Abstract
Regulated Ire1-dependent decay (RIDD) is a feedback mechanism in which the endoribonuclease Ire1 cleaves endoplasmic reticulum (ER)-localized mRNAs encoding secretory and membrane proteins in eukaryotic cells under ER stress. RIDD is artificially induced by chemicals that generate ER stress; however, its importance under physiological conditions remains unclear. Here, we demonstrate the occurrence of RIDD in filamentous fungus using Aspergillus oryzae as a model, which secretes copious amounts of amylases. α-Amylase mRNA was rapidly degraded by IreA, an Ire1 ortholog, depending on its ER-associated translation when mycelia were treated with dithiothreitol, an ER-stress inducer. The mRNA encoding maltose permease MalP, a prerequisite for the induction of amylolytic genes, was also identified as an RIDD target. Importantly, RIDD of malP mRNA is triggered by inducing amylase production without any artificial ER stress inducer. Our data provide the evidence that RIDD occurs in eukaryotic microorganisms under physiological ER stress.
Similar content being viewed by others
Introduction
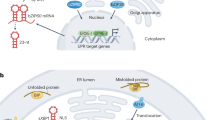
Eukaryotic mRNAs encoding secretory and membrane proteins (SMPs) are translated by ribosomes on the endoplasmic reticulum (ER). Misfolded SMPs accumulate in the ER and induce an unfolded protein response (UPR) to counteract ER stress by increasing the capacity for protein folding and degradation of misfolded SMPs1,2,3. In the UPR, the ER-localized transmembrane kinase/endoribonuclease Ire1 is activated to splice an intron of the pre-mRNA encoding Hac1/XBP1, a basic leucine zipper-type transcription factor. The functional transcription factor translated from this unconventionally spliced mRNA induces the expression of ER chaperone and ER-associated degradation (ERAD) genes to refold and degrade unfolded SMPs, respectively. The UPR pathway is well-conserved from yeast to humans, except for the fission yeast Schizosaccharomyces pombe4.
In addition to splicing Hac1/XBP1 mRNA, metazoan Ire1 cleaves ER-localized mRNAs to reduce the load of nascent polypeptides entering the ER5. Two mRNA fragments generated by Ire1 cleavage are rapidly degraded: the 5′-mRNA fragment without the poly(A) tail is degraded by the 3′–5′ exonuclease complex exosome in cooperation with the Ski2–Ski3–Ski8 (Ski) complex, and the 3′-mRNA fragment is degraded by the 5′–3′ exoribonuclease Xrn1. This novel mRNA degradation pathway is called regulated Ire1-dependent mRNA decay (RIDD). In 2006, RIDD was discovered in the fruit fly Drosophila melanogaster6 and is now widely found in mammalian cells, plants, S. pombe, and the human pathogenic yeast Candida glabrata7,8,9,10,11 but not in the budding yeast Saccharomyces cerevisiae.
Filamentous fungi produce large amounts of secretory hydrolytic enzymes; thus, many fungi are industrially used as enzyme sources12,13. Aspergillus oryzae is used for the production of sake, soy sauce, and soybean paste (miso) in Japan, and it secretes copious amounts of amylolytic and proteolytic enzymes required for the efficient utilization of starch and proteins, respectively, present in raw materials14,15,16. In Aspergillus fungi, the expression of amylolytic genes is induced by the transcriptional regulator AmyR in the presence of maltose15,16. In Aspergillus nidulans, isomaltose converted from maltose by the transglycosylation activity of secreted α-glucosidase induces AmyR activation17. However, the mechanism of AmyR activation in A. oryzae is more complex: maltose is incorporated by the maltose permease MalP and converted to isomaltose by the transglycosylation activity of the intracellular α-glucosidase MalT18. This unique conversion mechanism from maltose to isomaltose probably contributes to the high capacity of amylolytic enzyme production in A. oryzae. When A. oryzae produces amylolytic enzyme, UPR is induced and IreA (an ortholog of yeast Ire1) is essential for mycelial growth19.
The UPR in filamentous fungi, including A. oryzae, has been well studied under forced ER stress induced by the addition of chemicals such as dithiothreitol (DTT) and tunicamycin, as well as by forced high-level expression of secretory homologous or heterologous proteins20,21,22,23,24. In addition to UPR induction, the abundance of mRNAs encoding secretory proteins decreases when ER stress is induced by the addition of DTT in filamentous fungi24,25,26,27,28. The DTT-dependent decrease in the mRNA levels of genes encoding secretory hydrolases was proposed to be due to transcriptional repression and was termed repression under secretion stress (RESS)25,26. In addition, ER stress reduces the transcript level of amyR in Aspergillus niger and A. oryzae22,29,30. Therefore, in filamentous fungi, the decrease in mRNA levels induced by ER stress is believed to be due to transcriptional repression; however, reports on RIDD are lacking.
In this study, we demonstrated the presence of RIDD in filamentous fungi. Using a modified α-amylase gene with a mutation in the secretory signal sequence, we found that the DTT-induced decrease in the α-amylase mRNA level was dependent on its ER-associated translation and IreA in A. oryzae. In addition, we found that physiological ER stress induced by amylolytic enzyme production strongly induces RIDD of malP mRNA. Our data highlight the potential for A. oryzae, in which both UPR and RIDD are induced by physiological ER stress, to be a useful model for elucidating the physiological significance of the ER stress response in eukaryotic microorganisms.
