Abstract
Bromus tectorum L. is arguably the most successful invasive weed in the world. It has fundamentally altered arid ecosystems of the western United States, where it now found on an excess of 20 million hectares. Invasion success is related to avoidance of abiotic stress and human management. Early flowering is a heritable trait utilized by B. tectorum, enabling the species to temporally monopolize limited resources and outcompete the native plant community. Thus, understanding the genetic underpinning of flowering time is critical for the design of integrated management strategies. To study flowering time traits in B. tectorum, we assembled a chromosome scale reference genome for B. tectorum. To assess the utility of the assembled genome, 121 diverse B. tectorum accessions are phenotyped and subjected to a genome wide association study (GWAS). Candidate genes, representing homologs of genes that have been previously associated with plant height or flowering phenology traits in related species are located near QTLs we identified. This study uses a high-resolution GWAS to identify reproductive phenology genes in a weedy species and represents a considerable step forward in understanding the mechanisms underlying genetic plasticity in one of the most successful invasive weed species.
Similar content being viewed by others
Introduction
Bromus tectorum is the most abundant invasive weed in North America. In the western United States, it is estimated to infest 31.4% (210,000 km2) of the Great Basin1. It is most notorious for and especially problematic in non-cropland and rangelands of the intermountain west where B. tectorum invasion has altered intervals between fires from at least 60 years to less than 5 years2,3, exacerbating the degradation of ecosystems caused by climate change1,4. Bromus tectorum is also a damaging weed in agricultural crops, causing substantial yield loss in winter wheat (Triticum aestivum L.) across a large proportion of the western North American wheat producing region5.
The history of Bromus tectorum (L.) in North America is proposed to have begun as the arrival of a small number of founder genotypes from multiple introductions originating from a wide range of native habitats in Eurasia6. Due to multiple introductions of B. tectorum from Eurasia to North America the overall genetic diversity in Eurasia is higher than in North America but within populations the genetic diversity was higher in North America because the North American populations of B. tectorum are often a heterogenous mix of the introduced genotypes6. The seed likely arrived in animal bedding, grain contaminants, or it was imported intentionally as a potential forage7. Bromus tectorum, known as cheatgrass or downy brome, is predominantly a self-fertilizing, cleistogamous species, adapted to multiple ecosystems in its native range8. Although the species is distributed across North American ecosystems, the population has retained genetic signatures that trace back to ancestral populations and ecosystems6. Thus, the success of B. tectorum invasiveness is due in large part to the diversity of these original populations that exhibit substantial levels of plasticity in phenological traits. Plasticity allows individual B. tectorum plants to establish them-selves in a locale by being pre-adapted to local changes in the availability of in-season resources9, while populations of B. tectorum are composed of an assemblage of diverse genotypes, facilitating success in response to long term and local variation in climate10,11,12. Such adaptive variation in life cycle traits13,14 hinders management efforts in both natural and agricultural ecosystems. In short, B. tectorum individuals and populations express adaptive plasticity and, when expressed as earlier and variable reproductive phenology, allow B. tectorum to outcompete for limited resources at the expense of the native plant community9. Where it successfully invades, B. tectorum germinates and flowers early, facilitating access to limited resources, usually moisture, well before the native vegetation or crops can compete successfully for these resources15,16.
Phenotypic variation in B. tectorum for adaptive traits like the aforementioned flowering time, but also seed dormancy and vernalization are undoubtedly major drivers of B. tectorum’s highly successful invasion across a wide variety of North American ecosystems13,15,16,17. In model and crop Poaceae species, growth and flowering phenology is controlled by an array of light and temperature sensitive gene pathways. Unfortunately, little is currently known about the genetic basis of adaptive variation in B. tectorum. Orthologous genes, such as FT (flowering time), VRN1 (VERNALIZATION1), and VRN2 (VERNALIZATION2) are likely contributing factors18. As a member of the Pooideae subfamily in the Poaceae family, B. tectorum is closely related to the Triticeae tribe, which includes the agriculturally important species barley (Hordeum vulgare L.), wheat (Triticum aestivum L.) and rye (Secale cereale L.)19,20. Thus, extensive genomic resources, including well annotated genomes, in these sister taxa are available that will facilitate comparative genomics and gene discovery efforts in B. tectorum.
