Abstract
The predominant view in structure-based drug design is that small-molecule ligands, once bound to their target structures, display a well-defined binding mode. However, structural stability (robustness) is not necessary for thermodynamic stability (binding affinity). In fact, it entails an entropic penalty that counters complex formation. Surprisingly, little is known about the causes, consequences and real degree of robustness of protein-ligand complexes. Since hydrogen bonds have been described as essential for structural stability, here we investigate 469 such interactions across two diverse structure sets, comprising of 79 drug-like and 27 fragment ligands, respectively. Completely constricted protein-ligand complexes are rare and may fulfill a functional role. Most complexes balance order and disorder by combining a single anchoring point with looser regions. 25% do not contain any robust hydrogen bond and may form loose structures. Structural stability analysis reveals a hidden layer of complexity in protein-ligand complexes that should be considered in ligand design.
Similar content being viewed by others
Introduction
Biomolecular systems present a large number of degrees of freedom and must find a suitable balance between order and disorder. In the particular case of non-covalent complexes, they can exist in a continuum spectrum of possibilities, ranging from the lock-and-key model to extreme disorder1,2. While the importance of target flexibility is well-appreciated in drug discovery3, the flexibility of small-molecule ligands in their bound state has attracted much less attention. Detailed analyses reveal that ligands often retain residual mobility4,5,6. However, changes in binding mode are more the exception than the norm7,8 and ligand design based on rigid crystallographic geometries has been remarkably successful9. Explicit consideration of multiple binding modes is acknowledged as important for computational studies10, but invariably leads to more complex formalisms11,12. Perhaps for these reasons, little is known about the molecular mechanisms that control structural stability, to what extent do ligands preserve flexibility or what are the energetic and functional consequences of rigidity.
It is important to note that structural stability (robustness) is fundamentally different from thermodynamic stability (i.e., binding free energy; ∆Gbind). This is eloquently exemplified in the recent work by Borgia et al., where a protein-protein complex with picomolar affinity is shown to lack structure2. While ∆Gbind has been the center of attention of scientific research for decades, little attention has been paid to the factors that determine if a complex will be tight or loose. The source of structural robustness must be sought on sharp (and possibly transitory) energetic barriers that keep the atoms in their positions of equilibrium. Such hypothetical barriers, like the ones that determine binding kinetics, could have their origin in intramolecular (i.e., conformational rearrangement), bimolecular (e.g., repulsive transitional configurations) or many-body effects (e.g., desolvation)13. But they will only provide structural stability if the barriers are steep and located very close to the position of minimum energy. In that respect, hydrogen bonds (HBs) are ideal candidates because they have strict distance and angular dependencies14 and are one of the most frequent interaction types in protein-ligand complexes15. The contribution of HBs to ∆Gbind has been largely debated in the literature16,17,18,19,20. The current consensus is that it is highly variable and context dependent, but their contribution to thermodynamic stability is 1.8 kcal mol−1 at the most17. However, due to desolvation, the transitional penalty of breaking a HB can be much larger21. Indeed, we have shown that this is the case for water-shielded HBs, which can even act as kinetic traps22. More recently, we have also shown that formation of structurally robust intermolecular HBs at specific positions is a necessary condition for binding, and have developed a method to assess the robustness of individual HBs that is very effective in virtual screening applications23.
Here, we perform a systematic investigation of the possible role of HBs as structural anchors of protein-ligand complexes. We find that most complexes combine a robust anchoring point with more labile interactions, but cases of completely constricted and very loose complexes also exist. Our findings not only confirm a general role of HBs as source of structural stability, but also offer a new perspective to understand and design ligand-receptor complexes.
