. The light blue dash, yellow dot, and grey dash-dot lines indicate the mean number of infected vertices for homogeneous, heterogeneous and individual based mean-field approximations respectively. b Illustrates the probability distribution of the total number of infections computed from 100,000 stochastic simulations, each run until t = 1000 (grey shading). The corresponding probability distribution computed from the approximate lumping is illustrated in blue. c and d Illustrate the same as a and b, respectively, but for an Erdős–Rényi graph with N = 369 vertices and mean degree z = 20.
Low dimensional mean-field approximations can perform poorly on networks with heterogeneous structure (e.g. when hubs, clustering or communities are present), and Fig. 2a, b illustrate this. By way of contrast, we also present results for an Erdős–Rényi graph where the accuracy of mean-field approximations is better. Specifically, we chose a network uniformly at random from those with N = 369 vertices (the same size as the Haselmere 1m network) and mean degree z = 20 (i.e. selecting 3690 random edges—note this is the less common type of Erdős Rényi graph), and in Fig. 2c, d we illustrate results corresponding to those in Fig. 2a, b, respectively. In this case, the accuracy is significantly improved and our approach even appears marginally better than the other comparable mean-field theories illustrated. We obtain similar results if we average over many graphs sampled at random.
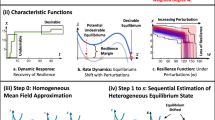
In Fig. 3 we compare our error bound with the error produced via stochastic simulations of the SISa model on four benchmark real-world networks, including the Haselmere 1m network34,35, a protein interaction network36,37,38, an autonomous-systems Internet network39 and a US power grid network40. For each network in Fig. 3, we compute stochastic simulations of SISa dynamics on the network with ambient infection rate α = 0.01, infection transmission rate β = 2(γ−α)(N−1)/(zN) and recovery rate γ = 1, which would give a stationary infected fraction of 0.5 in the corresponding SISa compartmental model. Half of the vertices are chosen uniformly at random to be initially infected and the number of infected vertices is computed after the process is approximately stationary. For each network, we compute the mean fraction of infected vertices from multiple realisations of the stochastic simulations. We also numerically compute solutions of the lumped system to find the lumped probability distribution x(t) with initial condition corresponding to the average number of infected vertices of the stationary stochastic simulations. The stochastic simulation error (solid black lines in Fig. 3) is the absolute magnitude of the difference between the mean fraction of infected vertices in the stochastic simulations and approximate lumping. We compare this with our bound on the approximate lumping error (red dashed lines in Fig. 3) by numerically integrating Eq. (5) using \({\widetilde{a}}_{i}^{+}{x}_{i}^{* }\). The long-term behaviour of the bound is comparable, i.e. the over-estimate is a similar amount, for different sizes of network and error. The results for these examples are representative of other real-world networks. To illustrate this, in Fig. 4 we compare the errors computed from stochastic simulations (horizontal axis) with the corresponding errors computed using our approximation and bound (vertical axis) for 18 real-world networks, including the four used in Fig. 3. These networks constitute a standard benchmark test-set, including networks with heterogeneous topology on which mean-field approximations vary in accuracy20. The circular and triangular markers correspond to the approximation and bound, respectively. The SISa parameter values used are the same as in Fig. 3, i.e. α = 0.01, β = 2(γ−α)(N−1)/(zN) and γ = 1. The legend indicates which network has been used and these are ordered from the smallest simulation error at the top (furthest left in the figure) to the largest at the bottom (furthest right in the figure). References for each network, as well as information about size and mean degree, are included in Supplementary Note 3. Figure 4 shows that for a range of benchmark real-world networks our approximation gives a good estimate of the magnitude of the mean error and our bound is informative, i.e. these are correlated with the error (Pearson correlation coefficient: 0.62, p-value < 0.01 [without karate: 0.86, p-value ≪ 0.01]) and in all cases give a value <1.
Comparison of the evolution of the mean-field approximation error y(t) over time t for the SISa model (solid black line), computed using stochastic simulations, with our theoretical bound (dashed red line) for four real-world networks. a Uses the Haselmere 1m network34,35, b uses a protein interaction network36,37,38, c uses an autonomous-systems Internet network39 and d uses a US power grid network40.
Conclusion
In summary, we have presented a mathematical foundation for mean-field approximations of a wide class of dynamical processes on networks that facilitates the quantification of approximation error. We have used approximate lumping to derive low-dimensional systems of equations directly from the exact master equation description, whose approximation error is minimal, in the sense that it is closest to an exact lumping, and can be quantified.