Here we report a high-quality, annotated reference genome for B. tectorum. We utilize the reference genome in a genome wide association study (GWAS) to identify candidate genes for reproductive phenology (days to first joint, days to first visible panicle, days to first ripe seed and number of tillers) and plant height—traits that are known to directly influence the success of B. tectorum as an invasive weed species. Genome wide association studies (GWAS) are particularly useful for dissecting complex traits in species where controlled crossing is not practical or possible21,22,23,24,25. Our study demonstrates a successful application of GWAS for the identification of QTL controlling heritable reproductive phenology traits in a weedy, highly invasive species. The identification of candidate gene targets controlling important climate and management adaptive characteristics underlying these QTL suggests how B. tectorum might respond to climate change, thus enabling the development enhanced and more reliable management practices for this highly invasive and problematic weed species.
Results and discussion
Genome assembly
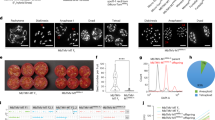
The Omni-C chromosomal assembly produced a near complete assembly for B. tectorum. The assembly was 2,482 megabases (Mb) in total length (Supplementary Table S1) with 1298 genes and 82% of the genome consisting of repetitive elements (Supplementary Table S2). The contig N50 was 19.4 Mb, the scaffold N50 was 357.4, and the resulting assembly contained 92.1% of the BUSCO genes with 259 gaps. The final assembly had a L50 of four and a L90 of seven corresponding to the seven chromosomes (x = 7) expected for members of the Pooideae in the Poeae, Aveneae, Bromeae and Triticeae tribes26.
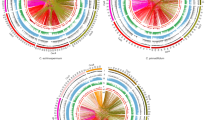
Based on CDS comparisons using MCScanX27, the seven largest scaffolds of the B. tectorum reference genome were found to have a one-to-one syntenic relationship with the seven chromosomes of the barley genome—not surprising given their phylogenetic proximity. As expected, synteny between B. tectorum and H. vulgare was highest in the chromosome arms where gene density is known to be high and lowest though the centromeric region where gene density is substantially reduced (Fig. 1). A translocation is present between chromosomes two and five where the first half of Bt5 has synteny with chromosome two in barley and the second half of Bt2 has synteny with chromosome 5 in barley (Fig. 1).
Phenotypic analysis
Broad Sense Heritability (Reliability), the mean, standard deviation, min, max, and correlation of Best Linear Unbiased Estimates (BLUEs) were calculated to characterize the variation in traits and obtain a measure of the effect of each genotype. The distributions of BLUEs for reproductive phenology traits indicated a bimodal distribution (Fig. 2), where more rapidly flowering and taller plants consisted of accessions from Washington and GRIN collections. In contrast, Montana accessions flowered later and were shorter (Supplementary Data 1, Supplementary Data 2 and Fig. S1). Broad sense heritability was high for all the traits and ranged from 0.94 for tiller number to 0.99 for days to first visible panicle (VPN) (Table 1). Genotypes had a wide range of BLUEs for all traits measured: plant height (PH) ranged from 36.5 to 89 cm, number of tillers ranged from 5.5 to 38.5, and days to first ripe seed (FRS) ranged from 43.2 to 112.5 d (Table 1). Spearman correlations between BLUEs of different reproductive phenology traits were all above 0.95 (Fig. 2). Reproductive phenology and PH traits were moderately negatively correlated, with Spearman correlations ranging from the −0.44 (PH and FRS) to −0.53 (PH and J1).
Histograms with density ticks and a smoothing line are on the diagonals. The upper diagonal contains the Spearman correlation between BLUEs of traits with “***” denoting a significant level (p < 0.0001) of correlation between traits, where N = 121 genotypes. The lower diagonal contains scatter plots between traits with center dot, a centroid with a standard deviation of 1 and locally estimated scatterplot smoothing (LOESS) smoothing curve.
Genome-wide association mapping for height, tiller number, and phenology traits
To identify regions of the genome associated with variation in adaptive traits, a GWAS was performed on 121 genotypes to find QTL for PH, VPN, days to first visible joint (J1), FRS, days to 50% ripe seed (AWN50) and tiller number using BLINK with three principal components and significance threshold of 0.05 after multiple testing correction. Nineteen QTLs were significantly (q < 0.05) associated with PH and reproductive phenology, with one QTL for PH (1), nine QTL for VPN (9), three QTL for J1 (3), nine QTL for FRS (9), and three QTL for AWN50 (3). A QTL on Bt6 (Bt6:8628087) was significant (q < 0.05) for all the reproductive phenology related traits except J1, where a second significant (q = 2.070E-07) QTL for J1 was at a nearby position (Bt6:8417488) on Bt6 (Table 2). The QTL on Bt1 (Bt1:276755111) was significant (q < 0.05) for J1, FRS, and AWN50 (Table 2). QTLs on Bt2 (Bt2:9403921) and Bt7 (Bt7:347158582) were significantly (q < 0.05) associated with VPN and FRS (Table 2).