Results
Robust hydrogen bonds are common in protein-ligand complexes
Using Dynamic Undocking (DUck), an MD-based computational procedure23, we have assessed the robustness of every HB in a set of 79 drug-like protein-ligand complexes from the Iridium Data Set24. Detailed information about the data set and the selection criteria is presented in Supplementary Methods and Supplementary Table 1. Each HB was pulled to a distance of 5 Å, according to the DUck protocol reported previously23,25. In this way, we obtain a work value (WQB) that reflects the cost of breaking each HB. In other words, the WQB value indicates if the interaction under investigation gives rise to a narrow (local) minimum in the free-energy landscape, and estimates its depth. Based on our previous research, we define HBs as robust (i.e., capable of providing structural stability) if WQB > 6 kcal mol−1, labile if WQB < 4 kcal mol−1 and medium otherwise.
The distribution of work values for the entire set of 345 HBs ranges from 0 to 26 kcal mol−1, with a of maximum probability in the 0–6 kcal mol−1 region and a gradual decrease thereafter (Fig. 1a). Noteworthy, more than half HBs (57.4%) are robust. In order to provide a critical assessment of these results, we have sought correlation with experimental observables and have also considered if WQB values might be dominated by the interaction energies. Larger WQB values imply a narrower minimum and, thus, restricted mobility, which should translate into a more localized electron density, that is, lower crystallographic B-factors. As B-factors are heavily influenced by the refinement methods used and their absolute values can be meaningless26,27, we have normalized the B-factor of the ligand atom that makes the hydrogen bond relative to the average B-factor of the whole ligand. Encouragingly, atoms forming HBs with larger WQB values tend to have lower relative B-factors (Supplementary Fig. 4). A second aspect to consider is whether DUck calculations merely reflects short-range protein-ligand interaction, or—as intended—it captures a global effect that considers enthalpic and entropic contributions from both the solute and the solvent. Lack of correlation between interaction energies and WQB confirms that the latter is true (Supplementary Fig. 5). Of particular interest is to assess the effect of charge reinforcement on HBs, as the energetic, entropic and solvation terms of neutral hydrogen bonds and salt bridges are drastically different28. We have classified all HBs into neutral, mixed (ionic-neutral) and salt bridges (Fig. 2, Supplementary Data 2). We find that salt bridges are only very slightly skewed towards more robust interactions than neutral HBs. The distributions were compared with two sample Kolmogorov-Smirnov statistical test, yielding p-value of 0.08. Mixed types are completely indistinguishable from neutral ones (p-value = 0.42). Unexpectedly, the maximal values are equal across all three categories. Theoretically, ionic species could provide even larger energetic barriers because their desolvation costs are much larger. We speculate that there may be no biological use for them, as the maximal WQB values observed here already ensure very robust and long-lived structures.
Frequency of robust HBs in protein-ligand complexes. Histograms of frequency of HBs by WQB value for: a all simulated HBs (345), c HBs in enzyme active sites (253), e HBs in the ligand binding site of nuclear receptors (27), g HBs in carbohydrate binding sites (95), i HBs in allosteric sites (25). Pie charts showing share of complexes with at least one robust HB (WQB > 6 kcal mol−1, pink), all labile HBs (WQB < 4 kcal mol−1, green) or intermediate situations (red) for: b all simulated complexes (79), d enzymes (56), f nuclear receptors (7), h carbohydrate binding site (14), j allosteric sites (8)
The distribution of robust HBs is rather inhomogeneous across complexes, as they have 2.5 on average, but a quarter of the complexes have none (Fig. 1b). Considering that structural stability is not a requisite for tight binding and that HBs may not the only mechanism capable of providing structural stability, it is striking that 75% of the complexes in this set are anchored through HBs. A further 14% of complexes present medium values and only in 9 cases (11%) all their HBs are labile (Supplementary Fig. 6). Two of those cases are very low affinity complexes. In the remaining cases, structural stability might be provided by other mechanisms or may be lacking (see examples in Supplementary Fig. 6). It is important to note that the level of structural stability reported here may be overestimated due to the composition of the data set, entirely derived from X-ray crystallography, a technique that relies on order to solve structures.
