Abstract
Metal-catalyzed chemical transformations performed at the cellular level bear great potential for the manipulation of biological processes. The complexity of the cell renders the use of transition metal chemistry difficult in cellular systems. The delivery of the reactive catalyst and the control of its spatial localization remain challenging. Here we report the surface functionalization of the unicellular eukaryote Chlamydomonas reinhardtii with a tailor-made artificial metalloenzyme for on-cell catalysis. The functionalized cells remain viable and are able to uncage a fluorogenic substrate on their surface. This work leverages cell surface engineering to provide live cells with new-to-nature reactivity. In addition, this operationally simple approach is not genetically encoded and thereby transient, which offers advantages with regard to temporal control, cell viability, and safety. Therefore, and as a feature, the movement of the functionalized cells can be directed by light (via phototaxis), allowing for the three-dimensional localization of catalysts by outside stimuli.
Similar content being viewed by others
Introduction
Over the past decades, the ability to introduce new-to-nature chemical reactions in living cells has emerged as a desirable goal, and as requirement, much progress has been achieved in the field of bioorthogonal chemistry1. The development of chemospecific biocompatible reactions enabled the labeling of biomolecules and the activation of probes at a cellular level, key to the study of biological processes. However, until recently, the set of available bioorthogonal transformations was mostly limited to bond-forming reactions for bioconjugations2,3,4,5,6,7,8,9,10. Technologies to perform chemical synthesis beyond ligations at a cellular level are still needed. Bond-breaking reactions bear a great potential in chemical biology and medical applications11. Despite constant progress in this field, the selective cleavage of chemical bonds without perturbing cells’ native chemical processes still remains a challenge. The introduction of transition metal complexes into cells has recently proven to be able to bring new reactivity to cellular systems and was efficiently used for the activation of probes and prodrugs in cellulo12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28. However, most metal complexes penetrate cells with poor site selectivity leading to potential undesired side-reactions. The lack of spatial control as well as control over the faith of the reactive metal center in the cell constitute a barrier to further medical applications.
For years, evolution has shaped and optimized enzymes, making possible a set of reactions crucial to life. On the other hand, chemists have developed numerous methods for the synthesis of molecules beneficial to mankind. Previous work showed that both of these worlds can be merged to complement the work of natural enzymes. Artificial metalloenzymes (ArMs hereafter), resulting from the incorporation of a reactive metal cofactor into a protein scaffold, combine the versatility of metal catalysts with the unique tri-dimensional environment of proteins. From olefin metathesis29 and cyclopropanation30 to C-H insertion31, recent advances in protein engineering and organometallic synthesis enabled a wide range of non-natural reactions to be performed in a biocompatible manner32. The artificial allylic deallylase used here, catalyzes the cleavage of allyl-carbamate protected amines. Examples of this versatile reaction taking place on E. coli and in mammalian cells have been recently reported33,34.
The cell surface is the site of numerous key biological processes. For this reason, the engineering of the surface of mammalian cells, bacteria, yeasts, and microalgae has gained increasing attention. Synthetic molecules bearing defined functional groups are currently exploited to remodel cell surfaces in a selective and rational manner.35 The modification of cell phenotypes by chemical surface engineering found applications in many areas including gene transfer36, proteomics37, cell therapy38, cell patterning39, or nucleic acid transfection40.
Surface functionalization of live cells with an artificial metalloenzyme could allow both reactivity and spatial control over the desired transformation. In this context, bacterial surface display has emerged as a promising technology for the development and the study of new reactions at a cellular level34,41. We reasoned that the functionalization of the cell surface of an eukaryote without relying on genetic means would represent an attractive alternative. Such approach might offer flexibility by allowing time control of the functionalization, being non-hereditary and operationally simple to perform.
In this study, we report the surface functionalization of live Chlamydomonas reinhardtii cells with a tailor-made artificial allylic deallylase for on-cell catalysis. The engineered cells are viable and possess the ability to catalyze a reaction that is new-to-nature.
Results
Cell surface functionalization of Chlamydomonas reinhardtii
For our proof of concept of on-cell catalysis, we selected the unicellular green alga Chlamydomonas reinhardtii (C. reinhardtii). The microalga, for which the full genome sequence has been obtained42, is widely used as model organism in cell and molecular biology43. The microscopic eukaryote has gained attention due to its ability to carry out photosynthesis. Its use has recently expanded from the study of fundamental processes to applied research in biofuels44, therapeutic protein production45, or wound healing46,47. Interested in leveraging further the potential of C. reinhardtii, our group recently reported its surface engineering for antibiotic delivery48. C. reinhardtii cells are spherical, 10 µm large, and feature two flagella which allow motility. Combined with its ability to sense light, locomotion renders the microalga phototactic49. The cell membrane of C. reinhardtii is surrounded by a multilayered cell wall composed of glycoproteins. Both covalent and non-covalent interactions between glycopeptide residues give rise to the cell wall structure50.
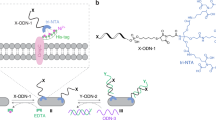
We envisioned to install the artificial metalloenzyme on the cell surface of C. reinhardtii by a covalent approach. Several types of chemical functionalizations have recently been reported. These techniques take advantage of reactive functional groups readily available at the cell surface, such as thiols or amines38,39,51,52. In our case, we hypothesized that amino acid residues bearing amino groups within the cell wall of C. reinhardtii might react with good electrophiles such as N-hydroxysuccinimide esters. To test this hypothesis, we first aimed at functionalizing the cell surface with succinimidyl ester 1. Cells were collected from the culture, washed from the culture medium, resuspended in phosphate-buffered saline (PBS), and incubated with carboxyfluorescein succinimidyl ester 1 for 1 h (Fig. 1a). After washing and resuspension in PBS, cells were analyzed by confocal laser scanning microscopy. As illustrated in Fig. 1b, fluorescence images revealed that 1 covalently reacts with C. reinhardtii as witnessed on the green channel (detection of fluorescein). Moreover, we were pleased to observe that the labeling was restricted to the cell surface as no fluorescence signal was detected from the cytoplasm. To confirm that the covalent labeling was indeed targeting the cell wall, we repeated the experiment with C. reinhardtii CC-406, a cell wall-deficient mutant which lacks major components of the cell wall. In the absence of a proper cell wall, only a faint labeling was detected (Supplementary Figure 1), confirming that the N-hydroxysuccinimide ester moiety is reacting with glycoprotein components of the cell wall of C. reinhardtii.