Plant height
We identified a QTL that was significantly (q = 1.360E-05, Table 2) associated with PH on Bt6:301800092. The QTL explained 16.9% (Table 2) of the phenotypic variation. The MAF at the PH QTL on Bt6 was 0.44, indicating both allelic states are common in our panel. Searching the area flanking the QTL associated with PH at Bt6:301800092 on both sides by up to 500 Kb revealed a homolog of Xanthine Dehydrogenase (XDH) 29 Kb from the QTL and a homolog of Indole-3-pyruvate monooxygenase YUCCA6 (YUC6) 242 Kb from the QTL (Table 3). XDH and YUC6 are promising candidate genes for the QTL associated with PH we identified on Bt6 as they both are well document to be associated with changes in PH, senescence, and response to drought28,29. In rice (Oryza sativa L.), overexpression of the XDH homolog led to increased PH while under-expression of the XDH homolog resulted in reduced PH28, indicating that homologs of XDH would be fitting candidate genes for PH. Indeed, in rice, a GWAS identified XDH as a candidate gene for coleoptile length in response to flooding30 indicating XDH homologs may be involved with stem elongation in grasses and thus PH. Furthermore, in Arabidopsis knock-out mutations of the XDH gene led to reduced PH31. Knock-out mutations of the YUC6 gene in Arabidopsis was found to increase auxin production in the shoots leading to reduced PH32. Further investigation, including gene expression experiments and/or knockout of the XDH gene homolog in B. tectorum, is needed to validate and further understand the underlying genetics controlling the large effect QTL for PH.
Unexpectedly, PH and reproductive phenology timing were negatively correlated, contradicting previous findings that PH and reproductive phenology were positively correlated (i.e., earlier flowering plants do not have as much time to grow)33,34,35. Lolium perenne (L.) was also found to have a negative genetic correlation between PH and flowering time36. The negative correlation was thought to be the result of selection imposed by grazing, where biological fit plants remained short until they were ready to flower at which point they elongated and flowered quickly37. Bromus tectorum may also be under similar grazing selection pressure38. Competition with crops could also select for taller, fast-growing phenotypes to facilitate competition for space.
Although only a single QTL was uncovered by the GWAS for PH in B. tectorum, there is likely many QTL of small effect that GWAS model did not have the statistical power to detect. When GWASs were used to uncover variants associated with complex diseases in humans, the QTLs identified only explained a fraction of the variation compared to what was expected based on heritability estimates39, becoming known as “the missing heritability problem”. Because the missing heritability problem is caused by a lack of power to detect causal variants and by inflated measures of heritability, increasing the sample size can partially resolve the heritability problem by increasing the power to detect causal variants. In Atlantic salmon (Salmo salar L.), an initial GWAS for age at maturity with a sample size of 1518 only detected a single QTL that explained 39% of the variation in age at maturity40. However, a subsequent GWAS study for age at maturity in S. salar with a sample size 11,166 uncovered the same QTL previously identified along with 115 other QTLs with smaller effect sizes and low MAFs41. If subsequent GWASs are performed for PH in B. tectorum, then substantially increasing the number of genotypes in the GWAS would likely uncover smaller effect loci contributing to PH.
Phenology traits
Flowering time is an important adaptive trait for ensuring the survival of plants in a broad range of climates, often driving local adaptation42,43,44. In predominantly self-fertilizing species, large effect QTLs control the variation in flowering time45,46, in contrast to maize where flowering time is controlled by many QTL of small effect47. Here, we phenotyped four traits (VPN, J1, FRS, AWN50) associated with flowering time to facilitate a GWAS to identify candidate reproductive phenology genes. Days to first visible panicle (VPN) was the first reproductive phenology stage observed and had the highest heritability at 0.99 (Table 1). The largest effect estimated for a significant (q = 3.630E-07) QTL for VPN was found at Bt7:70795764, explaining 16.3% of the variation in VPN with a MAF of 0.26. The eight-remaining significant (q < 0.05) QTL on Bt3 (Bt3:2025814), Bt4 (Bt4:382636922), Bt6 (Bt6:8628087), Bt7 (Bt7:347158582), Bt2 (Bt2:9403921), Bt3 (Bt3:383345273), Bt3 (Bt3:324738958) and Bt3 (Bt3:324739133) explained 4.3, 4.3, 2.9, 4.7, 3.3, 3.8, 3.4 and 3.4% of the phenotypic variation, respectively, with MAF ranging from 0.2 to 0.49 (Table 2). Small to moderate effect QTL were also underlying the emergence of panicles from the flag leaf in barley48 with the PVE of QTLs ranging from 1 to 13%.