Our approximation results in a ‘density dependent’ system from which even lower dimensional ODE approximations can be rigorously derived in the large N limit41,42,43. Note that the lumped transition rates which we have derived only characterise network structure in terms of the mean degree, so do not account for variations in topology that may affect the dynamics. However, there is scope to extend our approach to more accurate degree-based mean-field17 and high-accuracy approximate master equations18,31 through more fined-grained lumpings by considering finer partitions of vertices and states30. There may also be alternative methods to bound the error44, potentially making use of theory developed for operator semi-groups43. While we extend our approach to quadratic VSTMs in Supplementary Note 5, further generalisations to arbitrary nonlinear VSTMs, e.g. via their power series expansions, may be possible. For non-smooth VSTMs, such as threshold models, consideration of the averaging process of the infinitesimal generator may also facilitate the derivation of approximations. The approach developed in this paper could also be applied to other complex systems, e.g. a natural generalisation is to multilayer network structures45,46 via the supra-adjacency matrix representation. However, the details of the specific application are likely to be crucial and will inevitably influence the structure of the Markov chain state-space and hence how much our approach needs to be adapted to deal with these considerations.
The COVID-19 pandemic has brought epidemic modelling into the spotlight and variants of compartmental models have influenced policy: for example, the UK’s Scientific Advisory Group for Emergencies47 at the time of writing list stochastic transmission models48,49,50,51 as modelling inputs. Such models incorporate realistic features such as age structure and geography. However, the underlying contact network is difficult to obtain and we should consider the consequence of not accounting for this in our models. For example, Fig. 2a shows that mean-field approximations (which includes compartmental models) are a poor representation of the true dynamics. Thus varying infection rates to fit such models to data could distort their interpretation and hence the consequences of policy interventions.
Methods
Mathematical formulation
Let G = (V, E) denote a network with vertex set V and edge set E ⊂ V × V, where the number of vertices is N = ∣V∣. We consider dynamical processes on finite connected simple networks (i.e. undirected, unweighted and with no self-loops) described by continuous-time Markov chains where each vertex can be in one of a finite number M of vertex-states and the set of possible vertex-states is \({{{{{{{\mathcal{W}}}}}}}}=\{{{{{{{{{\mathcal{W}}}}}}}}}_{1},{{{{{{{{\mathcal{W}}}}}}}}}_{2},\ldots ,{{{{{{{{\mathcal{W}}}}}}}}}_{M}\}\). We use caligraphic variables to indicate vertex-states. The state-space of the Markov chain is the set of all permutations of N vertex-states chosen from \({{{{{{{\mathcal{W}}}}}}}}\) with repetition. This is equivalent to \({{\Omega }}={{{{{{{{\mathcal{W}}}}}}}}}^{V}\), i.e. the set of all functions from V to \({{{{{{{\mathcal{W}}}}}}}}\), and so if the network is in state S ∈ Ω then the vertex-state of vertex v ∈ V is S(v). Since the number of states in Ω is MN, we can enumerate the states in state-space so that \({{\Omega }}=\{{S}^{[1]},{S}^{[2]},\ldots ,{S}^{[{M}^{N}]}\}\).
We assume that the dynamics are governed by homogeneous SVT models, which includes models of spin systems, epidemics, opinion dynamics, diffusion of innovation and a variety of other social dynamics28,52. In a homogeneous SVT model, a vertex changes vertex-state at a rate that is a function of only the number of its neighbours in each vertex-state and the rate function is the same for all vertices. Furthermore, transitions only occur between pairs of states that differ in at most one vertex-state. We call such pairs of states transition pairs and use the notation \({S}^{[k]}\mathop{ \sim }\limits^{v}{S}^{[l]}\) to indicate that the states S[k] and S[l] form a transition pair with transition vertex v, i.e. if \({S}^{[k]}\mathop{ \sim }\limits^{v}{S}^{[l]}\) then S[k](v) ≠ S[l](v) and S[k](u) = S[l](u) for all u ≠ v. For vertex v and state S[k] let \({n}^{[k]}(v)=({n}_{1}^{[k]}(v),{n}_{2}^{[k]}(v),\ldots ,{n}_{M}^{[k]}(v))\), where \({n}_{m}^{[k]}(v)\) is the number of neighbours of v with vertex-state \({{{{{{{{\mathcal{W}}}}}}}}}_{m}\). For k ≠ l, the transition rate between states S[k] and S[l] in homogeneous single-vertex transition models is then given by
where \({f}_{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}({n}_{1},{n}_{2},\ldots ,{n}_{M})\ge 0\) is the VSTM, i.e. the rate that a vertex in vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) changes to vertex-state \({{{{{{{\mathcal{B}}}}}}}}\) if it has n1 neighbours in vertex-state \({{{{{{{{\mathcal{W}}}}}}}}}_{1}\), n2 neighbours in vertex-state \({{{{{{{{\mathcal{W}}}}}}}}}_{2}\), etc. We focus on VSTMs that are affine functions of n[k](v), given by (1). Most SVTs have VSTMs of this form28, although notable exceptions include non-zero temperature Ising-Glauber dynamics53, the nonlinear q-voter model54 and threshold models10. Nonlinear VSTMs are discussed further in Supplementary Note 5, where we present results for the quadratic case.
