Abstract
Mycobacterium tuberculosis Clp proteases are targeted by several antitubercular compounds, including cyclomarin A (CymA). CymA exerts its toxicity by binding to AAA + chaperone ClpC1. Here, we show that CymA can also bind a partial homologue of ClpC1, known as ClpC2, and we reveal the molecular basis of these interactions by determining the structure of the M. tuberculosis ClpC2:CymA complex. Furthermore, we show deletion of clpC2 in Mycobacterium smegmatis increases sensitivity to CymA. We find CymA exposure leads to a considerable upregulation of ClpC2 via a mechanism in which binding of CymA to ClpC2 prevents binding of ClpC2 to its own promoter, resulting in upregulation of its own transcription in response to CymA. Our study reveals that ClpC2 not only senses CymA, but that through this interaction it can act as a molecular sponge to counteract the toxic effects of CymA and possibly other toxins targeting essential protease component ClpC1 in mycobacteria.
Similar content being viewed by others
Introduction
Mycobacterium tuberculosis (Mtb), the aetiological agent of tuberculosis (TB), persists to this day as a major burden on global healthcare with multidrug-resistant and extensively drug-resistant strains compounding the difficulties in treating TB1,2,3. Consequently, it is necessary to resort to antibiotics that are more expensive and have lower efficacy with more side effects4,5,6. Therefore, novel antitubercular drugs together with a molecular understanding of their mechanism and cellular targets are needed to control the spread of TB.
Modular chaperone-protease complexes share a similar overall architecture consisting of a cylindrical proteolytic core that associates with a chaperone ring, thereby coordinating unfolding and proteolysis of specific protein substrates7,8,9,10. The caseinolytic protease (Clp) complexes are almost ubiquitous in bacteria, absent in only a few with minimal genomes11,12. In mycobacteria, the Clp chaperone-protease complexes feature a proteolytic core, ClpP, composed of both ClpP1 and ClpP2 subunits, that associates with an unfoldase, either ClpC1 or ClpX, to form the active ClpC1P or ClpXP proteases, respectively13,14,15. ClpC1 contains a mostly α-helical N-terminal domain (NTD) comprising the first 145 residues of ClpC1, followed by two AAA + (ATPases associated with various cellular activities) modules required for the energy-dependent substrate unfolding and translocation activity16. The NTD of ClpC1 and homologues associates with adaptor/regulatory proteins contributing to ClpC1P activity regulation and is critical for delivery of certain substrate classes17,18,19,20,21,22,23,24. In Mtb, ClpC1, ClpX and ClpP core subunits are all essential for cell viability, making the Clp system a promising drug target25,26,27.
Several compounds with antitubercular activity are known to target the mycobacterial Clp system28 and many of these compounds specifically target ClpC1 NTD, including lassomycin, ecumicin and cyclomarin A (CymA)29,30,31,32. CymA is a cyclic heptapeptide that is synthesised by the marine bacterium Streptomyces sp. CNB-98233,34,35. The compound is effective against mycobacteria but was found to show little activity against Staphylococcus aureus and Escherichia coli34. Biochemical and structural studies revealed CymA binds to the NTD of ClpC1 and it is this interaction that is responsible for CymA bactericidal activity32. In further studies, CymA was observed to enhance the ATPase and proteolytic activity as well as reduce substrate specificity of ClpC1 with subsequent deregulation of ClpC1P proteolytic activity leading to cell death21,36.
Mycobacteria also encode a ClpC1 partial homologue annotated as ClpC2. Given the sequence homology between ClpC1 NTD and ClpC2, we hypothesised CymA can also bind ClpC2. Through biochemical and structural studies, we show that ClpC2 presents a competitive binding site for CymA and is capable of sequestering CymA away from ClpC1. Furthermore, our in vivo investigations in Mycobacterium smegmatis show that the interaction between ClpC2 and CymA protects the organism against CymA-induced toxicity. In addition, we reveal that ClpC2 strongly upregulates its own transcription in response to CymA, shedding light on the regulatory mechanism by which mycobacteria counteract the toxic effects of ClpC1-targeting antibiotic compounds. Combined our results uncover and characterise a mycobacterial natural antibiotic resistance mechanism that senses and neutralises compounds targeting an essential protein degradation pathway.
Results
ClpC2 binds to the antitubercular compound CymA
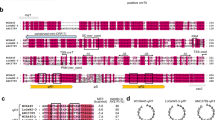
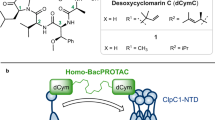
The bactericidal peptide CymA (Fig. 1a) binds to the NTD of ClpC121,32,34,35. In addition to ClpC1, many actinobacteria encode a ClpC1 partial homologue annotated as ClpC2 that shares sequence similarity with ClpC1 NTD (Fig. 1b). Given the resemblance between ClpC1 NTD and the ClpC2 C-terminal 160 residues (Supplementary Fig. 1), we hypothesised CymA can also interact with ClpC2. An alignment of the amino acid sequence of the Mtb ClpC1 NTD and the ClpC2 C-terminal region from several actinobacteria shows ClpC1 residues (F2, F80 and E89) critical for interaction with CymA are also conserved in ClpC2 (Supplementary Fig. 1)32.
a Structural formula of cyclomarin A (CymA). b Domain organisation of Mtb ClpC1 alongside its partial homologue ClpC2. ClpC1 NTD, N-terminal domain (cyan); AAA, ATPases Associated with diverse cellular Activities (shades of grey); MD, middle domain (black). The ClpC1 partial homologue, ClpC2, is coloured according to sequence homology to ClpC1 NTD: homologous region (cyan); non-homologous region (light blue). c, d Injection profiles and binding isotherms for ITC titrations of ClpC11–145 against CymA (c) and ClpC2 against CymA (d). The dissociation constant is indicated within each panel. Measurements were performed at 37 °C titrating aliquots of 50 μM ClpC11–145 (c) or ClpC2 (d) to 5 μM CymA. N and KD values are the mean ± standard deviation from three independent experiments.
To investigate the molecular interaction between ClpC2 and CymA, we heterologously expressed and purified Mtb ClpC2. The binding parameters between ClpC1 or ClpC1 NTD and CymA1, an amino-alcohol derivative of CymA, were previously analysed by isothermal titration calorimetry (ITC), producing dissociation constants (KD) values of 17.5 nM and 12 nM, respectively32. We used ITC to assess the affinity between CymA and ClpC2 and compared it to the affinity of the compound for ClpC1 NTD. For ClpC1 NTD and CymA, we observed a KD in the low nanomolar range (2.3 nM) in ITC measurements (Fig. 1c), which is comparable to the value observed for the amino-alcohol derivative (CymA1). We determined a similar affinity for the binding of CymA to ClpC2 with a KD of 2.2 nM and a stoichiometry of 1:1 (Fig. 1d). These data clearly indicate ClpC2, like ClpC1, binds tightly to CymA.
