Abstract
In rice (Oryza sativa) tissue culture, callus can be induced from the scutellum in embryo or from the vasculature of non-embryonic organs such as leaves, nodes, or roots. Here we show that the auxin signaling pathway triggers cell division in the epidermis of the scutellum to form an embryo-like structure, which leads to callus formation. Our transcriptome data show that embryo-, stem cell-, and auxin-related genes are upregulated during scutellum-derived callus initiation. Among those genes, the embryo-specific gene OsLEC1 is activated by auxin and involved in scutellum-derived callus initiation. However, OsLEC1 is not required for vasculature-derived callus initiation from roots. In addition, OsIAA11 and OsCRL1, which are involved in root development, are required for vasculature-derived callus formation but not for scutellum-derived callus formation. Overall, our data indicate that scutellum-derived callus initiation is regulated by an embryo-like development program, and this is different from vasculature-derived callus initiation which borrows a root development program.
Similar content being viewed by others
Introduction
Callus is a group of fast-dividing parenchyma cells, and many types of callus could be formed in tissue culture or upon wounding1,2,3. Studies in the dicot model plant Arabidopsis thaliana and the monocot model plant rice (Oryza sativa) indicated that callus could be induced from different regeneration-competent cells of explants in tissue culture. For example, the vascular adult stem cells may serve as regeneration-competent cells to initiate callus (hereafter described as vasculature-derived callus) in Arabidopsis4,5,6,7 and rice7, and the epidermal cells of the rice scutellum, which is the rice cotyledon in embryo, can also act as regeneration-competent cells responsible for the initiation of callus8,9 (hereafter described as scutellum-derived callus).
Initiation of the vasculature-derived callus borrows the root developmental pathway4,5,6,7,10,11,12,13,14,15,16. In Arabidopsis, lateral or adventitious roots are initiated from adult stem cells in vasculature, e.g., xylem pole pericycle cells in roots or procambium and some vascular parenchyma cells in leaves, and those adult stem cells are responsible for the initiation of vasculature-derived callus in tissue culture4,5,6,7,17. In rice, vasculature-derived callus can be initiated from outer bundle sheath cells in the immature region of leaves or from phloem-pole pericycle cells in roots7, and the phloem-pole pericycle cells also initiate lateral roots in rice root system formation.
The cellular structure of vasculature-derived callus resembles the root primordium (RP) and root apical meristem (RAM)6,7,13. Studies in Arabidopsis revealed that many genes related to RP/RAM are highly expressed in vascular-derived callus, including the WUSCHEL-RELATED HOMEOBOX (WOX) transcription factor family genes AtWOX5 and AtWOX7 (AtWOX5/7) and the APETALA2 (AP2)-like transcription factor family genes PLETHORA1 and 2 (AtPLT1/2) and AtPLT3/5/76,13,14,18,19. In rice, OsWOX5 is also highly expressed in vasculature-derived callus7.
In rice tissue culture, scutellum-derived callus is initiated from the epidermal cells of the scutellum8,9. Genetic, transcriptome, and epigenome analyses have revealed many gene networks in the formation of scutellum-derived callus in rice9,20,21,22,23. However, our knowledge on scutellum-derived callus formation in rice is still limited, and the developmental and molecular strategies adopted in scutellum-derived callus formation in comparison with vasculature-derived callus formation need to be explored. In this study, we propose that scutellum-derived callus initiation might borrow the embryo development program in rice.
Results
Auxin signaling pathway mediates scutellum-derived callus initiation
To study the role of auxin in scutellum-derived callus formation, we cultured mature seeds of wild-type rice on callus-inducing medium (CIM) with a high level of auxin. Many calli were visible to the naked eye at 20 days of culture (Fig. 1a). Overexpression of OsMicroRNA393a (35Spro:OsMIR393a) and OsMIR393b (35Spro:OsMIR393b), which can target and degrade the mRNA of the auxin receptor genes TRANSPORT INHIBITOR RESPONSE1 (OsTIR1) and AUXIN SIGNALING F BOX PROTEIN2 (OsAFB2)24, resulted in the loss of scutellum-derived callus formation on CIM25 (Fig. 1b, c; Supplementary Fig. 1). Similarly, double mutations in OsTIR1 and OsAFB224 also led to defects in scutellum-derived callus formation (Fig. 1d; Supplementary Fig. 1). Therefore, auxin could be the key hormone that triggers scutellum-derived callus formation in rice.
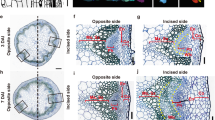
a–d Tissue culture of rice mature seeds from wild-type (WT) (a), 35Spro:OsMIR393a (b), 35Spro:OsMIR393b (c), and Ostir1 Osafb2 (d) on CIM for 20 d. We tested more than 90 wild-type seeds, and all of them formed scutellum-derived callus. We tested more than 30 seeds from 35Spro:OsMIR393a, 35Spro:OsMIR393b, or Ostir1 Osafb2, and none of them formed scutellum-derived callus. e–i Sections showing callus formation from scutellum (e) of wild-type rice seeds on CIM at t0 (f), 2 d (g), 5 d (e, h), and 7 d (i). See Supplementary Fig. 2 for details. j–o Sections showing cell division from scutellum of 35Spro:OsMIR393b (j–l) and Ostir1 Osafb2 (m–o) seeds on CIM at t0 (j, m), 2 d (k, n), and 7 d (l, o). Scale bars, 5 mm (a–d), 500 μm (e), 50 μm (f–o).
