Abstract
Cellulose is an important abundant renewable resource on Earth, and the microbial cellulose utilization mechanism has attracted extensive attention. Recently, some signalling molecules have been found to regulate cellulose utilization and the discovery of underlying signals has recently attracted extensive attention. In this paper, we found that the hydrogen sulfide (H2S) concentration under cellulose culture condition increased to approximately 2.3-fold compared with that under glucose culture condition in Ganoderma lucidum. Further evidence shown that cellulase activities of G. lucidum were improved by 18.2-27.6% through increasing H2S concentration. Then, we observed that the carbon repressor CreA inhibited H2S biosynthesis in G. lucidum by binding to the promoter of cbs, a key gene for H2S biosynthesis, at “CTGGGG”. In our study, we reported for the first time that H2S increased the cellulose utilization in G. lucidum, and analyzed the mechanism of H2S biosynthesis induced by cellulose. This study not only enriches the understanding of the microbial cellulose utilization mechanism but also provides a reference for the analysis of the physiological function of H2S signals.
Similar content being viewed by others
Introduction
Cellulose acts as the major component of plant cell walls as well as an abundant renewable carbohydrate1,2. Because resource is currently being exhausted, further effective utilization of cellulose is of great significance. Cellulose can be utilized by microorganisms as a low-cost and sustainable source of carbon1,3. Thus, microbial cellulose utilization is beneficial to the sustainable use of energy and carbon cycles in the biosphere, and deserves emphasis. The present work provided a fundamental understanding of microbial cellulose utilization components, including cellulases, regulating transcription factors and regulating signaling pathways. Enzymes with cellulose hydrolysis functions include three types, cellobiohydrolases (CBHs), endoglucanases (EGs) and β-glucosidases (Bgs)4,5. Some transcription factors have also been reported to play roles in cellulose utilization in filamentous fungi, such as the positive transcriptional activators, XlnR, ClrA and ClrB6,7,8,9, and the repressor, CreA3,10,11,12,13. Cellulase activity can be regulated by basal levels of enzyme production and transcription factors, which are widely recognized. However, the role of many underlying regulators involved in cellulose utilization remains to be explored.
To date, some signal transduction pathways have been reported to participate in the regulation of cellulase activities under different stimuli conditions with a variety of modes, such as the G protein signaling pathway, AMP-activated protein kinase (AMPK) signaling pathway, mitogen-activated protein kinase (MAPK) signaling pathway, Ca2+ signaling pathway and the cyclic AMP-dependent protein kinase A (cAMP-PKA) signaling pathway11,14,15,16,17,18,19. For example, members of MAPK signaling pathway were involved in cellulose utilization in Trichoderma reesei, such as the negatively regulatory, Tmk1 and Tmk2, and the positively regulatory, Tmk315,20. The Ca2+ signaling pathway was activated by cAMP to improve cellulase expression in T. reesei14,21. In Aspergillus nidulans, PKA indirectly phosphorylated at S319 of CreA under glucose culture condition, which resulted in inhibiting entry of CreA into the nucleus and reducing the transcriptional inhibition of cellulase19. Our previous study investigated the involvement of some signaling pathways in cellulose utilization in G. lucidum. For example, GlSwi6B, a member of MAPK signal transduction pathways, significantly increased the concentration of cytosolic Ca2+, thereby promoting the activities of cellulase and xylanase in G. lucidum17. Glsnf1, a member of AMP-activated protein kinase, improved cellulase activity by reducing the transcription level of the creA gene11. These works add new insights into our understanding of cellulose degradation and are of great significance for the effective utilization of cellulose as a renewable carbohydrate resource. Therefore, exploring novel signaling molecules that regulate cellulose utilization is important.
Hydrogen sulfide (H2S) is now recognized as an endogenous signaling gasotransmitter in various species22,23. Cystathionine β-synthase (CBS), which catalyzes the condensation of homocysteine and cysteine to produce H2S24. H2S biosynthesis can be induced by multiple stresses. In G. lucidum, CBS-synthesized H2S was induced by heat stress and inhibited the heat-induced secondary metabolism accumulation22. In wine-producing Saccharomyces cerevisiae, H2S biosynthesis was induced in the absence of assimilable nitrogen25, which suggests that H2S could be induced by nutrient deficiency. Furthermore, H2S exhibited multiple physiological functions in response to various stresses. For example, H2S acted as an antioxidant and antiapoptotic signal in animals26. H2S also acted as an antioxidant signal molecule to improve the growth rate of plants under Cu2+ and heat conditions27,28. Interestingly, cysteine supplementation reduced the furfural-induced accumulation of reactive oxygen species (ROS) and increased biomass during lignocellulosic utilization through increasing H2S concentration in Zymomonas mobilis29. These results implied a potential role of H2S in cellulose utilization, while direct evidence remains unavailable. Therefore, the mechanism of H2S biosynthesis and physiological function under cellulose culture conditions need to be explored.
Fungi are well-known organic-decomposing agents, especially mushrooms, which can use cellulose substrates1. G. lucidum is an important large basidiomycete with both medicinal value and economic value. Genome sequencing studies have shown that G. lucidum contains one of the largest sets of wood-breaking enzymes among basidiomycetes30. Therefore, G. lucidum is a good material for studying the regulatory mechanism of cellulose utilization. However, the role of H2S signaling pathways in cellulose utilization remains unclear. In this study, we found that the H2S concentration was increased under cellulose culture conditions. Increasing H2S concentration through pharmacological and genetic means enhanced cellulase activity. Furthermore, carbon repressor CreA inhibited the expression of cbs, a gene encoding the H2S synthetic enzyme, and the biosynthesis of H2S under cellulose culture conditions. Further research found that CreA binds to the cbs promoter at “CTGGGG”. Our study explored a novel signaling molecule, H2S, which promotes the cellulose utilization in G. lucidum, and analyzed the mechanism of cellulose-induced H2S biosynthesis. It was beneficial not only to the cultivation of G. lucidum but also to the utilization of the most abundant carbon resources in the biosphere.
Results
The intracellular concentration of H2S was increased by cellulose
To explore whether H2S signals respond to changes in the carbon source, H2S concentration was measured in Ganoderma lucidum under different carbon sources (glucose or microcrystalline cellulose) culture conditions. As shown in Fig. 1a, b, we observed that the fluorescence of H2S, measured by SF7-AM fluorescence probe, was significantly (p < 0.01) increased to ~2.3-folds under cellulose culture condition compared with that under glucose culture condition. This result shows that the intracellular H2S concentration of G. lucidum is increased by cellulose culture condition.