Splitting this analysis by the type of binding site (Fig. 1c–j, Supplementary Table 2) provides strong indication that the behavior is dictated by the nature of the receptor. The proportion of robust complexes increases to 82% in the case of enzyme active sites, which speaks about the need of keeping the substrate in place for efficient catalysis. Nuclear receptors form fewer HBs with their ligands, but most of them (78%) are robust and all ligands (100%) are well anchored. In this case, forming a rigid structure may be necessary to stabilize the AF2 co-regulatory protein binding surface in an optimal conformation for co-activator binding29. Carbohydrate binding sites, on the other hand, form many more HBs with their ligands, but a lower proportion of robust ones (46%). Finally, in the case of allosteric ligands, only 40% of complexes are robust, suggesting that these sites tend to yield looser complexes. As demonstrated in the case of HIV reverse transcriptase inhibitors (Fig. 3c), lack of robust HBs does not preclude tight binding. In fact, a multiplicity of binding modes might be beneficial to preserve binding affinity when the target is mutated, thus averting resistance30,31. While the distribution of HB strength between the four types of binding sites that we have defined is quite different (see Supplementary Tables 4 and 5 for statistical tests), individual cases can deviate from the norm (e.g., the allosteric ligand 1YV3 is extremely robust) and more examples will be needed to reach firm conclusion about site-dependence.
Structures of protein-ligand complexes that form potentially labile structures. Weak hydrogen bonds (WQB < 4 kcal mol−1) marked in green. a Complex of FXa with inhibitor RPR208707 (PDB id 1F0S; Ki = 18 nM) forms two direct, but labile, HBs with the protein. An additional water-mediated HB with the catalytic residues (yellow dotted lines) might provide structural stability. b An antibody that recognizes phosphocholine (PDB id 2MCP) forms two charge-reinforced but labile HBs. A cation-pi interaction (yellow dotted lines) might provide structural stability. c Reverse transcriptase inhibitor (PDB id 1JLA; IC50 = 6 nM) forms a single but labile HB with the protein. No other source of structural stability is apparent
Strong hydrogen bonds form fragment-sized structural anchors
To understand whether robust HBs originated from a single or multiple areas on a ligand, all HBs in each complex were clustered, based on their distance in space, into fragment-sized group of atoms (Supplementary Fig. 1). In the majority of complexes (62%) robust HBs were located in a single group, forming a strong structural anchor (Fig. 4, Supplementary Table 6). The concentration of robust interactions on a single site, allowing a some degree of movement to the other parts, minimizes the entropic costs and can be desirable from a binding affinity perspective6. Only 23% of ligands form two structural anchors on separate regions, though this is more common in the case of carbohydrate-binding proteins (Supplementary Table 7). Three exceptional ligands manage to form 3 distinct stable anchors. Interestingly, they have completely unrelated functions, chemical structures and physical properties but—at least in two of those cases—there is a possible functional explanation for the extreme robustness (Fig. 5).