To study the cellular origin and developmental progress during scutellum-derived callus initiation, we analyzed sections of wild-type mature seeds on CIM. Callus was initiated from the epidermal cells of the scutellum (Fig. 1e)8. The epidermal cell started to divide at 2 days of culture on CIM, resulting in an apical cell and a basal cell (Fig. 1f, g; Supplementary Fig. 2). The apical and basal cells continued to divide and form an embryo-like structure, in which the apical cell had formed a globular structure and the basal cell had formed a suspensor-like structure at 5 days of culture on CIM (Fig. 1h; Supplementary Fig. 2). Cell division continued in the globular structure, which gradually detached from the suspensor-like structure (Fig. 1i; Supplementary Fig. 2). The detached globular structure continued cell division to form the scutellum-derived callus (Fig. 1i; Supplementary Fig. 2).
Cell division occurred in the epidermal cells of 35Spro:OsMIR393b and the Ostir1 Osafb2 double mutant (Fig. 1j–o), but the dividing cells could not further develop to form the embryo-like structure (Fig. 1l, o). Therefore, no scutellum-derived callus could be formed and detached from the scutellum.
To confirm that the auxin signaling pathway is essential for rice scutellum-derived callus formation on CIM, we created the mutations in AUXIN RESPONSE FACTOR5 (OsARF5) by CRISPR/Cas9 (Supplementary Fig. 3). The Osarf5 mutants showed partially defective formation of scutellum-derived callus (Fig. 2).
a–c Phenotype of callus formation from the scutella of wild-type (a) Osarf5-18 (b) and Osarf5-38 (c) on CIM at 20 d. d Statistical analysis of scutellum-derived callus formation in wild-type, Osarf5-18, and Osarf5-38 on CIM at 20 d. The individual values are indicated by dots. The data are presented as mean values ± s.e.m. from four biological replicates (10 calli in each replicate). **P < 0.01 in two-sided Student’s t-test. Scale bars, 5 mm (a–c).
Analysis of the β-glucuronidase (GUS) marker line DR5pro:GUS showed that the auxin response was not activated in the scutellum before tissue culture (Supplementary Fig. 4a, b, f, g), and was highly active in the dividing cells in the epidermis of the scutellum (Supplementary Fig. 4c–e, h–j) and in the detached globular structure (i.e. scutellum-derived callus) (Supplementary Fig. 4e). Analysis of OsTIR1pro:GUS showed that OsTIR1 was expressed in the scutellum at all stages in tissue culture (Supplementary Fig. 4k–t).
Overall, these data indicate that the auxin signaling pathway is required for the proper cell division of the scutellum epidermal cell to initiate an embryo-like structure, which leads to the formation of scutellum-derived callus.
Initiation of scutellum-derived and vasculature-derived calli adopts different molecular strategies
We next tested whether scutellum-derived callus formation and vasculature-derived callus formation share similar genetic pathways in rice. Scutellum-derived callus showed positive staining of Sudan red (Fig. 3a), while vasculature-derived callus formed from the root explant did not (Fig. 3b), indicating that scutellum-derived callus but not vasculature-derived callus has the embryo-like cell identity that is abundant in lipids. The indole-3-acetic acid inducible11 (Osiaa11) mutant, which is defective in lateral root formation26, showed defective vasculature-derived callus formation from the primary root on CIM (Fig. 3c, d). The crown rootless1 (Oscrl1) mutant, which is defective in adventitious root formation from the node27, showed defective vasculature-derived callus formation from the node on CIM (Fig. 3e, f). However, normal scutellum-derived callus was able to be produced from the mature seeds of both Osiaa11 and Oscrl1 culture on CIM (Fig. 3g, h). In addition, the vasculature-derived callus marker gene OsWOX57, which is a root apical stem cell niche-related gene28,29, was not highly induced during the division of the scutellum epidermal cells in scutellum-derived callus initiation (Fig. 3i–k), while its expression was induced in the vasculature of the embryo to form vasculature-derived callus (Fig. 3l). These data indicate that rice vasculature-derived callus adopts OsIAA11/OsCRL1/OsWOX5-mediated root development program, while scutellum-derived callus does not.
a, b Sudan red staining of scutellum-derived callus from mature seeds (a) or vasculature-derived callus from root (b) on CIM at 10 d. c, d Primary roots of wild-type (c) and Osiaa11 (d) tissue-cultured on CIM for 20 d. e, f Nodes of wild-type (e) and Oscrl1 (f) tissue-cultured on CIM for 14 d. g, h Seeds of Osiaa11 (g) and Oscrl1 (h) tissue-cultured on CIM for 20 d. i–l In situ hybridization using anti-OsWOX5 probe in scutellum (i–k) and vasculature (l) of wild-type embryos in mature seeds at 2 d (j) or 5 d (k, l). Seeds cultured on N6 medium without hormone treatment for 2 d served as the control (i), because t0-seeds were dry and unsuitable for in situ hybridization. Some vasculatures in the embryo can initiate vasculature-derived callus with OsWOX5 expression (l). Scale bars, 1 mm (a, b), 5 mm (c–h), 50 μm (i–k), 500 μm (l).
Transcriptome framework of scutellum-derived callus formation
We carried out RNA-seq analysis using mature embryos of wild-type and 35Spro:OsMIR393b at 2, 5 and 7 days of culture on CIM, compared with the control (Supplementary Fig. 5). The analysis of wild-type transcriptome might reveal the genes involved in scutellum-derived callus formation, and the transcriptome comparison of the wild-type and 35Spro:OsMIR393b could indicate the genes regulated by the auxin signaling pathway during scutellum-derived callus formation.