Developmentally, days to first joint (J1) was the second reproductive phenology trait to occur. Three significant (q < 0.05) quantitative trait loci (QTL) were associated with J1. The QTL at Bt6 (Bt6:8417488; q = 2.070E−07) was 12 Kb from a homolog of Heading Date Repression 1 (HDR1) and explained 6.7% of the phenotypic variation with an MAF of 0.35 (Tables 2, 3). The QTL at Bt1 (Bt1:276755111; q = 3.870E−05) explained 8.1% of the phenotypic variation with a MAF of 0.2. The other QTL on Bt1 (Bt1:169814345; q = 1.909E−03) explained 4.2% of the phenotypic variation and had an MAF of 0.35. Although no previous studies have directly mapped genes for time of the visible first joint in a grass species, the high correlations between reproductive phenology traits (Fig. 2) indicate that the genetic architecture of J1 in B. tectorum should be like the genetic architecture of other reproductive phenology traits. A QTL mapping study in Panicum hallii (Vasey), a perennial primarily self-fertilizing grass species native to the Southwestern United States, revealed only two QTL controlling flowering time with PVEs of 6.4 and 7.449, indicating that moderate effect QTL like those found underlying J1 may be underlying reproductive phenology traits in wild populations of self-fertilizing grasses.
The next developmental stage was first ripe seed (FRS). Days to first ripe seed was associated with nine significantly (q < 0.05) associated QTL. The leading QTL located on Bt7 (Bt7:347158582; q = 1.360E−05) explained 7.2% of the phenotypic variation of FRS and was common (MAF = 0.29) (Table 2). The remaining QTLs on Bt7 (Bt7:347158582; q = 5.280E−09), Bt1 (Bt1:173453655; q = 1.582E−04), Bt1(Bt1:39144935; q = 1.786E−04), Bt3 (Bt3:48964562; q = 1.786E−04), Bt1 (Bt1:276755111; q = 2.823E−04), Bt2 (Bt2:9403921; q = 4.166E−04), Bt4 (Bt4:382201414; q = 1.026E−03), Bt3 (Bt3:4974562; q = 1.836E−03) and Bt6 (Bt6:8628087; q = 4.110E−02) explained 1.4%, 1.8%, 1.2%, 4.1%, 1.9%, 0.6%, 4.0% and 2.3% of the variation, respectively (Table 2). The MAF of QTL associated with FRS were lower than those identified for the other reproductive phenology traits, varying from 0.058 to 0.29.
In Erythranthe laciniata (A. Gray), a self-fertilizing dicot plant species, populations along an altitudinal gradient differed in their times to FRS with populations from low elevations having the earliest FRS dates and high-elevation populations having the latest FRS dates50. The differences in the FRS phenotype between populations of E. lacinata are likely driven by moderate to large effect QTL controlling other reproductive phenology trait with PVEs ranging from 9 to 39%51, exceeding the effect sizes identified for FRS in B. tectorum (Table 2). However, the QTL mapping study in E. laciniata included both genotypes that require and do not require vernalization51. The GWAS study presented here only included B. tectorum genotypes that required vernalization, thus major QTL associated with vernalization would not be present. The lack of variation in vernalization genes in the B. tectorum genotypes used for the GWAS could explain the absence of large effect QTL underlying FRS in B. tectorum.
The final developmental reproductive phenology trait to occur was AWN50, resulting in three significant associations (q < 0.05). The leading QTL on Bt1 (Bt1:276755111; q = 6.760E−05) explained 6.5% of the phenotypic variation and had a MAF of 0.2 (Table 2). The other two QTL were located on Bt5 (Bt5:348443767; q = 1.637E-04) and Bt6 (Bt6:8628087; q = 5.973E−03) explaining 5.6% and 4.8% of the phenotypic variation with MAFs of 0.37 and 0.2, respectively. Although no previous studies have mapped genes for AWN50, the high correlation with other reproductive phenology traits likely means that the genetic architecture underlying the traits will be similar.