Results
Rapid decrease in α-amylase mRNA level after adding DTT
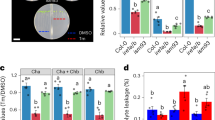
Studies have shown that ER stress elicited with chemicals such as DTT rapidly downregulates the expression of the major secreted hydrolytic enzymes in filamentous fungi, including A. oryzae28,30. To examine the selective repression of transcription of secretory protein-coding genes upon the addition of DTT in more detail, we monitored the mRNA levels of genes encoding secretory α-amylase (amyA, amyB, and amyC) and glucoamylase (glaA), together with the genes encoding the ER chaperone, BipA (bipA), and the cytoplasmic enolase (enoA) for a shorter period after DTT was added to the culture of the A. oryzae RIB40 strain grown in YPM medium (Fig. 1, left panels). A. oryzae RIB40 contains three copies of α-amylase genes, amyA, amyB, and amyC, the sequences of which are almost identical31. The levels of the amyA/B/C and glaA mRNAs decreased dramatically within 30 min after the addition of DTT. As expected, the expression level of bipA increased markedly upon adding DTT, confirming that the addition of DTT induced the UPR. In contrast, the enoA mRNA level was unaffected by DTT addition. DTT-induced downregulation of secreted protein expression is believed to result from transcriptional repression, which depends on the cis-elements in the promoter regions of their genes and decreased expression of trans-acting activators21,25,26. Accordingly, we substituted the promoter sequence of amyB with that of actin (actA), the expression of which was not affected by DTT treatment (Supplementary Fig. 1). The actAp-amyB DNA construct was introduced into an A. oryzae strain lacking all three copies of the intrinsic α-amylase genes (ΔamyA/B/C), and the amyB mRNA level was examined. Importantly, we found that the amyB mRNA level decreased, similar to that expressed from the amyB promoter (Fig. 1). These results suggested that a mechanism other than transcriptional repression was involved in the rapid decrease in the amyA/B/C mRNA levels caused by DTT addition.
Decrease in α-amylase mRNA level depended on the secretory signal sequence
Previous and present studies have shown a rapid decrease in mRNA levels in the DTT-treated condition, specifically of those encoding secretory proteins25,26. Therefore, we generated modified amyB with a mutation in the secretory signal sequence to investigate whether the reduction in the α-amylase mRNA level by DTT was dependent on ER targeting. In the modified amyB, termed amyB-FS, two nucleotides (A and T) in the second codon (ATG) were deleted and inserted near the end of the secretory signal sequence to introduce a frameshift in the signal peptide sequence of AmyB without compromising the overall mRNA structure (Fig. 2a). The amyB-FS gene is expected to encode a non-functional secretory signal sequence due to a frameshift mutation, as shown using SignalP-6.0 prediction (https://services.healthtech.dtu.dk/service.php?SignalP-6.0) (Fig. 2b). A plasmid harboring an intact or modified amyB was introduced into the chromosome of the ΔamyA/B/C strain. A decrease in the mRNA level of amyB-FS was not observed compared to that of intact amyB upon DTT treatment, although the amyB-FS mRNA was still gradually decreased (Fig. 2c). These results suggested that the decrease in the α-amylase mRNA level was dependent on its localization in the ER.
a The nucleotide sequences of the 5′-CDS region of the wild type (amyB) and the frameshift mutant (amyB-FS) of amyB and their deduced amino acid sequences. The underlines indicate the deleted and inserted nucleotides. b Prediction of signal peptide of the wild type and mutant AmyB using SignalP 6.0. The likelihood scores for the signal peptides were 0.9992 and 0 for wild type and mutant AmyB, respectively. Sec/SPI; secretory signal peptides transported by the Sec translocon and cleaved by signal peptidase I. The characters “n”, “h”, and “c” indicate the N-terminal region, the center hydrophobic region, and the C-terminal region of the signal peptide, respectively. c Northern blot analysis of transcripts prepared from the DTT-treated ΔamyA/B/C strain expressing the wild type or mutant amyB genes.
α-amylase mRNA was cleaved by IreA
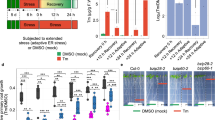
As the reduction in the α-amylase mRNA level upon DTT treatment was dependent on its targeting to the ER, we assumed that RIDD might be involved in this phenomenon. To test this hypothesis, we utilized a knockdown strain of ireA (nmtAp-ireA), the expression of which was repressed in the presence of thiamine under the control of the nmtA promoter, as ireA is an essential gene in A. oryzae23. Mycelia grown in the presence of thiamine were transferred to a maltose-containing medium (still containing thiamine) to induce α-amylase expression, followed by the addition of DTT to the medium. We found that the decrease in the amyA/B/C mRNA levels was markedly delayed by ireA repression (Fig. 3a), indicating that IreA plays an important role in reducing the amyA/B/C mRNAs under ER stress. We next confirmed whether suppression of the decline in the amyA/B/C mRNA levels was directly caused by ireA repression and not by the inefficient processing of the hacA mRNA. Toward this, we expressed artificially activated hacA (hacAi), which lacks the 20-base intron spliced from the hacA pre-mRNA by IreA, in the nmtAp-ireA strain23 and found that the expression of hacAi did not affect the IreA-dependent decrease in the amyA/B/C mRNA levels induced by DTT treatment (Fig. 3b). This indicated that the decrease in the amyA/B/C mRNA levels was controlled by IreA but not by the downstream events of HacA activation. If the amyA/B/C mRNAs are cleaved by IreA, the resulting mRNA fragments lacking a poly(A) tail are expected to be rapidly degraded by the 3′ to 5′ mRNA degradation pathway. To test this hypothesis, we generated a disruption mutant of an ortholog of the RNA helicase Ski2 (AoSki2), which is a component of the Ski complex mediating 3′ to 5′ mRNA degradation by the cytoplasmic exosome. Northern blot analysis using the 5′-terminal region of the amyA/B/C coding sequence (CDS) as the probe showed that the abundance of the short fragments of the amyA/B/C mRNAs increased noticeably after Aoski2 deletion, which is dependent on DTT treatment (Fig. 3c). Furthermore, the accumulation of short fragments was severely inhibited by ireA repression (Fig. 3c), indicating that the amyA/B/C mRNAs were cleaved in an IreA-dependent manner in response to the ER stress induced by DTT addition. Taken together, our observations suggested that the rapid decrease in the amyA/B/C mRNA levels upon DTT treatment was mainly mediated by RIDD.