Succinimidyl esters enable covalent labeling of the cell wall of C. reinhardtii. a Schematic representation of cell wall covalent labeling with 1. b Confocal laser scanning microscopy images of cells after treatment with PBS (control) or with 1 (0.65 mM), 1 h, PBS (treated). Green channel: detection of fluorescein. Red channel: detection of the chloroplast. Scale bar, 10 µm
Functionalization of C. reinhardtii with streptavidin
Encouraged by these results, we next employed the N-hydroxysuccinimide (NHS) ester strategy for C. reinhardtii functionalization with streptavidin (SAV), the host protein of our artificial metalloenzyme. The high affinity between the tetrameric protein isolated from Streptomyces avidinii and its ligand biotin is extensively used in the molecular life sciences. The tetrameric structure of streptavidin allows it to bind simultaneously to multiple biotinylated ligands. We envisioned to functionalize the cell surface with biotin anchor 2 in a first step (Fig. 2a). The NHS group would allow for covalent reaction with C. reinhardtii as demonstrated previously. The polyethylene glycol chain would act as a spacer and increase the water solubility of the anchor. The biotin moiety would provide the affinity to streptavidin and would allow us to install the desired protein on the cell surface. Cells were first washed with PBS, and incubated for 1 h with biotin anchor 2 (Fig. 2a). After washing the excess of anchor 2, cells were resuspended in PBS, incubated with streptavidin-Alexa Fluor 488 conjugate 3 for 20 min, washed, and imaged on a confocal laser scanning microscope. Different concentrations were screened for 3 (Supplementary Figure 2). As presented in Fig. 2b, a clear signal from the cell surface in the Alexa 488 channel could be detected, confirming functionalization with streptavidin. Once again, no signal was detected from cells’ cytoplasm, highlighting great cell wall selectivity. Control samples, consisting of cells incubated with streptavidin-Alexa 488 without prior functionalization with the biotin anchor 2, displayed no signal on the Alexa 488 channel. The absence of signal in these control samples confirmed that there is no unspecific binding of streptavidin to the microalga and that prior functionalization with biotin anchor 2 is a prerequisite. To further demonstrate that the functionalization is statistically significant, flow cytometry experiments were carried out. As displayed in Fig. 2c, previous results from confocal microscopy imaging were confirmed. Functionalization with streptavidin is statistically significant, and more than 99% of the treated cells displayed green fluorescence with a relatively narrow distribution of the fluorescence intensity (Supplementary Figures 3 and 4). Moreover, only cells treated with both anchor 2 and streptavidin-Alexa 488 conjugate 3 displayed green fluorescence, as depicted in Fig. 2d. These experiments further confirmed that the biotin anchor 2 is necessary for the functionalization with streptavidin, excluding non-specific interactions of streptavidin with C. reinhardtii. We then focused on assessing the viability of the functionalized cells. Propidium iodide, a red-fluorescent nuclear stain, was used to selectively stain non-viable cells. As illustrated in Fig. 2e, we were pleased to see that the functionalization with both the biotin anchor 2 (step 1) and streptavidin (step 2) had very small impact on the cell viability since the median fluorescence intensity of propidium iodide was very similar for all samples. A total of 77% of the treated cell population was functionalized and viable. The procedure described above thus allows for specific surface functionalization of C. reinhardtii with streptavidin, with preserved cell viability.
Surface functionalization of C. reinhardtii with streptavidin. a Schematic representation of the two-step procedure for surface immobilization of streptavidin. In a first step, cells are covalently functionalized with the biotin anchor 2 and then incubated with the streptavidin-Alexa 488 conjugate 3. b Confocal laser scanning microscopy images of cells after treatment with PBS (not treated), 3 (50 µg mL−1), 20 min, PBS (control) or 2 (0.65 mM), 1 h, PBS then 3 (50 µg mL−1), 20 min, PBS (treated). Green channel: detection of Alexa 488. Red channel: detection of the chloroplast. Images are shown as red and green channel overlay. Scale bar, 10 µm. c Flow cytometry analysis after functionalization. Representation of one experiment of n = 3 as a density plot, propidium iodide (Texas red channel) vs. Alexa 488 (FITC channel). d Efficiency of the functionalization with streptavidin-Alexa 488 conjugate: median Alexa 488 fluorescence intensity. e Cell viability after functionalization with streptavidin: median propidium iodide fluorescence intensity. d, e Data are represented as mean ± SD (n = 3) and analyzed by one-way ANOVA (post hoc Tukey test; *P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001; ****P ≤ 0.0001)
Development of an artificial allylic deallylase
After demonstrating the functionalization of live cells with the streptavidin host, focus was set on the reaction and the artificial metalloenzyme. We selected the cleavage of allyl-carbamate-protected amines, a reaction that is foreign to nature, as our target transformation. Amines are a versatile functional group present in a broad range of natural products as well as synthetic drugs. The deprotection of amino groups within active small molecules and biomolecules can be used for prodrug activation (Fig. 3a). Caging of a functional group in a molecule of interest can be used to lower its activity by masking an important moiety in the parent molecule. Once the caging group is cleaved off, the molecule regains its inherent biological activity. If prodrugs have been successfully used to enhance a drug’s permeability, distribution, or metabolism, the major challenge remains the usage of prodrugs to overcome toxicity issues.53 In particular, prodrugs might be a solution to lower adverse effects on healthy tissues in treatments relying on highly cytotoxic but poorly selective agents.54 In this particular case, a catalytic and spatial control of this uncaging process is crucial for therapeutic applications. As a model reaction for on-cell catalysis, we selected the uncaging of allyl-carbamate-protected aminocoumarin 4 by the organoruthenium complex 5, reported by Meggers and co-workers19 (Fig. 3b). The authors reported that ruthenium-based catalysts such as 5 could perform the catalytic cleavage of O-allyl carbamates in biologically relevant conditions. ProCoumarin 4 is water soluble and the parent free aminocoumarin 6 displays an increased fluorescence upon uncaging, allowing for straightforward monitoring of the reaction.