Distribution of complexes based on the number of structural anchors. Representative of each group is presented in the following image: a 0 anchors: Chitinase B with inhibitor (PDB id 1W1P; IC50 = 5 mM); b 1 anchor: Queuine tRNA-ribosyltransferase with inhibitor (PDB id 1N2V; Ki = 83 µM), c 2 anchors: Vitamin D3 receptor with calcipotriol (PDB id 1S19; Kd = 0.31 nM) and d 3 anchors: Uridylate kinase-AMP (PDB id 1UKZ). Weak hydrogen bonds (WQB < 4 kcal mol−1) marked in green, medium (4 ≤ WQB < 6 kcal mol−1) in yellow and strong (WQB ≥ 6 kcal mol−1) in red. e Pie chart presenting distribution of number of anchors across the data set. f Distribution of strength of HBs (WQB) versus the number of HBs per group of atoms. The box shows the quartiles and the whiskers show the rest of distribution excluding outliers. The swarmplot showing all data points is placed in top of the boxplot
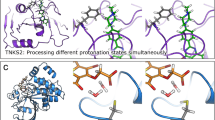
Structures of completely constricted complexes hint at potential functional role. Cases in the dataset where the ligand presents three structural anchors are: a Uridilate kinase with AMP (PDB id 1UKZ) where the base, ribose and phosphate of the nucleotide are forming three distinctive centers of interactions. Structural stability may be necessary for efficient catalysis. b Glucocorticoid receptor ligand-binding domain bound to dexamethasone (PDB id 1M2Z; Kd = 19 nM). The ligand has three regions that form robust interaction, well separated in space but located on the steroid core, thus behaving as a single rigid block. Structural stability may be necessary for agonistic response. c Influenza virus neuraminidase with inhibitor BCX-1812 (PDB id 1L7F; Ki single digit nM for various virus strains). Three different functional groups branching out of the pentane scaffold form robust interactions in this extremely polar and solvent exposed binding site. Weak hydrogen bonds (WQB < 4 kcal mol−1) marked in green, medium (4 ≤ WQB < 6 kcal mol−1) in yellow and strong (WQB ≥ 6 kcal mol−1) in red. Structural anchors are marked with shaded areas: green, red, and blue
The distribution of WQB per number of HBs in a local group (Fig. 4f) is suggestive of cooperative behavior. HBs in isolation usually do not form robust interactions (mean and median values: (4.7 ± 4.1) and 3.7 kcal mol−1, respectively), although in exceptional cases they can reach values above 10 kcal mol−1. By contrast, when three or more HBs cluster together, formation of robust complexes is the most common outcome (mean and median values: (9.4 ± 5.8) and 9.0 kcal mol−1, respectively). The HBs within these clusters present relatively similar WQB values (Supplementary Fig. 7), suggesting that they often behave in a concerted-like manner. This synergic and mutually dependent behavior not only ensures higher barriers to dissociation, but is also well-suited to provide selectivity, as small changes in the composition or geometry of one of the partners may result in large changes in magnitude of WQB (see example in Supplementary Fig. 8).
Protein-fragment complexes are more static than protein-ligand complexes
The observation that most drug-like ligands combine tightly bound regions with looser ones makes us wonder about fragment-sized ligands. Do they balance order and disorder in some other way (e.g., using fewer attachment points)? Or, perhaps, depending on the site they bind to, they are either dynamic or fully constrained? In order to answer these questions, we have extended our analysis with a set of 27 fragment-protein complexes (126 individual HBs) from the SERAPhiC dataset32. Strikingly, we find that fragments have an almost identical behavior to standard ligands, with 49% of robust HBs (2.3 per ligand) and 73% of ligands presenting at least one robust interaction. The distribution and maximal WQB values are also very similar (Fig. 6). This indicates that, proportionally, fragments are more static than standard ligands. This agrees with the observations that fragments have a more enthalpic binding33 and that they have a higher proportion of buried HBs34. It also justifies that, in spite of their low binding affinity, most fragments already have a well-defined binding mode that serves as a foundation from which to spread and catch additional interactions. However, not all fragments form robust interactions and we propose that these are less suitable as starting points because their binding mode can change, confounding structure-activity interpretation and rendering optimization more difficult. Indeed, fragments are known to change their binding mode when evolved into larger molecules7,35,36,37,38,39. These may be attempts at building on what is assumed to be a solid foundation but turns out to be unstable ground, a possibility that we shall investigate in the future. It should also be noted that the fraction of well-anchored fragments may be different for fragments hits that fail to crystallize. The overlap between X-ray crystallography and other biophysical screening methods can be rather low40 and progressing fragments that fail to crystallize is deemed difficult but worthwhile41.