a Northern blot analysis of the amyA/B/C mRNAs in ireA-repressing strain treated with DTT. After induction of α-amylase gene expression, DTT was added to the culture medium and the mycelia were harvested at the indicated times. b Northern blot analysis of the amyA/B/C mRNAs in hacA-expressing strains treated with DTT. c Northern blot analysis of the amyA/B/C mRNAs in WT, ΔAoski2, and ΔAoski2/nmtAp-ireA strains treated with DTT. The asterisks indicate short mRNA fragments.
mRNAs of α-amylase and malP were cleaved under physiological condition
Although DTT treatment noticeably increased the levels of the short fragments of the amyA/B/C mRNAs, they were detectable even before DTT treatment in the ΔAoski2 strain (Fig. 3c). As UPR is induced by amylolytic enzyme production in A. oryzae23, we hypothesized that the production of amylolytic enzymes alone can trigger RIDD in A. oryzae. To test this hypothesis, we examined the expression of amyA/B/C in the ΔAoski2 strain after incubation in liquid maltose medium without adding DTT. Short fragments of the amyA/B/C mRNAs were consistently observed along with the full length amyA/B/C mRNAs after adding maltose (Fig. 4a). When ireA expression was repressed in ΔAoski2 strain, accumulation of the short fragments of the amyA/B/C mRNAs was reduced, whereas the expression level of the full length amyA/B/C mRNAs was not affected (Fig. 4b, Supplementary Fig. 2). This suggested that IreA-dependent cleavage of the amyA/B/C mRNAs occurred even without DTT. Interestingly, we found that the malP mRNA, a prerequisite for the induction of amylolytic genes by maltose in A. oryzae, was degraded in an IreA-dependent manner. The short fragment of the malP mRNA was observed in the ΔAoski2 strain when the 5′-CDS of malP was used as a probe (Fig. 4a). The level of the full length malP mRNA was dramatically reduced between 60 and 90 min after maltose addition, whereas those of the shorter fragments were not. These results suggested that the full length malP mRNA was cleaved for degradation, and that the resulting shorter malP mRNA might transiently emerge and be degraded by cytoplasmic exosomes. When ireA expression was repressed in the ΔAoski2 strain, the decrease in the level of the full length malP mRNA was delayed, accompanied by the delayed appearance of the shorter mRNA (Fig. 4b), suggesting that the malP mRNA might be cleaved by IreA under the condition of amylolytic enzyme production. To confirm whether the malP mRNA cleavage correlated with ER stress, the malP mRNA was detected after DTT treatment of the ΔAoski2 strain. The short malP mRNA fragments were rapidly generated along with the short amyA/B/C mRNA fragments in the ΔAoski2 strain treated with DTT (Supplementary Fig. 3a), which was repressed by ireA repression (Supplementary Fig. 3b). Taken together, these results indicated that the malP mRNA is degraded in an IreA-dependent manner under ER stress.
a Northern blot analysis of the amyA/B/C and malP mRNAs in wild type and ΔAoski2 strains. The mycelia were harvested after incubating in maltose medium for the indicated times to extract total RNAs. The asterisks indicate short mRNA fragments. b Northern blot analysis of the amyA/B/C and malP mRNAs in ΔAoski2 and ΔAoski2/nmtAp-ireA strains. The mycelia were harvested after incubating in maltose medium containing 20 μM thiamine for the indicated times. The asterisks indicate short mRNA fragments.
RIDD of malP mRNA was induced by the production of amylolytic enzymes
We have previously shown that UPR induction by maltose can be avoided by deletion of the transcription factor AmyR, which induces the expression of amylolytic genes upon activation by maltose23. Therefore, we decided to verify whether malP mRNA cleavage, observed when cultured in maltose medium, could be induced by the production of amylolytic enzymes. Toward this, we examined the malP transcripts in the ΔAoski2ΔamyR double-disruption mutant. Northern blot analysis showed that the deletion of amyR repressed the appearance of the 5′-malP mRNA fragments, accompanied by a delayed decrease in the level of the full length malP mRNA in the ΔAoski2 strain (Fig. 5a). This suggested that physiological ER stress induced by amylolytic enzyme production triggers the RIDD of the malP mRNA. As AmyR regulates the expression of secretory α-glucosidases, which hydrolyze maltose to glucose, amyR disruption may affect malP transcription, which is repressed via carbon catabolite repression (CCR). To verify the effect of amyR disruption on the CCR, the malT mRNA encoding a cytoplasmic α-glucosidase, transcription of which is regulated cooperatively with malP32,33, was detected using the 5′-CDS of malT as a probe. Short fragments were not observed when the malT mRNA was analyzed in the ΔAoski2 strain (Fig. 5a), confirming that mRNAs encoding cytoplasmic proteins do not become targets of RIDD. Nevertheless, malT transcription decreased gradually over time and amyR disruption suppressed this decrease. This suggested that malT and malP transcription is repressed by CCR, which is triggered by glucose generated from maltose. Importantly, however, the level of the full length malP transcript decreased faster than that of malT in the ΔAoski2 strain. This suggested that after maltose drastically elevates the production of amylolytic enzymes, the number of malP transcripts is downregulated by two different mechanisms: RIDD and CCR.