Approximate lumping
To coarse-grain the network dynamics, we consider lumping of Markov chains55. An exact lumping Π = {Π1, Π2, …, Πr} is a partition of state-space that preserves the Markov property, a necessary and sufficient condition for which is that the sum of transition rates out of a state S[k] ∈ Πi into the cell Πj is the same for all states in the cell Πi. In matrix notation, this is equivalent to the existence of an r × r matrix q such that
where \({{{{{{{\bf{C}}}}}}}}\in {\{0,1\}}^{{M}^{N}\times r}\) is the collector matrix29 whose kjth component is
We call Eq. (7) the lumpability condition.
Note that q can be given explicitly by introducing the distributor matrix29\({{{{{{{\bf{D}}}}}}}}\in {{\mathbb{R}}}^{r\times {M}^{N}}\), whose ilth component is
Specifically, q = DQC satisfies the lumpability condition when Q commutes with CD28.
A lumping that does not satisfy the lumpability condition, and hence does not preserve the Markov property, is an approximate lumping29. Recall that we consider approximate lumping partitions based on sets of states that have the same number of vertices in each vertex-state and use the generator q = DQC even when the lumpability condition is violated. Motivated by the condition for an exact lumping (7), for a given matrix norm ∣∣ ⋅ ∣∣ we define the approximate lumping discrepancy as ∣∣QC−Cq∣∣. Note that QC−Cq is a matrix of size MN × r, which in the case of an exact lumping has all zero entries, thus the approximate lumping discrepancy measures how far (in terms of the specific norm used) the approximate lumping is from being an exact lumping. For this reason, we choose q to minimise the approximate lumping discrepancy.
We now give an outline of the proof of Theorem 2.1, i.e. that q = DQC minimises the approximate lumping discrepancy using the Frobenius norm. With the Frobenious norm ∣∣ ⋅ ∣∣F we have
Consequently \(\parallel {{{{{{{\bf{QC}}}}}}}}-{{{{{{{\bf{Cq}}}}}}}}{\parallel }_{{{{{{{{\rm{F}}}}}}}}}^{2}\) can be minimised by choosing qij to be the average of the sum of rates out of states in the ith level and into the jth level, i.e.
where \({\left({{N}\atop{{s}^{[i]}}}\right)}\) is short for the multinomial \({\left({{N}\atop{{s}_{1}^{[i]},{s}_{2}^{[i]},\ldots ,{s}_{M}^{[i]}}}\right)}\). This is exactly what is obtained if one uses the definitions of the collector and distributor matrices to compute (DQC)ij. A detailed proof of Theorem 2.1 is provided in the Supplementary Methods. Note that the q that minimises the approximate lumping discrepancy depends on the particular norm used; the Frobenius norm is advantageous because it results in an intuitive averaging process that is also analytically tractable.
For \({{{{{{{\mathcal{A}}}}}}}}\in {{{{{{{\mathcal{W}}}}}}}}\), let \({\nu }_{{{{{{{{\mathcal{A}}}}}}}}}\) be a vector of length M whose mth component is \({\nu }_{{{{{{{{\mathcal{A}}}}}}}}m}=0\) if \({{{{{{{\mathcal{A}}}}}}}}\ne {{{{{{{{\mathcal{W}}}}}}}}}_{m}\) and \({\nu }_{{{{{{{{\mathcal{A}}}}}}}}m}=1\) if \({{{{{{{\mathcal{A}}}}}}}}={{{{{{{{\mathcal{W}}}}}}}}}_{m}\). Then for SVT models, the only possible non-zero rates are between pairs of lumped states that satisfy \({s}^{[j]}={s}^{[i]}+{\nu }_{{{{{{{{\mathcal{B}}}}}}}}}-{\nu }_{{{{{{{{\mathcal{A}}}}}}}}}\), with \({{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}\in {{{{{{{\mathcal{W}}}}}}}}\) and \({{{{{{{\mathcal{A}}}}}}}}\ne {{{{{{{\mathcal{B}}}}}}}}\), i.e. a vertex switches from vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) to \({{{{{{{\mathcal{B}}}}}}}}\). It follows that the lumped states can also be ordered so that q is a quasi-birth–death process and hence q is tridiagonal by blocks.