Development of an artificial allylic deallylase for the activation of proCoumarin 4. a Cleavage of protected amines for prodrug activation. b Uncaging of allyl-carbamate-protected proCoumarin 4 as our model reaction. c Development of the artificial allylic deallylase 10 based on ruthenium complex 7 and biotinylated ligand 8. d In vitro screening of artificial allylic deallylases for the deprotection of proCoumarin 4. Reaction conditions: proCoumarin 4 (500 µM), ruthenium cofactor 9 (5 µM in DMF), purified streptavidin isoforms (10 µM, free biotin-binding sites), PBS (1×, pH 7.4), 0.5% DMF, 25 °C, 250 rpm shaking, 4 h. Fluorescence of product 6 was determined in a plate reader at λex. = 395 nm and λem. = 460 nm from triplicate reactions. Conversions were calculated by comparison with a standard curve of coumarin 6 (see Supplementary Figure 7 for details). Data are represented as mean ± SD (n = 3)
We thus synthesized biotinylated ligand 8 and reacted it with complex 7 to form the biotinylated ruthenium catalyst [CpRu(QA-Biot)(OH2)] 9 (QA = 2-quinolinecarboxylate). Incorporation into streptavidin afforded the artificial allylic deallylase (ADAse) 10 (Fig. 3c). In the deprotection of proCoumarin 4, the free ruthenium cofactor 9 revealed only modest activity. However, incorporated into streptavidin wild type (SAV WT) the yield of product 6 was increased threefold (Fig. 3d). We performed an in vitro mutant screening at positions S112X and K121X in close proximity of the active metal cofactor (Supplementary Figures 5 and 6). Overall, 34 single mutants were tested for ADAse activity. The best performing mutants at the positions S112X and K121X were S112M and K121R, respectively. We subsequently screened a selected set of double mutants: unfortunately, the double mutant S112M-K121R did not reveal a higher activity than the corresponding single mutant K121R (48% and 55% yield, respectively). However, the combination of mutant S112Y, which only displayed the fourth highest activity in the single mutant screening, with mutant K121R resulted in a highly active ADAse. Catalysis with the double mutant S112Y-K121R yielded a seven-fold higher concentration of coumarin product 6 than the corresponding wild-type ADAse (81% vs. 11% yield) after 4 h at RT.
Cell surface engineering with the ADAse
After achieving a good catalytic activity with the ADAse [CpRu(QA-Biot)(OH2)] · SAV S112Y-K121R, the focus was set on developing a procedure for its immobilization on the microalga. We planned to adapt the procedure described above for the cell surface functionalization with streptavidin (Fig. 4a). Cells were first washed in PBS, treated with the biotin anchor 2 (0.65 mM), and washed again. To ensure the availability of free SAV-binding sites, necessary for the surface functionalization, a 4:1 ratio of free SAV-binding site:cofactor was used (vs. a 2:1 ratio for the in vitro screening). In parallel, the Ru-cofactor 9 (2.5 µM) was prepared by incubating the biotinylated ligand 8 with 7 in a 1:1 ratio in DMF for 15 min and then incubated with the selected streptavidin S112Y-K121R mutant (10 µM free biotin-binding sites) to assemble the ADAse 10. Biotinylated cells were then incubated with 10 for 20 min, the buffer was removed, the cells were washed and resuspended in PBS (Fig. 4b). After the functionalization of C. reinhardtii with the ADAse, we were concerned about cell viability, as transition metals are known to be cytotoxic above certain concentrations. The cell viability was thus assessed by flow cytometry after functionalization, and the immobilization of ADAse 10 turned out to have no detectable impact on cells’ viability (Supplementary Figures 8 and 9).
On-cell catalysis with the artificial allylic deallylase. a Schematic representation of C. reinhardtii functionalization with the artificial metalloenzyme 10. Cells were first functionalized with the biotin anchor 2, step (1) and then with the artificial allylic deallylase 10, step (2). b Optimized workflow for the cell surface functionalization. c Uncaging of the proCoumarin 4 (400 µM) by the engineered cells. Only cells functionalized with biotin anchor 2 (0.65 mM), 1 h, PBS and then the ADAse 10: cofactor 9 (2.5 µM) and S112Y-K121R (10 µM, free biotin-binding sites) develop a signal. Activity is given as fluorescence intensity arising from 6 in arbitrary units after 18 h of reaction in PBS at 25 °C. d Reactivity of the functionalized cells is increasing with biotin anchor 2 concentration. Activity is given as fluorescence intensity arising from 6 in arbitrary units after 24 h of reaction in PBS at 25 °C, proCoumarin 4 (400 µM). (1) Wild-type streptavidin was used in this case
Activity of the engineered cells
With the functionalized and viable cells in hand, we then moved on to test their ability to perform the desired transformation. The reaction was initiated by addition of proCoumarin 4 at a concentration of 400 μM in PBS to the engineered cells. After 18 h, fluorescence of the uncaged coumarin 6 was measured (Fig. 4c). Reactivity could be unambiguously detected for cells functionalized with the selected ADAse 10. Untreated cells did not show any intrinsic catalytic activity. Cells incubated with Ru-cofactor 9 or the ADAse 10 without prior incubation with biotin anchor 2 were not active either. Demonstrating the effect of ligand-accelerated catalysis, cells functionalized with the WT ADAse showed a greatly reduced reactivity compared to the selected mutant, corroborating the results of the streptavidin mutants screening. These experiments confirmed: (i) that there is no C. reinhardtii-derived background activity; (ii) that there is no cellular uptake of the Ru-cofactor 9 nor the ADAse 10; (iii) that biotin anchor 2 is needed; (iv) that the assembly of the streptavidin and the cofactor 9 is responsible for the observed transformation.