a Northern blot analysis of the amyA/B/C and malP mRNAs in ΔAoski2 and ΔAoski2ΔamyR strains. Total RNAs were extracted from mycelia after incubating in maltose medium for the indicated time points. The asterisks indicate short mRNA fragments. b Growth of single and double disruption mutant strains of the Ski complex and amyR. Approximately 1 × 104 conidiospores of each strain were grown for 3 days at 30 °C on minimal agar media containing 1% glucose or maltose as a sole carbon source.
In fission yeast, ski2 deletion leads to increased susceptibility to tunicamycin, an antibiotic widely used to induce ER stress9. Interestingly, the growth of the Aoski2 disruptant was severely impaired on agar media containing maltose but not glucose (Fig. 5b). This carbon source-dependent growth defect was also observed in a disrupted strain of the Ski3 ortholog (AoSki3), another subunit of the Ski complex. amyR deletion rescued the growth defect of the ΔAoski2 and ΔAoski3 strains on maltose medium (Fig. 5b). This suggested that RIDD contributes to maintenance of cellular homeostasis in A. oryzae under conditions that produce amylolytic enzymes, although detailed information regarding the roles of the 3′–5′ mRNA decay in the production of amylolytic enzymes is not available.
Discussion
RIDD is a feedback mechanism that alleviates ER stress caused by the accumulation of misfolded proteins and/or the overload of secretory and membrane proteins in the ER. Treatment of cells with chemicals such as DTT and tunicamycin mimics the accumulation of misfolded proteins in the ER to induce RIDD in many eukaryotes. The decrease in the mRNA levels of secretory hydrolase-coding genes upon treatment with DTT is also observed in filamentous fungi, although it has been attributed to the repression of transcription, called RESS, of target genes. In this study, we showed that RIDD contributes to a DTT-induced decrease in the transcripts of secretory proteins in filamentous fungi. In addition, RIDD of the amyA/B/C and malP mRNAs occurred when amylolytic enzyme production was induced without chemical treatment. A schematic diagram showing the response to physiological ER stress induced by amylolytic enzyme production is illustrated in Fig. 6. To the best of our knowledge, this is the first study to demonstrate that RIDD occurs in eukaryotic microorganisms under natural physiological conditions.
a Expression of amylolytic genes is induced by maltose uptake through MalP, and the unfolded proteins resulting from the synthesis of nascent polypeptides from their mRNAs induce ER stress. b The resulting ER stress activates IreA, leading to HacA activation and RIDD. Thereafter, RIDD and the HacA-mediated UPR and RESS alleviate ER stress. In addition to RIDD and RESS, carbon catabolite repression (CCR) and MalP endocytosis induced by glucose generated from the maltose hydrolysis also contribute to the decrease in amyA/B/C and malP mRNAs. The correctly folded protein is represented by the three-dimensional structural image of α-amylase (PDB ID: 6TAA). The red arrows indicate the activation of transcription factors and CCR.
According to previous studies and our current study, filamentous fungi appear to utilize two mechanisms, RESS and RIDD, to reduce the excess load of secretory proteins in the stressed ER. However, in our experimental condition involving DTT treatment, RIDD might be the major route utilized to decrease the transcripts of secretory proteins in A. oryzae. This was substantiated by several lines of evidence: (i) the replacement of the transcriptional promoter of amyB with that of actA did not suppress the reduction in the amyB mRNA level; (ii) the frameshift mutation in the ER targeting signal of AmyB markedly suppressed the rapid decrease in its transcript; (iii) IreA, but not HacA, was required to decrease amyA/B/C mRNAs; (iv) degradation products of amyA/B/C mRNAs appeared in the ΔAoski2 strain. Although our findings indicated that RIDD plays a major role in the reduction of transcripts encoding secretory proteins in A. oryzae, the slower decrease in the amyB-FS mRNA levels upon DTT treatment reflects the RESS of the amyB mRNA. In other words, the decrease in amyB mRNA could depend completely on RESS unless amyB mRNA is not targeted to the ER. Genome-wide transcriptomic analyses revealed that the transcription of amyR was markedly downregulated in hacAi-expressing A. niger and A. oryza29,30. These observations implied that RESS is regulated by the UPR if the transcription of RESS targets is mediated by AmyR. As ireA-repressed cells expressing hacAi suppressed the reduction in the amyA/B/C mRNA levels treated with DTT, we speculated that RIDD and RESS might be regulated by different pathways. We propose that filamentous fungi use a dual mechanism for repressing the abundance of ER-localized mRNAs to alleviate ER stress. RIDD rapidly removes mRNAs localized in the ER, while RESS prevents de novo synthesis of specific mRNAs. Filamentous fungi can produce 100 g/l of secreted proteins34 and thus may have a unique dual ER stress response system to deal with ER stress induced by the production of high levels of secretory proteins.
