Abstract
The acquisition of 14N NMR spectra in solid samples is challenging due to quadrupolar couplings with magnitudes up to several MHz. This nucleus is nonetheless important as it is involved in the formation of essential secondary structures in biological systems. Here we report the structural study of the atomic environment of amide functions in polypeptides using magic-angle spinning NMR spectroscopy of the ubiquitous 14N isotope. The cyclic undecapeptide cyclosporin, in which only four hydrogen atoms are directly bound to nitrogen atoms, is chosen for illustration. Structural details of different environments can be revealed without resorting to isotopic enrichment. The network of inter- and intra-residue dipolar couplings between amide 14N nuclei and nearby protons can be probed and mapped out up to a tunable cutoff distance. Density functional theory calculations of NMR quadrupolar interaction tensors agree well with the experimental evidence and allow the unambiguous assignment of all four non-methylated NH nitrogen sites and neighboring proton nuclei.
Similar content being viewed by others
Introduction
Solid-state NMR is an invaluable tool to obtain structural details of arbitrary materials with atomic resolution. Inhomogeneous interactions such as anisotropic chemical shifts, dipolar and quadrupolar couplings reveal a wealth of information on the local environment on an atomic scale1. In contrast to diffraction techniques that rely on long-range order, NMR can provide valuable information about amorphous systems. Furthermore, incoherent dynamic processes such as local motions and chemical exchange effects can be identified2. Compositional and positional disorder in materials may also be investigated3. In contrast to NMR studies in solution, where the linewidths increase with the size of the dissolved molecule, solid-state NMR may be applied to proteins and assemblies without any restriction on mass, even in their native biological environments. Heterogeneous matrices such as bone tissue are also suitable for analysis with this powerful technique4.
In applications to biomolecules, it is common practice to enrich the 13C and 15N isotopes. The former nucleus is only 1.1% abundant, the remaining 98.9% 12C nuclei with spin quantum number I = 0 being NMR silent. The 15N isotope has an even lower natural abundance (0.4%), however, the highly abundant 14N isotope (99.6%) can be observed by NMR. Unfortunately, observation of this nucleus with I = 1 is particularly challenging due to (i) the absence of a favorable central transition mI = + 1/2 ↔ mI = –1/2 that occurs in half-integer quadrupolar spins (I > 1) and (ii) the broadening due to quadrupolar couplings which affects the two observable single-quantum transitions mI = ±1 ↔ mI = 0 to both first and second order, resulting in inhomogeneously broadened lineshapes which typically span several MHz. Therefore, the possibility of acquiring 14N spectra routinely would clearly revolutionize the use of NMR spectroscopy in systems where isotopic enrichment of 15N nuclei is very challenging, exceedingly expensive, or indeed impossible.
Over the past decade or so, a number of experimental methods have been proposed to acquire 14N NMR spectra in the solid state5,6,7,8. The intrinsic low sensitivity and efficiency of these approaches has limited most applications to relatively simple chemical systems such as single amino acids like glycine, histidine or alanine. More challenging systems that have been studied by 14N NMR techniques include di- or tripeptides such as AspAla, AlaAlaGly or GlyGlyGly9,10,11. Other structural investigations have explored the different modes of self-assembly of nucleosides such as guanosine12,13,14. Applications have also been reported on pharmaceutical polymorphs, cocrystals and amorphous dispersions15,16,17,18. However, the acquisition of 14N spectra of large biomolecules such as the GB3 protein (6 kDa) has required the use of dynamic nuclear polarization (DNP) techniques19. We have recently introduced a double cross-polarization (DCP) transfer 1H → 14N → 1H for the indirect detection of 14N spectra20. We have shown the method to be straightforward to implement and robust with respect to sample-specific parameters, and efficient for CQ values up to about 3 MHz.
The calculation of NMR parameters based on density functional theory (DFT)21 has emerged as an invaluable tool for spectral interpretation and assignment in both solid-state and solution NMR22,23,24,25,26,27,28. In particular, the combination of DFT methods and solid-state NMR techniques has become known as NMR crystallography29,30,31,32,33,34. The possibility of computing both isotropic and anisotropic interactions expected for a molecular model can be used to assess the agreement between experimental evidence and different structural hypotheses. These computational tools are particularly useful for the prediction of 14N NMR spectra since the wide frequency dispersion of the resonances and their inhomogeneous lineshapes (so-called “powder patterns”) are determined by both anisotropic chemical shifts and second-order quadrupolar interactions.