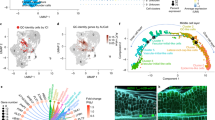
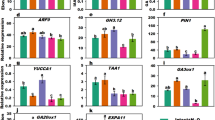
Weighted Correlation Network Analysis (WGCNA)30 was performed and nine modules were identified (Fig. 4a, b, Supplementary Data 1). The module eigengene (ME) turquoise showed a high correlation with scutellum-derived callus formation in the wild-type background but not in the 35Spro:OsMIR393b background (Fig. 4b), indicating the module turquoise might be controlled by the auxin signaling pathway during scutellum-derived callus formation. Gene Ontology (GO) enrichment (Supplementary Fig. 6) analyses showed that genes related to cell division as well as many other biological processes were enriched in the module turquoise. We identified 229 transcription factors in the top 30% hub genes of the module turquoise, including some key genes related to embryo regulation [e.g. the Arabidopsis LEAFY COTYLEDON1 (AtLEC1) homolog OsLEC1, the Arabidopsis ABSCISIC ACID INSENSITIVE3 (AtABI3) homolog VIVIPAROUS1 (OsVP1), and O. sativa LEC2 and FUSCA3 Like1 (OsLFL1)], stem cell regulation (OsWOX2/9c and OsPLTs), and auxin production, signaling, and transport [OsYUCCA1, OsIAAs, OsARFs, and OsPIN-FORMEDs (OsPINs)] (Fig. 4c, d). The RNA-seq data showed that expression levels of these key genes were gradually upregulated during scutellum-derived callus formation on CIM in the wild-type background but this upregulation was severely impaired in the 35Spro:OsMIR393b background (Fig. 4d). Reverse transcription-polymerase chain reaction (RT-PCR) analyses confirmed that OsLEC1, OsWOX2, and OsWOX9C were upregulated during scutellum-derived callus formation from wild-type rice seeds on CIM (Supplementary Fig. 7).
a RNA-seq and WGCNA analysis of scutella from wild-type and 35Spro:OsMIR393b at 2, 5, and 7 d cultured on CIM. Wild-type and 35Spro:OsMIR393b scutella cultured on N6 medium for 2 d without hormone treatment served as the control, because t0-seeds were dry and unsuitable for analysis. Genes with high correlations are grouped into a module, and different modules are indicated by different colors. b Correlation analyses between module eigengene (ME) values (ME turquoise, ME green, ME black, ME blue, ME brown, ME red, ME pink, ME yellow, ME gray) and each culture conditions (N6, CIM 2 days, CIM 5 days, CIM 7 days). ME value is the representative of the expression profile of all genes in the corresponding module. Numbers indicate the correlation coefficient, and the numbers in brackets indicate P values for the correlation. The ME turquoise showed a high correlation with scutellum-derived callus formation in the wild-type background but not in the 35Spro:OsMIR393b background. Therefore, the genes in module turquoise were used for further analysis in (c, d). c Bar chart analysis of the frequency of transcription factor families. d Relative expression levels of embryo-, stem cell-, and auxin-related genes. e Overlapped genes of upregulated genes in vasculature- or scutellum-derived callus formation and embryo-, leaf-, root-, or shoot-specific expression genes are identified. The percentage of overlapped gene numbers in tissue/organ-specific genes are shown by heatmap.
We further analyzed transcriptome correlation of callus and rice tissues or organs. We collected RNA-seq data of four rice tissues or organs (i.e. embryo31, leaf32, root33, and shoot33), and identified tissue/organ-specific expression genes. By comparison of upregulated genes from the RNA-seq data of vasculature-derived callus7 and scutellum-derived callus (data from this study) with those tissue/organ-specific expression genes, we found that root-specific genes were highly enriched in the upregulated genes during vasculature-derived callus formation, whereas embryo-specific genes were highly enriched in the upregulated genes during scutellum-derived callus formation (Fig. 4e).
OsLEC1 is required for scutellum-derived callus initiation
To confirm the auxin-activated genes detected from the RNA-seq data are required for scutellum-derived callus formation in rice, we selected the embryo-specific gene OsLEC1 for further analysis. The Oslec1 mutants31 showed partially defective scutellum-derived callus formation (Fig. 5a–g). Successive cell division of the scutellum epidermis was lost and the embryo-like structure could not be formed in the Oslec1 mutants in tissue culture (Fig. 5h–j). Overexpression of OsLEC1 also resulted in partially defective callus formation31, suggesting that the proper level of OsLEC1 expression might be important for scutellum-derived callus formation. The OsLEC1pro:GUS marker line31 indicated that the OsLEC1 promoter might be highly activated in the scutellum-derived callus (Supplementary Fig. 8a, b). The RT-PCR result showed that OsLEC1 was expressed during scutellum-derived callus formation but was not expressed during vasculature-derived callus formation from the primary root (Fig. 5k). In addition, the Oslec1 primary root could produce vasculature-derived callus (Fig. 5l). The study in AtLEC genes also showed that AtLECs are not required for organ regeneration from vasculature-derived callus in Arabidopsis34. qRT-PCR analysis revealed that the OsLEC1 expression level is significantly reduced in the Osarf5 mutants compared with the wild-type when mature seeds were cultured on CIM (Supplementary Fig. 8c), indicating that OsLEC1 might be upregulated by the OsARF5-mediated auxin signaling pathway35. Together, these data indicate that OsLEC1 is involved in auxin-mediated scutellum-derived callus formation from the rice scutellum but is not required for vasculature-derived callus formation.
a–f Rice seeds from wild-type (a, d), Oslec1-1 (b, e), and Oslec1-2 (c, f) tissue-cultured on CIM for 26 d. g Statistical analysis of scutellum-derived callus formation in wild-type, Oslec1-1, and Oslec1-2 on CIM at 16 d. Error bars show SEM with nine biological replicates (10 calli in each replicate). The individual values are indicated by dots. **P < 0.01 in two-sided Student’s t-test compared with the wild-type control. h–j Sections showing cell division of scutellum epidermis from wild-type (h), Oslec1-1 (i) and Oslec1-2 (j) seeds on CIM at 7 d. k RT-PCR analysis of OsLEC1 expression pattern in vasculature-derived callus and scutellum-derived callus (33 cycles). OsUBQ5 serves as the control (30 cycles). Numbers indicate the days cultured on CIM. The bands of OsUBQ5 and OsLEC1 are derived from different gels. The bands from vasculature-derived callus and scutellum-derived callus were separately collected from the same gel, and the experiments were performed under the same conditions. The predicted band sizes were indicated. Two biological repeats were analyzed and showed the same result. l Primary roots of Oslec1-1 tissue-cultured on CIM for 20 d, showing vasculature-derived callus formation. See Fig. 3c for the wild-type control. Scale bars, 1 cm (a–c), 5 mm (d–f), 50 μm (h–j), 1 mm (l).