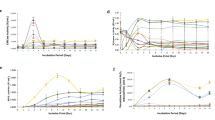
a, b Change in the H2S concentration was measured by SF7-AM fluorescence probe in wild-type (wt) strain under glucose or cellulose culture condition. The average fluorescence intensity values of all mycelia in the 6 photos were quantified. Scale bar = 100 μm. c Log2(foldchange) of genes expression level of putative H2S biosynthetic enzymes: L-cysteine desulfhydrase (lcd1 and lcd2), cysteine synthase (cs1 and cs2), cystathionine γ-lyase (cse1, cse2, cse3 and cse4), cystathionine β-synthase (cbs), 3-mercaptopyruvate sulfurtransferase (3-mst). The different letters indicate significant differences between the lines (“*“ means p < 0.05, “**“ means p < 0.01, “***“ means p < 0.001, according to Student’s t test). d, e Change in the H2S concentration was measured by SF7-AM fluorescence staining in wt, sicontrol, cbs-silenced and cbs-overexpressed strains under glucose and cellulose culture condition. Scale bar = 100 μm. f Relative expression level of cbs gene in wt, sicontrol, cbs-silenced and cbs-overexpressed strains, cultured under glucose and cellulose condition. The different letters indicate significant differences between the lines (p < 0.05, according to Duncan’s multiple range test).
The gene transcription levels of 10 putative H2S biosynthetic enzymes were also measured. As shown in the heatmap, the expression levels of lcd1, cse1 and cbs were significantly (p < 0.001) increased (Fig. 1c). Among them, the cbs expression level was significantly (p < 0.001) increased to ~13.3-fold under cellulose culture condition compared with that under glucose culture condition (Fig. 1c), which exhibits the most pronounced response to cellulose and implies that the cystathionine β-synthase (CBS) may be involved in regulating H2S biosynthesis under cellulose culture condition.
Then, a cbs gene overexpression vector was constructed (Supplementary Fig. 1a) and transfected into G. lucidum. Two cbs-overexpressed strains (cbs-oe8 and cbs-oe26) were selected because the relative cbs mRNA content increased ~3.8–4.4-fold (Supplementary Fig. 1b). The H2S concentration and the cbs expression level were detected in wild-type (wt), cbs-silenced, cbs-overexpressed and sicontrol strains under glucose or cellulose culture conditions. The fluorescence of H2S and the expression level of cbs gene in cbs-silenced strains significantly (p < 0.05) decreased by ~54.1–54.7% and 88.6–90.3% compared with that in the wt strain under cellulose culture condition, and significantly (p < 0.05) increased in cbs-overexpressed strains by ~70.5–76.3% and 148.7–164.6% (Fig. 1d, e, f), which indicated that CBS promoted H2S biosynthesis in G. lucidum under cellulose culture condition. In addition, cellulose culture condition result in a significant (p < 0.05) increase in the fluorescence of H2S in cbs-silenced strains compared with glucose culture conditions to ~1.3-folds, which was less than that in the wt strain, and no significant (p > 0.05) change in the expression level of cbs gene (Fig. 1d, e, f).
These results suggest that cellulose promotes the H2S biosynthesis in G. lucidum, and CBS might be one of the main H2S biosynthetic enzymes under cellulose culture condition.
H2S enhanced cellulase activity and cellulose utilization in G. lucidum
To explore the effect of H2S signal on cellulose utilization, cellulase activity was measured in the presence of sodium hydrosulfide (NaHS, a H2S donor) and hypotaurine (HT, a H2S scavenger), at a concentration that can significantly alter the intracellular H2S content in G. lucidum (Supplementary Fig. 2). As shown in Fig. 2a, the addition of NaHS increased cellulase activity, and the promoting effect increased gradually with increasing NaHS concentration. The addition of 60 μM NaHS significantly (p < 0.001) increased cellulase activity by ~27.6% compared with no treatment (Fig. 2a). The addition of HT inhibited cellulase activity, and the inhibitory effect was more evident with increasing HT concentration (Fig. 2b). HT (2 mM) significantly (p < 0.001) reduced cellulase activity by ~18.6% compared with no treatment (Fig. 2b). Pharmacological experiments suggest that H2S enhances cellulase activity in G. lucidum under cellulose culture condition.
a, b Endocellulase (CMCase) activities in wt strain under cellulose culture condition in the presence of sodium hydrosulfide (NaHS, H2S donor) and hypotaurine (HT, H2S scavenger). c, d Endocellulase (CMCase) activities in wt, sicontrol, cbs-silenced and cbs-overexpressed strains under cellulose culture condition in the in presence of NaHS and HT. e, f The growth length of wt, sicontrol, cbs-silenced and cbs-overexpressed strains after 15 days cultivated on wood chips. The red line marked the position of the wt strain grown on wood chips for 15 days. The different letters indicate significant differences between the lines (“**“ means p < 0.01, “***“ means p < 0.001, according to Student’s t test).
To explore the potential influence of H2S biosynthesized by CBS on cellulose utilization, cellulase activity was measured in wt, cbs-silenced, cbs-overexpressed and sicontrol strains in the presence of NaHS and HT. Cellulase activity in cbs-silenced strains significantly (p < 0.001) reduced by ~31.9–32.2% compared with that in the wt strain (Fig. 2c). NaHS addition significantly (p < 0.01) increased cellulase activity of cbs-silenced strains by ~27.1–28.2% compared with under cellulose culture condition alone (Fig. 2c). Cellulase activity in cbs-overexpressed strains significantly (p < 0.001) increased by ~18.2–18.7% compared with that in the wt strain (Fig. 2d). HT addition significantly (p < 0.001) reduced cellulase activity in cbs-overexpressed strains by ~17.8-18.6% with under only cellulose culture condition (Fig. 2d). To demonstrate the potential influence of CBS on cellulose utilization of G. lucidum, wood chips were used as a main carbon source to cultivate wt, cbs-silenced, cbs-overexpressed and sicontrol strains for 15 days, and the growth length were measured. As shown in Fig. 2e, f, the growth length of cbs-silenced strains significantly (p < 0.001) decreased by ~53.4–56.1% compared with wt strain, while cbs-overexpressed significantly (p < 0.01) increased by ~14.4–16.4%. These results suggest that CBS-synthesized H2S improves the cellulase activity and cellulose utilization.
These combined results suggest that H2S improves the cellulase activity of G. lucidum under cellulose culture condition.