In this paper, we present the structural study of a biologically-relevant polypeptide by means of 14N NMR spectroscopy at a very fast magic-angle spinning (MAS) rate νR = 100 kHz35,36,37,38,39. In particular, the non-methylated amide nitrogen sites in a powdered polycrystalline sample of cyclosporin, a well-known immunosuppressing drug40,41, are investigated by means of DCP 1H-14N-1H through-space correlation experiments. This 1.2 kDa cyclic undecapeptide represents one of the largest molecules so far investigated by means of 14N NMR methods and should open the way for solid-state studies of proteins of several tens of kDa. Through-space dipolar couplings between 14NH nitrogen atoms and directly-bound 1HN protons can be readily identified. Spatial proximities between proton-carrying amide nitrogen atoms and remote backbone or side-chain protons of surrounding residues can also be determined by tuning the length of the polarization-transfer interval. The full network of heteronuclear 14N-1H dipolar couplings can thus be characterized. DFT calculations of chemical shifts and quadrupolar couplings support the experimentally observed spectral features. Precious indications about the mean frequencies and inhomogeneous lineshapes of 14N resonances for different environments can be drawn from computations of NMR parameters. In particular, for a given 14N environment, the mean frequencies are predominantly determined by the magnitude of the quadrupolar constant CQ, via the isotropic part of the second-order quadrupolar interaction, while isotropic chemical shifts are almost negligible.
Results
Solid-state NMR
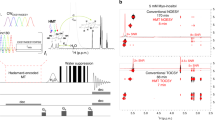
The structure of cyclosporin reported in the literature42, initially obtained by X-ray diffraction techniques, was optimized with planewave-pseudopotential DFT methods by means of the VASP code43,44,45. The resulting geometry is shown in Fig. 1(a). The sequence of residues in this undecapeptide is shown in (b). The four H-bonds (three transannular and one of 3 → 1 type) involving the four non-methylated NH sites are indicated by dashed lines. These H-bonds result in a substantially rigid conformation in both crystalline state and in solution42. The proton spectrum of a powdered sample of cyclosporin acquired with a 90°-τ-180°-τ- echo in a field B0 = 20.0 T (ν0 (1H) = 850.0 MHz, ν0 (14N) = 61.4 MHz) and at a spinning rate νR = 100 kHz (rotor period τR = 10 μs) is shown in Fig. 2(a). High fields and ultra-fast spinning rates improve the resolution of proton spectra and allow the separation of distinct environments by virtue of the increased spread of the chemical shifts and more efficient averaging of homonuclear dipolar couplings. Figure 1(a) also shows, in red and vertically displaced for clarity, a spectrum obtained after initial selective presaturation46 of the aliphatic region (dashed black arrow). The expansion shows how presaturation leaves the four amide HN protons that appear in the range 7–10 ppm substantially unperturbed. Shoulders associated with all three peaks towards lower frequencies suggest that the sample may be inhomogeneous and possibly contain a minor phase or polymorph. Attenuation of the intense signals of aliphatic protons that are not directly bound to any N atoms allows one to acquire the 1H-14N correlation spectrum as shown in Fig. 2(b), which is substantially free of any t1 noise artifacts that appear due to imperfect cancellation of intense signals if one does not use any presaturation. This spectrum was acquired with contact pulses for cross polarization of length τCP = 18τR = 180 μs, so that transfers over short distances in non-methylated NH amide groups are favored. The vertical 14N (ω1) dimension shows the full spectral width of 1/τR = νR = 100 kHz (1629 ppm for 14N at B0 = 20.0 T). It is worth bearing in mind that the 4 amide HN protons represent a fraction of only 4/111 = 3.6% with respect to the full “bath” of 111 protons in this sample, without considering possible water molecules involved in hydration. This is much lower than in 14N MAS studies previously reported which utilized samples such as glycine, alanine or histidine, where the analogous ratios are 3/5, 3/7 and 4/9 (60, 43 and 44%), respectively. Nonetheless, a clear correlation between the amide protons and nitrogen sites can be identified in Fig. 2(b) near ca. 300 ppm in the vertical 14N dimension.