Discussion
Auxin is the key hormone that triggers initiation of both scutellum-derived callus and vasculature-derived callus. However, there are some differences between the two types of calli (see the model in Fig. 6).
First, the two types of calli initiate from different regeneration-competent cells. In Arabidopsis, vasculature-derived callus initiates from the xylem-pole pericycle cells in roots4,5,6 or procambium and some vascular parenchyma cells in leaf7,17. In rice, vasculature-derived callus initiates from phloem-pole pericycle cells in roots or outer bundle sheath cells in the immature region of leaves7. In rice, the epidermal cells of the scutellum may serve as the regeneration-competent cells to initiate scutellum-derived callus8. Overall, vascular adult stem cells are responsible for vasculature-derived callus initiation, while epidermis of embryo could be the primary source to initiate scutellum-derived callus.
Second, the two types of calli show different developmental morphologies. The tissue structure of vasculature-derived callus resembles RP/RAM6,10,11 and comprises three cell layers, i.e. the outer cell layer similar to epidermis and lateral root cap, the middle cell layer with quiescent center-like identity, and the inner cell layer similar to vascular initials in the RAM13,19. Initiation of scutellum-derived callus resembles embryogenesis. The epidermis of scutellum divides and developments into an embryo-like morphology with a globular structure and a suspensor-like structure.
Third, scutellum-derived and vasculature-derived calli may require some different molecular and developmental programs for their initiation. In Arabidopsis, vasculature-derived callus initiation requires root-organogenesis-related genes, e.g. AtLBD16, AtWOX5, AtPLTs, and AtIAA1410,12,13,14,16,18,36. In rice, OsCRL1 (the AtLBD16 homologue), OsWOX5 (the AtWOX5 homologue), and OsIAA11 (the AtIAA14 homologue) are also required for vasculature-derived callus formation7,16. In contrast, initiation of scutellum-derived callus does not require some of the root-organogenesis-related genes in rice (e.g. OsCRL1 and OsIAA11). Instead, genes involved in embryo development are required in scutellum-derived callus formation in rice (e.g. OsLEC1). Overall, vasculature-derived callus initiation borrows root development program, whereas scutellum-derived callus initiation might borrow the embryo development program.
The developmental similarities of scutellum-derived callus and somatic embryogenesis need to be further explored. Somatic embryogenesis might also borrow some pathways from embryo development. In Arabidopsis, somatic embryogenesis can be initiated from the epidermis in the junction of the shoot apical meristem and the cotyledon in embryo controlled by a high level of auxin37. In Arabidopsis, somatic embryogenesis essentially involves two groups of genes38. The first group consists of stem cell-related genes, such as AtWUS39,40,41, AtWOX237, BABYBOOM (AtBBM, also known as AtPLT4)42,43, and EMBRYOMAKER (AtEMK, also known as AtPLT5)44. The second group consists of embryo-related genes, including AtLEC1 encoding the HAP3 subunit of the CCAAT-binding transcription factor45,46,47, the B3-domain genes AtLEC248,49,50,51, FUSCA3 (AtFUS3)51,52,53, AtABI347, and the MADS-box transcription factor gene AGAMOUS-LIKE15 (AtAGL15)54,55,56,57. Some homologues of these genes were upregulated and involved in scutellum-derived callus in rice. It will be interesting to compare the initiation processes of scutellum-derived callus and somatic embryogenesis in the future.
There are many types of calli induced in response to wounding or in tissue culture1,2,3,58,59. It will be interesting to identify more types of cells that can serve as regeneration-competent cells to initiate callus, to analyze how auxin can induce different types of calli from different types of explants, to test whether/how different types of calli can undergo organ regeneration and/or somatic embryogenesis, to study whether/how different types of calli can be converted to each other, and to see whether different types of calli share some common molecular pathways (e.g. the stem cell-related pathway) during their initiation in the future. For example, scutellum-derived and vasculature-derived calli might share some common regulatory pathways20,21, and PLT genes were upregulated during initiation of the two types of calli (Fig. 4c, d)6,13,14,19,21. The answers to these questions will improve our understanding of callus formation in the future.
Methods
Plant materials
Oryza sativa L. ssp. Japonica ‘Nipponbare’, ‘Taichung 65’, and Indica ‘Kasalath’ were used as the wild-type lines. The 35Spro:MIR393a, 35Spro:MIR393b, Ostir1 Osafb2, Oslec1-1, Oslec1-2, DR5pro:GUS, and OsTIR1pro:GUS transgenic plants (Nipponbare background) as well as Oscrl1 (Taichung 65 background) and Osiaa11 (Kasalath background) have been described in previous studies23,27,31,60,61. To generate OsARF5 mutants, OsARF5-specific sequences (5′-AGTATCTGGGTCTGCCTGAT-3′ and 5′-GGAATATCTTCTCTGCGGCA-3′) were used as the targets for Cas9 to mutate OsARF5 by CRISPR/Cas962.