ClpC2 dimerises via its N-terminal domain while the C-terminal domain binds CymA
To obtain a molecular understanding of ClpC2 and its interaction with CymA, we set up crystallisation experiments with ClpC2 alone and in the presence of CymA. We determined the structure de novo using the programme ARCIMBOLDO37 for ab initio phasing, since molecular replacement failed, and refined it to ~2.0 Å resolution. Due to proteolysis in the crystallisation drop, the determined structure included ClpC2 residues 4–72 (ClpC24–72), the region of ClpC2 without homology to ClpC1, but not the C-terminal portion of ClpC2 (Table 1, Fig. 2a). ClpC24–72 forms a dimer in the asymmetric unit, with each monomer featuring a four-helix bundle (Fig. 2b). Analysis of the structure using PDBePISA38 strongly suggests dimerisation between the two N-terminal fragments with a total buried surface area of 1068 Å2. The intermolecular interactions within the dimerisation interface are primarily mediated by highly conserved hydrophobic residues and two salt bridges established by the D46-R57 pair (Fig. 2b). To confirm that the interface observed in the crystal structure reflects a homodimer interaction also occurring in solution, a ClpC2 R57A variant was generated and both ClpC2 WT and ClpC2 R57A were analysed by size-exclusion chromatography. Compared to the elution positions of size standard proteins, ClpC2 WT (monomeric molecular weight 26 kDa) appears to exist in an equilibrium between monomer and dimer at this concentration whereas the ClpC2 R57A variant elution profile corresponds to the monomeric species (Supplementary Fig. 2).
a Crystal structure of the N-terminal domain of ClpC2 (ClpC24–72). b ClpC24–72 forms a homodimer in the crystallographic asymmetric unit (left panel) with each monomer coloured in light blue or light teal. In the right panel is a close-up view of the dimeric interface between ClpC2 monomers coloured according to the conservation of residues. Dashed lines suggest electrostatic interactions. c Overall structure of ClpC2 C-terminal domain (ClpC294–252; cyan) bound to CymA (orange). d The unbiased mFo-DFc Fourier map of CymA at 3.0 σ was calculated following molecular replacement using PDB 3WDC as a search model excluding ligands. e ClpC294–252 (cyan) bound to CymA (orange) is superimposed onto ClpC1 NTD bound to CymA (PDB 3WDC; grey) showing a root mean square deviation (RMSD) of 2.0 Å. f CymA (orange, stick representation) bound to ClpC294–252 (surface representation), coloured according to hydrophobicity. g Detailed molecular interactions between CymA and ClpC2 C-terminal domain. Shown in stick representation are the residues of ClpC2 (cyan) that are in close proximity to CymA (orange) with suggested hydrogen bond interactions represented by dashed lines and water molecules represented as red spheres. h A cartoon model of ClpC2 dimer colour-coded according to the presented ClpC2 crystal structures.
To also obtain structural information on the ClpC2 C-terminal, CymA-binding domain, we then constructed a ClpC2 truncation variant covering ClpC2 residues 94–252 that are homologous to ClpC1 NTD based on the sequence alignment (see Supplementary Fig. 1). This ClpC2 truncation variant (ClpC294–252) was successfully co-crystallised with CymA and the structure of the complex was determined by molecular replacement using the ClpC1 NTD structure (PDB 3WDC) as a search model. The final atomic model of ClpC294–252 bound to CymA was refined to a resolution of 1.43 Å (Table 1, Fig. 2c, d). Like the ClpC1 NTD, ClpC294–252 forms two helical repeats and it binds CymA in an analogous fashion (Fig. 2e)32. The binding of CymA to ClpC294–252 mainly stems from hydrophobic interactions that are largely mediated by the two highly conserved phenylalanine residues (F96 and F172; Fig. 2f, g). The two residues also contribute polar contacts through their backbone amides to the carbonyls of the N-reverse prenylated tryptophan and alanine of CymA with additional polar contacts provided by E181 and Q94 (Fig. 2g). Taken together, the structures we have solved in this study suggest ClpC2 dimerises via its N-terminal domain with the C-terminal domain harbouring the CymA-binding site (Fig. 2h).
ClpC2 protects against CymA-induced toxicity
Like Mtb, Mycobacterium smegmatis (Msm), which is used as a model system for studying Mtb biology, is also sensitive to CymA and it encodes a ClpC2 homologue. Therefore, we turned to Msm to analyse the contribution of ClpC2 to cell viability in the absence and presence of CymA. We generated a clpC2 knockout strain (ΔclpC2) in Msm which displayed growth behaviour similar to the parent strain (WT) under standard growth conditions (Fig. 3a). However, upon addition of sub-MIC50 concentrations of CymA32, ΔclpC2 exhibited diminished growth compared to the parent strain, and the WT phenotype was restored in a clpC2 complementary strain (comclpC2 WT) (Fig. 3a). To assess whether these effects originate from CymA-binding to ClpC2, we complemented the ΔclpC2 strain with a CymA-binding-impaired variant. Previously, F2 was identified as a key residue for CymA-binding to Mtb ClpC132, and our structural studies show the equivalent residue in Mtb ClpC2 (F96) contributing to CymA-binding (Fig. 2g). Therefore, we complemented ΔclpC2 with clpC2 F99A (the Msm equivalent to F96), and we observed attenuated growth upon exposure to CymA (Fig. 3a). The observed phenotype suggests ClpC2 protects mycobacteria from CymA-induced toxicity by directly binding the toxic compound.
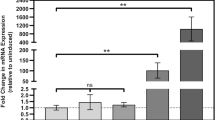
a A clpC2 knockout (ΔclpC2) strain was prepared in Msm. The ΔclpC2 (red diamonds), parent (WT, black squares) as well as ΔclpC2 strains complemented with clpC2 WT (comclpC2 WT, grey circles) or complemented with clpC2 F99A (comclpC2 F99A, blue triangles) were grown to exponential phase, then diluted to an OD600 of 0.005 before monitoring the OD600 of the cultures grown in either the absence or presence of 150 nM CymA. Error bars depict the standard deviation from three biological replicates. b The effect of CymA on cellular ClpC2 protein levels was analysed by Western blot. Msm culture was grown to an OD600 of 0.5 in 7H9 medium at 37 °C before adding 150 nM CymA. Cells were harvested, then lysed at the indicated time points following addition of CymA. RpoB is included as a loading control. c The counteractive effect of ClpC2 in a CymA-stimulated FITC-casein degradation assay. The proteolytic activity of 0.5 µM ClpC1 hexamer/0.8 µM ClpP1P2 tetradecamer was stimulated by 2 µM CymA and the rate of degradation was determined following the addition of increasing concentrations of ClpC2. The rate of degradation is normalised so that 0% stimulation is the proteolytic activity in the absence of CymA and 100% stimulation is equivalent to the maximum CymA-induced stimulation above basal activity. Error bars represent standard deviation from three independent experiments. d Quantitative Western blot was employed to analyse the ratio of ClpC1 (open circles) to ClpC2 (filled squares) following addition of CymA. Msm culture was grown to an OD600 of 0.5 in 7H9 medium at 37 °C before adding 150 nM CymA. Samples were harvested at the indicated time points following addition of CymA and ClpC1 or ClpC2 proteins quantified by immunoblotting in reference to a protein standard curve using purified ClpC1 and ClpC2 protein. ClpC1 and ClpC2 protein levels are normalised relative to ClpC1 at timepoint 0 h. Representative blots are shown in Supplementary Fig. 3. Error bars depict standard deviation from biological triplicates. e Expression levels of clpC2 in the absence and presence of 150 nM CymA as analysed by RT-qPCR. Msm cultures were grown to an OD600 of 0.5 followed by a one-hour incubation in either the absence or presence of 150 nM CymA. clpC2 gene expression was normalised to the housekeeping gene rpoB. Error bars represent standard deviation of technical triplicates from three biological replicates. Statistical analysis by two-tailed unpaired Student’s t test; P value = 0.0015.
ClpC2 is upregulated in response to CymA
To understand further the ClpC2 response to CymA, we explored the effect of CymA on intracellular ClpC2 protein levels in WT Msm. We added sub-MIC50 concentrations of CymA to mid-log phase Msm cultures and followed the protein levels by Western blot. While the RpoB loading control remains constant throughout, no band for ClpC2 is detected in the Western blot in the absence of CymA, and the signal increases by several orders of magnitude from a faint band at 1 hour post CymA addition to 8 h following addition of CymA (Fig. 3b).