Y1H screening for regulators of the cbs gene identifies diverse transcription factors
To further understand the mechanism of cellulose induced H2S biosynthesis, cbs promoter was analyzed. The intervening region between the cbs gene and upstream gene is very short (551 bp). Therefore, a total potential cbs promoter (+1 to −551 pb) was used as bait in a yeast one-hybrid (Y1H) library screen. The Y1H assay indicated that a total of 13 putative transcription factors (TFs) may directly bind to the cbs promoter (Supplementary Table 1). These putative TFs might regulate cbs gene transcription and H2S biosynthesis in G. lucidum under different conditions. Among them, three TFs, had been reported in G. lucidum, were observed: CreA (mediated cellulose utilization of G. lucidum11), GCN4 and SKO1 (mediated nitrogen utilization of G. lucidum31), and four highly conserved TFs were observed: TFIIB, MCM1, Xbp1, and Crz1 (Fig. 3a and Supplementary Table 1). To explore the regulatory mechanism of cbs transcription and H2S biosynthesis under cellulose culture condition in G. lucidum, CreA, a classical carbon catabolite repressor, was selected for further study.
a The result of highly conservative transcription factors (TFs) selected by a yeast one-hybrid (Y1H) library screen (n = 3). b Diagram of cbs promoter sequence. The black boxes indicate the binding sequence of CreA. c Y1H assay verification between CreA and cbs promoter. d Electrophoretic mobility shift assay (EMSA) for binding action of purified CreA or nuclear extraction with cbs promoter.
CreA could bind to the cbs promoter
To further determine the binding effect of CreA to the cbs promoter, Y1H assay and EMSA were performed. The Y1H assay indicated that compared with the negative control (pGADT7), Y1HGold yeast transformed pGADT7-CreA, developed obvious colonies (Fig. 3c). The EMSA result showed that compared with lanes without protein addition, an obvious binding complex band was observed in lanes with purified His-CreA protein addition (Fig. 3d). These binding effects were competitively inhibited by unlabeled cold probes (Fig. 3d). Gradually decreased binding complex bands were observed between 5’ biotin-labeled probes of the cbs promoter and purified His-CreA protein in lanes with a decrease in protein concentration (100 mM, 50 mM, 20 mM) (Fig. 3d). These results suggest that CreA can bind to the cbs promoter.
To further explore the binding site of CreA in the cbs promoter, the cbs promoter was analyzed, and observed a typical CreA binding site, “CTGGGG”, at -232 bp to -226 bp (Fig. 3b). Then, two GG nucleotides in the middle of the conserved binding domain were mutated to AA nucleotides as a previous study32. As shown in results of the Y1H assay, compared with the positive control (CBS-AbAi), Y1HGold yeast, transformed CBS-Mut-AbAi, did not develop significant colonies (Fig. 3c). The EMSA showed that compared with the positive control (non-mutated 5’ biotin-labeled probe), the mutated 5’ biotin-labeled probe could not bind with purified CreA protein (Fig. 3d). These results indicate that CreA can bind to the cbs promoter at “CTGGGG”.
CreA inhibits cbs gene expression and reduces H2S biosynthesis
To explore the regulatory role of CreA in the CBS/H2S signaling pathway in G. lucidum, a CreA-overexpress vector was constructed (Supplementary Fig. 3a) and transfected into G. lucidum. Two creA-overexpressed strains (creA-oe6 and creA-oe8) were selected for further analyses. The relative mRNA content level significantly (p < 0.001) increased about 5.7–6.0-fold and the relative protein content level significantly (p < 0.001) increased by ~52.7–54.5% in creA-overexpressed strains compared with the wt strain (Supplementary Fig. 3b, Supplementary Fig. 6, Fig. 4a, b). The relative mRNA and protein content levels in creA-silenced strains, previously established22, significantly (p < 0.001) reduced by ~74.8–75.6% and 32.6–33.7% compared with those in the wt strain, respectively (Supplementary Fig. 3b, Supplementary Fig. 6, Fig. 4a, b).
a, b Immunoblot analysis of CreA proteins in wt, sicontrol, creA-silenced and creA-overexpressed strains, cultured on CYM medium for 7 days. c Relative expression level of cbs gene in wt, sicontrol, creA-silenced and creA-overexpressed strains, cultured on CYM medium for 7 days. d, e Change in the H2S concentration was measured by SF7-AM fluorescence staining in wt, sicontrol, creA-silenced and creA-overexpressed strains, cultured on CYM medium for 7 days, measured by SF7-AM. Scale bar = 100 μm. The different letters indicate significant differences between the lines (“**“ means p < 0.01, “***“ means p < 0.001, according to Student’s t test).
Transcription levels of the cbs gene and fluorescence levels of H2S were measured in wt, creA-silenced, creA-overexpressed and sicontrol strains. Compared with that in the wt strain, the transcription level of the cbs gene in creA-silenced strains significantly (p < 0.001) increased to ~2.1-folds and the fluorescence of H2S was significantly (p < 0.01) increased by ~47.5–49.2% (Fig. 4c, d, e). The transcription level of the cbs gene and the fluorescence of H2S in creA-overexpressed strains significantly (p < 0.01) decreased by ~44.3–48.3% and 29.1–30.3%, respectively (Fig. 4c, d, e). These results indicate that CreA reduces cbs gene transcription levels and intracellular H2S biosynthesis.
CreA inhibits cbs gene expression and reduces H2S biosynthesis under cellulose culture condition
To explore the role of CreA in cbs expression and H2S biosynthesis, the binding effect of CreA to the cbs promoter was measured under glucose and microcrystalline cellulose culture condition. From the results of ChIP-qPCR, we found that cellulose culture condition significantly (p < 0.05) reduced the binding of CreA to the cbs promoter in the wt strain by ~51.1% compared with glucose culture condition (Supplementary Fig. 4 and Fig. 5a), indicating that cellulose culture conditions inhibited the DNA binding activity to the cbs promoter of CreA. In addition, under cellulose culture condition, the binding of CreA to the cbs promoter was significantly (p < 0.05) reduced by ~56.9–64.4% in creA-silenced strains and significantly (p < 0.05) increased to ~3.9–4.2-fold in creA-overexpressed strains compared with that in the wt strain (Fig. 5a). Then, transcription levels of the cbs gene and fluorescence levels of H2S were measured in the wt, creA-silenced, creA-overexpressed and sicontrol strains. Under cellulose culture condition, the transcription level of the cbs gene in creA-silenced strains was significantly (p < 0.05) increased to ~2.0-folds compared with that in the wt strain, and the fluorescence of H2S was significantly (p < 0.05) increased to ~1.8-folds (Fig. 5b, c, d). The transcription level of the cbs gene in creA-overexpressed strains under cellulose culture condition was significantly (p < 0.05) reduced by ~85.9–86.5% compared with that in the wt strain under, and the fluorescence of H2S was significantly (p < 0.05) reduced by ~68.8–70.8% (Fig. 5b, c, d). These results indicate that cellulose inhibit the DNA binding activity to the cbs promoter of CreA, thereby enhancing cbs transcription and H2S biosynthesis under cellulose culture conditions.
a ChIP assays for relative binding action with cbs promoter in wt, sicontrol, creA-silenced and creA-overexpressed strains, cultured under glucose and cellulose condition. b Relative expression level of cbs gene in wt, sicontrol, creA-silenced, creA-overexpressed and creA-cbs-silenced strains, cultured under glucose and cellulose condition. c, d Change in the H2S concentration was measured by SF7-AM fluorescence staining in wt, sicontrol, creA-silenced, creA-overexpressed and creA-cbs-silenced strains, cultured under glucose and cellulose condition. Scale bar = 100 μm. The different letters indicate significant differences between the lines (p < 0.05, according to Duncan’s multiple range test).