Maltose incorporated via MalP is transglycosylated to isomaltose by the cytoplasmic α-glucosidase MalT to activate AmyR, thereby inducing amylolytic enzymes in A. oryzae18,33,35. We have previously shown that the addition of glucose to the media represses the expression of malT and malP32 and accelerates the degradation of MalP via endocytosis35. These findings suggested that MalP-mediated maltose uptake is an upstream event in the regulation of amylolytic gene expression. Interestingly, in addition to catabolite repression for optimizing starch metabolism, RIDD targets the malP transcript for overcoming the stress caused by protein overload in the ER. This observation is consistent with that of our previous report showing that the induction of amylolytic enzyme production by maltose induces the UPR19. These results suggested that the RIDD of the malP transcript might function as part of a feedback control to avoid further loading of amylolytic enzymes into the ER. Loss of the Ski complex, which accumulates mRNA remnants from the cleavage by IreA, caused severe growth inhibition in the maltose medium. Although the mechanism of toxicity remains elusive, clearance of mRNA remnants may be important for the maintenance of cellular homeostasis in A. oryzae.
In the present study, we identified the amyA/B/C and malP transcripts as targets of RIDD under physiological conditions. However, the efficiency of cleavage by IreA may differ; malP mRNA may be more susceptible to cleavage than amyA/B/C mRNAs. Although the underlying reason is not known, this might be attributed to the specificity of IreA for the sequence and the secondary structure of the RNAs. Mammalian Ire1a specifically cleaves the CUGCAG sequence with a stem-loop structure36. In filamentous fungi, an intron within HAC1/hacA pre-mRNA is cleaved by Ire1/IreA in the C(U/C)GCAG sequence of the stem-loop structure37. This suggests that the CUGCAG sequence is also a target of cleavage by IreA in filamentous fungi. A single CUGCAG sequence is present within the 5′-region of both amyA/B/C and malP mRNAs. In particular, the CUGCAG sequence in malP mRNA was located in the loop region of the predicted large stem-loop structure (Supplementary Fig. 4). In contrast, the appearance of multiple short fragments of both amyA/B/C and malP mRNAs suggested that these mRNAs have more than one cleavage site. Further experiments are required to determine the cleavage sites of these mRNAs. More examples of RIDD targets will also help us understand the structural features underscoring the cleavage specificity of IreA in filamentous fungi. We found that the growth of the Ski complex-deficient strains was inhibited not only by maltose and starch as carbon sources but also by xylose and xylan, inducers of xylanolytic and cellulolytic gene expression38 (Supplementary Fig. 5), suggesting that A. oryzae may harbor more targets of physiological RIDD. The identification of a wide range of RIDD targets and their cleavage sites will provide a comprehensive understanding regarding the mechanisms and roles of RIDD in filamentous fungi. The cleavage sites of the RIDD targets in most eukaryotes remain unclear39. Information obtained from studies on filamentous fungi will also be helpful for understanding RIDD in eukaryotic cells.
RIDD function has been well studied in metazoans40,41. In particular, RIDD is involved in insulin secretion via the degradation of insulin mRNA under high-glucose conditions42,43, indicating that RIDD plays an important role under physiological conditions. In microorganisms, the molecular mechanism and role of the UPR have been well studied in S. cerevisiae, whereas RIDD was not found in this yeast. In contrast, recent studies revealed the molecular mechanism of RIDD in S. pombe, although this yeast lacks an HAC1 ortholog. To the best of our knowledge, this is the first study to show that RIDD occurs under physiological conditions in eukaryotic microorganisms. In addition, this study underscores the potential for A. oryzae to be a useful model for elucidating the physiological significance of the ER stress response in eukaryotic microorganisms since both UPR and RIDD are actuated in this filamentous fungus under physiological ER stress condition. Further analysis of RIDD in A. oryzae will improve our understanding of its functions in microorganisms and other eukaryotes.
Methods
Strains and media
A. oryzae NS4 (niaD−, sC−)44 and the ΔligD::loxP pyrG− strain (ΔligD::loxP, sC−, niaD−, pyrG−)45, derived from the wild type strain, A. oryzae RIB40 (National Research Institute of Brewing Stock Culture), were used as recipient strains for gene disruption. The A. oryzae strains used in this study are listed in Supplementary Table 1. Escherichia coli DH5α was used for the construction and propagation of plasmid DNAs. A. oryzae was grown in YPM medium (1% yeast extract, 1% Bacto Peptone, and 2% maltose) or Czapek–Dox (CD) medium (0.5% (NH4)2SO4, 0.05% KCl, 0.2% KH2PO4, 0.05% MgSO4, trace amounts of FeSO4, ZnSO4, CuSO4, MnSO4, Na2B4O7, and (NH4)6Mo7O24, and 1% sugar) supplemented with 0.003% methionine, if required. Glycerol and maltose (1%) were used as carbon sources to not induce and induce amylolytic gene expression, respectively. HIPOLYPEPTON (0.1%; Nihon Pharmaceutical Co., Ltd., Tokyo, Japan) was added to promote growth when A. oryzae strains were pre-cultured in CD medium. For the DTT treatment, 20 mM DTT was added directly to the cultures.