CymA was shown to have a stimulatory effect on ClpC1-dependent degradation of the model substrate casein21,36. Given the strong upregulation of ClpC2 in vivo, we decided to investigate the impact of ClpC2 on a CymA-stimulated degradation assay using fluorescein isothiocyanate-conjugated casein (FITC-casein). Adding increasing concentrations of ClpC2 counteracted the stimulatory effect of CymA on casein degradation (Fig. 3c), suggesting ClpC2 can sequester CymA, preventing it from binding ClpC1 and thus from modulating ClpC1 activity. In order to protect ClpC1 by competing for CymA, ClpC2 would have to reach cellular concentrations in excess of ClpC1. To compare the in vivo protein levels of ClpC1 and ClpC2, we quantified ClpC1 and ClpC2 protein levels at various time points following addition of CymA by quantitative Western blot. ClpC1 levels remained fairly constant throughout the 8-hour timeframe. In contrast, ClpC2 levels changed considerably starting at undetectable levels immediately after addition of CymA and rising to 84-fold in excess of ClpC1 around 8 h after addition of CymA (Fig. 3d, Supplementary Fig. 3). This suggests Msm can produce substantial quantities of ClpC2 well in excess of ClpC1 to sequester the toxic CymA.
Considering the large upregulation in ClpC2 protein, we next analysed the effect of CymA on ClpC2 transcription. Therefore, we grew Msm cultures in either the absence or presence of CymA and extracted total RNA for analysis of clpC2 transcript levels by RT-qPCR. Comparison of relative transcript levels reveals a 263-fold increase in clpC2 mRNA following exposure to sub-MIC50 concentrations of CymA for one-hour (Fig. 3e). Given the dramatic increase in clpC2 expression, we directed our attentions to the influence of CymA on clpC2 transcription.
ClpC2 acts as a transcription factor to repress its own expression
We employed a DNA pull-down assay based on the protocol reported in Chaparian and van Kessel39 to identify the transcription factor responsible for the upregulation of ClpC2 in response to CymA (Supplementary Fig. 4a). The DNA probe used in the pull-down assay encompassed 300 bp of the DNA sequence upstream of the Msm clpC2 start codon to ensure it included the putative promoter region. As a control, a stretch of intragenic Msm DNA sequence of similar length was used. Msm lysate was prepared from cultures grown in either the presence or absence of CymA and served as the source for the unknown transcription factor. Proteins retained with the DNA through affinity purification were subjected to tryptic digestion and analysed by liquid chromatography followed by tandem mass spectrometry (LC-MS/MS). Approximately 800 proteins were detected (Supplementary Data 1) many of which are suspected false positives. One of the proteins detected in the DNA pull-down assay was ClpC2 itself. Surprisingly, ClpC2 also was detected in the pull-down carried out in the absence of CymA (Supplementary Fig. 4b), where it was even slightly enriched. This was very unexpected considering the comparatively low ClpC2 protein levels we measured in absence of CymA (Fig. 3b, d), and suggested a potential role for ClpC2 in regulating its own expression.
To investigate how ClpC2 might autoregulate its expression, both WT and ΔclpC2 strains were transformed with an integrative plasmid that encodes the luciferase (luc) gene under the control of the clpC2 promoter (see Supplementary Fig. 5). The effect of CymA on luc transcript levels was then followed by RT-qPCR for both strains. In agreement with Fig. 3e, luciferase expression in the modified WT culture was significantly higher in the presence of CymA compared to in the absence of CymA (Fig. 4a). This pattern of luciferase expression was not observed for the modified ΔclpC2 cultures (Fig. 4a). Here, luciferase expression levels in the absence of CymA are already at the high level of expression that the modified WT cultures only reached in the presence of CymA. These results suggest that ClpC2 is required for repression of its own expression under standard growth conditions and that this transcriptional repression is lifted following the addition of CymA. The most direct role ClpC2 could play to achieve this effect is by acting as a transcriptional repressor of its own promoter.
a The luciferase gene under the control of the clpC2 promoter was introduced into the genomes of both WT and ΔclpC2 strains via an integrative vector (see Supplementary Fig 5). Cells were harvested following the addition of CymA and the total RNA was extracted in preparation for luciferase transcript levels analysis by RT-qPCR. Luciferase transcript levels are normalised to the housekeeping gene rpoB and expression levels are expressed relative to the wild-type strain grown in the absence of CymA. Error bars depict standard deviation from three biological replicates with statistical significance determined by two-tailed unpaired Student’s t test. b Alignment of DNA sequences immediately upstream of the clpC2 open reading frame in a selection of actinobacteria. The start codon for clpC2 and homologues is shaded in light green. Pseudopalindromic sequence regions upstream of the start codon are shaded in light blue. The sequence logo is displayed above the alignment. Mtb, Mycobacterium tuberculosis; Msm, Mycobacterium smegmatis; Mma, Modestobacter marinus; Ser, Saccharopolyspora erythraea; Rer, Rhodococcus erythropolis; Gsi, Gordonia sihwensis; Nas, Nocardia asteroids. c ClpC2 DNA-binding activity was analysed by electrophoretic mobility shift assay (EMSA). 1–25 nM ClpC2 was added to 0.5 nM DNA that includes the putative clpC2 operator sequence. Data are representative of three independent experiments. d The binding of DNA to ClpC2 was analysed in the absence (filled squares) and presence (open circles) of 1 µM CymA. Gel band intensity was quantified by densitometry to determine the fraction of DNA bound to ClpC2. Representative gels are shown in Supplementary Fig. 8. Error bars represent the standard deviation of three independent experiments.
We analysed the DNA sequence upstream of the clpC2 open reading frame (ORF) in several actinobacterial organisms to determine the clpC2 operator DNA sequence and identified a conserved pseudopalindromic sequence one to two nucleotides upstream of the clpC2 ORF, which takes the form of CGTCAAC-n5-GTTGACG; where n is any nucleotide (Fig. 4b). The same palindromic sequence is repeated further upstream of the clpC2 ORF in many of the actinobacterial organisms analysed, and together these operator sites overlap with the putative −35 and −10 elements of the clpC2 promoter (Supplementary Fig. 6a). Additionally, the positioning of the −35 and −10 elements with respect to the clpC2 ORF suggests clpC2 transcripts are leaderless.
CymA lifts transcriptional repression
The putative clpC2 operator DNA sequence was incorporated into a DNA probe used in an electrophoretic mobility shift assay (EMSA). Titrating ClpC2 to the DNA probe produces a gel band shift confirming ClpC2 can bind the DNA (Fig. 4c). In the actinobacterial genomes containing two consecutive clpC2 operator sites (e.g., Msm), ClpC2 can bind both sites simultaneously presumably leading to more stringent regulation of clpC2 expression (Supplementary Fig. 6b). Furthermore, specificity for the clpC2 operator sequence was demonstrated with the addition of either nonspecific or specific competitor DNA to labelled clpC2 operator DNA (Supplementary Fig. 7a and 7b). The palindromic nature of the operator sites suggests the high-affinity interaction with DNA observed in Fig. 4c involves the ClpC2 dimer. Analysis of DNA binding by the dimerisation-deficient ClpC2 R57A and ClpC2 CTD variants in EMSA experiments provided further support for the importance of ClpC2 dimer formation for the high-affinity interaction with the clpC2 operator site (Supplementary Fig. 7c and 7d). To explore the effects of CymA on DNA-binding, we performed the EMSA across a broader ClpC2 concentration range in the absence and the presence of CymA (Fig. 4d, Supplementary Fig. 8). CymA had a noticeable inhibitory effect on the interaction between ClpC2 and its operator site, providing the explanation for the lifting of transcriptional repression observed in Figs. 3e and 4a.