We next studied the influence of the biotin anchor 2 concentration on the reactivity. Cells were treated with 2 at three different concentrations (0.65, 1.6, and 3.2 mM), maintaining the concentration of Ru-cofactor 9 and the streptavidin host S112Y-K121R constant to 2.5 and 10 µM, respectively. As illustrated in Fig. 4d, increased biotin anchor concentration led to higher activity and the uncaging of proCoumarin 4 was limited to cells engineered with the ADAse, albeit in low yields (3 to 4%). One can indeed imagine that an increased incubation concentration of 2 leads to denser biotinylation of the cell surface, and thus to a greater amount of ADAse 10 on the cell surface. Since the Ru-cofactor 9 and the streptavidin host concentrations were maintained constant, these results also indicate that after incubation with biotin anchor 2 at 0.65 mM, cells take up less than 2.5 µM of the ADAse 10. These experiments also reveal that a five-fold increase in concentration of the biotin anchor 2 does not lead to a five-fold increase in activity. This suggests that, as expected, a limited amount of ADAse 10 can be installed on the cell surface.
After demonstrating the catalytic activity of the engineered cells, focus was set on preserving the cell viability. Because the previous reaction conditions resulted in a decreased number of viable cells after catalysis, efforts were directed towards identifying reaction conditions with a reduced impact on cell viability. We hypothesized that prolonged exposure to PBS instead of C. reinhardtii culture medium (Kuhl medium) could be a reason for the decreased cell viability after catalysis. Additional investigations on the reaction time and reaction medium revealed that PBS has indeed a negative influence on cell viability (Supplementary Figure 10). Under optimized conditions (15 h reaction time in Kuhl medium), we were pleased to see that the engineered cells could significantly uncage the proCoumarin 4 (Fig. 5a) with a preserved cell viability albeit with lower efficiency (Fig. 5b and Supplementary Figure 11).
On-cell catalysis with preserved cell viability. Functionalization conditions: 2 (0.65 mM), 1 h, PBS; cofactor 9 (2.5 µM), streptavidin S112Y-K121R (10 µM free biotin-binding sites). a Uncaging of the proCoumarin 4 (400 µM, Kuhl medium) by the functionalized cells. Activity is given as fluorescence intensity coming from 6 in arbitrary units after 15 h at 25 °C. b Cell viability after reaction determined by flow cytometry with propidium iodide. a, b Data are represented as mean ± SD (n = 3) and analyzed by one-way ANOVA (post hoc Tukey test; *P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001; ****P ≤ 0.0001). (1) Wild-type streptavidin was used in this case
To further evaluate the impact of the functionalization on the cell viability, we challenged the phototactic ability of the cells after functionalization. C. reinhardtii CC-124, a strain with negative phototaxis ability55 (capacity to move away from the light source) was functionalized with the ADAse as previously described and subjected to a phototaxis assay. The functionalized cells still displayed negative phototaxis after functionalization (Fig. 6) as cells moved away from the green LED over the course of 15 min in a Petri dish. However, as the phototatic ability of C. reinhardtii depends on a variety of parameters (intensity of the light source, medium, growth phase), consistent movement was difficult to obtain.
Cells functionalized with the ADAse 10 maintain a phototactic response. C. reinhardtii CC-124 was functionalized as previously described and transferred to a Petri dish. a Picture taken before illumination with a green LED. b Picture taken after illumination with a green LED from one side of the dish for 15 min. Scale bar, 1 cm
Discussion
In summary, we have developed a system to endow a cell with the ability to catalyze on its surface a transformation that is new-to-nature. The approach developed here does not rely on the biological machinery of the target cell but takes advantage of chemical surface functionalization to introduce the artificial metalloenzyme. To the best of our knowledge, this study represents the first example of chemical surface engineering of a eukaryotic cell with a tailor-made artificial metalloenzyme. The artificial allylic deallylase introduced on the cell surface functions as a “chemical prosthesis” that provides the cell with new chemical capabilities. The streptavidin possesses a dual function and allows for both shielding of the metal center from the biological environment and cell surface immobilization of the metalloenzyme. The approach described here seems complementary to the strategy relying on the transformation of native membrane receptors into metalloenzymes, recently reported by Ghattas et al.56. However, introducing a fully artificial metalloenzyme based on the biotin-streptavidin technology has the advantage of providing more versatility (a single host protein for a variety of biotinylated cofactors) and the possibility to optimize the host protein by mutagenesis. Implementing new chemistry in cellular systems from the outside can be beneficial. It gives the advantage of closely controlling the site of the introduced abiotic reactive center and thus the site of the reaction to be catalyzed. This could open the door to spatial control over the process of prodrug uncaging that may prove beneficial for medical applications. The principle developed here could be expanded to surface engineering of mammalian cells. In the context of immunotherapy and cell therapy, the cell surface plays a crucial role, and the important efforts made in synthetic biology could benefit from new reactivity being implemented on the cell membrane. As an example, with the use of selective surface engineering methods (i.e. metabolic glycoengineering) to introduce the biotin moiety, one could develop further the approach discussed here to selectively equip cancer cells with such chemical prosthesis. The target cells would then be equipped to uncage prodrugs in an orthogonal and selective fashion on their surface. Giving the potential applicability of the concept introduced here, our aim is to expand this approach in order to provide new chemistry to modulate cellular systems from the outside-in.
Methods
General
5(6)-Carboxyfluorescein N-hydroxysuccinimide ester, Phosphate-Buffered Saline (Gibco, pH 7.4, 1 × ) and Prolong® Diamon Antifade Mountant were purchased from Thermo Fisher Scientific. Tris(acetonitrile)cyclopentadienylruthenium(II) hexafluorophosphate and propidium iodide were purchased from Sigma Aldrich. NHS-PEG12-biotin was purchased from BroadPharm. Dimethylformamide was purchased form Acros Organics. Streptavidin-Alexa Fluor 488 conjugate was purchased from Invitrogen. Cell counting slides were purchased from Kova International. All reagents were used without further purification. PBS buffer and Kuhl medium were strained over a Filtropur BT 50 0.45 mm filter (Sarstedt) prior to use.