Structure of cyclosporin and scheme depicting the sequence of residues. a DFT-optimized structure of cyclosporin. Nitrogen, oxygen, carbon and proton atoms are shown in blue, red, grey and white, respectively. Hydrogen bonds are represented by dashed lines. b Schematic representation of the structure of (a) where the relevant residues are shown. H-bonds are depicted by dashed lines. The two parallel dashed lines indicate an antiparallel β-pleated sheet structure. The D-Ala8 residue has an absolute (R) configuration, and there is a cis-amide linkage between Me-Leu9 and Me-Leu10
1H 1D and 1H-14N 2D NMR spectra of cyclosporin in the solid state. a Proton MAS NMR spectrum of cyclosporin acquired with a rotor-synchronized 90°-τ-180°-τ-echo in a static field B0 = 20.0 T (850 MHz for 1H, 61.4 MHz for 14N) at a spinning rate of νR = 100 kHz (black). In red, proton spectrum obtained with selective presaturation of the methyl region by means of a weak rf-field ν1 = 125 Hz applied for a duration τP = 1 s at the frequency indicated by a dashed vertical arrow. The black and red spectra are displaced vertically for clarity. The four amide protons HN in the 7–10 ppm region are highlighted in the expansion. b Two-dimensional 1H → 14N → 1H double cross-polarization (DCP) spectrum of cyclosporin acquired with selective presaturation and short contact pulses of length τCP = 18τR = 180 μs. 400 transients were averaged for each of the 256 rotor-synchronized t1 increments with a recovery delay of 0.2 s, resulting in a total experimental time of ca. 34 h. No line broadening was applied in (a), whereas 100 and 500 Hz were applied to direct and indirect dimensions of (b), respectively
Short-range correlations
Figure 3(a) shows an expansion of the 2D spectrum where three correlation peaks can be easily resolved in the horizontal 1H dimension, whereas the four NH signals appear substantially overlapped in the vertical 14N dimension. In order to unravel the overlapping 2D correlations and achieve spectral assignment, DFT calculations of NMR parameters were performed using the CASTEP code47. Figure 3(b) shows the resulting simulated 2D spectrum based on proton and nitrogen shifts as well as quadrupolar couplings predicted by CASTEP. The signals from separate sites are highlighted in different colors (Abu2 in green, Ala8 in orange, Ala7 in red, and Val5 in blue.) A few vertical cross sections were extracted from the 2D spectra in (a) and (b) as indicated by arrows. Considering that homogeneous signal decay does not allow subtle inhomogeneous features to be resolved, the simulated linewidths agree well with the experimental cross sections. More specifically, (i) the four NH correlations are indeed expected to overlap in the vertical 14N dimension and (ii) the size of the second-order quadrupolar broadening is correctly estimated to ca. 5 kHz by DFT. The experimental vertical cross sections do not allow one to resolve any second-order features of the 14N lineshapes. This is consistent with our earlier work20 and can be attributed to (i) the non-uniform efficiency of the CP transfer over the inhomogeneous lineshapes, and (ii) homogeneous signal decay in the evolution interval.
Comparison between experimental and DFT-calculated 1H-14N 2D correlation spectra. a Expansion of the amide region of the 2D spectrum of Fig. 2(b). The arrows indicate the positions in the proton spectrum where the 14N cross sections shown below were extracted. b Simulated spectrum analogous to that of (a) resulting from isotropic chemical shifts and quadrupolar parameters calculated by DFT methods. The 2D correlations and extracted 14N second-order lineshapes of the four distinct NH sites are highlighted with different colors. The simulated numbers of points were 32 and 128, for direct and indirect dimensions, respectively. The corresponding line broadenings were 350 and 300 Hz
Long-range correlations
Figure 4 shows a 1H-14N correlation spectrum acquired with longer CP contact pulses of length τCP = 60τR = 600 μs. When compared to that of Fig. 2(a), this clearly shows long-distance correlations between the NH sites and remote protons. This 1H-14N correlation spectrum, in analogy to routine 1H-15N correlation techniques that can be used for I = 1/2 nuclei, supplies 2D contour ridges which can map interatomic proximities beyond one-bond connectivities. The molecular structure of Fig. 1, optimized by DFT methods, can be very helpful to assist the assignment of through-space correlations. By inspection of the internuclear distances in this structure, one finds that the 14N nuclei of NH groups have intra-residue dipole-dipole couplings with HN, Hα, Hβ and Hγ side-chain protons of ca. 8, 1, 1 and 0.5 kHz, respectively. Other inter-residue couplings ranging from ca. 0.5 down to 0.15 kHz may also be identified. Relevant intra- and inter-residue distances shorter than 3.3 Å are given in Table 1. The assignment of long-range correlations based on the dipolar couplings in Table 1, supported by proton chemical shifts calculated by DFT, is highlighted by dashed magenta rectangles in the experimental spectrum of Fig. 4. The Hα protons are clearly distinguished near 5 ppm in the horizontal 1H dimension. The strongest correlations are observed for the most shielded methyl protons of Val5, which are in proximity of both intra- (Val5) and inter-residue (Ala7) nitrogen sites. The (Hβ)3 groups of Ala7 and Ala8 give rise to strong correlations with intra-residue NH nitrogen sites. Other clear correlations can be assigned to (Hβ)2 of Abu2 and Hγ of Bmt1. The NCH3 protons of Leu4 can transfer polarization to the NH group of Val 5, the NCH3 protons of Val11 are in contact with the NH sites of both Ala7 and Ala8, and the NCH3 protons of Sar3 are in spatial proximity with the NH site of Abu2. Also in agreement with the internuclear distances extracted from the structure optimized by DFT, one does not observe any correlations between the four NH sites and the side chains of Leu4, Leu9 and Leu10. No evidence could be found for any methylated NCH3 sites in the vertical 14N dimension. We ascribe this finding to poor polarization transfer, possibly due to inefficient spin locking for 14N sites with CQ values that should be in excess of ca. 4 MHz according to DFT. The full set of relevant dipolar proton-nitrogen couplings utilized for the assignment of the spectrum of Fig. 4 is summarized in Table 1.