Frequency of robust HBs in protein-fragment complexes. a Histogram of frequency of HBs by WQB value for all simulated HBs (124) in SERAPhiC dataset. b Pie chart showing share of complexes with at least one robust HB (WQB > 6 kcal mol−1, pink), all labile HBs (WQB < 4 kcal mol−1, green) or intermediate situations (red) for all protein-fragment complexes (26)
Structural stability is a consequence of binding free energy and desolvation
Finally, we want to consider what is the origin of the free energy barrier that causes structural stability. Knowing that a HB has a large WQB value can be likened to knowing the koff of a compound without knowing the kon nor ∆Gbind: larger values may indicate that it has a higher transition state (if ∆Gbind remains the same; Fig. 7a), that the complex is thermodynamically more stable (if kon remains the same; Fig.7b), or a combination thereof. In this data set, we find that anchoring sites often correspond to binding hot spots. This is indeed the case for all kinases and proteases, which have a well-known binding hot spot (Supplementary Table 2, Supplementary Fig. 1), as well as for most fragments. In such cases, ∆Gbind must be a component of WQB, but there is no correlation between both magnitudes (Supplementary Fig. 9), as already noted23. Thus, we conclude that WQB must be largely dominated by a transitory dissociation penalty. The origin of this penalty can be explained by a physical decoupling between HB rupture and resolvation, as described for water-shielded hydrogen bonds22. In support of this view, several studies of the reverse event have identified desolvation of the binding pocket as the rate-limiting step in ligand association21,42,43. Indeed, solvent exposed HBs invariably lead to low WQB values (but note that they can be thermodynamically stable)44, whereas water-shielding is a necessary but not sufficient condition of robust HBs (Supplementary Fig. 10).
Ways of achieving structural robustness. a Idealized representation of two dissociation pathways for complexes with the same ∆Gbind and different desolvation costs. The images above the blue curve shows the state of the system in bound, transition and unbound state of complex with well shielded, stable hydrogen bond. The images below the green curve show analogous images for the complex with solvent exposed hydrogen bond. b Likewise for three complexes with the same desolvation cost but different ∆Gbind. The images represent complexes with excellent shape complementarity that form (above the blue curve) or don’t form (below the red curve) favorable hydrogen-bonding pairs. The black dashed line marks the energy cutoff that classifies bond as structurally stable. c Example of a complex with high dissociation cost due to extreme water-shielding. d Example of a complex with high dissociation cost due to a tight network of multiple HBs. Weak hydrogen bonds (WQB < 4 kcal mol−1) marked in green and strong (WQB ≥ 6 kcal mol−1) in red
Discussion
Taken together, our results show that structural stability is a common property of protein-ligand complexes, but not an universal one. Cases of loose complexes, while relatively rare (10-20%), can be found even in a dataset originating exclusively from X-ray crystallography, a technique that requires structural homogeneity of the sample. The proportion could be larger amongst ligands that fail to crystallize. The level of residual mobility is also larger and more common than the static X-ray structures lead to think, as also concluded by a recent independent study4. In fact, most complexes balance order and disorder by combining a firm anchor with more relaxed peripheral interactions. Depending on the nature of the ligand and the binding site, each complex adopts a particular degree of robustness, that ranges from the very tight (e.g., nuclear receptor agonists) to the very loose (e.g., HIV-RT allosteric inhibitors). Each one of these solutions entails important consequences that have, so far, been neglected in drug design. First of all, a firm anchor provides a framework from which to grow and capture additional interactions, and the preservation of a common binding mode helps interpreting structure-activity relationships. This is particularly important for fragments as starting points for lead discovery. Secondly, structural robustness can have functional implications, particularly in the case of receptors, where flexibility has been linked to the agonist/antagonist response29,45. Thirdly, structural stability implies an entropic penalty and must be balanced to avoid loss of potency6,46. Finally, the deep and narrow energetic minima that cause rigidity also imply large penalties for small recognition defects, thus increasing the fidelity of the recognition event. This has been shown for protease-substrate pairs47 and HIV-protease inhibitors48. In conclusion, this work opens up the possibility of understanding and designing structural robustness in ligand-receptor complexes. We suggest that robustness analysis, which can help understand and control the level of mobility, should be an essential part of ligand design, not least because rigid parts demand more precise complementarity than flexible ones. Qualitatively, a visual inspection can reveal water-shielded HBs (Fig. 7c) and HB clusters (Fig. 7d), which are tell-tale signs of robustness. Quantitatively, DUck simulations offer an inexpensive and automated protocol to calculate WQB. While HBs appear to be the most common means of achieving structural robustness, other interaction types (e.g., cation-pi, water-mediated HBs, halogen bonds) should be considered in the future.