CreA inhibits the positive regulation of CBS/H2S in cellulose utilization of G. lucidum
To explore the role of H2S and CreA in cellulose utilization of G. lucidum, a creA-cbs-silenced plasmid was constructed (Supplementary Fig. 5a). Two creA-cbs-silenced strains (creA-cbs-i23 and creA-cbs-i27) were selected because of ~53.4–55.7% and 53.3–58.2% significant (p < 0.01) reduction in the relative mRNA content levels of the creA and cbs genes, respectively (Supplementary Fig. 5b, c). The transcription level of the cbs gene and the fluorescence of H2S in creA-cbs-i23 and creA-cbs-i27 strains under cellulose culture condition significantly (p < 0.01) reduced by ~85.6–86.0% and 74.1–77.5% compared with those in creA-silenced strains, respectively, but no significant (p > 0.05) difference was observed compared with those in cbs-silenced strains (Fig. 5b, c, d).
Cellulase activity in the wt, creA-silenced, creA-overexpressed, creA-cbs-silenced and sicontrol strains was measured in the presence of NaHS and HT treatment. Cellulase activity in creA-silenced strains was significantly (p < 0.05) increased by ~54.6–58.1% compared with that in the wt strain, and significantly (p < 0.05) decreased by ~35.6–38.2% in creA-overexpressed strains (Fig. 6a). This result suggested that CreA inhibits cellulase activity. When creA-overexpressed strains treated with NaHS, the cellulase activity significantly (p < 0.05) increased by ~27.6–29.6% compared with that no treatment (Fig. 6a). Cellulase activity in creA-cbs-silenced strains were significantly (p < 0.001) reduced by ~59.7–61.7% compared with that in creA-silenced strains, but there was no significant difference compared with that in cbs-silenced strains (Fig. 6a). These results suggest that the silencing of both creA and cbs can inhibit the creA silencing-induced decrease in cellulase activity.
a CMCase activities in wt, sicontrol, creA-silenced, creA-overexpressed, creA-cbs-silenced and cbs-silenced strains under cellulose culture condition in the presence of NaHS. b, c The growth length of wt, sicontrol, creA-silenced, creA-overexpressed and creA-cbs-silenced strains after 15 days cultivated on wood chips. The red line marked the position of the wt strain grown on wood chips for 15 days. The different letters indicate significant differences between the lines (“*“ means p < 0.05, “**“ means p < 0.01, “***“ means p < 0.001, according to Student’s t test).
Then, the growth rates of the wt, creA-silenced, creA-overexpressed, creA-cbs-silenced and sicontrol strains on wood chips were further tested. As shown in the Fig. 6b, c, the growth rates of the creA-overexpressed and creA-cbs-silenced strains were significantly (p < 0.05) reduced by ~52.0–54.4% and 51.4–53.1% compared to that of the wt strain, while the growth rate of the creA-silenced strain was significantly (p < 0.05) increased by ~22.1–23.1%. These results suggest that CreA reduces cellulose utilization of G. lucidum by inhibiting CBS/H2S signaling pathway.
Discussion
Cellulose utilization contributes to the carbon cycle in the biosphere. Microbial cellulose utilization is widely used in both agriculture and industry17,33. Exploring the potential molecular regulatory mechanisms of cellulose utilization in Ganoderma lucidum, a medicinal and edible basidiomycete, is not only be crucial to understanding the mechanism of mushroom growth and development, but also provides a theoretical basis for other cellulose-utilizing microorganisms. The efficiency of cellulose utilization dependent on the regulation of transcription factors and the activity of cellulase is widely recognized. However, recent studies have found that some signaling pathways could also take part in the regulation of cellulose utilization, such as Ca2+, cAMP and MAPK signals14,15,17. Therefore, the underlying molecular regulatory mechanism of cellulose utilization in G. lucidum deserves emphasis and awaits further investigation. In this paper, we depict the promoting effect of hydrogen sulfide (H2S) on the activity of cellulase in G. lucidum for the first time. Furthermore, H2S biosynthesis was negatively regulated by the carbon repressor transcription factor, CreA. Our results provided new insights into the mechanism of microbial cellulose utilization.
In this work, we observed that H2S improved the cellulose utilization in G. lucidum. Before this work, the role of other signaling molecules on cellulase activity and its mechanism were also reported, such as cAMP and Ca2+14,17,21,34. For instance, cAMP and Ca2+ improved the expression of cellulase in Trichoderma reesei14,21. Increasing cytosolic Ca2+ concentration promoted the activities of cellulase and xylanase in G. lucidum17. The silence of PoLaeA2 and PoLaeA3, key global regulators, leads to a reduction in cytosolic Ca2+ content and cellulase activity in Pleurotus ostreatus34. In addition, H2S has also been widely reported to have complex interactions with various signal molecules. For example, H2S increases cAMP level in resting cells, while decreases cAMP level when adenylyl cyclases are activated35. Additionally, H2S also acts as a regulator of the Ca2+ channels to regulate the intracellular Ca2+ concentration in various animal cells with different physiological effects36,37,38. H2S enhance the tolerance of plants to drought stress by activating Ca2+ signaling39. Our previous work demonstrated that H2S leads to a decrease in cytosolic Ca2+ level in G. lucidum under heat stress22. Therefore, whether H2S improve cellulase activity under cellulose culture conditions through interactions with other signaling pathways remains to be further studied.