We now give an outline of the proof of Theorem 2.2 by illustrating how we derive the elements of q from the full Markov chain description. Consider the case where qij corresponds to a vertex changing from vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) to \({{{{{{{\mathcal{B}}}}}}}}\), so \({s}^{[j]}={s}^{[i]}+{\nu }_{{{{{{{{\mathcal{B}}}}}}}}}-{\nu }_{{{{{{{{\mathcal{A}}}}}}}}}\). In Eq. (8), for each state S[k] ∈ Πi we sum the rates into Πj to get (QC)kj. As assumed, these non-zero rates are associated with vertices in vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) changing to \({{{{{{{\mathcal{B}}}}}}}}\). Thus we can go through each vertex in S[k] that is in vertex-state \({{{{{{{\mathcal{A}}}}}}}}\), count the number of its neighbours that are in each of the vertex-states to compute the transition rate (1), and sum these up. Equation (8) then averages these over all states in Πi. Our key insight is that rather than summing over states as Eq. (8) suggests, we can achieve the same total by summing over vertices and the possible states of neighbours.
For a vertex v with degree dv, the number of states in Πi where vertex v is in vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) and has n = (n1, n2, …, nM) neighbours in each of the vertex-states is
where we have used our generalised multinomial notation, indicated by the presence of vectors in the denominators, e.g. \({\left({{{d}_{v}}\atop{n}}\right)}={\left({{{d}_{v}}\atop{{n}_{1},{n}_{2},\ldots ,{n}_{m}}}\right)}\). The transition rate of a vertex from vertex-state \({{{{{{{\mathcal{A}}}}}}}}\) to \({{{{{{{\mathcal{B}}}}}}}}\) is given by Eq. (1). To compute qij we sum these rates over all N vertices and all possible values of n, and divide by the number of states to get
where the sum over n∣dv denotes a sum over all possible values of n such that n1 + n2 + ⋯ + nm = dv.
We deal with the \({\delta }_{0}^{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}\) and \({\delta }_{m}^{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}{n}_{m}\) terms separately. Using a generalisation of the Vandermonde indentity (see the Supplementary Methods for details), the sum with the constant term \({\delta }_{0}^{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}\) is
where we have assumed, without loss of generality, that the first index of the lumped state, \({s}_{1}^{[i]}\), corresponds to the vertex-state \({{{{{{{\mathcal{A}}}}}}}}\). For the \({\delta }_{m}^{{{{{{{{\mathcal{A}}}}}}}},{{{{{{{\mathcal{B}}}}}}}}}{n}_{m}\) terms, again using the generalised Vandermonde identity, we have
Substituting Eqs. (10) and (11) into Eq. (9), after some cancellation, yields Eq. (3). A detailed proof of Theorem 2.2 is included in the Supplementary Methods.
Error analysis of binary-state dynamics with a stationary distribution
We now focus on binary-state dynamics where there are two vertex-states, hence M = 2. Examples of binary-state dynamics include the SIS and voter models28 and in Supplementary Note 2 we provide a classification of the different types of binary-state dynamics. Consequently, we suppose that the set of vertex states is \({{{{{{{\mathcal{W}}}}}}}}=\{{{{{{{{\mathcal{S}}}}}}}},{{{{{{{\mathcal{I}}}}}}}}\}\) and refer to vertex-state \({{{{{{{\mathcal{S}}}}}}}}\) as ‘susceptible’ and vertex-state \({{{{{{{\mathcal{I}}}}}}}}\) as ‘infected’; an infection corresponds to a susceptible vertex becoming infected and a recovery corresponds to an infected vertex becoming susceptible. We can partition the state-space of binary-state dynamics into levels so that the ith level, Πi contains all states that have i infected vertices, for i = 0, 1, …, N, i.e. Π = {Π0, Π1, …, ΠN}. It follows that the approximate lumping generator q is tridiagonal and QC−Cq is tridiagonal by blocks of column vectors of varying size. For 0 ≤ i < N, the column vectors of QC−Cq just above the diagonal correspond to infections and we denote these by
Thus \({A}_{{{{\Pi }}}_{i}}\) captures the difference between the sum of infection rates out of states in level i into level i + 1, and the mean qi,i+1. Note that we use the subscript Πi to illustrate that the variable is a vector over the states in Πi. Similarly, for 0 < i ≤ N, the column vectors of QC−Cq just below the diagonal correspond to recoveries and we denote these by
so \({B}_{{{{\Pi }}}_{i}}\) captures the differences between the recovery rates out of level i into level i−1, and the mean. We then have
where the zero entries indicate appropriately sized vectors of zeroes.