Although phenotypic Spearman correlations between reproductive phenology traits are all above 0.95 (Fig. 2), and there is overlap in the QTLs detected (Table 2), the number of associated QTL detected varied from three to nine for reproductive phenology traits (Table 2). The most likely explanation is that slight differences in the phenotype are great enough to change the p-values for association of QTL with a trait. In maize, a study revealed nine QTLs for growing degree days (GDD) to silk while one of the QTL identified for GDD to silk was the only QTL identified by the GWAS for GDD to tassel52, indicating that it is possible for correlated reproductive phenology traits to yield different number of QTL. Furthermore, the GWAS study presented here uses the GWAS method BLINK, a multi-locus iterative GWAS model53. The process of BLINK iteratively updating the model and recalculating p-values means that if the traits are differing, even by a small amount, that different sets of QTLs could be selected early on to build the statistical model used for significance testing thus leading to substantially different sets of QTL being identified for the two correlated traits.
Survey of candidate genes associated with phenology
Seventeen genes were identified as punitive candidates for reproductive phenology traits, based on their proximity to QTL (within a 500 Kb interval, spanning 250 Kb on either side of the SNP) and previous functional characterization related to specific phenologies. The most promising of these genes is a homolog of HDR1. The proximity of a homolog of HDR1 with QTL from all the maturity-related trait analyzed suggests that HDR1 influences maturation in B. tectorum. In O. sativa, HDR1 promotes Heading date 1 (Hd1) and represses Early heading date 1 (Ehd1) delaying flowering54. Knockout mutations or RNA interference of HDR1 resulted in rice plants that flowered 30 days earlier in long day light conditions, making HDR1 a promising candidate gene for reproductive phenology traits54.
Multiple candidate genes were identified for several reproductive phenology-associated QTL, indicating multiple genes may underlie reproductive phenology in B. tectorum. The QTL on Bt2 (9403921) associated with VPN and FRS was 13, 29 and 133 Kb from homologs of ABC transporter B family member 19 (ABCB19), Cullin-3A (CUL3A), and Gibberellin 20 oxidase 2 (GA20OX2), respectively. Loss of function mutations in ABCB19, CUL3A, and GA20OX2 led to longer flowering times, indicating all three promote advancement of reproductive phenology55,56,57. Coincidentally, a gene near another QTL associated with FRS is known to interact with GA20OX2. The QTL on Bt1 (173453655) associated with FRS was 78Kb from orthologs of FAR1-RELATED SEQUENCE 5 (FRS5), a punitive transcription factor regulating far-red light control of development58, associated with adaptation to photoperiod59, and dehydration-responsive element-binding protein 1 F (DREB1F), a putative transcription factor, when upregulated, represses gibberellic acid (GA) biosynthesis catalyzed by GA20OX genes. The upregulation of DREB1F causing shorter plants with delayed flowering60, indicates that DREB1F and GA20OX can act in an epistatic manner to control reproductive phenology. A likelihood ratio test of the interaction between the FRS QTLs on Bt2 (9403921) and Bt1 (173453655) revealed epistasis is likely (X2 = 3.71 df = 1, p = 0.054), indicating DREB1F and GA20OX are interacting to control FRS in an epistatic manner. The epistatic interaction also indicates further that DREB1F and GA20OX are likely the genes actively controlling FRS at the QTL on Bt2 (9403921) and Bt1 (173453655).
The QTL associated with VPN on Bt3 (2025814) was 120Kb, 177Kb and 240Kb from homologues of BTB/POZ and MATH domain-containing protein 1 (BPM1), BTB/POZ and MATH domain-containing protein 2 (BPM2) and PHYTOCHROME-DEPENDENT LATE-FLOWERING (PHL), respectively. PHL triggers flowering under long-day conditions by repressing the phytochrome b (PHYB) and the constans (CO) genes61. BPM1 and BPM2 are transcription factors of the BTB/POZ and MATH domain-containing protein (BPM) gene family and are involved in regulating flowering time by making proteins that are part of the Cullin E3 ubiquitin ligase complexes that include CUL3A62. A likelihood ratio test did not detect epistasis (X2 = 0.02, df = 1, p = 0.901) between the QTL on Bt2 (9403921) near CUL3A and the QTL on Bt3 (2025814) near BPM1 and BPM2.