Long-range correlation 1H-14N 2D NMR spectrum. Two-dimensional 1H → 14N → 1H DCP spectrum analogous to that of Fig. 2(b) but acquired with longer contact pulses of length τCP = 60τR = 600 μs. One-bond distance correlations for the four NH sites are assigned according to the DFT calculations and highlighted in red. Long-range correlations between 14N and nearby intra- and inter-residue protons are highlighted in magenta. These were assigned according to the dipolar contacts in Table 1. In all, 420 transients were averaged for each of the 64 rotor-synchronized t1 increments with a recovery delay of 0.2 s, resulting in a total experimental time of ca. 9 h
The atomic details of the four NH sites are shown in Fig. 5. The correlation between the experimental 1H shifts of Fig. 4, assigned on the basis of internuclear 1H-14N distances of Table 1, and those calculated by CASTEP, is shown in Fig. 6. A correlation coefficient R2 = 0.995 is obtained, with a root mean square error of 0.3 ppm, thus further supporting our assignment and the good agreement between our experimental and calculated data.
1H-14N dipolar networks of non-methylated amide groups in cyclosporin. a Inter- and intra-residue proton nuclei within 3.3 Å of the NH nitrogen-14 site of Abu2 as extracted from the DFT optimized structure of cyclosporin. b–d Analogous environments for Val5 (b), Ala7 (c) and Ala8 (d) residues, respectively. Nitrogen and proton atoms are shown in blue and dark grey, respectively, whereas all other atoms are in transparent light grey for clarity
Correlation between experimental and DFT-calculated 1H isotropic chemical shifts. The experimental 1H shifts of the 2D spectrum of Fig. 4, assigned on the basis of the 1H-14N internuclear distances of Table 1, are plotted versus those calculated by CASTEP. A correlation coefficient R2 = 0.995 is obtained, with root mean square error of 0.3 ppm. The error bars of the experimental data are set to 2×lb = 600 Hz. Numerical values are given in Supplementary Table 1
Quadrupolar parameters
Figure 7 presents simulated 14N spectra based on the chemical shifts and quadrupolar parameters calculated by DFT. The NH and NCH3 sites are shown in blue and red, respectively. Analogous spectra where only the isotropic shift interactions were taken into account appear on the right-hand side, displayed by dashed lines with the same color coding. The anisotropic part of the chemical shift was not included since it is fully averaged out by fast spinning (νR = 100 kHz). These simulations allow one to appreciate how the second-order quadrupolar interaction does not only produce inhomogeneous broadening, but also results in significant isotropic shifts of the centers of gravity (first moments) of the powder patterns. Importantly, the DFT calculations show that the NH and NCH3 amide environments tend to fall in two distinct spectral regions. The NMR parameters for all 11 14N nuclei and for the relevant 1H nuclei of the 4 NH sites are given in Table 2. All non-methylated NH sites feature very similar electric field gradients (EFGs), with a narrow dispersion of quadrupolar coupling constants CQ = –3.64 ± 0.12 MHz. Similarly, the NCH3 sites are associated with narrowly distributed quadrupolar couplings CQ = –4.27 ± 0.09 MHz. Furthermore, the asymmetry parameters in the two environments are also distinct, with average values ηQ = 0.38 ± 0.04 for NH sites and ηQ = 0.13 ± 0.02 for NCH3 sites.
Simulations of anisotropic 14N lineshapes. One-dimensional 14N NMR spectra of cyclosporin based on the chemical shift and quadrupolar parameters calculated by DFT. NH and NCH3 sites are shown in blue and red, respectively. Analogous spectra which consider only the isotropic shifts are shown by dashed lines with the same colour coding. The vertical scale for these latter spectra was reduced by a factor 20 to compensate for the narrow linewidths. The line broadening for anisotropic and isotropic simulations were 300 and 100 Hz, respectively. Horizontal bars in the lower part of the figure indicate the ranges of isotropic displacements due to the quadrupolar and chemical shift interactions, in continuous and dashed lines, respectively, assuming the averaged NMR parameters in Table 2. The overall difference in displacement between the average NH and NCH3 sites due to both shift and quadrupolar interactions is shown by a black bar
The isotropic part of the second-order quadrupolar interaction is proportional to CQ2/ν0 and results in shifts of the first moments of the inhomogeneous lineshapes that, for low-γ nuclei with large CQ values such as 14N, can significantly exceed the range of the isotropic chemical shifts. For the two single-quantum transitions mI = ±1 ↔ mI = 0 of a I = 1 spin, the isotropic quadrupolar shift is:48
For the average quadrupolar parameters calculated by DFT for cyclosporin in Table 2, Eq. (1) yields νiso(Q) = 16.9 and 22.4 kHz at 20.0 T for the NH and NCH3 environments, respectively. In other words, the isotropic quadrupolar interaction leads to a relative displacement of the 14N signals of the NH and NCH3 sites by 5.5 kHz. In contrast, if one considers the isotropic part of the chemical shift interaction (which determines the dispersion of 15N signals in solids and solutions), one finds that the average isotropic shifts of the NH and NCH3 sites in cyclosporin are δiso = 6.6 and 6.2 kHz, i.e., a difference of only ca. 0.4 kHz. The ranges of the average isotropic displacements for 14N sites in cyclosporin are indicated by horizontal continuous and dashed bars in Fig. 7 for the shifts νiso(Q) and δiso, respectively, with the same color coding. A continuous black bar indicates the overall difference in displacement between the average NH and NCH3 sites due to both shift and quadrupolar interactions. These considerations imply that, since similar 14N environments lead to narrowly distributed CQ values, one can expect extensive overlap of lineshapes for chemically similar N environments (say, all NH sites), but a good separation between different chemical environments (say, between NH and NCH3 sites) by virtue of their different quadrupolar interactions. This observation is promising for future applications, as the resolution required to distinguish similar chemical environments can be supplied, as in the case at hand, in the horizontal 1H dimension of the 2D spectrum, where the resolution is expected to improve further at higher fields and faster spinning rates. Furthermore, these results suggest that the general lower accuracy of DFT-calculated isotropic shifts of 15N atoms when compared to that of 1H and 13C species49,50, may be reconsidered when dealing with 14N isotopes in the solid state, as both isotropic displacement νiso(Q) and anisotropic linewidth due to second-order quadrupolar interactions may largely exceed differences in isotropic chemical shifts.