In our previous work, we found that the silencing of creA led to an increase in transcription levels of cellulase genes and cellulase activity in G. lucidum under cellulose culture condition11. In this study, we observed that the cellulase activity in the creA-cbs silenced strains were significantly decreased than that of the wt strain (Fig. 6a). These results imply an important regulatory role of CBS-synthesized H2S in cellulase activity. Previous studies have reported that H2S affect protein function by a variety of potential mechanisms. Firstly, H2S could directly regulate the activity of proteins by persulfidation. For example, H2S regulates autophagy under endoplasmic reticulum stress in Arabidopsis by persulfidating ATG18a, a core autophagy component40. Secondly, H2S could regulate transcription factors by persulfidation, thereby affect the transcription expression of multiple genes downstream of the transcription factor. For instance, H2S involve in the regulation of cucurbitacin C synthesis in cucumber by increasing the persulfidation level of His-Csa5G156220 and His-Csa5G157230 (transcription factors) and transcriptionally activate Csa6G088690 (a key synthetase for CuC generation)41. Thirdly, H2S also could regulate the upstream regulator of transcription factors by persulfidation, thereby indirectly affect the downstream transcription expression of multiple genes. For example, H2S regulates ABA signaling by persulfidating SnRK2.6, promoting the activity of SnRK2.6 and its interaction with a transcription factor acting downstream of ABA signaling, ABF2, in guard cells42. In addition, these various mechanisms by which intracellular H2S affects physiological processes often exist simultaneously. For instance, in plants, H2S not only regulated transcription levels of drought-responsive genes but also induced the persulfidation level of proteins, which involved in cellular response to oxidative stress, hydrogen peroxide catabolism and so on, to involve in drought stress responses43. Therefore, further research is needed to clarify regulatory mechanisms of H2S on cellulase activity in G. lucidum.
Intracellular H2S biosynthesis was regulated by some transcription factors and induced by stress44. For example, transcription factor specificity protein 1 (Sp1) can bind to the promoter of both cse and cbs gene, two H2S biosynthetic enzymes, to regulate H2S biosynthesis in animals45,46. Nuclear factor (NF)-Y and upstream stimulatory factor 1 (USF-1) were also involved in the regulation of cbs promoter activity in HepG2 cells47. The transcription factor OsNACL35 increased in H2S concentration by directly upregulating the expression of OsDCD1 by binding to the promoter of OsDCD1 gene under salinity stress48. The regulation of transcription levels of H2S biosynthetic enzymes had been reported in animals and plants, but less so in microorganisms. In this study, yeast one-hybrid (Y1H) library screening results revealed the underlying transcription factors of the cbs gene, including CreA (mediated cellulose utilization of G. lucidum11), GCN4 and SKO1 (mediated nitrogen utilization of G. lucidum31), and four putative transcription factors (TFIIB, MCM1, Xbp1, Crz1). We further revealed that CreA combined with the cbs promoter, transcriptionally provoked cbs gene expression and improved H2S biosynthesis under cellulose culture condition by pharmacological experiments and genetic experiments. The roles of other transcription factors need to be further studied. The potential binding of multiple transcription factors to the cbs gene implied that H2S may be induced by a variety of growing conditions in G. lucidum, and providing unlimited possibilities for the physiological function of H2S signals under stress.
CreA, a carbon catabolite repressor, was previously reported to directly bind to the gene promoter of cellulolytic enzymes, inhibit the transcription of related genes, and regulate cellulose utilization in filamentous fungi, such as T. reesei, Aspergillus nidulans, G. lucidum and Neurospora crassa3,11,12,49. The transcriptomics analysis and the secretome analysis in N. crassa were observed that CreA repressed the expression level of genes, encoding enzymes involved in the utilization of alternative carbon sources, and cellulase activity under cellulose culture condition50. In T. reesei, CreA was considered as a regulator of the glucose assimilation rate51. Recent research has also shown that CreA could bind not only to the promoter of cellulose related genes, but also to the promoter of other genes to play a primary role in diverse physiological processes, such as carbon metabolism, secondary metabolism, iron homeostasis, oxidative stress response, development, N-glycan production, unfolded protein response, and nutrient and ion transport in A. nidulans52. Beyond carbon metabolism regulation, studies in various fungal species indicate additional CreA functions. For example, the ΔcreA mutants of Aspergillus flavus53, Magnaporthe oryzae54, and Beauveria bassiana55 are affected in development and virulence, while repression of genes with functions in nitrogen uptake, development, chromatin remodeling, and the mediator complex depends on a functional CreA in T. reesei51. In Aspergillus fumigatus, the ΔcreA mutant impacts growth, fitness, and virulence56. In this study, we found that CreA negatively regulated cbs expression by binding to the cbs promoter, and inhibiting H2S biosynthesis, thereby reducing cellulase activity. Additionally, under repressing carbon sources culture condition, such as cellulose, the transcriptional level of CreA was decreased32,49. Therefore, CBS/H2S signaling pathway was provoked under cellulose culture condition in G. lucidum. H2S is a signaling molecule with multiple biological functions, and these results also implied that CreA may play a greater biological role through the regulation of H2S biosynthesis.
In this study, we revealed that H2S signals were promoted by activating cbs transcription level under cellulose culture condition (Fig. 7). Then, we observed that H2S improved cellulase activity in G. lucidum (Fig. 7). Furthermore, we observed that CreA could bind to the promoter of cbs gene and reduce the transcriptional level of cbs gene (Fig. 7). In the case of repressing carbon sources, such as cellulose culture condition, CreA mRNA levels were decreased32,49. Therefore, cellulose reduced the inhibitory effect of CreA on the transcription level of cbs gene, thereby activating CBS/H2S signaling pathway under cellulose culture condition. These results explored a novel signaling molecule, H2S, which participate in the regulation of cellulose utilization. Our study not only provides insight into the response mechanism of H2S signals in exposure to cellulose in G. lucidum but also benefits to the utilization of the most abundant energy resources in the biosphere.
Methods
Strains and culture conditions
The wild-type (wt) Ganoderma lucidum strain (obtained from Shanghai Academy of Agricultural Science) was used in a previous work22. The G. lucidum strains used in this experiment: sicontrol, cbs-silenced and cbs-overexpressed strains were established previously22; creA-silenced strains were established previously11; creA-overexpressed strains were established as described in Supplementary Fig. 1, creA-cbs-silenced strains were established as described in Supplementary Fig. 3.
The culture conditions of G. lucidum was used as previous works22,57. For wood chips culture, G. lucidum were cultured on edible mushroom cultivation material (60% wood chip, 20% cottonseed hull, 18% wheat bran, 1% sucrose and 1% gypsum) at 28 °C and for 15 days. For other detection, G. lucidum were cultured in glucose-containing nutrient-rich liquid CYM medium at 28°C and 150 rpm for 5 days, and then collected and changed into single carbon source (1% glucose or microcrystalline cellulose) liquid MCM medium (0.46% KH2PO4, 0.05% MgSO4-7H2O, 0.5% (NH4)2SO4, 2 ml/l trace elements) at 28°C and 150 rpm for 2 days.