Construction of plasmids and strains
Plasmids encoding wild type and mutant amyB were constructed as follows: The A. oryzae amyB was amplified using polymerase chain reaction (PCR) with oligonucleotides, oTAKA381 and oTAKA382, and then inserted into HindIII/XbaI-digested pUC-niaD46 using the In-Fusion Snap Assembly master mix (Takara Bio, Shiga, Japan) to generate pUC-niaD-amyB. amyB-FS was obtained using QuikChange site-directed mutagenesis with pUC-niaD-amyB as the template DNA and the oligonucleotides YS3 and YS4. A plasmid expressing amyB under the control of the actA promoter was constructed as follows. The vector DNA containing niaD and amyB was PCR-amplified with pUC-niaD-amyB as the template DNA and oligonucleotides, vector-F and vector-R. The resulting vector fragment was assembled with the actA promoter PCR-amplified with the oligonucleotides, PactA-F and PactA-R using In-Fusion Snap Assembly master mix to generate pUC-niaD-PactA-amyB. These plasmids were linearized using NsiI and transformed in A. oryzae ΔamyA/B/C cells.
The plasmid for Aoski2 (AO0900110006369) disruption was constructed as follows: A DNA fragment harboring approximately 1.0 kb upstream sequence of Aoski2 was amplified using PCR and the ski2upsenKpnI and ski2upantiBamHI primers. The amplified fragment was digested with KpnI and BamHI and inserted into KpnI/BamHI-digested pUSC44, yielding pUSC-ski2up. A DNA fragment harboring approximately 1.0 kb downstream sequence of Aoski2 was amplified using the ski2downsenPstI and ski2downantiPstI primers. The amplified fragment was digested with PstI and inserted into PstI-digested pUSC-ski2up, yielding pUSC-∆ski2.
The plasmid for Aoski3 (AO090005001388) disruption was constructed as follows: A DNA fragment harboring approximately 1.0 kb upstream sequence of the Aoski3 5′-region was amplified using the ski3upsenBamHI and ski3upantiXbaI primers. The amplified fragment was digested with BamHI and XbaI and inserted into the BamHI/XbaI-digested pUSC, yielding pUSC-ski3up. A DNA fragment harboring the Aoski3 3′-region was also amplified using the ski3downsenPstI and ski3downantiSphI primers. The amplified fragment was digested with PstI and SphI, and inserted into PstI/SphI-digested pUSC-ski3up, yielding pUSC-∆ski3.
The plasmid for the replacement of the ireA upstream region with the nmtA promoter using the A. nidulans orotidine-5′-decarboxylase gene (pyrG) as the selectable marker was constructed as follows. The DNA fragment of the nmtA promoter associated with the ireA coding region was obtained from PstI/SphI-digested psCPnmtAireA19. The obtained DNA fragment was replaced with the ireA downstream region excised from PstI/SphI-digested p∆ireA::pyrG19 using two distinct ligation reactions, yielding pnmtAp-ireA::pyrG. The DNA fragment obtained from the SpeI/SphI-digested pnmtAp-ireA::pyrG was used to transform the ΔAoski2 pyrG− strain.
The DNA fragment for amyR disruption was obtained from the plasmid p∆amyR::pyrG, as described previously19.
The nucleotide sequences of all primers used for gene disruption are shown in Supplementary Table 2.
Fungal transformation
DNA fragments for gene disruption were introduced into A. oryzae using the polyethylene glycol (PEG)-protoplast method47, and Yatalase (Takara Bio Inc.) was used for protoplast preparation. To complement pyrG deficiency, plasmid pUC/pyrG/niaD48 was introduced into the ΔAoski2 and ΔAoski3 mutants using ΔligD::loxP pyrG− as the host strain.
Fungal cultivation
To examine the reduction in α-amylase mRNA levels after DTT addition, the WT (RIB40) and ΔamyA/B/C strains expressing the WT and mutant amyB, respectively, were grown in YPM medium at 30 °C for 24 h and treated with 20 mM DTT.
To examine the involvement of IreA in the DTT-induced reduction of α-amylase mRNA levels, the WT (ΔligD::loxP pyrG::niaD) and nmtAp-ireA strains were pre-cultured in CD (glycerol) medium containing 0.1% HIPOLYPEPTON and 20 μM thiamine at 30 °C for 24 h, and the mycelia were resuspended in CD (maltose) medium containing 20 μM thiamine. After 60 min-incubation at 30 °C, DTT was added to the medium to a final concentration of 20 mM.
To examine the amyA/B/C and malP mRNA levels after maltose addition without DTT treatment, the WT and ΔAoski2 strains were pre-cultured in CD (glycerol) medium containing 0.1% HIPOLYPEPTON at 30 °C for 24 h, and the mycelia were resuspended in CD (maltose) medium.
Northern blot analysis
A. oryzae mycelia cultured in the liquid medium were harvested over time using an open-ended pipette and Miracloth (Merck Millipore, Billerica, MA, USA), washed with distilled water, and then frozen in liquid nitrogen. The mycelia were ground into a fine powder in liquid nitrogen using a mortar and pestle. The powdered mycelia were suspended in ISOGEN (Nippon Gene Co., Ltd., Tokyo, Japan) and total RNA was extracted according to the manufacturer’s instructions. Northern blot analysis was performed as described previously49. Digoxigenin (DIG)-labeled DNA fragments were synthesized using PCR with the primers shown in Supplementary Table 3 and were used as probes.
Statistics and reproducibility
All experiments were repeated at least twice independently as biological replications and the results are reproducible. All attempts at replication were successful.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Mori, K. Signalling pathways in the unfolded protein response: development from yeast to mammals. J. Biochem. 146, 743–750 (2009).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Adams, C. J., Kopp, M. C., Larburu, N., Nowak, P. R. & Ali, M. M. U. Structure and molecular mechanism of ER stress signaling by the unfolded protein response signal activator IRE1. Front. Mol. Biosci. 6, 11 (2019).