Cell culture
Chlamydomonas reinhardtii (11–32b) was purchased from EPSAG (Experimentelle Phykologie und Sammlung von Algenkulturen der Universität Göttingen) and grown in Kuhl medium (1011.1 mg L−1 KNO3, 621 mg L−1 NaH2PO4·H2O, 89 mg L−1 Na2HPO4·2H2O, 246.5 mg L−1 MgSO4·7H2O, 14.7 mg L−1 CaCl2·2H2O, 6.95 mg L−1 FeSO4·7 H2O (Fe-EDTA complex), 1 mL L−1 micronutrient solution (61.0 mg L−1 H3BO3, 169.0 mg L−1 MnSO4·H2O, 287.0 mg L−1 ZnSO4·7H2O, 2.5 mg L−1 CuSO4·5H2O, 12.5 mg L−1 (NH4)6Mo7O24·4H2O) for 1 L of medium) at 20 °C under cycles of medium light (30 V) for 12 h and dark for 12 h. The cell wall-deficient mutant (CC-406 cw15 mt-) was purchased from the Chlamydomonas Resource Center, University of Minnesota and grown in Tris Minimal medium (2.42 mg L−1 Triz-ma® base, 25 mL salt solution (15.0 mg L−1 NH4Cl, 4 g L−1 MgSO4·7H2O, 2.0 g L−1 CaCl2·2H2O), 0.375 mL phosphate solution (28.8 g L−1 K2HPO4, 14.4 g L−1 KH2PO4), 1 mL Fe-EDTA complex (6.95 mg L−1 FeSO4·7H2O)) at 20 °C under cycles of medium light (30 V) for 12 h and dark for 12 h. Chlamydomonas reinhardtii CC-124 was purchased from the Chlamydomonas Resource Center, University of Minnesota and grown in TAP medium (20 mL of 1 M Tris base, 1 mL of glacial acetic acid, 10 mL of solution A (40.0 g L−1 NH4Cl, 50 g L−1 CaCl2·2H2O, 10 g L−1 MgSO4·7H2O), 1.0 mL phosphate solution (108 g L−1 K2HPO4, 56 g L−1 KH2PO4), 10 mL Tris-acetate solution (242 g L−1 Tris base, 100 mL L−1 glacial acetic acid), 1 mL Hutner’s trace elements solution (50.0 g L−1 EDTA-Na2, 11.14 g L−1 H3BO3, 22.0 g L−1 ZnSO4·7H2O, 5.1 g L−1 MnCl2, 5.0 g L−1 FeSO4, 1.6 g L−1 CoCl2, 1.6 g L−1 CuSO4·5H2O, 1.1 g L−1 (NH4)6Mo7O24·4H2O)) for 1 L of medium, at 20 °C under cycles of medium light (30 V) for 12 h and dark for 12 h.
Labeling of C. reinhardtii with 5(6)-carboxyfluorescein
Microalgae samples were taken from the culture and transferred in Eppendorf tubes. The cell density was determined with a counting grid and adjusted to 3 × 106 cells mL−1. The samples were then centrifuged at 700 RCF for 5 min to afford the algae pellet. The supernatant (950 μL) was removed and PBS (950 μL) was added. Samples were vortexed, centrifuged at 700 RCF for 5 min, the supernatant (950 μL) was removed, and replaced by fresh PBS (950 μL). This washing procedure was repeated three times. For the last repetition, PBS (950 μL) was removed, the freshly prepared solution of 5(6)-Carboxyfluorescein N-hydroxysuccinimide (0.65 mM in PBS) was added and samples were vortexed. The samples were then incubated in a thermoshaker at 360 rpm for 40 min and at 700 rpm for 20 min at 25 °C. After incubation, samples were washed according to the following procedure: samples were first centrifuged at 700 RCF for 5 min. The supernatant (950 μL) was removed and replaced with fresh PBS (950 μL). Samples were vortexed and centrifuged again. This washing procedure was repeated six times.
Functionalization of C. reinhardtii with streptavidin
Microalgae samples were taken from the culture, strained, and transferred in Eppendorf tubes. The cell density was determined with a counting grid and adjusted to 1 × 106 cellsmL−1. The samples were then centrifuged at 700 RCF for 5 min to afford the algae pellet. The supernatant (950 µL) was removed and PBS (950 µL) was added. Samples were vortexed, centrifuged at 700 RCF for 5 min, the supernatant (950 µL) was removed and replaced by fresh PBS (950 µL). This washing procedure was repeated three times. For the last repetition, PBS (950 µL) was removed, the freshly prepared solution of biotin anchor 2 (0.65 mM in PBS) was added, and samples were vortexed. The samples were incubated in a thermoshaker at 360 rpm for 40 min and at 700 rpm for 20 min at 25 °C. After incubation, samples were washed according to the following procedure. Samples were first centrifuged at 700 RCF for 5 min. The supernatant (950 µL) was removed and replaced with fresh PBS (950 µL). Samples were vortexed and centrifuged again. This washing procedure was repeated six times. A freshly prepared solution of streptavidin-Alexa Fluor 488 conjugate (50 µg mL−1 in PBS) was added, the samples were vortexed, and then shaken in a thermoshaker at 360 rpm and 25 °C for 20 min. After incubation, samples were washed three times according to the procedure described above.
Microscopy imaging
Coverslips were coated with a 0.1% gelatin solution for 15 min and air dried. On drop of algae suspension was applied on the coverslip and the water was evaporated without the cells being dried out. A fixation solution (9% formaldehyde, 3% glutaraldehyde, 0.1% NaCl) was mixed 1:1 with Kuhl medium, placed on the cells, and left for 5 min at room temperature. The fixation solution was rinsed away with Kuhl medium and the sample was mounted with Prolong® Diamon Antifade Mountant. A Leica SP5 was used with the following parameters: argon laser, 20%; resolution, 8 bits; room temperature; objective, HC PL APO lambda blue 63.0 × 1.40 OIL UV; Leica AF acquisition software. For details on laser and gain see Supplementary Table 1.