Experimental 14N linewidths
The 2D spectrum shown in Figs. 2(b), 3(a) results from the acquisition of 256 rotor-synchronized t1 time increments. The indirect acquisition time is therefore t1max = 256 × 10 μs = 2.56 ms, which allows for an indirect digital resolution of ca. 400 Hz. However, the inhomogeneous 14N lineshapes experimentally observed are on the order of 5 kHz, in full agreement with our DFT calculations. Considering that second-order features are difficult to identify due to non-uniform cross-polarization transfer for different crystallite orientations, homogeneous losses and the extensive overlap expected for similar chemical environments, one may conclude that the experimental times for the acquisition of spectra such as that of Fig. 2(b) may be substantially reduced without any loss of structural information. In order to prove this concept, the indirect 14N projection of the spectrum of Fig. 2(b) is compared in Supplementary Figure 1 to that resulting from Fourier transformation of only 160 and 128 t1 time increments. One can appreciate that the signal intensity is conserved, while the reduction of digital resolution has little effect on the observed projections. Moreover, the identical resolution in the direct 1H dimension grants for substantially identical 2D correlation contours. Further experimental proof of this concept is given by the spectrum of Fig. 4, where only 64 t1 time increments were acquired.
Discussion
This paper presents the structural study of amide functions in polypeptides like cyclosporin by means of solid-state 14N NMR spectroscopy. Double cross polarization (DCP) 1H → 14N → 1H offers sufficient sensitivity to detect all four NH environments in a sample where protons bound to nitrogen atoms constitute less than 4% of the full proton bath. The use of long contact pulses allows one to monitor spatial proximities beyond immediate neighbors. Therefore, structural details similar to those derived from conventional solution-state NMR techniques on 15N-enriched samples can be obtained in natural 14N abundance in the solid state. Spurious signals rising from the large proton bath can be efficiently suppressed by weak selective presaturation of the aliphatic region. Depending on the desired resolution in the 14N dimension, satisfactory spectra can be obtained in about 10 h when spinning at 100 kHz at 20 T. The structural details unveiled by these experiments are in agreement with interatomic distances obtained with DFT methods. The full set of dipolar 14N-1H couplings involving amide nitrogen nuclei can be ascertained. Calculations of chemical shifts and quadrupolar couplings agree very well with the experimental evidence. Some overlap is expected and indeed observed for the four NH environments. The seven methylated NCH3 sites are also predicted to overlap in part. The signals of the NH and NCH3 sites, however, do not overlap because of distinct quadrupolar interactions that result in different second-order isotropic shifts. The use of very high fields above 23 T (above 1 GHz for 1H, i.e., above 72 MHz for 14N) would lead to a significant reduction of second-order quadrupole interactions, therefore lead to narrower signals, and possibly allow indirect detection of methylated NCH3 sites by our DCP method. Furthermore, fast spinning beyond 100 kHz would further reduce broadening by proton-proton couplings and lead to line-narrowing in the 1H dimension of the 2D correlation spectra. This study should open the way for solid-state 14N NMR studies of proteins, nucleic acids and other biomolecules in natural abundance.