In order to rationalise such a mechanism of CymA-dependent derepression on the molecular level, we turned again to the structure of ClpC294–252-CymA to identify the putative site for DNA binding. We observed a highly conserved surface region in the ClpC2 C-terminal domain (Fig. 5a) that coincides with a positively charged region (Fig. 5b). Contributing to the high degree of conservation and charge is a prominent helix (residues 174-189) that features two conserved arginines (R185 and R189; Fig. 5c). Therefore, to explore the potential for this region of ClpC2 to bind DNA, we tested a ClpC2 R185A/R189A variant in an EMSA experiment and observed the double mutant is unable to bind DNA as efficiently as ClpC2 WT (Fig. 5d). As R185 and R189 are conserved in Msm ClpC2 (R188 and R192), we analysed the clpC2 mRNA levels of ΔclpC2 complemented with either clpC2 R188A/R192A (comclpC2 R188A/R192A) or clpC2 R57A (comclpC2 R57A), the mutant that exhibited impaired dimerisation behaviour (Supplementary Fig. 2). Both, complementation with clpC2 R188A/R192A and with clpC2 R57A showed increased expression levels of 2.0-fold and 4.6-fold, respectively, compared to complementation with ClpC2 WT (Supplementary Fig. 9a). The knockout strain complemented with clpC2 R188A/R192A showed similar growth behaviour as the knockout strain complemented with clpC2 WT, indicating that the defect in repression of clpC2 does not interfere with the protective effect of ClpC2 against CymA toxicity and also does not lead to a growth defect in rich medium under standard culture conditions (Supplementary Fig. 9b). Interestingly, although the arginine double-mutant can clearly still bind CymA, as evidenced by the lack of growth impairment observed with this mutant in the presence of CymA, the putative DNA-binding site is quite close to and might overlap partially with the CymA-binding site (Fig. 5a, b). Therefore, we hypothesise CymA either prevents DNA binding directly by sterically occluding the DNA-binding site, or CymA induces an alternative configuration of ClpC2 C-terminal domains that is incapable of interacting efficiently with DNA.
a Surface representation of ClpC294–252-CymA coloured according to residue conservation produced using ConSurf60,61,62. CymA is coloured in orange and shown in stick representation. b Structure of ClpC294–252 depicting surface charge and shown in the same orientation as in a. c Close-up of the suggested DNA recognition helix shown in the forefront. Arginines suggested to contribute to DNA-binding are shown in stick representation. d Comparison of ClpC2 WT and ClpC2 R185A/R189A double mutant in an electrophoretic mobility shift assay. ClpC2 variants were titrated against 0.5 nM DNA probe and loaded onto a 12% polyacrylamide gel. Data are representative of three independent experiments.
Expression of clpC2 is also induced under heat stress
In order to investigate if other stressors might alleviate clpC2 repression, we tested clpC2 mRNA levels under heat shock conditions. Inducing heat shock by increasing the temperature to 45 °C caused an increase in clpC2 expression of ~twofold (Supplementary Fig. 10a). This suggests that stresses affecting ClpC2 stability might also lead to derepression and hence upregulation of ClpC2. ClpC1 is known to degrade unstructured model substrates like casein that can serve as a mimic for heat-destabilised proteins. Given the homology between the ClpC2 CTD and ClpC1 NTD, it is possible that ClpC2 might also bind disordered proteins. We, therefore, tested the effect of an excess of ClpC2 on ClpC1-dependent casein degradation. In the presence of ClpC2, FITC-casein degradation is slowed with a proportion of the substrate population protected from degradation, implying that at high micromolar concentrations, ClpC2 can bind FITC-casein and prevent degradation of the substrate by the ClpC1P complex (Supplementary Fig. 10b). These results demonstrate that ClpC2 shares some protein binding partners with ClpC1.
Discussion
Mtb Clp proteases are considered to be promising drug targets with both inhibition and activation being deleterious to the organism. As such, these proteases have already been implicated in the mechanism of several natural products exhibiting mycobactericidal activity28,29,40. However, relatively little is known about the defensive response of the bacteria to the relevant antibacterial compounds. Here we report a role for the ClpC1 partial homologue ClpC2 in mediating against CymA-induced toxicity in mycobacteria. Furthermore, we can ascribe a mechanism by which ClpC2 binds DNA and represses its own expression under standard growth conditions. When presented with CymA, ClpC2 binds to CymA, hindering the interaction of ClpC2 with its own operator site and leading to loss of repression and subsequent upregulation of ClpC2. This upregulation of ClpC2 increases the capacity to bind free CymA, reducing the CymA available to interact with ClpC1 and therefore alleviating CymA-induced toxicity (Fig. 6).
ClpC2 binds to its own operator site (clpC2o) performing the role of a transcriptional repressor under standard growth conditions. Upon exposure to CymA, both ClpC1 and ClpC2 bind CymA leading to both aberrant ClpC1P activity as well as the lifting of clpC2 gene repression resulting in the upregulation of ClpC2. The increased ClpC2 protein levels bind the available CymA, preventing the toxic consequences of CymA interaction with ClpC1.
Our ITC titrations and ClpC1P in vitro degradation assay suggest ClpC2 can bind CymA as well as compete with ClpC1 for the binding of CymA (Figs. 1d and 3c). However, CymA is only one of several antitubercular peptides that are known to bind to ClpC1 NTD. Co-crystal structures of ClpC1 NTD with ecumicin and rufomycin revealed overlapping binding sites with CymA, despite opposing effects on ClpC1P proteolytic activity30,31,41,42. Based on our findings, it appears plausible that other cyclic peptides may also bind to ClpC2. These compounds are generally of natural origin and are likely encountered by ClpC2-expressing actinobacteria in their natural habitats. Therefore, one possible explanation that fits with the data we present is that ClpC2 has the role to protect ClpC1 from these compounds by providing a competitor pool of binding sites for the toxins, titrating them away from interaction with ClpC1. Alternative sources for toxic ClpC1 interactors might be bacteriophages, as phage-encoded proteins were previously reported to deregulate protein degradation by binding to the NTD of the ClpC1 homologue in Bacillus subtilis43. Furthermore, ClpC2 may help to regulate the delivery of endogenous proteins to the Clp complexes by competing for specific substrates or substrate adaptors that usually dock at the ClpC1 NTD. For example, by inhibiting the degradation of specific substrates, ClpC2 might prevent adverse downstream cellular effects upon exposure to certain stressors. Our results with the model substrate casein support this notion and suggest a role for ClpC2 under heat stress, where it might act as a “holdase” chaperone to alleviate ClpC1 overload.
The upregulation of ClpC2 was previously observed in several transcriptional profiling studies into regulation during nutrient starvation, lysosomal stress and in persister/latent cells in Mtb44,45,46,47,48,49. In our study, we observed significant upregulation of ClpC2 in response to CymA and to a milder degree also under heat shock. With the finding that ClpC2 acts as a transcriptional repressor and accounts for the significant upregulation of ClpC2, we have identified the transcriptional mechanism responsible for the regulation of clpC2 gene expression in this and other studies.
The analysis of the Mtb ClpC2 structures that we present here suggests DNA interaction occurs via ClpC2 CTD and that the putative DNA-binding site is directly adjacent to or even partially overlapping the CymA-binding site, interfering with the binding of ClpC2 to the clpC2 promoter and ultimately leading to derepression of the clpC2 gene. The structural data suggest that CymA either prevents access of DNA to the DNA-binding site sterically or by triggering a conformational change in the ClpC2 C-terminal domains. Which signal might trigger the derepression of clpC2 in other stress and adaptive responses requires further investigation. It is possible that secondary metabolites could act in place of CymA-binding, eliciting similar conformational changes in the C-terminal ClpC2 domains.
A similar distribution of positive charges as in the putative DNA-binding region of ClpC2 is observed in the corresponding region of ClpC1, prompting us to test ClpC1 in an EMSA experiment using the DNA probe containing the clpC2 operator site. ClpC1 exhibited an affinity with a KD in the low micromolar range, which constitutes at least two orders of magnitude lower affinity than exhibited by ClpC2 (Supplementary Fig. 11a). Furthermore, ClpC1 shows little sequence specificity for the clpC2 operator site (Supplementary Fig. 11b, 11c). This is in accordance with previous reports of unspecific interaction with DNA for the ClpC1 functional homologue, E. coli ClpA, which was suggested to play a role in the degradation of certain replication initiator proteins50,51.
Taken together, we show ClpC2 presents an alternative binding site for CymA and that this, combined with the pronounced upregulation of ClpC2, helps to protect the cell from CymA-induced toxicity. Our work contributes to the growing understanding of the Clp complexes as drug targets and offers insight into the complexities of CymA-induced toxicity.