Transcription level detection
The transcription level of cbs and creA were detected by RT-qPCR according to method described previously22. Total RNAiso Plus (TaKaRa, Dalian, China) was used to extract total RNA and cDNA was reverse transcribed using a PrimeScript RT reagent kit (TaKaRa, Dalian, China). 18S rRNA was used as a reference to analyze the transcription levels of cbs and creA. The oligonucleotide primers used are listed in the Supplementary Table 2. The relative transcription levels of genes were determined using the 2−ΔΔCT method.
Endogenous H2S concentration detection
H2S concentration in G. lucidum was detected after incubation with sulfidefluor-7 acetoxymethyl ester (SF7-AM) as described in a previous study22,58. The average fluorescence intensity values of all mycelia in each photo were quantified. Zeiss Axio Imager A1 fluorescence microscope was used to fluorescence image for all samples under the same microscopy settings. The average fluorescence intensity of mycelium was analyzed using ZEN software.
Endocellulase activity (CMCase) detection
Endocellulase activity was detected according to previously described methods11. The culture supernatants were collected for endocellulase activity (CMCase) assays. In brief, citric acid buffer (50 mM, pH 4.8) and 2% (W/V) sodium carboxymethyl cellulose (CMC-Na) were added and the reaction was carried out at 50 °C for 30 min, and then detected optical density at 540 nm.
Yeast one-hybrid assay
A yeast one-hybrid (Y1H) assay was performed using the matchmaker gold yeast one-hybrid library screening system (Clontech, China). The target sequence of cbs promoter (+1 to −551pb) was cloned and inserted into the pAbAi vector. The CBS-AbAi vector was digested with BstBI enzyme (TAKARA, China) and then transformed into yeast stains. Strains were grown on SD/-Ura media for 2 days and positive transformants were selected by PCR. Next, the minimal inhibitory concentration of aureobasidin A (AbA) was confirmed as follows. Y1HGold (CBS-AbAi) were suspended in 0.9% NaCl (OD600: 0.002) and dotted onto SD/-Ura medium with AbA (0, 100, 150, 200, 300, 400, 500, 600, 800, and 1000 ng ml−1) for 3 days. AbA (600 ng ml−1) completely suppressed the growth of yeast strains and was used for subsequent experiments.
The Y1H screen assay was performed as previously described59. The cDNA library (pGADT7 vector) was conducted by SMART cDNA production technology (oebiotech, China). Then, screen the cDNA library by cotransformation and transformed yeast stains are plated on SD/-Leu/+AbA to select for colonies.
For the transcriptional activation test, the full-length complementary cDNA of creA was cloned and inserted into the pGADT7 vector. The constructed plasmids (pGADT7-CreA) were then transformed into Y1HGold (CBS-AbAi) on SD/-Leu/+AbA medium for 3 days, and the positive transformants and were chosen and dotted as described above.
Expression and purification of CreA
CreA protein was expressed and purified as previously described11. The creA-28a plasmid was transformed into E. coli BL21 (DE3), and protein expression was induced by 1 mM isopropyl β-D-thiogalactopyranoside (IPTG) at 28 °C for 4 h. Protein verification was performed using SDS-PAGE.
Electrophoretic mobility shift assay
An electrophoretic mobility shift assay (EMSA) was performed as previously described46,59. Both forward (5′-TGGCTGGGGC-3′) and reverse (5′-GCCCCAGCCA-3′) primers of cbs promoter were biotin labeled at the 5’ end. The mutated primers were 5′-TGGCTGTTGC-3′ (forward) and 5′-GCAACAGCCA-3′ (reverse) biotin labeled at the 5’ end. For EMSA analysis, the biotin-labeled primers were incubated with purified CreA protein (10 μg, 5 μg, 2 μg, respectively) in 5 μl binding buffer [10 mM Tris-HCl (pH 8.0), 1 mM DTT, 0.1 mM EDTA, 50 mM KCl and 5% glycerol]. In addition, the mixture without protein was used as a negative control.
Chromatin immunoprecipitation assay
Chromatin immunoprecipitation assay (ChIP) was performed as previously described46. In brief, cross-linking of DNA with proteins was performed with 1% formaldehyde. The nuclei were isolated, and sheared chromatin was prepared by sonication. The sheared chromatin was immunoprecipitated with rabbit polyclonal anti-CreA antibodies and rabbit serum (D601019, Sangon, China, negative control). Another aliquot of sheared chromatin without incubation with antibodies was prepared as input. As shown in Sigure S2, DNA, isolated from immunoprecipitation, was detected by PCR with primers for the cbs promoter region were 5′-TTGACGCGGACGGACAT-3′ (forward) and 5′-GGGAAGATGGTGGCAGAA-3′ (reverse). The relative binding efficiency of CreA to the cbs promoter was detected by RT-qPCR.
Western blotting
Western blotting was performed as previously described11. Briefly, proteins from mycelia samples were separated in a 12% (w/v) SDS-PAGE gel and transferred to polyvinylidene difluoride membranes (Bio-Rad). The primary antibodies used to detect specific proteins in this report were anti-CreA (1:2000, rabbit polyclonal), anti-Actin (1:2000, mouse; CMCTAG) and anti-Histone3 (1:2000, AT0005, CMCTAG). ImageJ software was used to quantify the densities of the bands.
Statistical analysis
The statistical analyses were performed using GraphPad Prism 8.0.2 (GraphPad Software, San Diego, CA, USA). All experimental data shown in this article were carried out in three independent samples to ensure that trends and relationships observed in cultures were reproducible. The error bars indicate the standard deviation from the mean of triplicates. The data were analyzed using Student’s t test or Duncan’s multiple range test. The p < 0.05 was considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files) or are available from the corresponding author on reasonable request. The source data underlying the graphs in the figure are shown in Supplementary Data 1. Uncropped western blots are in Supplementary Fig. 6.
References
Lynd, L. R., Weimer, P. J., van Zyl, W. H. & Pretorius, I. S. Microbial cellulose utilization: fundamentals and biotechnology. Microbiol. Mol. Biol. Rev. 66, 506–577 (2002).
Lynd, L. R., Wyman, C. E. & Gerngross, T. U. Biocommodity engineering. Biotechnol. Prog. 15, 777–793 (1999).
Lichius, A., Seidl-Seiboth, V., Seiboth, B. & Kubicek, C. P. Nucleo-cytoplasmic shuttling dynamics of the transcriptional regulators XYR1 and CRE1 under conditions of cellulase and xylanase gene expression in Trichoderma reesei. Mol. Microbiol. 94, 1162–1178 (2014).