To simplify the error computation we assume that the initial distribution of the full Markov chain is stationary so that X(t) = X*, whose kth component is \({X}_{k}^{* }\). We also use \({X}_{{{{\Pi }}}_{i}}^{* {{{{{{{\rm{T}}}}}}}}}={({X}_{k}^{* })}_{{S}^{[k]}\in {{{\Pi }}}_{i}}\) to denote the vector of stationary probabilities of states in Πi. Hence we find that
where
The σi contain information about the full system and therefore cannot be directly computed for typical systems of interest, i.e. when the size of the full state-space is beyond what can be stored in computer memory.
We now consider the equilibrium solutions of Eqs. (2) and (4) in turn. For binary-state dynamics, our lumped approximation is a birth–death process, where a birth corresponds to an infection and a death corresponds to a recovery. Thus we can write
where the rates λi and μi are finite and positive. The analytical expression for the stationary distribution \({x}^{* }={({x}_{0}^{* },{x}_{1}^{* },\ldots ,{x}_{N}^{* })}^{{{{{{{{\rm{T}}}}}}}}}\) of such a birth–death process can be found in standard texts, e.g. Kijima27, but we reproduce it here in order to introduce notation that we will use when we derive the equilibrium of the error ODEs (4). The stationary distribution x* solves the recursion relation
which has solution
where ϕ0 = 1, for i > 0
and
Similar to the lumped dynamics, the equilibrium of the error ODEs (4), \({y}^{* }={({y}_{0}^{* },{y}_{1}^{* },\ldots ,{y}_{N}^{* })}^{{{{{{{{\rm{T}}}}}}}}}\), satisfies the system of equations
where 0 < i < N. It follows that the solution solves the recursion
Since both X* and x* are probability distributions, their elements sum to one and thus the sum of \({y}_{i}^{* }\) is zero. Consequently for i > 0 we find
where ψ0 = 0, for i > 0
and
By substituting Eq. (13) into the definition of the mean error \({\bar{y}}^{* }\), given by Eq. (6), we find
where
and \({\bar{x}}^{* }={\sum }_{i = 0}i{x}_{i}^{* }\) is the stationary mean number of infected vertices. Thus we have split the calculation of \({\bar{y}}^{* }\) into terms σi, which depend on the full Markov chain (and hence must be approximated), and terms ρi, which depend on the lumped system (and hence can be computed). Moreover, using the definition of \({\bar{x}}^{* }\) and Φ, it is straightforward to prove that ρi > 0 for all i, which suggests an intuitive bound on the absolute value of the stationary mean error given by
Example: error approximation for the SISa model
We now consider results for the SISa model32, where the VSTM has infection rate \({f}_{{{{{{{{\mathcal{S}}}}}}}},{{{{{{{\mathcal{I}}}}}}}}}({n}_{1},{n}_{2})=\alpha +\beta {n}_{1}\), recovery rate \({f}_{{{{{{{{\mathcal{I}}}}}}}},{{{{{{{\mathcal{S}}}}}}}}}({n}_{1},{n}_{2})=\gamma\) and \({f}_{{{{{{{{\mathcal{S}}}}}}}},{{{{{{{\mathcal{S}}}}}}}}}={f}_{{{{{{{{\mathcal{I}}}}}}}},{{{{{{{\mathcal{I}}}}}}}}}=0\). We derive bounds on the ∣σi∣ terms for the SISa model, which with Eq. (15) allow us to bound \(| {\bar{y}}^{* }|\). We also consider approximations of the σi terms, which with Eq. (14) allow us to approximate \({\bar{y}}^{* }\). Using Eq. (8), for the SISa model we find for S[k] ∈ Πi that
where
Note that \({n}_{{{\mbox{SI}}}}^{[k]}\) is the number edges that connect susceptible vertices with infected vertices (hereon referred to as SI edges) in the state S[k]. Our formula for \({\left({{{{{{{\bf{QC}}}}}}}}\right)}_{k,i+1}\) above for the SISa model follows from the fact that there are N−i susceptible vertices, and summing how many infected neighbours each has is equivalent to counting the number of SI edges. It follows that
so the entry in \({A}_{{{{\Pi }}}_{i}}\) corresponding to the state S[k] is proportional to the difference between the number of SI edges in state S[k] and the average of the number of SI edges in states in the ith level. A similar calculation shows that (QC)k,i−1 = γi and hence \({B}_{{{{\Pi }}}_{i}}=0\) for all i, i.e. the total recovery rate of a state in the SISa model is the same for all states in the same level. Thus for the SISa model \({\sigma }_{i}={A}_{{{{\Pi }}}_{i}}^{{{{{{{{\rm{T}}}}}}}}}{X}_{{{{\Pi }}}_{i}}^{* }\), hence if \({a}_{i}^{+}=\mathop{\max }\nolimits_{{S}^{[k]}\in {{{\Pi }}}_{i}}| {A}_{{{{\Pi }}}_{i}}|\) then
Determining \({a}_{i}^{+}\) and the sum of probabilities in the ith level would allow us to bound the absolute value of the mean error, but this may be intractable in practice because it requires knowledge of the full Markov chain. Thus to obtain a bound on the stationary absolute mean error of the SISa model, we use an approximation for \({a}_{i}^{+}\), denoted by \({\widetilde{a}}_{i}^{+}\), and then assume that \({\widetilde{a}}_{i}^{+}{x}_{i}^{* }\ge | {A}_{i}^{{{{{{{{\rm{T}}}}}}}}}{X}_{{{{\Pi }}}_{i}}^{* }|\). In Supplementary Note 3 we show that while this assumption does not always hold, we typically obtain an informative bound regardless.