Materials and methods
Bacterial strains and growth conditions
Mycobacterium smegmatis MC2155 SMR5 and derived strains were routinely grown at 37 °C with shaking in DIFCO™ Middlebrook7H9 broth (BD) supplemented with 0.05% Tween 80 and 0.2% glycerol. The antibiotics apramycin (50 µg/ml) or hygromycin B (100 µg/ml) were added as needed. CymA (dissolved in DMSO) was added to the media as indicated and the DMSO concentration was matched in the relevant control experiments. The DMSO concentration of cultures was maintained at less than 1% in all instances.
Protein expression and protein purification
All proteins were expressed in E. coli Rosetta (DE3) by isopropyl-β-D-thiogalactopyranosid (IPTG) induction in LB media at 20 °C overnight. Cells were harvested then resuspended in lysis buffer (50 mM HEPES-KOH pH 7.5, 300 mM KCl, 5 mM MgCl2, DNaseI, 1 mM PMSF, 1× Roche protease inhibitor cocktail) and lysed using a Microfluidizer M110-L (Microfluidics). Lysate was clarified by centrifugation in a SS34 rotor (ThermoScientific) at 20,000 rpm for 1 h at 4 °C. Untagged Mtb ClpC2 WT, ClpC2 R57A and ClpC2 R185A/R189A variants were precipitated by adding (NH4)2SO4 to 35% saturation followed by resolubilisation and dialysis into 50 mM HEPES-KOH pH 7.5, 75 mM KCl, 2 mM EDTA, 10% glycerol then subjected to anion exchange chromatography (Source Q column) using a salt gradient of 0.1–1 M KCl. Fractions containing ClpC2 were selected following SDS-PAGE and adjusted to 50 mM HEPES-KOH pH 7.5, 25% (NH4)2SO4, 150 KCl, 10% glycerol and further purified by hydrophobic interaction chromatography using a Butyl Sepharose 4 Fast Flow column with a 25–0% (NH4)2SO4 gradient. ClpC2 fractions were pooled and loaded onto a Superdex 75 16/600 column (GE Healthcare) for further purification in buffer GF (50 mM HEPES-KOH pH 7.5, 150 mM KCl, 10% glycerol). Far-UV CD spectra of the ClpC2 variants were recorded and compared to the spectrum of ClpC2 WT to ensure the variants are folded (Supplementary Fig. 12).
Lysate of untagged Mtb ClpC1 was diluted and pH-adjusted to 50 mM HEPES-KOH pH 7.1, 75 mM KCl, 2 mM EDTA, 10% glycerol and loaded onto a Fast Flow Q anion exchange column and purified using a salt gradient of 0.1–0.5 M KCl. Fractions containing ClpC1 were pooled and ClpC1 precipitated by adding (NH4)2SO4 to 40% saturation and then resolubilised and dialysed into 50 mM HEPES-KOH pH 7.1, 75 mM KCl, 2 mM EDTA, 10% glycerol. Soluble ClpC1 was again subjected to anion exchange chromatography (Source Q column) using a salt gradient of 0.1–0.5 M KCl followed by (NH4)2SO4 precipitation (40% saturation). ClpC1 protein was resuspended and dialysed into 50 mM HEPES-KOH pH 7.1, 150 mM KCl, 10% glycerol and loaded onto a HiLoad Superdex 200 26/600 column (GE Healthcare).
Recombinantly expressed ClpP1 and ClpP2 with C-terminal tetra-histidine tags (His4) were each purified by immobilised metal affinity chromatography (IMAC) using Ni2+- IMAC Sepharose 6 FF resin (GE Healthcare) followed by further purification on a Superose 6 column (GE Healthcare) in 50 mM HEPES-KOH pH 7.5, 300 mM NaCl, 10% glycerol. ClpP1P2 N-terminal propeptide processing was achieved by overnight incubation of equimolar concentrations of ClpP1-His4 and ClpP2-His4 at room temperature in 25 mM HEPES-KOH pH 7.5, 100 mM KCl, 5 mM MgCl2, 0.1 mM EDTA, 10% glycerol in the presence of 1 mM benzyloxycarbonyl-l-leucyl-l-leucinal (PeptaNova). Processed ClpP1P2 was further purified by gel filtration using a Superdex 200 column.
Both ClpC1 residues 1-145 (ClpC1 NTD) and ClpC2 residues 94-252 (ClpC294–252) were expressed with an N-terminal deca-histidine tag (His10) followed by both the tobacco etch virus (TEV) and human rhinovirus 3c protease (3c) cleavage sites. His10-tev-3c-ClpC1 NTD and His10-tev-3c-ClpC294–252 were first purified by IMAC using Ni2+- IMAC Sepharose 6 FF resin and both proteins cleaved overnight at 4 °C with TEV protease. ClpC1 NTD and ClpC294–252 were further purified by Ni2+-NTA chromatography and the flow-through was collected and loaded onto a Superdex 75 16/600 column in buffer GF. All proteins were flash-frozen in liquid nitrogen and stored at −20 °C.
Circular dichroism
Far-UV CD spectra of ClpC2 WT and ClpC2 variants R57A and R185A/R189A were recorded on a JASCO J-810 CD spectropolarimeter at a concentration of 8 µM. The spectra were measured at 23 °C in 50 mM Na2HPO4-NaH2PO4 (pH 7.4), 150 mM KCl in a 0.1 cm quartz cuvette. The recorded spectra were averages of 10 measurements and were corrected for buffer background.
Isothermal titration calorimetry
ClpC1 NTD and ClpC2 were dialysed into ITC buffer (50 mM HEPES-KOH pH 7.5, 150 mM KCl, 9% DMSO, 1 mM EDTA) and their concentrations were determined by measuring absorbance at 280 nm. CymA was diluted into ITC buffer and care was taken to maintain the same concentration of DMSO as for the dialysed proteins. 50 μM (monomer concentration) of either ClpC1 NTD or ClpC2 was titrated into 5 μM CymA (1.4 ml) at 37 °C beginning with a 2 μl injection followed by 39 injections of 5 μl injections at intervals of 180 seconds using a VP-ITC instrument (MicroCal). Data were analysed using Origin v.7.0383 (OriginLab) with a one-site binding model.
Degradation assay
Degradation of FITC-casein (Sigma) was carried out in 50 mM HEPES-KOH pH 7.1, 150 mM KCl, 20 mM MgCl2, 10% glycerol by 0.5 μM ClpC1 hexamer and 0.8 μM ClpP1P2 tetradecamer with the addition of 0.4 mM activator peptide benzyloxycarbonyl-L-leucyl-L-leucinal (PeptaNova), 0.5 mM DTT, 1 U/ml creatine phosphokinase, 20 mM phosphocreatine, 10 mM ATP at 37 °C. The CymA-stimulated degradation of FITC-casein (0.5 μM) was performed in the presence of 2 μM CymA and 0–18 μM ClpC2. Whereas the effect of an excess of ClpC2 on the degradation of FITC-casein (10 μM) was carried out in the presence of 100 μM ClpC2. Fluorescence intensity increases upon degradation of FITC-casein as a consequence of fluorophore dequenching was monitored using the BioTek Synergy 2 Plate Reader with a 485/20 nm band-pass emission filter and a 528/20 nm band-pass excitation filter. Rates of degradation were determined from the initial slopes and used to determine the CymA-induced stimulation of ClpC1P activity at the different concentrations of ClpC2.
Protein crystallisation
Both ClpC2 (full length) and ClpC294–252 were dialysed into 20 mM HEPES-KOH pH 7.5, 20 mM KCl before crystallisation by sitting drop vapour diffusion. Full-length ClpC2 was added to the crystallisation drop and took ~31 days to form crystals in 25% w/v PEG 3350, 0.1 M Tris-HCl pH 8.5. Crystals were soaked in cryo-protectant (35% w/v PEG 3340, 0.1 M Tris-HCl pH 8.5) prior to flash-freezing in liquid nitrogen. CymA was added 1.1-fold in molar excess to ClpC294–252 and the complex concentrated to 12 mg/ml. The ClpC294–252-CymA complex crystallised in 100 mM sodium acetate-HCl pH 4.5, 30% (w/v) PEG 8 K, 200 mM LiSO4. Crystals were allowed to grow for five days then soaked in cryo-protectant (100 mM sodium acetate-HCl pH 4.5, 30% (w/v) PEG 8 K, 200 mM LiSO4, 15% v/v glycerol) and flash frozen in liquid nitrogen.