Tissue culture
For time-lapse histological observations of callus induction from the scutellum, sterile seeds of wild type or mutants were placed on N6 medium (0.5 g l–1 MES, 3% w/v sucrose, and 0.4% w/v phytagel, pH 5.8) or CIM medium (N6 basal medium supplemented with 10 μM 2, 4-dichlorophenoxyacetic acid, pH 5.8) and cultured in a growth chamber under a 16-h light, 28 °C/8-h dark, 24 °C cycle16. The scutellum was detached as described previously23 and fixed at 4 °C in 2.5% v/v glutaraldehyde in phosphate buffer (0.1 M, pH 7.0). Scutella collected from dry seeds and fixed directly in 2.5% v/v glutaraldehyde served as the t0 control.
For vasculature-derived callus induction from the primary root, seeds of Kathalath and the Osiaa11 mutant were sterilized, soaked in water overnight at 37 °C, sown on Murashige and Skoog (MS) medium63, and then grown vertically in a growth chamber for 3 days under a 16-h light, 28 °C/8-h dark, 26 °C cycle. The primary roots of 3-d-old seedlings were cut and cultured on N6 medium or CIM.
For vasculature-derived callus induction from the stem base (the node), seeds of Taichung 65 and the Oscrl1 mutant were sterilized and then cultured on MS medium for 3 days under a 16-h light, 28 °C/8-h dark, 26 °C cycle. The 3-d-old seedings were cut and cultured on CIM.
Three independent repeats were analyzed to confirm the results of phenotype analysis.
In situ hybridization and Sudan red staining
In situ hybridization was performed as described previously using the probe for OsWOX57. Sudan red staining was performed according to the published protocol64.
Histochemical detection of GUS activity
For β-glucuronidase (GUS) staining, tissues were incubated in X-Gluc solution overnight at 37 °C as described previously61, and then fixed in pre-chilled 2.5% v/v glutaraldehyde in phosphate buffer (0.1 M, pH 7.0) overnight before being photographed and/or sectioned.
Semi-thin sectioning
Semi-thin sectioning was performed as described previously65. Briefly, tissues were fixed overnight at 4 °C in 2.5% v/v glutaraldehyde in phosphate buffer (0.1 M, pH 7.0), and then washed three times in phosphate buffer (0.1 M, pH 7.0) for 15 min at each step. The samples were dehydrated through an ethanol series (30, 50, 70, 80, 90, 95, and 100% ethanol, 15 min each), then immersed in absolute acetone for 20 min. After substitution with a 1:1 mixture of absolute acetone and Spurr’s resin mixture for 1 h, the tissues were transferred to a 1:3 mixture of absolute acetone and Spurr’s resin mixture for 3 h, and then embedded in pure Spurr’s resin mixture overnight. The samples were heated at 70 °C for more than 9 h. The embedded samples were sectioned at 2-μm thickness using a rotary microtome (Leica) and stained or not stained (for GUS-stained tissues) by basic fuchsin before observations using a Nikon Eclipse 80i microscope equipped with a Nikon C-C phase contrast turret condenser (Nikon).
Paraffin sectioning
For paraffin sectioning, the scutella of Nipponbare seeds cultured on N6 medium or CIM were fixed in ethanol-acetic acid (3:1, v/v), dehydrated through an ethanol-xylene series, stained with safranin, and embedded in paraffin. Then, 10-μm sections were cut with a microtome, mounted on slides covered with gelatin, deparaffinized in xylene, and rehydrated through an ethanol series before observations using a Nikon Eclipse 80i microscope equipped with a Nikon C-C phase contrast turret condenser (Nikon).
Scanning electron microscope (SEM)
Plant tissues were fixed overnight at 4 °C in 2.5% glutaraldehyde in 0.1 M phosphate buffer (pH 7.0). Then the samples were prepared for SEM according to the previous method31 in Bio-ultrastructure Analysis Lab of Analysis Centre of Agrobiology and Environmental Sciences, Zhejiang University.
RNA-seq analysis
For scutella collection, seeds of wild-type rice (Nipponbare) and 35Spro:MIR393b were separately cultured on N6 medium (2 d, the control) and CIM (2, 5, 7 d). For each sample, scutella from 10 seeds were used to extract total RNA. Library construction and deep sequencing were carried out using the Illumina HiSeq 4000 platform (LC Bio, Hangzhou, China). The RNA-seq analysis (LC Bio, Hangzhou, China) was carried out using the reference genome in the Rice Annotation Project (RAP) database (https://rapdb.dna.affrc.go.jp/index. html) by the HISAT package v2.066. Sequence-dependent bias and amplification noise were removed using UMI-tools v1.0.067. The mapped reads for each sample were assembled using StringTie v1.3.468. The differentially expressed mRNAs and genes were selected with the following criteria: log2 (fold change) >1 or log2 (fold change) < −1, and with statistical significance (p < 0.05) using the R package – edgeR69.
WGCNA analysis were performed as previously described30. The parameters of WGCNA program were as follows: gene expression > 1; soft threshold = 1 (estimate value); deep split = 2; min module size = 30; merge cut height = 0.1. In each module, the genes with eigengene-based connectivity value (|KME|) > 0.9 and topological overlap measure (TOM) value > 0.25 were regarded as hub genes.
The list of transcription factors in Oryza sativa subsp. japonica was from Plant Transcription Factor Database (http://planttfdb.gao-lab.org/)70. The GO analysis and heat-map of differentially expressed genes normalized by Z-score was constructed using tools in Omicstudio (https://www.omicstudio.cn/login).