Ilmén, M., Saloheimo, A., Onnela, M. L. & Penttilä, M. E. Regulation of cellulase gene expression in the filamentous fungus Trichoderma reesei. Appl. Environ. Microbiol. 63, 1298–1306 (1997).
Wu, M. H., Kao, M. R., Li, C. W., Yu, S. M. & Ho, T. D. A unique self-truncation of bacterial GH5 endoglucanases leads to enhanced activity and thermostability. BMC Biol. 20, 137 (2022).
Stricker, A. R., Mach, R. L. & de Graaff, L. H. Regulation of transcription of cellulases- and hemicellulases-encoding genes in Aspergillus niger and Hypocrea jecorina (Trichoderma reesei). Appl. Microbiol. Biotechnol. 78, 211–220 (2008).
Xia, C. et al. Introduction of heterologous transcription factors and their target genes into Penicillium oxalicum leads to increased lignocellulolytic enzyme production. Appl. Microbiol. Biotechnol. 103, 2675–2687 (2019).
Craig, J. P., Coradetti, S. T., Starr, T. L. & Glass, N. L. Direct target network of the Neurospora crassa plant cell wall deconstruction regulators CLR-1, CLR-2, and XLR-1. mBio 6, e01452–01415 (2015).
Huberman, L. B., Coradetti, S. T. & Glass, N. L. Network of nutrient-sensing pathways and a conserved kinase cascade integrate osmolarity and carbon sensing in Neurospora crassa. Proc. Natl Acad. Sci. USA 114, E8665–e8674 (2017).
Li, Z. et al. Synergistic and dose-controlled regulation of cellulase gene expression in Penicillium oxalicum. PLoS Genet. 11, e1005509 (2015).
Hu, Y. et al. In Ganoderma lucidum, Glsnf1 regulates cellulose degradation by inhibiting GlCreA during the utilization of cellulose. Environ. Microbiol. 22, 107–121 (2020).
Lin, H., Hildebrand, A., Kasuga, T. & Fan, Z. Engineering Neurospora crassa for cellobionate production directly from cellulose without any enzyme addition. Enzym. Microbial Technol. 99, 25–31 (2017).
Hildebrand, A., Szewczyk, E., Lin, H., Kasuga, T. & Fan, Z. Engineering Neurospora crassa for improved cellobiose and cellobionate production. Appl. Environ. Microbiol. 81, 597–603 (2015).
Chen, Y. et al. cAMP activates calcium signalling via phospholipase C to regulate cellulase production in the filamentous fungus Trichoderma reesei. Biotechnol. Biofuels 14, 62 (2021).
Wang, M. et al. Role of Trichoderma reesei mitogen-activated protein kinases (MAPKs) in cellulase formation. Biotechnol. Biofuels 10, 99 (2017).
Tisch, D., Kubicek, C. P. & Schmoll, M. New insights into the mechanism of light modulated signaling by heterotrimeric G-proteins: ENVOY acts on gna1 and gna3 and adjusts cAMP levels in Trichoderma reesei (Hypocrea jecorina). Fungal Genet. Biol. 48, 631–640 (2011).
Lian, L. D. et al. GlSwi6 positively regulates cellulase and xylanase activities through intracellular Ca2+ signaling in Ganoderma lucidum. J. Fungi 8, https://doi.org/10.3390/jof8020187 (2022).
Liu, P. et al. Enhanced cellulase production by decreasing intercellular pH through H+-ATPase gene deletion in Trichoderma reesei RUT-C30. Biotechnol. Biofuels 12, 195 (2019).
Ribeiro, L. F. C. et al. Comprehensive analysis of Aspergillus nidulans PKA phosphorylome identifies a novel mode of CreA regulation. mBio 10, https://doi.org/10.1128/mBio.02825-18 (2019).
Wang, M. et al. A mitogen-activated protein kinase Tmk3 participates in high osmolarity resistance, cell wall integrity maintenance and cellulase production regulation in Trichoderma reesei. PloS One 8, e72189 (2013).
Chen, Y. et al. N,N-dimethylformamide induces cellulase production in the filamentous fungus Trichoderma reesei. Biotechnol. Biofuels 12, 36 (2019).
Tian, J. L. et al. Hydrogen sulfide, a novel small molecule signalling agent, participates in the regulation of ganoderic acids biosynthesis induced by heat stress in Ganoderma lucidum. Fungal Genet. Biol. 130, 19–30 (2019).
Zhao, W., Zhang, J., Lu, Y. & Wang, R. The vasorelaxant effect of H2S as a novel endogenous gaseous K(ATP) channel opener. EMBO J. 20, 6008–6016 (2001).
Landry, A. P., Roman, J. & Banerjee, R. Structural perspectives on H2S homeostasis. Curr. Opin. Struct. Biol. 71, 27–35 (2021).
Jiranek, V., Langridge, P. & Henschke, P. A. Regulation of hydrogen sulfide liberation in wine-producing Saccharomyces cerevisiae strains by assimilable nitrogen. Appl. Environ. Microbiol. 61, 461–467 (1995).
Jha, S., Calvert, J. W., Duranski, M. R., Ramachandran, A. & Lefer, D. J. Hydrogen sulfide attenuates hepatic ischemia-reperfusion injury: role of antioxidant and antiapoptotic signaling. Am. J. Physiol. Heart Circ. Physiol. 295, H801–806, (2008).
Zhang, H. et al. Hydrogen sulfide promotes wheat seed germination and alleviates oxidative damage against copper stress. J. Integr. Plant Biol. 50, 1518–1529 (2008).
Christou, A., Filippou, P., Manganaris, G. A. & Fotopoulos, V. Sodium hydrosulfide induces systemic thermotolerance to strawberry plants through transcriptional regulation of heat shock proteins and aquaporin. BMC Plant Biol. 14, 42 (2014).
Yan, X. et al. Cysteine supplementation enhanced inhibitor tolerance of Zymomonas mobilis for economic lignocellulosic bioethanol production. Bioresour. Technol. 349, 126878 (2022).
Liu, D. et al. The genome of Ganoderma lucidum provides insights into triterpenes biosynthesis and wood degradation. PloS One 7, e36146 (2012).
Lian, L. et al. GCN4 enhances the transcriptional regulation of AreA by interacting with SKO1 to mediate nitrogen utilization in Ganoderma lucidum. Appl. Environ. Microbiol. 88, e0132222 (2022).