Methods
Solid-state NMR
The sample of cyclosporin A was purchased from Sigma-Aldrich and used without further purification. All DCP MAS spectra were recorded at the CEMHTI CNRS laboratory in Orleans, on a wide-bore Bruker 850 spectrometer (20.0 T) with an Avance-III console, using 0.7 mm ZrO2 rotors containing ca. 1 mg cyclosporin spinning at νR = 100 kHz in a double-resonance MAS probe. The proton rf-field strength was ν1 = 179 kHz (90° pulse width τp = 1.4 μs.) The constant rf-field strengths during both 1H → 14N → 1H cross-polarization intervals were ν1(1H) = 53 and ν1(14N) = 83 kHz, whereas a weak rf field ν1(1H) = 125 Hz was utilized for selective presaturation during an interval τp = 1 s. The lengths of the contact pulses were τcp = 180 and 600 μs, for the spectra of Figs. 2 and 4, respectively. Recovery delays of 0.2 and 1 s were used for experiments with and without presaturation, respectively. Phase cycling for coherence pathway selection was used on the 14N channel only, with phases ϕ1 = {0 2}, ϕ2 = {0 0 2 2} for the first and second contact pulses in the 14N channel, respectively, with a receiver phase ϕrec = {0 2 2 0}. The chemical shifts were referenced with respect to adamantane for 1H and NH4Cl for 14N. The magic angle was adjusted using KBr as usual.
DFT calculations
All DFT calculations were performed at the gamma point using the PBE51 functional and a planewave energy cutoff of 50 Ry. The geometry optimization was performed with VASP 5.4.143,44,45 on a single molecule in a 25 × 25 × 25 Å cubic cell, requiring forces acting on all atoms to be smaller than 0.02 eV/Å. The electronic cores were described with the projector augmented wave method52. The resulting structure is superposed onto that obtained by X-ray diffraction in Supplementary Figure 2. NMR parameters were computed for the optimized geometries using CASTEP47 as implemented in the Materials Studio suite, version 8.0, available on the national supercomputing facility CINES. The gauge-including projector augmented wave (GIPAW) algorithm was employed to reconstruct all-electron wave functions in the presence of a magnetic field53,54. Cartesian coordinates of the optimized structure of cyclosporin utilized in this study and all calculated NMR parameters are given in Supplementary Tables 2 and 3.
NMR simulations
The simulations of the NMR spectra based on DFT parameters were carried out with Simpson55 and involved rotor-synchronized detection of the FID assuming an initial state comprising only in-phase coherence ρ(0) = Ix. Since the choice of reference shieldings is arbitrary, the offset of the carrier frequency νrf in these simulations was adapted to match the experimental results. The simulated 2D spectrum of Fig. 3(b) represents pure adsorptive ideal correlations between ideal 14N and 1H lineshapes, and results from the sum of four individual 2D spectra associated with each of the four HN pairs. Each of these individual 2D spectra was constructed by forming the product Re[S(ω1)] Re[S(ω2)], where the column vector S(ω1) is the spectrum for an ideal 14N second-order quadrupolar lineshape simulated with Simpson assuming NMR parameters calculated with CASTEP, and the row vector S(ω2) is a Lorentzian lineshape simulated with Simpson and associated with a 1H spin with an isotropic shift given by CASTEP calculations.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Duer, M. J. Solid-State NMR Spectroscopy: Principles and Applications. (Blackwell Science, UK, 2002).
Ernst, R. R., Bodenhausen, G. & Wokaun, A. Principles of Nuclear Magnetic Resonance in One and Two Dimensions. (Clarendon Press, Oxford, 1987).
Mitchell, M. R., Carnevale, D., Orr, R., Whittle, K. R. & Ashbrook, S. E. Exploiting the chemical shielding anisotropy to probe structure and disorder in ceramics: 89Y MAS NMR and first-principles calculations. J. Phys. Chem. C. 116, 4273–4286 (2012).
Chow, W. Y. et al. NMR spectroscopy of native and in vitro tissues implicates polyADP ribose in biomineralization. Science 344, 742–746 (2014).
Gan, Z. Measuring amide nitrogen quadrupolar coupling by high-resolution 14N/13C NMR correlation under magic-angle spinning. J. Am. Chem. Soc. 128, 6040–6041 (2006).
Cavadini, S., Lupulescu, A., Antonijevic, S. & Bodenhausen, G. Nitrogen-14 NMR spectroscopy using residual dipolar splittings in solids. J. Am. Chem. Soc. 128, 7706–7707 (2006).
O’Dell, L. A. Direct detection of nitrogen-14 in solid-state NMR spectroscopy. Prog. Nucl. Magn. Reson. Spectrosc. 59, 295–318 (2011).
Cavadini, S. Indirect detection of nitrogen-14 in solid-state NMR spectroscopy. Prog. Nucl. Magn. Reson. Spectrosc. 56, 46–77 (2010).
Tatton, A. S., Bradley, J. P., Iuga, D. & Brown, S. P. 14N–1H Heteronuclear multiple-quantum correlation magic-angle spinning NMR spectroscopy of organic solids. Z. Phys. Chem. 226, 1187–1203 (2012).
Cavadini, S., Abraham, A., Ulzega, S. & Bodenhausen, G. Evidence for dynamics on a 100 ns time scale from single- and double-quantum nitrogen-14 NMR in solid peptides. J. Am. Chem. Soc. 130, 10850–10851 (2008).