Microscopy processing
Images were processed using Fiji (ImageJ) software by adjusting the brightness and contrast. All acquired images are 8-bit images with a full display range of 0–255. Overlay images have not been corrected after merging the channels. For details see Supplementary Table 2.
Instrument parameters for flow cytometry
For the detection of chlorophyll, a 488 nm laser was used with 695/40 emission filters. For the detection of Alexa Fluor 488, a 488 nm laser was used with 530/30 emission filters. For the detection of propidium iodide, a 561 nm laser was used with 610/20 emission filters.
Flow cytometry analysis of functionalization efficiency
Samples were centrifuged for 5 min at 2000 RCF. The supernatant was removed and replaced by fresh Kuhl Medium. A volume corresponding to two million cells (based on previous cell density measures) was placed to 5 mL tubes and the volume completed to a total volume of 600 μL with Kuhl medium and kept in the dark. The resulting cell suspensions were then analyzed on a BD LSR Fortessa SORP (BD Biosciences). A 488 nm laser was used for the excitation of Alexa Fluor 488 and a FITC (530/30) filter for the detection of the emitted fluorescence signal (peak area).
Flow cytometry analysis of cell viability
Samples were centrifuged for 5 min at 2000 RCF. The supernatant was removed and replaced by fresh Kuhl Medium. A volume corresponding to two million cells (based on previous cell density measures) was placed to 5 mL tubes and the volume completed to a total volume of 600 μL with Kuhl medium. Dead cells for positive control were heated to 90 °C for 20 min. Propidium iodide (2.5 mM in DMSO, 1.8 μL) was added, and the sample was vortexed. The resulting cell suspensions was then analyzed on a BD LSR Fortessa SORP (BD Biosciences). A 561 nm laser was used for the excitation of propidium iodide and a PE-Texas Red (610/20) filter for the detection of the emitted fluorescence signal (peak area).
Processing of flow cytometry data
All flow cytometry data recorded as described above were processed with FlowJo.
Synthesis of the ruthenium cofactor 9
Ruthenium cofactor 9 was prepared in situ by mixing solutions of [CpRu(MeCN)3]PF6 7 and biotinylated ligand 8 in a 1:1 ratio in DMF. Ligand 8 was prepared as described by Heinisch et al.34. ProCoumarin 4 and product 6 were prepared as described by Meggers et al.19.
Expression of streptavidin isoforms
Streptavidin isoforms were expressed in E. coli BL21(DE3) cells containing pET11b_SAV plasmids and purified as described elsewhere57. The number of free biotin-binding sites was determined by titration with biotin-4-fluorescein58.
Uncaging of proCoumarin 4 with the ADAse
The following stock solutions were prepared: (i) PBS (1×, pH 7.4) containing 50 mM Na2HPO4/NaH2PO4 and 0.9% NaCl, (ii) streptavidin isoforms (2 mM free biotin-binding sites in water), (iii) ruthenium cofactor 9 (1 mM; prepared by mixing 2 mM solutions of [CpRu(MeCN)3]PF6 7 and biotinylated ligand 8 in DMF in a 1:1 ratio), and (iv) coumarin substrate 4 (10 mM in water). Catalysis reactions were performed in flat 96-well plates at a reaction volume of 200 μL and a substrate concentration of 500 μM. PBS buffer was filled into a 96-well plate followed by the addition of streptavidin and ruthenium cofactor stock solutions. The resulting solution was incubated at room temperature for 10 min (to allow streptavidin to bind to the biotinylated cofactor). Substrate stock solution was added and the plate was sealed with an aluminum cover and incubated under the given conditions. An aliquot of the reaction mixture (3 μL) was then diluted with water (147 μL) in a new flat black 96-well plate. Fluorescence of the product 6 was determined in a plate reader at λex. = 395 nm/λem. = 460 nm and compared with a standard curve of the product (Supplementary Figure 6).
Functionalization of C. reinhardtii with the ADAse
Microalgae samples were taken from the culture, strained, and transferred in Eppendorf tubes. The cell density was determined with a counting grid and adjusted to 4 × 106 cells mL−1. The samples were then centrifuged at 2000 RCF for 5 min to afford the algae pellet. The supernatant (950 µL) was removed and PBS (950 µL) was added. Samples were vortexed, centrifuged at 2000 RCF for 5 min, the supernatant (950 µL) was removed and replaced by fresh PBS (950 µL). This washing procedure was repeated three times. For the last repetition, PBS (950 µL) was removed, the freshly prepared solution of biotin anchor 2 (0.65 mM in PBS) was added, and samples were vortexed. The samples were incubated in a thermoshaker at 360 rpm for 40 min and at 700 rpm for 20 min at 25 °C. A stock solution of the ruthenium cofactor 9 was prepared by incubating tris(acetonitrile)cyclopentadienylruthenium(II)hexafluorophosphate 7 (1 mM in DMF) and the biotinylated ligand 8 (1 mM in DMF) in a 1:1 ratio for 15 min. The artificial metalloenzyme was then assembled by incubating streptavidin (10 µM, free biotin-binding sites) with the ruthenium cofactor (2.5 µM) in PBS in a total volume of 180 µL for 15 min. This solution was then transferred to the cells in a total volume of 250 μL. Samples were shaken on a thermoshaker at 350 rpm and 25 °C for 20 min. After incubation, samples were washed according to the following procedure. Samples were first centrifuged at 2000 RCF for 5 min. The supernatant (250 μL) was removed and replaced with fresh PBS (950 μL). Samples were vortexed and centrifuged again. This washing procedure was repeated three times.
Optimized on-cell uncaging of the proCoumarin 4
Cells were first functionalized with the artificial allylic deallylase and washed as described above. ProCoumarin 4 (400 µM in Kuhl medium, 200 µL total volume) was added to the microalgae pellet. The samples were then incubated in the dark on a thermoshaker at 950 rpm and 25 °C for 15 h. Samples were then centrifuged at 2000 RCF for 5 min. The reaction supernatant was removed and filtered through a PTFE 0.2 μm syringe filter. The filtered reaction supernatant (30 μL) and PBS buffer (120 μL) were placed in a 96-well plate and analyzed on a plate reader (λex. = 395 nm, λem. = 460 nm).