Strauss, J. et al. The function of CreA, the carbon catabolite repressor of Aspergillus nidulans, is regulated at the transcriptional and post-transcriptional level. Mol. Microbiol. 32, 169–178 (1999).
Lev, S. & Horwitz, B. A. A mitogen-activated protein kinase pathway modulates the expression of two cellulase genes in Cochliobolus heterostrophus during plant infection. Plant Cell 15, 835–844 (2003).
Zhang, G. et al. Functional roles of laeA-like genes in fungal growth, cellulase activity, and secondary metabolism in Pleurotus ostreatus. J. Fungi 8, https://doi.org/10.3390/jof8090902 (2022).
Cao, X. et al. The role of hydrogen sulfide in cyclic nucleotide signaling. Biochem. Pharmacol. 149, 20–28 (2018).
Kimura, Y. et al. Polysulfides are possible H2S-derived signaling molecules in rat brain. FASEB J. 27, 2451–2457 (2013).
Nagai, Y., Tsugane, M., Oka, J. & Kimura, H. Hydrogen sulfide induces calcium waves in astrocytes. FASEB J. 18, 557–559 (2004).
Castro-Piedras, I. & Perez-Zoghbi, J. F. Hydrogen sulphide inhibits Ca2+ release through InsP3 receptors and relaxes airway smooth muscle. J. Physiol. 591, 5999–6015 (2013).
Jin, Z. et al. Hydrogen sulfide interacting with abscisic acid in stomatal regulation responses to drought stress in Arabidopsis. Plant Physiol. Biochem. 62, 41–46 (2013).
Aroca, A., Yruela, I., Gotor, C. & Bassham, D. C. Persulfidation of ATG18a regulates autophagy under ER stress in Arabidopsis. Proc. Natl Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2023604118 (2021).
Liu, Z. et al. The role of H2S in low temperature-induced cucurbitacin C increases in cucumber. Plant Mol. Biol. 99, 535–544 (2019).
Chen, S. et al. Hydrogen sulfide positively regulates abscisic acid signaling through persulfidation of SnRK2.6 in guard cells. Mol. Plant 13, 732–744 (2020).
Jurado-Flores, A., Aroca, A., Romero, L. C. & Gotor, C. Sulfide promotes tolerance to drought through protein persulfidation in Arabidopsis. J. Exp. Bot. 74, 4654–4669 (2023).
Wang, R. Physiological implications of hydrogen sulfide: a whiff exploration that blossomed. Physiol. Rev. 92, 791–896 (2012).
Sen, N. et al. Hydrogen sulfide-linked sulfhydration of NF-κB mediates its antiapoptotic actions. Mol. Cell 45, 13–24 (2012).
Wu, N., Siow, Y. L. & O, K. Ischemia/reperfusion reduces transcription factor Sp1-mediated cystathionine beta-synthase expression in the kidney. J. Biol. Chem. 285, 18225–18233 (2010).
Ge, Y., Konrad, M. A., Matherly, L. H. & Taub, J. W. Transcriptional regulation of the human cystathionine beta-synthase-1b basal promoter: synergistic transactivation by transcription factors NF-Y and Sp1/Sp3. Biochem. J. 357, 97–105 (2001).
Sun, Y. et al. A NAC transcription factor from ‘Sea Rice 86’ enhances salt tolerance by promoting hydrogen sulfide production in rice seedlings. Int. J. Mol. Sci. 23, https://doi.org/10.3390/ijms23126435 (2022).
Ries, L. N., Beattie, S. R., Espeso, E. A., Cramer, R. A. & Goldman, G. H. Diverse regulation of the CreA carbon catabolite repressor in Aspergillus nidulans. Genetics 203, 335–352 (2016).
Sun, J. & Glass, N. L. Identification of the CRE-1 cellulolytic regulon in Neurospora crassa. PloS One 6, e25654 (2011).
Portnoy, T. et al. The CRE1 carbon catabolite repressor of the fungus Trichoderma reesei: a master regulator of carbon assimilation. BMC Genom. 12, 269 (2011).
Chen, Y. et al. Carbon catabolite repression governs diverse physiological processes and development in Aspergillus nidulans. mBio 13, e0373421 (2021).
Fasoyin, O. E. et al. Carbon catabolite repression gene creA regulates morphology, aflatoxin biosynthesis and virulence in Aspergillus flavus. Fungal Genet. Biol. 115, 41–51 (2018).
Hong, Y. et al. Carbon catabolite repressor MoCreA is required for the asexual development and pathogenicity of the rice blast fungus. Fungal Genet. Biol. 146, 103496 (2021).
Luo, Z., Qin, Y., Pei, Y. & Keyhani, N. O. Ablation of the creA regulator results in amino acid toxicity, temperature sensitivity, pleiotropic effects on cellular development and loss of virulence in the filamentous fungus Beauveria bassiana. Environ. Microbiol. 16, 1122–1136 (2014).
Beattie, S. R. et al. Filamentous fungal carbon catabolite repression supports metabolic plasticity and stress responses essential for disease progression. PLoS Pathog. 13, e1006340 (2017).
Xu, W. et al. Mitochondrial pyruvate carrier regulates the lignocellulosic decomposition rate through metabolism in Ganoderma lucidum. FEMS Microbiol. Lett. 368, https://doi.org/10.1093/femsle/fnab088 (2021).
Li, K. et al. Hydrogen Sulfide Regulates Glucose Uptake in Skeletal Muscles via S-Sulfhydration of AMPK in Muscle Fiber Type-Dependent Way. J Nutr. 153, 2878–2892 (2023).
Zhu, J. et al. Dual functions of AreA, a GATA transcription factor, on influencing ganoderic acid biosynthesis in Ganoderma lucidum. Environ. Microbiol. 21, 4166–4179 (2019).
Acknowledgements
This work was supported by: China Agriculture Research System (project number CARS20), the National Natural Science Foundation of China (project numbers 31972059 and 32272787), and the Postgraduate Research and Practice Innovation Program of Jiangsu Province (project number KYCX220709).
Author information
Authors and Affiliations
Contributions
J.S. and M.W.Z. designed the study. J.S., J.Q., H.L., L.Z., and X.H. carried out experiments and analyzed data. L.S., J.Z., R.L., A.R., and M.W.Z. provided supervisor oversight. J.S. wrote the manuscript. All authors gave input and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Peer review
Peer review information
Communications Biology thanks Pedro Gonçalves and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: David Favero. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shangguan, J., Qiao, J., Liu, H. et al. The CBS/H2S signalling pathway regulated by the carbon repressor CreA promotes cellulose utilization in Ganoderma lucidum. Commun Biol 7, 466 (2024). https://doi.org/10.1038/s42003-024-06180-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06180-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.