Jarvis, J. A., Haies, I. M., Williamson, P. T. F. & Carravetta, M. An efficient NMR method for the characterisation of 14N sites through indirect 13C detection. Phys. Chem. Chem. Phys. 15, 7613–7620 (2013).
Webber, A. L. et al. Identifying guanosine self assembly at natural isotopic abundance by high-resolution 1H and 13C solid-state NMR spectroscopy. J. Am. Chem. Soc. 133, 19777–19795 (2011).
Reddy, G. N. M., Marsh, A., Davis, J. T., Masiero, S. & Brown, S. P. Interplay of noncovalent interactions in ribbon-like guanosine self-assembly: an NMR crystallography study. Cryst. Growth Des. 15, 5945–5954 (2015).
Reddy, G. N. M., Malon, M., Marsh, A., Nishiyama, Y. & Brown, S. P. Fast magic-angle spinning three-dimensional NMR experiment for simultaneously probing H—H and N—H proximities in solids. Anal. Chem. 88, 11412–11419 (2016).
Tatton, A. S. et al. Probing intermolecular interactions and nitrogen protonation in pharmaceuticals by novel 15N-edited and 2D 14N-1H solid-state NMR. Cryst. Eng. Comm. 14, 2654–2659 (2012).
Tatton, A. S. et al. Probing hydrogen bonding in cocrystals and amorphous dispersions using 14N–1H HMQC solid-state NMR. Mol. Pharm. 10, 999–1007 (2013).
Veinberg, S. L. et al. Natural abundance 14N and 15N solid-state NMR of pharmaceuticals and their polymorphs. Phys. Chem. Chem. Phys. 18, 17713–17730 (2016).
Maruyoshi, K. et al. Identifying the intermolecular hydrogen-bonding supramolecular synthons in an indomethacin–nicotinamide cocrystal by solid-state NMR. Chem. Commun. 48, 10844–10846 (2012).
Jarvis, J. A. et al. Measurement of 14N quadrupole couplings in biomolecular solids using indirect-detection 14N solid-state NMR with DNP. Chem. Commun. 53, 12116–12119 (2017).
Carnevale, D., Ji, X. & Bodenhausen, G. Double cross polarization for the indirect detection of nitrogen-14 nuclei in magic angle spinning NMR spectroscopy. J. Chem. Phys. 147, 184201 (2017).
Parr, R. G. & Yang, W. Density-Functional Theory of Atoms and Molecules. (Oxford Univ. Press, Oxford, 1989).
Bagno, A., Rastrelli, F. & Saielli, G. Predicting 13C NMR spectra by DFT calculations. J. Phys. Chem. A 107, 9964–9973 (2003).
Bonhomme, C. et al. First-principles calculation of NMR parameters using the gauge including projector augmented wave method: a chemist’s point of view. Chem. Rev. 112, 5733–5779 (2012).
Ashbrook, S. E. & McKay, D. Combining solid-state NMR spectroscopy with first-principles calculations – a guide to NMR crystallography. Chem. Commun. 52, 7186–7204 (2016).
Carnevale, D., del Amo, V., Philp, D. & Ashbrook, S. E. Detecting solid-state reactivity in 10-hydroxy-10,9-boroxophenanthrene using NMR spectroscopy. Tetrahedron 66, 6238–6250 (2010).
Castro, M. et al. Molecular modeling, multinuclear NMR, and diffraction studies in the templated synthesis and characterization of the aluminophosphate molecular sieve STA-2. J. Phys. Chem. C. 114, 12698–12710 (2010).
Carnevale, D., Ashbrook, S. E. & Bodenhausen, G. Solid-state NMR measurements and DFT calculations of the magnetic shielding tensors of protons of water trapped in barium chlorate monohydrate. RSC Adv. 4, 56248–56258 (2014).
Banach, E., Invernizzi, C., Baudin, M., Neier, R. & Carnevale, D. Columnar self-assembly of N,N′,N′′-trihexylbenzene-1,3,5-tricarboxamides investigated by means of NMR spectroscopy and computational methods in solution and the solid state. Phys. Chem. Chem. Phys. 19, 5525–5539 (2017).
Harris, R. K. NMR crystallography: the use of chemical shifts. Solid State Sci. 6, 1025–1037 (2004).
Elena, B., Pintacuda, G., Mifsud, N. & Emsley, L. Molecular structure determination in powders by NMR crystallography from proton spin diffusion. J. Am. Chem. Soc. 128, 9555–9560 (2006).
Taulelle, F. Fundamental Principles of NMR Crystallography (eMagRes, John Wiley & Sons, 13 Ltd, 2007).
Salager, E., Stein, R. S., Pickard, C. J., Elena, B. & Emsley, L. Powder NMR crystallography of thymol. Phys. Chem. Chem. Phys. 11, 2610–2621 (2009).
Reddy, G. N. M. et al. An NMR crystallography study of the hemihydrate of 2′, 3′-O-isopropylidineguanosine. Solid. State Nucl. Magn. Reson. 65, 41–48 (2015).