Data availability
All data generated during the current study are available as a part of the Supplementary Information in the form of sdf and mol2 files.
References
Tompa, P. & Fuxreiter, M. Fuzzy complexes: polymorphism and structural disorder in protein-protein interactions. Trends Biochem. Sci. 33, 2–8 (2008).
Borgia, A. et al. Extreme disorder in an ultrahigh-affinity protein complex. Nature 555, 61–66 (2018).
Cozzini, P. et al. Target flexibility: an emerging considertaion in drug discovery. J. Med. Chem. 51, 6237–6255 (2008).
Van Zundert, G. C. P. et al. QFit-ligand reveals widespread conformational heterogeneity of drug-like molecules in X-ray electron density maps. J. Med. Chem. 61, 11183–11198 (2018).
Klebe, G. Applying thermodynamic profiling in lead finding and optimization. Nat. Rev. Drug Discov. 14, 95–110 (2015).
Glas, A., Wamhoff, E.-C., Kruger, D. M., Rademacher, C. & Grossmann, T. N. Increased conformational flexibility of a macrocycle–receptor complex contributes to reduced dissociation rates. Chem. Eur. J. 23, 16157–16161 (2017).
Malhotra, S. & Karanicolas, J. When does chemical elaboration induce a ligand to change its binding mode? J. Med. Chem. 60, 128–145 (2017).
Kuhnert, M. et al. Tracing binding modes in hit-to-lead optimization: chameleon-like poses of aspartic protease inhibitors. Angew. Chem. Int. Ed. 54, 2849–2853 (2015).
Sliwoski, G., Kothiwale, S., Meiler, J. & Lowe, E. W. Computational methods in drug discovery. Pharmacol. Rev. 66, 334–395 (2014).
Mobley, D. L. & Dill, K. A. Binding of small-molecule ligands to proteins: ‘what you see’ is not always ‘what you get’. Structure 17, 489–498 (2009).
Kaus, J. W. et al. How to deal with multiple binding poses in alchemical relative protein-ligand binding free energy calculations. J. Chem. Theory Comput. 11, 2670–2679 (2015).
Gill, S. C. et al. Binding modes of ligands using enhanced sampling (BLUES): rapid decorrelation of ligand binding modes via nonequilibrium candidate Monte Carlo. J. Phys. Chem. B 122, 5579–5598 (2018).
Pan, A. C., Borhani, D. W., Dror, R. O. & Shaw, D. E. Molecular determinants of drug–receptor binding kinetics. Drug Discov. Today 18, 667 (2013).
Bissantz, C., Kuhn, B. & Stahl, M. A medicinal chemist’s guide to molecular interactions. J. Med. Chem. 53, 5061–5084 (2010).
Ferreira de Freitas, R. & Schapira, M. A systematic analysis of atomic protein–ligand interactions in the PDB. Med. Chem. Commun. 8, 1970–1981 (2017).
Fersht, A. R. The hydrogen bond in molecular recognition. Trends Biochem. Sci. 12, 301–304 (1987).
Pace, C. N. Energetics of protein hydrogen bonds. Nat. Struct. Mol. Biol. 16, 681–682 (2009).
Pace, C. N. et al. Contribution of hydrogen bonds to protein stability. Protein Sci. 23, 652–661 (2014).