Wu, H., Ng, B. S. H. & Thibault, G. Endoplasmic reticulum stress response in yeast and humans. Biosci. Rep. 34, e00118 (2014).
Maurel, M., Chevet, E., Tavernier, J. & Gerlo, S. Getting RIDD of RNA: IRE1 in cell fate regulation. Trends Biochem. Sci. 39, 245–254 (2014).
Hollien, J. & Weissman, J. S. Decay of endoplasmic reticulum-localized mRNAs during the unfolded protein response. Science 313, 104–107 (2006).
Han, D. et al. IRE1α kinase activation modes control alternate endoribonuclease outputs to determine divergent cell fates. Cell 138, 562–575 (2009).
Hollien, J. et al. Regulated Ire1-dependent decay of messenger RNAs in mammalian cells. J. Cell Biol. 186, 323–331 (2009).
Kimmig, P. et al. The unfolded protein response in fission yeast modulates stability of select mRNAs to maintain protein homeostasis. eLife 1, e00048 (2012).
Mishiba, K. et al. Defects in IRE1 enhance cell death and fail to degrade mRNAs encoding secretory pathway proteins in the Arabidopsis unfolded protein response. Proc. Natl Acad. Sci. USA 110, 5713–5718 (2013).
Miyazaki, T., Nakayama, H., Nagayoshi, Y., Kakeya, H. & Kohno, S. Dissection of Ire1 functions reveals stress response mechanisms uniquely evolved in Candida glabrata. PLoS Pathog. 9, e1003160 (2013).
Meyer, V. Genetic engineering of filamentous fungi — progress, obstacles and future trends. Biotechnol. Adv. 26, 177–185 (2008).
Druzhinina, I. S. & Kubicek, C. P. “Chapter Two—Familiar Stranger: Ecological Genomics of the Model Saprotroph and Industrial Enzyme Producer Trichoderma reesei Breaks the Stereotypes” in Advances in Applied Microbiology (eds. Sariaslani, S. & Gadd, G. M.) pp. 69–147 (Academic Press, 2016).
Machida, M., Yamada, O. & Gomi, K. Genomics of Aspergillus oryzae: learning from the history of koji mold and exploration of its future. DNA Res. 15, 173–183 (2008).
Gomi, K. Regulatory mechanisms for amylolytic gene expression in the koji mold Aspergillus oryzae. Biosci. Biotechnol. Biochem. 83, 1385–1401 (2019).
Tanaka, M. & Gomi, K. Induction and repression of hydrolase genes in Aspergillus oryzae. Front. Microbiol. 12, 677603 (2021).
Murakoshi, Y., Makita, T., Kato, M. & Kobayashi, T. Comparison and characterization of α-amylase inducers in Aspergillus nidulans based on nuclear localization of AmyR. Appl. Microbiol. Biotechnol. 94, 1629–1635 (2012).
Ichikawa, T. et al. Crucial role of the intracellular α-glucosidase MalT in the activation of the transcription factor AmyR essential for amylolytic gene expression in Aspergillus oryzae. Biosci. Biotechnol. Biochem. 85, 2076–2083 (2021).
Tanaka, M., Shintani, T. & Gomi, K. Unfolded protein response is required for Aspergillus oryzae growth under conditions inducing secretory hydrolytic enzyme production. Fungal Genet. Biol. 85, 1–6 (2015).
Ohno, A., Maruyama, J., Nemoto, T., Arioka, M. & Kitamoto, K. A carrier fusion significantly induces unfolded protein response in heterologous protein production by Aspergillus oryzae. Appl. Microbiol. Biotechnol. 92, 1197–1206 (2011).
Yokota, J. et al. Cellular responses to the expression of unstable secretory proteins in the filamentous fungus Aspergillus oryzae. Appl. Microbiol. Biotechnol. 101, 2437–2446 (2017).
Kwon, M. J. et al. The transcriptomic fingerprint of glucoamylase over-expression in Aspergillus niger. BMC Genom. 13, 701 (2012).
Guillemette, T. et al. “Chapter One—Methods for Investigating the UPR in Filamentous Fungi” in Methods in Enzymology, The Unfolded Protein Response and Cellular Stress (eds. Part B. & Conn, P. M.), pp. 1–29 (Academic Press, 2011).
Sims, A. H. et al. Transcriptome analysis of recombinant protein secretion by Aspergillus nidulans and the unfolded-protein response in vivo. Appl. Environ. Microbiol. 71, 2737–2747 (2005).
Pakula, T. M. et al. The effects of drugs inhibiting protein secretion in the filamentous fungus Trichoderma reesei. Evidence for down-regulation of genes that encode secreted proteins in the stressed cells. J. Biol. Chem. 278, 45011–45020 (2003).
Al-Sheikh, H. et al. Endoplasmic reticulum stress leads to the selective transcriptional downregulation of the glucoamylase gene in Aspergillus niger. Mol. Microbiol. 53, 1731–1742 (2004).
Guillemette, T. et al. Genomic analysis of the secretion stress response in the enzyme-producing cell factory Aspergillus niger. BMC Genom. 8, 158 (2007).
Wang, B. et al. Survey of the transcriptome of Aspergillus oryzae via massively parallel mRNA sequencing. Nucleic Acids Res. 38, 5075–5087 (2010).
Carvalho, N. D. et al. Genome-wide expression analysis upon constitutive activation of the HacA bZIP transcription factor in Aspergillus niger reveals a coordinated cellular response to counteract ER stress. BMC Genom. 13, 350 (2012).