We now describe how we obtain \({\widetilde{a}}_{i}^{+}\). Note that \({a}_{i}^{+}\) arises from the state in level i with either the largest or smallest number of SI edges. We refer to these states as the max and min SI states respectively. Finding the max SI states is equivalent to the Max-Cut problem, which is NP complete33. Finding the min SI states is also difficult because one needs to identify maximal cliques, which is also NP complete56. Because of this, we settle instead for estimates based on a greedy algorithm that starts from the state with all susceptible vertices and sequentially chooses a susceptible vertex to become infected that introduces the largest or smallest number of SI edges.
The algorithm is as follows. For binary-state dynamics in which vertices are either susceptible or infected, we iterate from level 0 to \(\lfloor \frac{N}{2}\rfloor\), picking a new vertex at each level to switch from susceptible to infected. There is only one state in level 0, in which all vertices are susceptible, so this is the state identified by the algorithm at the 0th level. Suppose that at the ith level the state S[k] is identified by the algorithm, then for each susceptible vertex v in S[k], we compute the number of infected neighbours \({n}_{1}^{[k]}(v)\) and the number of susceptible neighbours \({n}_{2}^{[k]}(v)\). We then pick the vertex with the largest difference \({n}_{1}^{[k]}(v)-{n}_{2}^{[k]}(v)\) (which may be negative) to be infected, and this is the state that the algorithm identifies for the i + 1th level. If there are multiple such vertices then we pick the one with the lowest index. This last step ensures our algorithm is deterministic, although to destroy possible correlations between vertex degrees and their labels, it may be necessary initially to randomise the vertex labelling. In binary-state dynamics there is a symmetry about \(\lfloor \frac{N}{2}\rfloor\), by switching susceptible vertices to infected and infected to susceptible, which preserves the number of SI edges. We apply this symmetry to the states selected so far to determine the states in levels above \(\lfloor \frac{N}{2}\rfloor\). Clearly one could perform a more extensive search, but our goal is to have an algorithm that scales well with the number of vertices. A nearly identical process can be used to identify a state in each level with a low number of SI edges by selecting the vertex with the smallest difference \({n}_{1}^{[k]}(v)-{n}_{2}^{[k]}(v)\) to become infected.
For level i, we use \({\widetilde{n}}_{i}^{+}\) and \({\widetilde{n}}_{i}^{-}\) to denote the maximum and minimum number of SI edges found by this algorithm, respectively. We also attempt to approximate σi with \({a}_{i}^{* }{x}_{i}^{* }\), where
This gives a measure of the skew of the distribution of the number of SI edges in each state in the same level.
Data availability
The networks and derived data are available in the Research Data Leeds Repository57.
Code availability
The code used to produce the derived data and figures is available in the Research Data Leeds Repository57.
References
Newman, M. Networks (Oxford University Press, 2018).
Kiss, I. Z., Miller, J. C. & Simon, P. L. Mathematics of Epidemics on Networks (Springer, 2017).
Pastor-Satorras, R., Castellano, C., Van Mieghem, P. & Vespignani, A. Epidemic processes in complex networks. Rev. Mod. Phys. 87, 925 (2015).
Galam, S. Minority opinion spreading in random geometry. Eur. Phys. J. B 25, 403 (2002).
Sood, V. & Redner, S. Voter model on heterogeneous graphs. Phys. Rev. Lett. 94, 178701 (2005).
Sznajd-Weron, K. & Sznajd, J. Opinion evolution in closed community. Int. J. Mod. Phys. C 11, 1157 (2000).
Bass, F. M. A new product growth for model consumer durables. Manag. Sci. 15, 215 (1969).
Mellor, A., Mobilia, M., Redner, S., Rucklidge, A. M. & Ward, J. A. Influence of Luddism on innovation diffusion. Phys. Rev. E 92, 012806 (2015).