Gao, J., Bosco, D. A., Powers, E. T. & Kelly, J. W. Localized thermodynamic coupling between hydrogen bonding and microenvironment polarity substantially stabilizes proteins. Nat. Struct. Mol. Biol. 16, 684–690 (2009).
Nick Pace, C., Martin Scholtz, J. & Grimsley, G. R. Forces stabilizing proteins. FEBS Lett. 588, 2177–2184 (2014).
Mondal, J., Friesner, R. A. & Berne, B. J. Role of desolvation in thermodynamics and kinetics of ligand binding to a kinase. J. Chem. Theory Comput. 10, 5696–5705 (2014).
Schmidtke, P., Javier Luque, F., Murray, J. B. & Barril, X. Shielded hydrogen bonds as structural determinants of binding kinetics: application in drug design. J. Am. Chem. Soc. 133, 18903–18910 (2011).
Ruiz-carmona, S. et al. Dynamic undocking and the quasi-bound state as tools for drug discovery. Nat. Chem. 9, 201–206 (2017).
Warren, G. L., Do, T. D., Kelley, B. P., Nicholls, A. & Warren, S. D. Essential considerations for using protein-ligand structures in drug discovery. Drug Discov. Today 17, 1270–1281 (2012).
Majewski, M., Ruiz-Carmona, S. & Barril, X. in Rational Drug Design: Methods and Protocols (eds. Mavromoustakos, T. & Kellici, T. F.) 195–215 (Springer, New York, 2018).
Kleywegt, G. J. & Jones, T. A. in Methods in Enzymology 208–230 (Elsevier, 1997).
Parthasarathy, S. & Murthy, M. R. N. On the correlation between the main-chain and side-chain atomic displacement parameters (B values) in high-resolution protein structures. Acta Crystallogr. Sect. D. Biol. Crystallogr. 55, 173–180 (1999).
Barril, X., Aleman, C., Orozco, M. & Luque, F. J. Salt bridge interactions: stability of the ionic and neutral complexes in the gas phase, in solution, and in proteins. Proteins Struct. Funct. Bioinforma. 32, 67–79 (1998).
Mayer-Wrangowski, S. C. & Rauh, D. Monitoring ligand-induced conformational changes for the identification of estrogen receptor agonists and antagonists. Angew. Chem. Int. Ed. 54, 4379–4382 (2015).
Das, K. et al. Roles of conformational and positional adaptability in structure-based design of TMC125-R165335 (Etravirine) and related non-nucleoside reverse transcriptase inhibitors that are highly potent and effective against wild-type and drug-resistant HIV-1 variants. J. Med. Chem. 47, 2550–2560 (2004).
Lee, W. G., Chan, A. H., Spasov, K. A., Anderson, K. S. & Jorgensen, W. L. Design, conformation, and crystallography of 2-naphthyl phenyl ethers as potent anti-HIV agents. ACS Med. Chem. Lett. 7, 1156–1160 (2016).
Favia, A. D., Bottegoni, G., Nobeli, I., Bisignano, P. & Cavalli, A. SERAPhiC: a benchmark for in silico fragment-based drug design. J. Chem. Inf. Model. 51, 2882–2896 (2011).
Ferenczy, G. G. & Keseru, G. M. Thermodynamics of fragment binding. J. Chem. Inf. Model. 52, 1039–1045 (2012).
Giordanetto, F., Jin, C., Willmore, L., Feher, M. & Shaw, D. E. Fragment hits: what do they look like and how do they bind? J. Med. Chem. 62, 3381–3394 (2019).
Drwal, M. N. et al. Structural insights on fragment binding mode conservation. J. Med. Chem. 61, 5963–5973 (2018).
Mpamhanga, C. P. et al. One scaffold, three binding modes: novel and selective pteridine reductase 1 inhibitors derived from fragment hits discovered by virtual screening. J. Med. Chem. 52, 4454–4465 (2009).