Zhou, B. et al. Identification of functional cis-elements required for repression of the Taka-amylase A gene under secretion stress in Aspergillus oryzae. Biotechnol. Lett. 37, 333–341 (2015).
Hunter, A. J., Jin, B. & Kelly, J. M. Independent duplications of α-amylase in different strains of Aspergillus oryzae. Fungal Genet. Biol. 48, 438–444 (2011).
Hasegawa, S., Takizawa, M., Suyama, H., Shintani, T. & Gomi, K. Characterization and expression analysis of a maltose-utilizing (MAL) cluster in Aspergillus oryzae. Fungal Genet. Biol. 47, 1–9 (2010).
Suzuki, K. et al. Distinct mechanism of activation of two transcription factors, AmyR and MalR, involved in amylolytic enzyme production in Aspergillus oryzae. Appl. Microbiol. Biotechnol. 99, 1805–1815 (2015).
Ward, O. P. Production of recombinant proteins by filamentous fungi. Biotechnol. Adv. 30, 1119–1139 (2012).
Hiramoto, T. et al. Endocytosis of a maltose permease is induced when amylolytic enzyme production is repressed in Aspergillus oryzae. Fungal Genet. Biol. 82, 136–144 (2015).
Oikawa, D., Tokuda, M., Hosoda, A. & Iwawaki, T. Identification of a consensus element recognized and cleaved by IRE1α. Nucleic Acids Res. 38, 6265–6273 (2010).
Saloheimo, M., Valkonen, M. & Penttilä, M. Activation mechanisms of the HACI-mediated unfolded protein response in filamentous fungi. Mol. Microbiol. 47, 1149–1161 (2003).
Noguchi, Y. et al. Genes regulated by AoXlnR, the xylanolytic and cellulolytic transcriptional regulator, in Aspergillus oryzae. Appl. Microbiol. Biotechnol. 85, 141–154 (2009).
Le Thomas, A. et al. Decoding non-canonical mRNA decay by the endoplasmic-reticulum stress sensor IRE1α. Nat. Commun. 12, 7310 (2021).
Huang, S., Xing, Y. & Liu, Y. Emerging roles for the ER stress sensor IRE1α in metabolic regulation and disease. J. Biol. Chem. 294, 18726–18741 (2019).
Coelho, D. S. & Domingos, P. M. Physiological roles of regulated Ire1 dependent decay. Front. Genet. 5, 76 (2014).
Lipson, K. L. et al. Regulation of insulin biosynthesis in pancreatic beta cells by an endoplasmic reticulum-resident protein kinase IRE1. Cell Metab. 4, 245–254 (2006).
Eletto, D., Eletto, D., Boyle, S. & Argon, Y. PDIA6 regulates insulin secretion by selectively inhibiting the RIDD activity of IRE1. FASEB J. 30, 653–665 (2016).
Yamada, O., Lee, B. R. & Gomi, K. Transformation system for Aspergillus oryzae with double auxotrophic mutations, niaD and sC. Biosci. Biotechnol. Biochem. 61, 1367–1369 (1997).
Mizutani, O., Masaki, K., Gomi, K. & Iefuji, H. Modified Cre-loxP recombination in Aspergillus oryzae by direct introduction of Cre recombinase for marker gene rescue. Appl. Environ. Microbiol. 78, 4126–4133 (2012).
Tanaka, M., Ito, K., Matsuura, T., Kawarasaki, Y. & Gomi, K. Identification and distinct regulation of three di/tripeptide transporters in Aspergillus oryzae. Biosci. Biotechnol. Biochem. 85, 452–463 (2021).
Gomi, K., Iimura, Y. & Hara, S. Integrative transformation of Aspergillus oryzae with a plasmid containing the Aspergillus nidulans argB gene. Agric. Biol. Chem. 51, 2549–2555 (1987).
Ichinose, S., Tanaka, M., Shintani, T. & Gomi, K. Improved α-amylase production by Aspergillus oryzae after a double deletion of genes involved in carbon catabolite repression. Appl. Microbiol. Biotechnol. 98, 335–343 (2014).
Tanaka, M., Tokuoka, M., Shintani, T. & Gomi, K. Transcripts of a heterologous gene encoding mite allergen Der f 7 are stabilized by codon optimization in Aspergillus oryzae. Appl. Microbiol. Biotechnol. 96, 1275–1282 (2012).
Acknowledgements
We thank Dr. Osamu Mizutani for providing the ΔligD::loxP pyrG− mutant strain. This study was supported by JSPS KAKENHI (grant numbers 09J06211 and 16K15084), the JSPS Core-to-Core Program (Advanced Research Networks) titled “Establishment of international agricultural immunology research-core for a quantum improvement in food safety”, and an NISR Research Grant from the Noda Institute for Scientific Research, Japan. We would like to thank Editage (www.editage.com) for English language editing.
Author information
Authors and Affiliations
Contributions
M.T., K.G., and T.S. designed the study; M.T., S.S., S.Z., J-I.Y., and Y.S. performed the study; M.T., K.G., Y.K., Y.Y., and T.S. analyzed the data; M.T., K.G., and T.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Kai Heimel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Meritxell Requilme and Christina Karlsson Roesnthal. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tanaka, M., Zhang, S., Sato, S. et al. Physiological ER stress caused by amylase production induces regulated Ire1-dependent mRNA decay in Aspergillus oryzae. Commun Biol 6, 1009 (2023). https://doi.org/10.1038/s42003-023-05386-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05386-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.