Data collection, structure determination, model building and refinement
X-ray diffraction data sets were collected at beamlines X06SA and X06DA of the Swiss Light Source (Paul Scherrer Institute, Villigen, Switzerland) and were indexed and integrated using XDS52. Data merging and scaling was carried out using the programme AIMLESS53 from the CCP4 suite54. The phase problem of ClpC24–72 was solved using ARCIMBOLDO37 with default settings and search of nine copies of a helix containing 14 residues. The output poly-Ala model was then subject to auto-building using AutoBuild software in Phenix55 provided with the ClpC2 protein sequence. The output atomic model was further improved by iterative model building in COOT56 and refinement in Phenix.refine57. The crystal structure of ClpC294–252-CymA was determined by molecular replacement with the programme Phaser58 using the previously solved structure of Mtb ClpC1 NTD (PDB 3WDC) as a search model excluding any ligand. The atomic model was first improved by AutoBuild and then by iterative model building in COOT and refinement in Phenix.refine. Statistics of both structures are summarised in Table 1. All figures were generated with PyMol (The PyMol Molecular Graphics System, version 2.4.1, Schrodinger, LLC) or UCSF Chimera (version 1.15).
Generation of Msm clpC2 knockout strain
A suicide plasmid for the markerless deletion of clpC2 was constructed by amplifying 1500 bp upstream and 1500 bp downstream of clpC2 ORF in Msm genomic DNA using primers listed in Supplementary Table 1a and assembling the PCR fragments into a XmnI digested pGOAL19 plasmid (Addgene plasmid #20190) using NEBuilder HiFi DNA Assembly (New England Biolabs). Following construction of the suicide plasmid, 200 μl Msm electrocompetent cells were transformed by electroporation with 1 μg UV-irradiated plasmid in a 0.2 cm cuvette (BioRad) and a pulse of 2.5 kV applied using a BioRad GenePulser. Electroporated cells were allowed to recover for 5 h at 37 °C in 7H9 and then plated on 7H10 agar plates with added hygromycin. Once colonies began to appear, the agar was underlayed with 0.4% X-Gal to allow for the identification of single-crossover colonies (SCOs). SCOs (blue colonies) were selected and grown in 7H9 at 37 °C before plating on 7H10 agar plates supplemented with 2% (w/v) sucrose followed by continued incubation at 37 °C then underlaying the agar with 0.4% X-Gal. Putative double-crossover colonies were identified by their white colour and screened by PCR for the complete deletion of clpC2.
Growth curves
Msm cultures were diluted to an OD600 of 0.005 in 7H9. Each culture was split between two flasks: CymA (150 nM) was added to one flask and the equivalent volume of DMSO (0.03%) was added to the other. The cultures were grown at 37 °C with shaking (200 rpm) and the culture growth monitored by periodically measuring the OD600.
Western blot analysis
Primary antibodies used for Western blotting are the polyclonal α-ClpC1 (1:10000 dilution) and α-ClpC2 (1:10000 dilution) antibodies raised in rabbits against Mtb ClpC1 and ClpC2, respectively (Eurogentec) and the commercially available monoclonal α-RpoB antibody (BioLegend, 1:10000 dilution, clone 8RB13) raised in mouse. Secondary antibodies were horseradish peroxidase (HRP)-conjugated α-rabbit (abcam, 1:10000 dilution, ab6721) and α-mouse (abcam, 1:10000 dilution, ab6789) antibody. CymA (150 nM) was added to a Msm culture (OD600 of ~0.5) grown in 7H9 media. Cells were harvested by centrifugation at the indicated time points and resuspended in 800 µl 50 mM HEPES-KOH, 150 mM KCl, 1 mM PMSF, 1× cOmplete, EDTA-free, protease inhibitor cocktail (Roche). Resuspended cells were lysed in 2 ml screw cap tubes containing 500–700 mg 0.15 mm zirconium oxide beads (NextAdvance) using a Minilys personal homogeniser (Bertin Technologies). The lysate was clarified by centrifugation for 10 minutes at 20,000 × g and the total protein concentration measured using a NanoDrop™ One spectrophotometer (ThermoScientific). 50 μg protein was separated on a 15% polyacrylamide gel and transferred to a PVDF Immobilon-P membrane (Merck) using a BioRad Trans-Blot SD semidry transfer cell at 20 V for 60 minutes. The PVDF membrane was first blocked for 1 hour with 1% PVP-40 in PBS-T (20 mM Na2HPO4, 2 mM KH2PO4, 137 mM NaCl, 2.7 mM KCl, 0.05% Tween 20) then incubated with the primary antibody in PBS-T with 1% PVP-40 before washing three times in PBS-T and applying the secondary antibody in PBS-T with 1% PVP-40 for 1 hour. The membrane was washed four times for 5 minutes each in PBS-T and the HRP-conjugated secondary antibodies detected using the ECL substrate (BioRad Clarity Western Substrate). Blots were imaged on an Amersham Imager 600 (GE Healthcare).
Quantitative Western blot analysis
Msm cultures were grown to OD600 of 0.5 and 150 nM CymA (or equivalent volume of DMSO) added to the cultures. At the indicated time points 20 OD equivalents of cell culture were pelleted and the cells were lysed following the above protocol. The total protein concentration in the cleared lysates was determined by Bradford Assay and their volume adjusted to a nominal concentration of 2 mg/ml. From here onwards the procedure is the same as stated in the protocol for Western blotting above. Protein levels present at the indicated time points were determined densitometrically from the blots using GelAnalyzer 19.1 by comparison to a standard curve prepared from purified ClpC1 and ClpC2. The ClpC1 and ClpC2 protein levels are reported relative to ClpC1 levels present at the 0 h timepoint.
RT-qPCR
Msm cultures were grown in 7H9 at 37 °C to the stated OD600 before application of the stressor followed by continued incubation for the indicated time. Cells were harvested by centrifugation and then lysed in 2 ml screw cap tubes containing 500–700 mg 0.15 mm zirconium oxide beads using a Minilys personal homogeniser. The lysate was cleared by centrifugation at 20,000 × g for 10 minutes and the total RNA purified using a SPLIT RNA Extraction kit (Lexogen) and following the manufacturer’s instructions. Removal of DNA was achieved using the Turbo DNase kit (ThermoFisher Scientific) and the RNA concentration measured using the NanoDrop™ One spectrophotometer. 100 ng RNA was reverse transcribed to complementary DNA (cDNA) using iScript Select cDNA Synthesis kit (BioRad). Primers for qPCR were designed using Primer3 software59 and are listed in Supplementary Table 1b. qPCR was performed using the Kapa Sybr Fast qPCR kit (Roche) in a BioRad CFX96 thermocycler with an initial 5 minutes at 95 °C step followed by 40 cycles of 95 °C for 10 seconds, 60 °C for 10 seconds, 72 °C for 12 seconds. Data were analysed using BioRad CFX Maestro Software. Relative gene expression was determined using the 2-ΔΔCt method and with rpoB or sigA as the reference gene.