Casale, E. et al. Fragment-based hit discovery and structure-based optimization of aminotriazoloquinazolines as novel Hsp90 inhibitors. Bioorg. Med. Chem. 22, 4135–4150 (2014).
Han, X. et al. Discovery of potent and selective CDK8 inhibitors through FBDD approach. Bioorg. Med. Chem. Lett. 27, 4488–4492 (2017).
Forster, A. B. et al. The identification of a novel lead class for phosphodiesterase 2 inhibition by fragment-based drug design. Bioorg. Med. Chem. Lett. 27, 5167–5171 (2017).
Schiebel, J. et al. Six biophysical screening methods miss a large proportion of crystallographically discovered fragment hits: a case study. ACS Chem. Biol. 11, 1693–1701 (2016).
Erlanson, D. A., Davis, B. J., Jahnke, W. & Box, G. Perspective fragment-based drug discovery: advancing fragments in the absence of crystal structures. Cell Chem. Biol. 26, 9–15 (2019).
Dror, R. O. et al. Pathway and mechanism of drug binding to G-protein-coupled receptors. Proc. Natl Acad. Sci. USA 108, 13118–13123 (2011).
Schuetz, D. A. et al. Ligand desolvation steers on-rate and impacts drug residence time of heat shock protein 90 (Hsp90) Inhibitors. J. Med. Chem. 61, 4397–4411 (2018).
Ciulli, A., Williams, G., Smith, A. G., Blundell, T. L. & Abell, C. Probing hot spots at protein-ligand binding sites: a fragment-based approach using biophysical methods. J. Med. Chem. 49, 4992–5000 (2006).
Ghanouni, P. et al. Functionally different agonists induce distinct conformations in the G protein coupling domain of the β2Adrenergic receptor. J. Biol. Chem. 276, 24433–24436 (2001).
Brandt, T. et al. Congeneric but still distinct: how closely related trypsin ligands exhibit different thermodynamic and structural properties. J. Mol. Biol. 405, 1170–1187 (2011).
Fuchs, J. E. et al. Cleavage entropy as quantitative measure of protease specificity. PLoS Comput. Biol. 9, e1003007 (2013).
Shen, Y., Radhakrishnan, M. L. & Tidor, B. Molecular mechanisms and design principles for promiscuous inhibitors to avoid drug resistance: lessons learned from HIV-1 protease inhibition. Proteins Struct. Funct. Bioinforma. 83, 351–372 (2015).
Acknowledgements
We thank C. Galdeano and F. J. Luque for helpful discussions and paper revision. The research was funded under the EU Horizon 2020 program, Marie Skłodowska-Curie grant agreement No. 675899 (FRAGNET). Spanish Ministerio de Economia (SAF2015-68749-R). Catalan government (2014 SGR 1189). Barcelona Supercomputing Center (computational resources).
Author information
Authors and Affiliations
Contributions
X.B. designed the project; M.M. performed calculations; S.R.C. contributed new analytic tools; M.M. and X.B. analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Majewski, M., Ruiz-Carmona, S. & Barril, X. An investigation of structural stability in protein-ligand complexes reveals the balance between order and disorder. Commun Chem 2, 110 (2019). https://doi.org/10.1038/s42004-019-0205-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-019-0205-5
This article is cited by
-
Design, synthesis, and in silico studies of novel di-(2-aryl hydrozonopropanal) arene derivatives as potent anticancer for targeting A2AR and LRP6 in HCT116 cell
Medicinal Chemistry Research (2024)
-
Reporting the anti-neuroinflammatory potential of selected spondias mombin flavonoids through network pharmacology and molecular dynamics simulations
In Silico Pharmacology (2024)
-
Predicting Plasmodium falciparum kinase inhibitors from antimalarial medicinal herbs using computational modeling approach
In Silico Pharmacology (2023)
-
Mode of action of teixobactins in cellular membranes
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.