Tatton, A. S. et al. Improving confidence in crystal structure solutions using NMR crystallography: the case of β-Piroxicam. Cryst. Growth Des. 18, 3339–3351 (2018).
Kobayashi, T. et al. Study of intermolecular interactions in the corrole matrix by solid-state NMR under 100 kHz MAS and theoretical calculations. Angew. Chem. Int. Ed. 52, 14108–14111 (2013).
Lamley, J. M. et al. Solid-state NMR of a protein in a precipitated complex with a full-length antibody. J. Am. Chem. Soc. 136, 16800–16806 (2014).
Nishiyama, Y. et al. Studies of minute quantities of natural abundance molecules using 2D heteronuclear correlation spectroscopy under 100 kHz MAS. Solid. State Nucl. Magn. Reson. 66–67, 56–61 (2015).
Sternberg, U. et al. 1H line width dependence on MAS speed in solid state NMR – comparison of experiment and simulation. J. Magn. Reson. 291, 32–39 (2018).
Zhang, R., Mroue, K. H. & Ramamoorthy, A. Proton-based ultrafast magic angle spinning solid-state NMR spectroscopy. Acc. Chem. Res. 50, 1105–1113 (2017).
Wenger, R. M. Synthesis of cyclosporine. Total syntheses of ‘cyclosporin A‘ and ‘cyclosporin H‘, two fungal metabolites isolated from the species Tolypocladium inflatum GAMS. Helv. Chim. Acta 67, 502–525 (1984).
Eléna, B., Hediger, S. & Emsley, L. Correlation of fast and slow chemical shift spinning sideband patterns under fast magic-angle spinning. J. Magn. Reson. 160, 40–46 (2003).
Loosli, H.-R. et al. Peptide conformations. Part 31. The conformation of cyclosporin a in the crystal and in solution. Helv. Chim. Acta 68, 682–704 (1985).
Kresse, G. & Hafner, J. Ab initio molecular dynamics for liquid metals. Condens. Matter Mater. Phys. 47, 558–561 (1993).
Kresse, G. & Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Comput. Mater. Sci. 6, 15–50 (1996).
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169–11186 (1996).
Robertson, A. J., Pandey, M. K., Marsh, A., Nishiyama, Y. & Brown, S. P. The use of a selective saturation pulse to suppress t1 noise in two-dimensional 1H fast magic angle spinning solid-state NMR spectroscopy. J. Magn. Reson. 260, 89–97 (2015).
Segall, M. D. et al. First-principles simulation: ideas, illustrations and the CASTEP code. J. Phys. Condens. Matter 14, 2717–2744 (2002).
Samoson, A. Satellite transition high-resolution NMR of quadrupolar nuclei in powders. Chem. Phys. Lett. 119, 29–32 (1985).
Zhu, T., He, X. & Zhang, J. Z. H. Fragment density functional theory calculation of NMR chemical shifts for proteins with implicit solvation. Phys. Chem. Chem. Phys. 14, 7837–7845 (2012).
Swails, J., Zhu, T., He, X. & Case, D. A. AFNMR: automated fragmentation quantum mechanical calculation of NMR chemical shifts for biomolecules. J. Biomol. NMR 63, 125–139 (2015).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Kresse, G. & Joubert, D. From ultrasoft pseudopotentials to the projector augmented-wave method. Phys. Rev. B 59, 1758–1775 (1999).
Pickard, C. J. & Mauri, F. All-electron magnetic response with pseudopotentials: NMR chemical shifts. Phys. Rev. B 63, 245101 (2001).
Yates, J. R., Pickard, C. J. & Mauri, F. Calculation of NMR chemical shifts for extended systems using ultrasoft pseudopotentials. Phys. Rev. B 76, 024401 (2007).
Bak, M., Rasmussen, J. T. & Nielsen, N. C. SIMPSON: a general simulation program for solid-state NMR spectroscopy. J. Magn. Reson. 147, 296–330 (2000).
Acknowledgements
The authors are indebted to Dr Vincent Sarou-Kanian and Dr Franck Fayon (CNRS Orléans) for assistance with the experiments and helpful discussions. This work was supported by the CNRS, the IR-RMN-THC network of the CNRS, the European Research Council (ERC contract ‘dilute para-water’, grant agreement number 339754), and the French “Equipements d’Excellence” programme Paris-en-Resonance, contract ANR-10-EQPX-09. Computer simulations were performed thanks to the CINES and TGCC (Grant 2017-GENCI Project A0010807364).
Author information
Authors and Affiliations
Contributions
D.C. designed the study, conducted the experiments, analyzed the data and wrote the manuscript, B.G. performed DFT calculations, G.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Carnevale, D., Grosjean, B. & Bodenhausen, G. Dipolar couplings in solid polypeptides probed by 14N NMR spectroscopy. Commun Chem 1, 73 (2018). https://doi.org/10.1038/s42004-018-0072-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0072-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.