Melnik, S., Ward, J. A., Gleeson, J. P. & Porter, M. A. Multi-stage complex contagions. Chaos 23, 013124 (2013).
D. J., W. A simple model of global cascades on random networks. Proc. Natl Acad. Sci. USA 99, 5766 (2002).
Baronchelli, A., Felici, M., Loreto, V., Caglioti, E. & Steels, L. Sharp transition towards shared vocabularies in multi-agent systems. J. Stat. Mech. 2006, P06014 (2006).
Bonabeau, E., Theraulaz, G. & Deneubourg, J.-L. Phase diagram of a model of self-organizing hierarchies. Physica A 217, 373 (1995).
Castelló, X., Eguíluz, V. M. & San Miguel, M. Ordering dynamics with two non-excluding options: bilingualism in language competition. New J. Phys. 8, 308 (2006).
Axelrod, R. The dissemination of culture: a model with local convergence and global polarization. J. Confl. Resolut. 41, 203 (1997).
Castellano, C., Marsili, M. & Vespignani, A. Nonequilibrium phase transition in a model for social influence. Phys. Rev. Lett. 85, 3536 (2000).
Vazquez, F. & Eguíluz, V. M. Analytical solution of the voter model on uncorrelated networks. N. J. Phys. 10, 063011 (2008).
Pastor-Satorras, R. & Vespignani, A. Epidemic spreading in scale-free networks. Phys. Rev. Lett. 86, 3200 (2001).
Gleeson, J. P. High-accuracy approximation of binary-state dynamics on networks. Phys. Rev. Lett. 107, 68701 (2011).
Fennell, P. G. & Gleeson, J. P. Multistate dynamical processes on networks: analysis through degree-based approximation frameworks. SIAM Rev. 61, 92 (2019).
Gleeson, J. P., Melnik, S., Ward, J. A., Porter, M. A. & Mucha, P. J. Accuracy of mean-field theory for dynamics on real-world networks. Phys. Rev. E 85, 026106 (2012).
Gomez-Gardenes, J., Latora, V., Moreno, Y. & Profumo, E. Spreading of sexually transmitted diseases in heterosexual populations. Proc. Natl Acad. Sci. USA 105, 1399 (2008).
Chatterjee, S. & Durrett, R. Contact processes on random graphs with power law degree distributions have critical value 0. Ann. Probab. 37, 2332 (2009).
Boguná, M., Castellano, C. & Pastor-Satorras, R. Nature of the epidemic threshold for the susceptible–infected–susceptible dynamics in networks. Phys. Rev. Lett. 111, 068701 (2013).
Pellis, L. et al. Eight challenges for network epidemic models. Epidemics 10, 58 (2015).
Sánchez-García, R. J. Exploiting symmetry in network analysis. Commun. Phys. 3, 1 (2020).
Ward, J. A. Dimension-reduction of dynamics on real-world networks with symmetry. Proc. R. Soc. A 477, 20210026 (2021).
Kijima, M. Markov Processes for Stochastic Modeling, Vol. 6 (CRC Press, 1997).
Ward, J. A. & López-García, M. Exact analysis of summary statistics for continuous-time discrete-state Markov processes on networks using graph-automorphism lumping. Appl. Netw. Sci. 4, 108 (2019).
Buchholz, P. Exact and ordinary lumpability in finite Markov chains. J. Appl. Probab. 31, 59 (1994).
Großmann, G. & Bortolussi, L. Reducing spreading processes on networks to Markov population models. In International Conference on Quantitative Evaluation of Systems (eds Parker, D. & Wolf, V.) 292–309 (Springer, 2019).
Gleeson, J. P. Binary-state dynamics on complex networks: Pair approximation and beyond. Phys. Rev. X 3, 021004 (2013).
Hill, A. L., Rand, D. G., Nowak, M. A. & Christakis, N. A. Infectious disease modeling of social contagion in networks. PLoS Comput. Biol. 6, e1000968 (2010).
Garey, M. R. & Johnson, D. S. Computers and Intractability (Freeman, 1979).
Klepac, P., Kissler, S. & Gog, J. Contagion! the BBC Four pandemic–the model behind the documentary. Epidemics 24, 49 (2018).
Kissler, S. M., Klepac, P., Tang, M., Conlan, A. J. & Gog, J. R. Sparking “The BBC Four Pandemic”: leveraging citizen science and mobile phones to model the spread of disease. Preprint at bioRxiv https://doi.org/10.1101/479154 (2020).
Colizza, V., Flammini, A., Maritan, A. & Vespignani, A. Characterization and modeling of protein–protein interaction networks. Physica A 352, 1 (2005).