Transcriptome sequencing data of the rice embryo (GSE179838)31, the rice leaf (GSE157400)32, the rice root (GSE217725)33, the rice shoot (GSE217725)33, and vasculature-derived callus from rice leaf explants (GSE86869)7 were published previously. The sequencing data were filtered using fastp v0.23.271 with default parameters and aligned to the Oryza sativa genome (IRGSP-1.0) using hisat2 v2.2.172, and gene expression was quantified using featurecounts v2.0.172. To identify tissue/organ-specific expression genes, the mean TPM values among different replicates for each tissue/organ were used, and the maximum mean TPM value was selected as the representative for multiple time points. A specificity measure (SPM)73 threshold of 0.90 was applied to determine tissue/organ-specific expression genes. We identified 767 embryo-specific genes, 474 leaf-specific genes, 616 root-specific genes, and 133 shoot-specific genes. In the transcriptome data of vasculature- or scutellum-derived callus, the genes that were upregulated during the induction process on CIM at a 1.5-fold change threshold were collected. For the transcriptome with replicates, differential gene expression was determined using Deseq2 v1.36.074 with an adjusted p-value threshold of <0.05; for the transcriptome without replicates, fold change was calculated based on TPM values. The proportion of upregulated genes among tissue/organ-specific expression genes was calculated for both transcriptomes and visualized using a heatmap generated by pheatmap v1.0.12 (Kolde R, 2019. pheatmap: Pretty Heatmaps. R package version 1.0.12, https://CRAN.R-project.org/package=pheatmap).
RT-PCR and qRT-PCR
RT-PCR and qRT-PCR were performed as previously described11,61. UBIQUITIN5 (OsUBQ5) served as the control. The primers used for PCR are listed in Supplementary Table 1.
Statistics and reproducibility
Two-sided Student’s t-test was used in this study for statistical analysis. For callus weight analysis, four (Fig. 2d) or nine (Fig. 5g) biological replicates were performed. For qRT-PCR analysis, three biological replicates were performed. Details of statistics and reproducibility are described in figure legends or method.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data and genetic materials used in this study are available from the corresponding authors upon request. The RNA-seq data obtained in this study have been deposited in the Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE179594, and can be accessed at http://xulinlab.cemps.ac.cn/. Sequence data from this article can be found in the Rice Annotation Project (https://rapdb.dna.affrc.go.jp/index.html) under the following accession numbers: OsTIR1 (Os05g0150500), OsAFB2 (Os04g0395600), OsIAA11 (Os03g0633500), OsCRL1 (Os03g0149000), OsPIN1a (Os06g0232300), OsPIN1c (Os11g0137000), OsPIN1d (Os12g0133800), OsYUCCA1 (Os01g0645400), OsIAA4 (Os01g0286900), OsIAA5 (Os01g0675700), OsIAA26 (Os09g0527700), OsARF4 (Os01g0927600), OsARF5 (Os02g0141100), OsARF8 (Os02g0628600), OsARF12 (Os04g0671900), OsARF13 (Os04g0690600), OsARF16 (Os06g0196700), OsARF19 (Os06g0702600), OsARF23 (Os11g0523800), OsWOX2 (Os01g0840300), OsWOX9C (Os05g0564500), OsPLT1 (Os04g0653600), OsPLT2 (Os06g0657500), OsPLT5 (Os01g0899800), OsPLT6 (Os11g0295900), OsPLT7 (Os03g0770700), OsPLT9 (Os03g0232200), OsVP1 (Os01g0911700), OsLEC1 (Os02g0725700), and OsLFL1 (Os01g0713600). Uncropped gel images are available in Supplementary Fig. 9. Numerical source data in Figs. 2d, 4c, 5g, and Supplementary Fig. 8c are shown in Supplementary Data 2.
References
Ikeuchi, M., Sugimoto, K. & Iwase, A. Plant callus: mechanisms of induction and repression. Plant Cell 25, 3159–3173 (2013).
Ikeuchi, M. et al. Molecular mechanisms of plant regeneration. Annu. Rev. Plant Biol. 70, 377–406 (2019).
Xu, L. & Huang, H. Genetic and epigenetic controls of plant regeneration. Curr. Top. Dev. Biol. 108, 1–33 (2014).
Che, P., Lall, S. & Howell, S. H. Developmental steps in acquiring competence for shoot development in Arabidopsis tissue culture. Planta 226, 1183–1194 (2007).
Atta, R. et al. Pluripotency of Arabidopsis xylem pericycle underlies shoot regeneration from root and hypocotyl explants grown in vitro. Plant J. 57, 626–644 (2009).
Sugimoto, K., Jiao, Y. & Meyerowitz, E. M. Arabidopsis regeneration from multiple tissues occurs via a root development pathway. Dev. Cell 18, 463–471 (2010).
Hu, B. et al. Divergent regeneration-competent cells adopt a common mechanism for callus initiation in angiosperms. Regeneration 4, 132–139 (2017).
Abe, T. & Futsuhara, Y. Genotypic variability for callus formation and plant regeneration in rice (Oryza sativa L.). Theor. Appl. Genet. 72, 3–10 (1986).
Bevitori, R., Popielarska-Konieczna, M., Dos Santos, E. M., Grossi-De-sá, M. F. & Petrofeza, S. Morpho-anatomical characterization of mature embryo-derived callus of rice (Oryza sativa L.) suitable for transformation. Protoplasma 251, 545–554 (2013).
Fan, M., Xu, C., Xu, K. & Hu, Y. LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res. 22, 1169–1180 (2012).
He, C., Chen, X., Huang, H. & Xu, L. Reprogramming of H3K27me3 is critical for acquisition of pluripotency from cultured Arabidopsis tissues. PLoS Genet. 8, e1002911 (2012).
Liu, J. et al. The WOX11-LBD16 pathway promotes pluripotency acquisition in callus cells during de novo shoot regeneration in tissue culture. Plant Cell Physiol. 59, 734–743 (2018).
Zhai, N. & Xu, L. Pluripotency acquisition in the middle cell layer of callus is required for organ regeneration. Nat. Plants 7, 1453–1460 (2021).