Statistical analyses
Prism 7 was used for data analysis and representation. Statistical analysis was performed by one-way analysis of variance (ANOVA) with post hoc Tukey test (for multiple comparison) and P-values <0.05 were considered statistically significant. The results were deemed significant at 0.01 < *P ≤ 0.05, very significant at 0.001 < **P ≤ 0.01, extremely significant at 0.0001 < ***P ≤ 0.001 and extremely significant at ****P ≤ 0.0001.
Data availability
The authors declare that all data supporting the findings of this study are available either in the paper and in the Supplementary Information or from the authors upon reasonable request.
References
Bertozzi, C. R. A decade of bioorthogonal chemistry. Acc. Chem. Res. 44, 651–653 (2011).
Prescher, J. A. & Bertozzi, C. R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005).
Sletten, E. M. & Bertozzi, C. R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Boyce, M. & Bertozzi, C. R. Bringing chemistry to life. Nat. Methods 8, 638–642 (2011).
Debets, M. F. et al. Bioconjugation with strained alkenes and alkynes. Acc. Chem. Res. 44, 805–815 (2011).
Thirumurugan, P., Matosiuk, D. & Jozwiak, K. Click chemistry for drug development and diverse chemical-biology applications. Chem. Rev. 113, 4905–4979 (2013).
Grammel, M. & Hang, H. C. Chemical reporters for biological discovery. Nat. Chem. Biol. 9, 475–484 (2013).
Patterson, D. M., Nazarova, L. A. & Prescher, J. A. Finding the right (bioorthogonal) chemistry. ACS Chem. Biol. 9, 592–605 (2014).
King, M. & Wagner, A. Developments in the field of bioorthogonal bond forming reactions—past and present trends. Bioconjug. Chem. 25, 825–839 (2014).
Shih, H.-W., Kamber, D. N. & Prescher, J. A. Building better bioorthogonal reactions. Curr. Opin. Chem. Biol. 21, 103–111 (2014).
Li, J. & Chen, P. R. Development and application of bond cleavage reactions in bioorthogonal chemistry. Nat. Chem. Biol. 12, 129–137 (2016).
Streu, C. & Meggers, E. Ruthenium-induced allylcarbamate cleavage in living cells. Angew. Chem. Int. Ed. 45, 5645–5648 (2006).
Yusop, R. M., Unciti-Broceta, A., Johansson, E. M. V., Sánchez-Martín, R. M. & Bradley, M. Palladium-mediated intracellular chemistry. Nat. Chem. 3, 239–243 (2011).
Unciti-Broceta, A., Johansson, E. M. V., Yusop, R. M., Sánchez-Martín, R. M. & Bradley, M. Synthesis of polystyrene microspheres and functionalization with Pd0 nanoparticles to perform bioorthogonal organometallic chemistry in living cells. Nat. Protoc. 7, 1207–1218 (2012).
Weiss, J. T. et al. Extracellular palladium-catalysed dealkylation of 5-fluoro-1-propargyl-uracil as a bioorthogonally activated prodrug approach. Nat. Commun. 5, 3277 (2014).
Li, J. et al. Palladium-triggered deprotection chemistry for protein activation in living cells. Nat. Chem. 6, 352–361 (2014).
Weiss, J. T. et al. Development and bioorthogonal activation of palladium-labile prodrugs of gemcitabine. J. Med. Chem. 57, 5395–5404 (2014).
Sánchez, M. I., Penas, C., Vázquez, M. E. & Mascareñas, J. L. Metal-catalyzed uncaging of DNA-binding agents in living cells. Chem. Sci. 5, 1901–1907 (2014).
Völker, T., Dempwolff, F., Graumann, P. L. & Meggers, E. Progress towards bioorthogonal catalysis with organometallic compounds. Angew. Chem. Int. Ed. 53, 10536–10540 (2014).
Tonga, G. Y. et al. Supramolecular regulation of bioorthogonal catalysis in cells using nanoparticle-embedded transition metal catalysts. Nat. Chem. 7, 597–603 (2015).
Völker, T. & Meggers, E. Transition-metal-mediated uncaging in living human cells-an emerging alternative to photolabile protecting groups. Curr. Opin. Chem. Biol. 25, 48–54 (2015).
Tomás-Gamasa, M., Martínez-Calvo, M., Couceiro, J. R. & Mascareñas, J. L. Transition metal catalysis in the mitochondria of living cells. Nat. Commun. 7, 12538 (2016).
Rubio-Ruiz, B., Weiss, J. T. & Unciti-Broceta, A. Efficient palladium-triggered release of Vorinostat from a bioorthogonal precursor. J. Med. Chem. 59, 9974–9980 (2016).
Khan, I., Agris, P. F., Yigit, M. V. & Royzen, M. In situ activation of a doxorubicin prodrug using imaging-capable nanoparticles. Chem. Commun. 52, 6174–6177 (2016).
Indrigo, E., Clavadetscher, J., Chankeshwara, S. V., Lilienkampf, A. & Bradley, M. Palladium-mediated in situ synthesis of an anticancer agent. Chem. Commun. 52, 14212–14214 (2016).
Völker, T. & Meggers, E. Chemical activation in blood serum and human cell culture: improved ruthenium complex for catalytic uncaging of alloc-protected amines. Chembiochem 18, 1083–1086 (2017).
Vidal, C., Tomás-Gamasa, M., Destito, P., López, F. & Mascareñas, J. L. Concurrent and orthogonal gold(I) and ruthenium(II) catalysis inside living cells. Nat. Commun. 9, 1913 (2018).
Martínez-Calvo, M. et al. Intracellular deprotection reactions mediated by palladium complexes equipped with designed phosphine ligands. ACS Catal. 8, 6055–6061 (2018).
Jeschek, M. et al. Directed evolution of artificial metalloenzymes for in vivo metathesis. Nature 537, 661–665 (2016).
Srivastava, P., Yang, H., Ellis-Guardiola, K. & Lewis, J. C. Engineering a dirhodium artificial metalloenzyme for selective olefin cyclopropanation. Nat. Commun. 6, 7789 (2015).