Colizza, V., Flammini, A., Serrano, M. A. & Vespignani, A. Detecting rich-club ordering in complex networks. Nat. Phys. 2, 110 (2006).
Network Data DIP [Network Data]. Protein interaction network of the yeast Saccharomyces cerevisae extracted with different experimental techniques and collected at the Database of Interacting Proteins (accessed Nov 2020) (http://dip.doe-mbi.ucla.edu/); https://sites.google.com/site/cxnets/research22.
Network Data CAIDA AS Internet [Network Data]. The CAIDA Autonomous System Relationships Dataset (accessed Jun 2008) http://www.caida.org/data/active/as-relationships; https://www.caida.org/data/request_user_info_forms/as_relationships.xml.
Network Data Power grid network [Network Data]. An undirected, unweighted network representing the topology of the Western States Power Grid of the United States (accessed Nov 2020); http://www-personal.umich.edu/m̃ejn/netdata/power.zip.
Kurtz, T. G. Solutions of ordinary differential equations as limits of pure jump markov processes. J. Appl. Probab. 7, 49 (1970).
Kurtz, T. Limit theorems for sequences of jump markov processes. J. Appl. Probab. 8, 344 (1971).
Ethier. S. N. & Kurtz, T. G. Markov Processes: Characterization and Convergence, Vol. 282 (John Wiley & Sons, 2009).
Hoffmann, K. H. & Salamon, P. Bounding the lumping error in markov chain dynamics. Appl. Math. Lett. 22, 1471 (2009).
Boccaletti, S. et al. The structure and dynamics of multilayer networks. Phys. Rep. 544, 1 (2014).
Kivelä, M. et al. Multilayer networks. J. Complex Netw. 2, 203 (2014).
Scientific Advisory Group for Emergencies. Scientific evidence supporting the government response to coronavirus (covid-19) (accessed May 2021); https://www.gov.uk/government/collections/scientific-evidence-supporting-the-government-response-to-coronavirus-covid-19.
Danon, L., Brooks-Pollock, E., Bailey, M. & Keeling, M. A spatial model of COVID-19 transmission in England and Wales: early spread, peak timing and the impact of seasonality. Phil. Trans. R. Soc. B 376, 20200272 (2021).
Danon, L., House, T. & Keeling, M. J. The role of routine versus random movements on the spread of disease in Great Britain. Epidemics 1, 250 (2009).
Kucharski, A. J. et al. Early dynamics of transmission and control of COVID-19: a mathematical modelling study. Lancet Infect. Dis. 20, 553 (2020).
Dureau, J., Kalogeropoulos, K. & Baguelin, M. Capturing the time-varying drivers of an epidemic using stochastic dynamical systems. Biostatistics 14, 541 (2013).
Ward, J. A. & Evans, J. A general model of dynamics on networks with graph automorphism lumping. In International Conference on Complex Networks and their Applications (eds Aiello, L. et al.) 445–456 (Springer, 2018).
Glauber, R. J. Time-dependent statistics of the Ising model. J. Math. Phys. 4, 294 (1963).
Castellano, C., Muñoz, M. A. & Pastor-Satorras, R. Nonlinear q-voter model. Phys. Rev. E 80, 041129 (2009).
Kemeny, J. G. & Snell, J. L. Finite Markov Chains (Springer-Verlag, 1960).
Karp, R. M. Reducibility among combinatorial problems. In Complexity of Computer Computations (eds Miller, R. E., Thatcher, J. W. & Bohlinger, J. D.) 85–103 (Springer, 1972).
Ward, J. A. Benchmark testing networks and figure files [Dataset] (Research Data Leeds Repository); https://doi.org/10.5518/1076 (2022).
Acknowledgements
A.T. acknowledges support from the Engineering and Physical Sciences Research council, and Jaywing Intelligence. P.L.S. acknowledges support from the Hungarian Scientific Research Fund, OTKA (grant no. 135241) and from the Ministry of Innovation and Technology NRDI Office within the framework of the Artificial Intelligence National Laboratory Programme. R.P.M. acknowledges support from the UK Research and Innovation Future Leaders Fellowship grant no. MR/S032525/1.
Author information
Authors and Affiliations
Contributions
J.A.W. and P.L.S. conceived the research. All authors (J.A.W., A.T., P.L.S., R.P.M.) undertook the research. J.A.W. carried out the numerical simulations and produced the figures. All authors contributed to discussing the results and writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Physics thanks Dong-Chao Guo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ward, J.A., Tapper, A., Simon, P.L. et al. Micro-scale foundation with error quantification for the approximation of dynamics on networks. Commun Phys 5, 71 (2022). https://doi.org/10.1038/s42005-022-00834-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-022-00834-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.