Kareem, A. et al. PLETHORA genes control regeneration by a two-step mechanism. Curr. Biol. 25, 1017–1030 (2015).
Sugimoto, K., Gordon, S. P. & Meyerowitz, E. M. Regeneration in plants and animals: dedifferentiation, transdifferentiation, or just differentiation? Trends Cell Biol. 21, 212–218 (2011).
Guo, F. et al. Callus initiation from root explants employs different strategies in rice and Arabidopsis. Plant Cell Physiol. 59, 1782–1789 (2018).
Liu, J. et al. WOX11 and 12 are involved in the first-step cell fate transition during de novo root organogenesis in Arabidopsis. Plant Cell 26, 1081–1093 (2014).
Kim, J.-Y. et al. Epigenetic reprogramming by histone acetyltransferase HAG1/AtGCN5 is required for pluripotency acquisition in Arabidopsis. EMBO J. 37, e98726 (2018).
Zhai, N., Pan, X., Zeng, M. & Xu, L. Developmental trajectory of pluripotent stem cell establishment in Arabidopsis callus guided by a quiescent center-related gene network. Development 150, dev200879 (2023).
Shim, S. et al. Transcriptome comparison between pluripotent and non-pluripotent calli derived from mature rice seeds. Sci. Rep. 10, 21257 (2020).
Zhao, N. et al. Systematic analysis of differential H3K27me3 and H3K4me3 deposition in callus and seedling reveals the epigenetic regulatory mechanisms involved in callus formation in rice. Front. Genet. 11, 1–16 (2020).
Indoliya, Y. et al. Decoding regulatory landscape of somatic embryogenesis reveals differential regulatory networks between japonica and indica rice subspecies. Sci. Rep. 6, 23050 (2016).
Zhang, H. et al. OsHDA710-mediated histone deacetylation regulates callus formation of rice mature embryo. Plant Cell Physiol. 61, 1646–1660 (2020).
Guo, F. et al. Functional analysis of auxin receptor OsTIR1/OsAFB family members in rice grain yield, tillering, plant height, root system, germination, and auxinic herbicide resistance. N. Phytol. 229, 2676–2692 (2021).
Xia, K. et al. OsTIR1 and OsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS ONE 7, e30039 (2012).
Zhu, Z.-X. et al. A gain-of-function mutation in OsIAA11 affects lateral root development in rice. Mol. Plant 5, 154–161 (2012).
Inukai, Y. et al. Crown rootless1, which is essential for crown root formation in rice, is a target of an AUXIN RESPONSE FACTOR in auxin signaling. Plant Cell 17, 1387–1396 (2005).
Zeng, M. et al. Stem cell lineage in body layer specialization and vascular patterning of rice root and leaf. Sci. Bull. 61, 847–858 (2016).
Sarkar, A. K. et al. Conserved factors regulate signalling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 446, 811–814 (2007).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 559 (2008).
Guo, F. et al. Rice LEAFY COTYLEDON1 hinders embryo greening during the seed development. Front Plant Sci. https://doi.org/10.1101/2021.08.18.456739 (2022).
Yang, D. et al. Transcriptome analysis of rice response to blast fungus identified core genes involved in immunity. Plant. Cell Environ. 44, 3103–3121 (2021).
Liu, J., Jie, W., Shi, X., Ding, Y. & Ding, C. Transcription elongation factors OsSPT4 and OsSPT5 are essential for rice growth and development and act with APO2. Res. Sq. Prepr. https://doi.org/10.21203/rs.3.rs-2549283/v1 (2023).
Gaj, M. D., Zhang, S., Harada, J. J. & Lemaux, P. G. Leafy cotyledon genes are essential for induction of somatic embryogenesis of Arabidopsis. Planta 222, 977–988 (2005).
Wójcikowska, B. & Gaj, M. D. Expression profiling of AUXIN RESPONSE FACTOR genes during somatic embryogenesis induction in Arabidopsis. Plant Cell Rep. 36, 843–858 (2017).
Shang, B. et al. Very-long-chain fatty acids restrict regeneration capacity by confining pericycle competence for callus formation in Arabidopsis. Proc. Natl Acad. Sci. USA 113, 5101–5106 (2016).
Wang, F.-X. et al. Chromatin accessibility dynamics and a hierarchical transcriptional regulatory network structure for plant somatic embryogenesis. Dev. Cell 54, 742–757.e8 (2020).
Salaün, C., Lepiniec, L. & Dubreucq, B. Genetic and molecular control of somatic embryogenesis. Plants 10, 1467 (2021).
Mordhorst, A., Hartog, M., El Tamer, M., Laux, T. & de Vries, S. Somatic embryogenesis from Arabidopsis shoot apical meristem mutants. Planta 214, 829–836 (2002).
Zuo, J., Niu, Q.-W., Frugis, G. & Chua, N.-H. The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J. 30, 349–359 (2002).
Su, Y. H. et al. Auxin-induced WUS expression is essential for embryonic stem cell renewal during somatic embryogenesis in Arabidopsis. Plant J. 59, 448–460 (2009).
Boutilier, K. et al. Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14, 1737–1749 (2002).
Horstman, A. et al. The BABY BOOM transcription factor activates the LEC1-ABI3-FUS3-LEC2 network to induce somatic embryogenesis. Plant Physiol. 175, 848–857 (2017).
Tsuwamoto, R., Yokoi, S. & Takahata, Y. Arabidopsis EMBRYOMAKER encoding an AP2 domain transcription factor plays a key role in developmental change from vegetative to embryonic phase. Plant Mol. Biol. 73, 481–492 (2010).
Lotan, T. et al. Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93, 1195–1205 (1998).
Kwong, R. W. et al. LEAFY COTYLEDON1-LIKE defines a class of regulators essential for embryo development. Plant Cell 15, 5–18 (2003).