Dydio, P. et al. An artificial metalloenzyme with the kinetics of native enzymes. Science 354, 102–106 (2016).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Okamoto, Y. et al. A cell-penetrating artificial metalloenzyme regulates a gene switch in a designer mammalian cell. Nat. Commun. 9, 1943 (2018).
Heinisch, T. et al. E. coli surface display of streptavidin for directed evolution of an allylic deallylase. Chem. Sci. 9, 5383–5388 (2018).
Stephan, M. T. & Irvine, D. J. Enhancing cell therapies from the outside in: cell surface engineering using synthetic nanomaterials. Nano Today 6, 309–325 (2011).
Lee, J. H. et al. Engineering novel cell surface receptors for virus-mediated gene transfer. J. Biol. Chem. 274, 21878–21884 (1999).
Hanson, S. R. et al. Tailored glycoproteomics and glycan site mapping using saccharide-selective bioorthogonal probes. J. Am. Chem. Soc. 129, 7266–7267 (2007).
Stephan, M. T., Moon, J. J., Um, S. H., Bersthteyn, A. & Irvine, D. J. Therapeutic cell engineering with surface-conjugated synthetic nanoparticles. Nat. Med. 16, 1035–1041 (2010).
Hsiao, S. C. et al. Direct cell surface modification with DNA for the capture of primary cells and the investigation of myotube formation on defined patterns. Langmuir 25, 6985–6991 (2009).
O’Brien, P. J., Elahipanah, S., Rogozhnikov, D. & Yousaf, M. N. Bio-orthogonal mediated nucleic acid transfection of cells via cell surface engineering. ACS Cent. Sci. 3, 489–500 (2017).
Grimm, A. R. et al. A whole cell E. coli display platform for artificial metalloenzymes: poly(phenylacetylene) production with a rhodium–nitrobindin metalloprotein. ACS Catal. 8, 2611–2614 (2018).
Merchant, S. S. et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318, 245–250 (2007).
Harris, E. H. Chlamydomonas as a model organism. Annu. Rev. Plant Physiol. Plant Mol. Biol. 52, 363–406 (2001).
Rosenberg, J. N., Oyler, G. A., Wilkinson, L. & Betenbaugh, M. J. A green light for engineered algae: redirecting metabolism to fuel a biotechnology revolution. Curr. Opin. Biotechnol. 19, 430–436 (2008).
Tran, M. et al. Production of unique immunotoxin cancer therapeutics in algal chloroplasts. Proc. Natl Acad. Sci. USA 110, E15–E22 (2013).
Schenck, T. L. et al. Photosynthetic biomaterials: a pathway towards autotrophic tissue engineering. Acta Biomater. 15, 39–47 (2015).
Chávez, M. N. et al. Towards autotrophic tissue engineering: photosynthetic gene therapy for regeneration. Biomaterials 75, 25–36 (2016).
Kerschgens, I. P. & Gademann, K. Antibiotic algae by chemical surface engineering. Chembiochem 19, 439–443 (2017).
Stavis, R. L. & Hirschberg, R. Phototaxis in Chlamydomonas reinhardtii. J. Cell Biol. 59, 367–377 (1973).
Miller, D. H., Lamport, D. T. A. & Miller, M. Hydroxyproline heterooligosaccharides in Chlamydomonas. Science 176, 918–920 (1972).
Sarkar, D. et al. Chemical engineering of mesenchymal stem cells to induce a cell rolling response. Bioconjug. Chem. 19, 2105–2109 (2008).
Terai, T. et al. Rational development of caged-biotin protein-labeling agents and some applications in live cells. Chem. Biol. 18, 1261–1272 (2011).
Huttunen, K. M., Raunio, H. & Rautio, J. Prodrugs—from serendipity to rational design. Pharmacol. Rev. 63, 750–771 (2011).
Mahato, R., Tai, W. & Cheng, K. Prodrugs for improving tumor targetability and efficiency. Adv. Drug Deliv. Rev. 63, 659–670 (2011).
Ueki, N. et al. Eyespot-dependent determination of the phototactic sign in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 113, 5299–5304 (2016).
Ghattas, W. et al. Receptor-based artificial metalloenzymes on living human cells. J. Am. Chem. Soc. 140, 8756–8762 (2018).
Köhler, V. et al. OsO4 streptavidin: a tunable hybrid catalyst for the enantioselective cis-dihydroxylation of olefins. Angew. Chem. Int. Ed. 50, 10863–10866 (2011).
Kada, G., Falk, H. & Gruber, H. J. Accurate measurement of avidin and streptavidin in crude biofluids with a new, optimized biotin–fluorescein conjugate. Biochim. Biophys. Acta 1427, 33–43 (1999).
Acknowledgements
We acknowledge the Center for Microscopy and Image Analysis (ZMB) and the Flow Cytometry Facility of the University of Zurich for training and maintenance of the instruments. We acknowledge the Swiss National Science Foundation as part of the NCCR Molecular Systems Engineering, the University of Basel and the University of Zurich for funding.
Author information
Authors and Affiliations
Contributions
M.S. and K.G. designed the study, M.S. performed the cell surface functionalization and the on-cell uncaging experiments. F.S. performed the synthesis of the catalysts and substrates, and screened the streptavidin mutants. M.S., F.S., T.R.W. and K.G. evaluated the data, wrote the manuscript, and have given approval for its final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Szponarski, M., Schwizer, F., Ward, T.R. et al. On-cell catalysis by surface engineering of live cells with an artificial metalloenzyme. Commun Chem 1, 84 (2018). https://doi.org/10.1038/s42004-018-0087-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0087-y
This article is cited by
-
Nanoparticle-modified microrobots for in vivo antibiotic delivery to treat acute bacterial pneumonia
Nature Materials (2022)
-
Engineering and emerging applications of artificial metalloenzymes with whole cells
Nature Catalysis (2021)
-
Biocompatibility and therapeutic potential of glycosylated albumin artificial metalloenzymes
Nature Catalysis (2019)
-
An artificial metalloenzyme biosensor can detect ethylene gas in fruits and Arabidopsis leaves
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.