DNA pull-down assay
Msm was grown in 7H9 to an OD600 of 0.5 before addition of 1 µM CymA (or DMSO for the control Msm culture) and incubated for a further 2 h. Cells were pelleted by centrifugation, resuspended in BS/THES (44.3% THES, 20% BS) with added cOmplete protease inhibitor cocktail (Roche) and lysed by bead beating with a Minilys personal homogeniser and 500–700 mg 0.15 mm zirconium oxide beads in 2 ml screw cap tubes. Lysate was sonicated with a Misonix Sonicator 3000 with three cycles of 15 seconds sonication followed by 1 minute incubation on ice and the lysate clarified by centrifugation at 20,000 × g for 20 minutes. The clarified lysate was diluted to 90 ODs/ml and incubated with streptavidin-conjugated agarose beads for 1 hour at room temperature and the precleared lysate separated from the agarose beads by centrifugation at 700 × g for 1 minute. The DNA probe was prepared by amplifying 300 bp upstream clpC2 (or intragenic DNA sequence for the control probe) using biotinylated primers (Supplementary Table 1c). 40 µg DNA was immobilised on 1 mg Dynabeads MyOne Streptavidin C1 (Invitrogen) using a magnetic stand and washed twice with B/W buffer (10 mM Tris-HCl pH 7.5, 1 mM EDTA, 2 M NaCl) followed by three washes with TE buffer (0.5 M Tris-HCl pH 8, 1 mM EDTA) and two washes with BS/THES. The DNA-bound beads were incubated for 30 minutes with 10 µg/ml salmon sperm DNA in BS/THES. DNA-bound beads were resuspended in 750 µl fresh BS/THES containing 50 µg/ml salmon sperm DNA to which 200 µl precleared lysate was added and the mixture incubated for 90 minutes at room temperature with agitation. The beads were washed five times with 500 µl BS/THES followed by a wash with 25 mM Tris-HCl pH 7.5, 100 mM NaCl and then a wash with 25 mM Tris-HCl pH 7.5, 200 mM NaCl. Proteins bound to the immobilised DNA were recovered at the Functional Genomics Center Zurich by incubation with 500 ng trypsin in digestion buffer (10 mM Tris pH 8.2, 2 mM CaCl2) at 60 °C for 30 minutes. The supernatant was collected following trypsin digest and combined with the supernatant produced from incubation of the beads with 0.1% TFA/ 50% acetonitrile then dried and dissolved in 20 μl ddH2O + 0.1% formic acid in preparation for liquid chromatography-tandem mass spectrometry (LC-MS/MS). LC-MS/MS was carried out using a nanoAcquity UPLC coupled to a Q-Exactive mass spectrometer (Thermo) and data analysed using the Mascot search engine (Matrixscience) where spectra were searched against Mycobacterium smegmatis (Swissprot) for the identification of proteins. Relative amounts of ClpC2 co-purified and detected with either the control DNA or clpC2 promoter DNA sequence were determined from spectral counts.
Electrophoretic mobility shift assay
DNA probes were synthesised by Microsynth AG and are listed in Supplementary Table 1d. To prepare double-stranded DNA, complementary strands were annealed by heating equimolar concentrations to 95 °C in a thermocycler and slowly cooled to 25 °C by reducing the temperature by 1 °C every minute. DNA-binding reactions were prepared by mixing the DNA probe with the indicated concentrations of protein in 5 mM HEPES-KOH pH 7.5, 30 mM KCl, 1% Glycerol (v/v), 10 mM MgCl2 and incubated at room temperature for 20 minutes before loading onto a 12% polyacrylamide gel. To analyse the effects of CymA on DNA-binding, 1 µM CymA was added to the binding reaction and the DMSO concentration matched for the control without CymA and the gel band density was quantified using GelAnalyzer 19.1 software.
Statistics and reproducibility
Statistical analysis and graphs were prepared using GraphPad Prism 9.3.1 software. Data are presented as mean ± standard deviation (SD) where relevant and the number of replicates is described in the respective figure legend. P values <0.05 were considered significant and statistical tests used are stated in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates and structure factors have been deposited in the Protein Data Bank database under the accession codes 8ADA (ClpC24–72) and 8AD9 (ClpC294–252-CymA). All other data are either contained in the paper or in the associated supplementary information files. Source data underlying graphs and uncropped images of gels and blots presented in this paper are provided in Supplementary Data 2.
References
Jassal, M. & Bishai, W. R. Extensively drug-resistant tuberculosis. Lancet Infect. Dis. 9, 19–30 (2009).
Portelli, S., Phelan, J. E., Ascher, D. B., Clark, T. G. & Furnham, N. Understanding molecular consequences of putative drug resistant mutations in Mycobacterium tuberculosis. Sci. Rep. 8, https://doi.org/10.1038/s41598-018-33370-6 (2018).
Chakaya, J. et al. Global tuberculosis report 2020 - reflections on the global TB burden, treatment and prevention efforts. Int. J. Infect. Dis. 113, S7–S12 (2021).
Kashongwe, I. M. et al. Outcomes and adverse events of pre- and extensively drug-resistant tuberculosis patients in Kinshasa, Democratique Republic of the Congo: a retrospective cohort study. Plos One 15, e0236264 (2020).
Ghazy, R. M. et al. A systematic review and meta-analysis of the catastrophic costs incurred by tuberculosis patients. Sci. Rep. 12, https://doi.org/10.1038/s41598-021-04345-x (2022).
WHO. Global Tuberculosis Report 2020. (2020).
Larsen, C. N. & Finley, D. Protein translocation channels in the proteasome and other proteases. Cell 91, 431–434 (1997).
Striebel, F., Kress, W. & Weber-Ban, E. Controlled destruction: AAA(+) ATPases in protein degradation from bacteria to eukaryotes. Curr. Opin. Struct Biol. 19, 209–217 (2009).
Horwich, A. L., Weber-Ban, E. U. & Finley, D. Chaperone rings in protein folding and degradation. Proc. Natl. Acad. Sci. USA 96, 11033–11040 (1999).
Weber-Ban, E. U., Reid, B. G., Miranker, A. D. & Horwich, A. L. Global unfolding of a substrate protein by the Hsp100 chaperone ClpA. Nature 401, 90–93 (1999).
Kress, W., Maglica, Z. & Weber-Ban, E. Clp chaperone-proteases: structure and function. Res. Microbiol. 160, 618–628 (2009).
Yu, A. Y. H. & Houry, W. A. ClpP: A distinctive family of cylindrical energy-dependent serine proteases. FEBS Lett. 581, 3749–3757 (2007).
Benaroudj, N., Raynal, B., Miot, M. & Ortiz-Lombardia, M. Assembly and proteolytic processing of mycobacterial ClpP1 and ClpP2. BMC Biochem. 12, https://doi.org/10.1186/1471-2091-12-61 (2011).
Akopian, T. et al. The active ClpP protease from M. tuberculosis is a complex composed of a heptameric ClpP1 and a ClpP2 ring. EMBO J. 31, 1529–1541 (2012).
Leodolter, J., Warweg, J. & Weber-Ban, E. The Mycobacterium tuberculosis ClpP1P2 protease interacts asymmetrically with its ATPase partners ClpX and ClpC1. Plos One 10, e0125345 (2015).
Kar, N. P., Sikriwal, D., Rath, P., Choudhary, R. K. & Batra, J. K. Mycobacterium tuberculosis ClpC1: characterization and role of the N-terminal domain in its function. FEBS J. 275, 6149–6158 (2008).
Dougan, D. A., Reid, B. G., Horwich, A. L. & Bukau, B. ClpS, a substrate modulator of the ClpAP machine. Mol. Cell 9, 673–683 (2002).
Dougan, D. A., Weber-Ban, E. & Bukau, B. Targeted delivery of an ssrA-tagged substrate by the adaptor protein SspB to its cognate AAA plus protein ClpX. Mol. Cell 12, 373–380 (2003).
Lau, J., Hernandez-Alicea, L., Vass, R. H. & Chien, P. A phosphosignaling adaptor primes the AAA+ protease ClpXP to drive cell cycle-regulated proteolysis. Mol. Cell 59, 104–116 (2015).
Kirstein, J. et al. Adaptor protein controlled oligomerization activates the AAA+ protein ClpC. EMBO J. 25, 1481–1491 (2006).
Maurer, M. et al. Toxic activation of an AAA plus protease by the antibacterial drug cyclomarin A. Cell Chem. Biol. 26, 1169 (2019).