Kagaya, Y. et al. LEAFY COTYLEDON1 controls seed storage protein genes through its regulation of FUSCA3 and ABSCISIC ACID INSENSITIVE3. Plant Cell Physiol. 46, 399–406 (2005).
Stone, S. L. et al. LEAFY COTYLEDON2 encodes a B3 domain transcription factor that induces embryo development. Proc. Natl Acad. Sci. USA 98, 11806–11811 (2001).
Stone, S. L. et al. Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: Implications for somatic embryogenesis. Proc. Natl Acad. Sci. USA 105, 3151–3156 (2008).
Braybrook, S. A. et al. Genes directly regulated by LEAFY COTYLEDON2 provide insight into the control of embryo maturation and somatic embryogenesis. Proc. Natl Acad. Sci. USA 103, 3468–3473 (2006).
Curaba, J. et al. AtGA3ox2, a key gene responsible for bioactive gibberellin biosynthesis, is regulated during embryogenesis by LEAFY COTYLEDON2 and FUSCA3 in Arabidopsis. Plant Physiol. 136, 3660–3669 (2004).
Luerßen, H., Kirik, V., Herrmann, P. & Miséra, S. FUSCA3 encodes a protein with a conserved VP1/ABI3-like B3 domain which is of functional importance for the regulation of seed maturation in Arabidopsis thaliana. Plant J. 15, 755–764 (1998).
Gazzarrini, S., Tsuchiya, Y., Lumba, S., Okamoto, M. & McCourt, P. The transcription factor FUSCA3 controls developmental timing in Arabidopsis through the hormones gibberellin and abscisic acid. Dev. Cell 7, 373–385 (2004).
Harding, E. W., Tang, W., Nichols, K. W., Fernandez, D. E. & Perry, S. E. Expression and maintenance of embryogenic potential is enhanced through constitutive expression of AGAMOUS-Like 15. Plant Physiol. 133, 653–663 (2003).
Zheng, Q., Zheng, Y. & Perry, S. E. AGAMOUS-like15 promotes somatic embryogenesis in Arabidopsis and soybean in part by the control of ethylene biosynthesis and response. Plant Physiol. 161, 2113–2127 (2013).
Zheng, Y., Ren, N., Wang, H., Stromberg, A. J. & Perry, S. E. Global identification of targets of the Arabidopsis MADS domain protein AGAMOUS-Like15. Plant Cell 21, 2563–2577 (2009).
Wang, H., Caruso, L. V., Downie, A. B. & Perry, S. E. The embryo MADS domain protein AGAMOUS-like 15 directly regulates expression of a gene encoding an enzyme involved in gibberellin metabolism. Plant Cell 16, 1206–1219 (2004).
Ikeuchi, M. et al. Wounding triggers callus formation via dynamic hormonal and transcriptional changes. Plant Physiol. 175, 1158–1174 (2017).
Iwase, A. et al. The AP2/ERF transcription factor WIND1 controls cell dedifferentiation in Arabidopsis. Curr. Biol. 21, 508–514 (2011).
Bian, H. et al. Distinctive expression patterns and roles of the miRNA393/TIR1 homolog module in regulating flag leaf inclination and primary and crown root growth in rice (Oryza sativa). N. Phytol. 196, 149–161 (2012).
Guo, F. et al. The miR393a/target module regulates seed germination and seedling establishment under submergence in rice (Oryza sativa L.). Plant. Cell Environ. 39, 2288–2302 (2016).
Xie, K., Minkenberg, B. & Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl Acad. Sci. USA 112, 3570–3575 (2015).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Aichinger, E. et al. CHD3 proteins and polycomb group proteins antagonistically determine cell identity in Arabidopsis. PLoS Genet. 5, e1000605 (2009).
Liu, H. et al. ARL1, a LOB-domain protein required for adventitious root formation in rice. Plant J. 43, 47–56 (2005).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Jin, J. et al. PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 45, D1040–D1045 (2017).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Xiao, S.-J., Zhang, C., Zou, Q. & Ji, Z.-L. TiSGeD: a database for tissue-specific genes. Bioinformatics 26, 1273–1275 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank Chuanzao Mao (Zhejiang University, China) and Makoto Matsuoka (Nagoya University, Japan) for providing plant materials. We thank Weizhen Hu (Agricultural Experiment Station of Zhejiang University) for the management of the greenhouse. We appreciate the kind help from Weilan Wang, Junying Li, Nianhang Rong, and Li Xie in the Bio-ultrastructure analysis Laboratory of the Analysis Centre of Agrobiology and Environmental Sciences, Zhejiang University. We thank Hanmin Chen (Zhejiang University) for his kind help in the study. This work was supported by grants from the National Natural Science Foundation of China (31971932/32225007/32201834), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB27030103), Science Foundation of Zhejiang Province (Grant no. LGN21C130006), China Agriculture Research System (CARS-05-05A), Research Startup Funding from Hainan Institute of Zhejiang University (NO.0201-6602-A12202), and the Key Research Program of CAS (QYZDB-SSW-SMC010).
Author information
Authors and Affiliations
Contributions
F.G., L.X., and H.B. designed the research. F.G. and H.W. performed sectioning and in situ hybridization. F.G., G.L., G.C., W.L., H.Z., D.L., and C.Z., performed tissue culture, genetic analysis, and bioinformatic analysis. F.G., N.H., M.Z., Y.S., P.J.S., L.X., and H.B. analyzed and discussed the data. F.G., L.X., and H.B. wrote the manuscript with the help from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Duncan Coleman, Shuang Wu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: David Favero.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, F., Wang, H., Lian, G. et al. Initiation of scutellum-derived callus is regulated by an embryo-like developmental pathway in rice. Commun Biol 6, 457 (2023). https://doi.org/10.1038/s42003-023-04835-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04835-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.