Roman-Hernandez, G., Hou, J. Y., Grant, R. A., Sauer, R. T. & Baker, T. A. The ClpS adaptor mediates staged delivery of n-end rule substrates to the AAA plus ClpAP protease. Mol. Cell 43, 217–228 (2011).
Ziemski, M., Leodolter, J., Taylor, G., Kerschenmeyer, A. & Weber-Ban, E. Genome-wide interaction screen for Mycobacterium tuberculosis ClpCP protease reveals toxin-antitoxin systems as a major substrate class. FEBS J. 288, 111–126 (2021).
Trentini, D. B. et al. Arginine phosphorylation marks proteins for degradation by a Clp protease. Nature 539, 48 (2016).
Raju, R. M. et al. Mycobacterium tuberculosis ClpP1 and ClpP2 function together in protein degradation and are required for viability in vitro and during infection. Plos Pathog. 8, https://doi.org/10.1371/journal.ppat.1002511 (2012).
Winther, K. S., Brodersen, D. E., Brown, A. K. & Gerdes, K. VapC20 of Mycobacterium tuberculosis cleaves the sarcin-ricin loop of 23S rRNA. Nat. Commun. 4, 2796 (2013).
Griffin, J. E. et al. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. Plos Pathog. 7, https://doi.org/10.1371/journal.ppat.1002251 (2011).
Lupoli, T. J., Vaubourgeix, J., Burns-Huang, K. & Gold, B. Targeting the proteostasis network for mycobacterial drug discovery. ACS Infect. Dis. 4, 478–498 (2018).
Gavrish, E. et al. Lassomycin, a ribosomally synthesized cyclic peptide, kills mycobacterium tuberculosis by targeting the ATP-dependent protease ClpC1P1P2. Chem. Biol. 21, 509–518 (2014).
Wolf, N. M. et al. Structure of the N-terminal domain of ClpC1 in complex with the antituberculosis natural product ecumicin reveals unique binding interactions. Acta Crystallogr. D 76, 458–471 (2020).
Wolf, N. M. et al. High-resolution structure of cipc1-rufomycin and ligand binding studies provide a framework to design and optimize anti-tuberculosis leads. ACS Infect. Dis. 5, 829–840 (2019).
Vasudevan, D., Rao, S. P. S. & Noble, C. G. Structural basis of mycobacterial inhibition by cyclomarin A. J. Biol. Chem. 288, 30883–30891 (2013).
Renner, M. K. et al. Cyclomarins A-C, new antiinflammatory cyclic peptides produced by a marine bacterium (Streptomyces sp.). J. Am. Chem. Soc. 121, 11273–11276 (1999).
Schmitt, E. K. et al. The natural product cyclomarin kills mycobacterium tuberculosis by targeting the ClpC1 subunit of the caseinolytic protease. Angew. Chem. Int. Ed. Engl. 50, 5889–5891 (2011).
Burstner, N. et al. Gift from nature: cyclomarin a kills mycobacteria and malaria parasites by distinct modes of action. Chembiochem 16, 2433–2436 (2015).
Taylor, G. et al. Antibacterial peptide CyclomarinA creates toxicity by deregulating the Mycobacterium tuberculosis ClpC1/ClpP1P2 protease. J. Biol. Chem. 298, 102202, (2022).
Rodriguez, D. D. et al. Crystallographic ab initio protein structure solution below atomic resolution. Nat. Methods 6, 651–U639 (2009).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Chaparian, R. R. & van Kessel, J. C. Promoter pull-down assay: a biochemical screen for DNA-binding proteins. Methods Mol. Biol. 2346, 165–172 (2021).
Gopal, P. et al. Pyrazinamide triggers degradation of its target aspartate decarboxylase. Nat. Commun. 11, https://doi.org/10.1038/s41467-020-15516-1 (2020).
Gao, W. et al. The cyclic peptide ecumicin targeting ClpC1 is active against mycobacterium tuberculosis in vivo. Antimicrob. Agents. Chemother. 59, 880–889 (2015).
Choules, M. P. et al. Rufomycin targets ClpC1 proteolysis in mycobacterium tuberculosis and M. abscessus. Antimicrob. Agents Chemother. 63, https://doi.org/10.1128/AAC.02204-18 (2019).
Mulvenna, N. et al. Xenogeneic modulation of the ClpCP protease of Bacillus subtilis by a phage-encoded adaptor-like protein. J. Biol. Chem. 294, 17501–17511 (2019).
Lin, W. W. et al. Transcriptional profiling of mycobacterium tuberculosis exposed to in vitro lysosomal stress. Infect. Immun. 84, 2505–2523 (2016).
Salina, E. G., Mollenkopf, H. J., Kaufmann, S. H. E. & Kaprelyants, A. S. M-tuberculosis gene expression during transition to the “non-culturable” state. Acta Naturae 1, 73–77 (2009).
Salina, E. G. et al. Potassium availability triggers Mycobacterium tuberculosis transition to, and resuscitation from, non-culturable (dormant) states. Open Biol. 4, https://doi.org/10.1098/rsob.140106 (2014).
Keren, I., Minami, S., Rubin, E. & Lewis, K. Characterization and transcriptome analysis of Mycobacterium tuberculosis persisters. Mbio 2, https://doi.org/10.1128/mBio.00100-11 (2011).
Torrey, H. L., Keren, I., Via, L. E., Lee, J. S. & Lewis, K. High persister mutants in Mycobacterium tuberculosis. Plos One 11, https://doi.org/10.1371/journal.pone.0155127 (2016).
Betts, J. C., Lukey, P. T., Robb, L. C., McAdam, R. A. & Duncan, K. Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol. Microbiol. 43, 717–731 (2002).
Kubik, S., Wegrzyn, K., Pierechod, M. & Konieczny, I. Opposing effects of DNA on proteolysis of a replication initiator. Nucleic Acids Res. 40, 1148–1159 (2012).
Bury, K., Wegrzyn, K. & Konieczny, I. Handcuffing reversal is facilitated by proteases and replication initiator monomers. Nucleic Acids Res. 45, 3953–3966 (2017).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D 66, 133–144 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution. Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Bailey, S. The Ccp4 suite - programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Terwilliger, T. C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D 64, 61–69 (2008).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Mccoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Koressaar, T. et al. Primer3_masker: integrating masking of template sequence with primer design software. Bioinformatics 34, 1937–1938 (2018).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010).
Celniker, G. et al. ConSurf: using evolutionary data to raise testable hypotheses about protein function. Isr. J. Chem. 53, 199–206 (2013).
Acknowledgements
We are grateful to Novartis for supplying Cyclomarin A for this study. We thank A. Cléry and J. Scheuermann for the initial help with ITC instrumentation and for the use of their ITC instrument, respectively. We would also like to thank the Functional Genomics Center Zurich for the acquisition of mass spectrometry data as well as the Protein Crystallisation Center Zurich for screening crystal conditions. In addition, we thank the Paul Scherrer Institute for the provision of synchrotron radiation beamtime and the staff at beamlines PXI (X06SA) and PXIII (X06DA) of the Swiss Light Source (SLS) for their assistance. This work was supported by an Eidgenössische Technische Hochschule Zürich research grant (ETH-40 16-1) to E.W.B.
Author information
Authors and Affiliations
Contributions
G.T. and E.W.B. conceived and planned the study. G.T., H.C. and J.L. performed the experiments. G.T., H.C., J.L., C.G. and E.W.B. analysed the data. G.T. and H.C. prepared the figures. G.T. and E.W.B. drafted the manuscript. G.T., H.C., J.L., C.G. and E.W.B. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Byung-Gil Lee and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Gene Chong.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Taylor, G., Cui, H., Leodolter, J. et al. ClpC2 protects mycobacteria against a natural antibiotic targeting ClpC1-dependent protein degradation. Commun Biol 6, 301 (2023). https://doi.org/10.1038/s42003-023-04658-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04658-9
This article is cited by
-
Protein quality control: from molecular mechanisms to therapeutic intervention—EMBO workshop, May 21–26 2023, Srebreno, Croatia
Cell Stress and Chaperones (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.