A QTL on Bt3 (324738958) associated with VPN was located 64Kb, 113Kb and 189Kb from homologs of UPSTREAM of FLC (UFC), Ultraviolet-B receptor 8 (UVR8) and Early Flowering 7 (VIP2), respectively. UVR8, VIP2 and UFC are involved with the regulation of Flowering Locus C (FLC) in A. thaliana63,64,65, suggesting one mechanism maintaining genetic variation in B. tectorum flowering time is the regulation of FLC expression. Homologs of four genes in the FHY3/FAR1 gene family were near QTL found associated with maturity traits indicating the FHY3/FAR1 gene family is part of a mechanism maintaining variation in maturity traits in B. tectorum. Homologues of FHY3 and FRS6 were found 147 and 149 Kb from a QTL on Bt7 (Table 3), respectively. A homologue of FRS7 was near the QTL on Bt3 associated with VPN (Table 3) FRS5 discussed earlier is also member of the FHY3/FAR1 gene family. The FHY3/FAR1 gene family is comprised of 14 homologous genes, regulating transcription as a response to far red light58. FHY3, FRS6, FRS7 have been demonstrated to regulate flowering time in A. thaliana58,66,67 making members of the FHY3/FAR1 good candidate genes for maturity traits.
In addition, a Cytokinin Dehydrogenase 2 (CKX2) homolog was identified as a candidate gene for the QTL at Bt3:48964562 associated with FRS. CKX2 catalyzes the oxidation of cytokinins68. Cytokinins have been shown to regulate flowering time via transcriptional activation of Twin Sister of FT (TST)69 and overexpression in members of the Cytokinin Dehydrogenase (CKX) gene family have shown to delay flowering time in long day conditions68 indicating that the CKX2 homolog could be controlling maturity traits through the regulation of cytokinins in B. tectorum.
Implications of the reference genome and phenology GWAS
The reference quality genome we have assembled is an invaluable resource for understanding the fundamental genetic controls that have facilitate one of the most successful invasive weeds in North America. Understanding the genetic basis of adaptive traits in B. tectorum will lead to improved management strategies. Using the genome we explored the genetic underpinnings of reproductive phenology traits in B. tectorum, revealing pathways and mechanisms contributing to adaptive plasticity which directly contributes to the species invasive spread across a wide range of environments. Indeed, our GWAS uncovered genetic mechanisms contributing to plasticity within the species. We identified QTL for maturity traits that were near candidate genes responsible for controlling the photoperiod pathway, plant hormone regulation, and transcription factors triggered by far-red or UV light.
Our study indicates that not only is the genome very similar to barley (Fig. 1), but the adaptive control of reproductive phenology also closely mirrors barley. Both domesticated and wild barley are adapted to latitudinal clines, where reproductive phenology is controlled by genes responding to environmental cues70. Our identification of a GA20OX ortholog as a candidate gene for reproductive phenology corroborates previous work, where GA20OX loss of function was found to reduce PH and delay flowering35,71,72. Interestingly, GA20OX orthologs are semi-dwarfing genes implicated in the green revolution73. The semi-dwarfing gene we identified as a candidate gene could also explain the negative correlation between reproductive phenology traits and PH. In addition, wild barley was found to adapt using variation in photoperiod response genes70, supporting our findings where the FHY3/FAR1 photoperiod receptor gene family was implicated in controlling reproductive phenology.
Although GA20OX is a known controller of PH, its homolog was not identified by the GWAS as associated with PH. Plant height is an adaptive trait with large effect QTL conserved for maintaining phenotypic variation within in and between species of Poaceae74. However, GWASs for PH in predominantly self-fertilizing grass species have revealed variable effect sizes of the leading QTLs (QTL with highest PVE for a trait in a GWAS) between species, ranging from 1 to 23%75,76,77, demonstrating that the genetic architecture of plant height varies among self-fertilizing grass species. Our study indicates that B. tectorum uses at least one moderate to large effect locus to control PH as we only identified a single locus explaining 16.9% of the phenotypic variation. If mutations in B. tectorum are slightly deleterious then it would be expected that B. tectorum would need to use a small number of large effect QTL to facilitate local adaptation in the presence of divergent stabilizing selection78. Thus, the identification of the single moderate to large effect locus for PH may indicate the presence of deleterious mutations and divergent stabilizing selection between locales in B. tectorum, although further experiments would be needed to validate the selection and deleterious mutations imposed on B. tectorum. The homologs of XDH and YUC6 as candidate genes reflected the involvement of stress response and hormonal regulator genes controlling PH as found in other phylogenetically proximal grasses, such as wheat28, barley29, and oats30.
Our study indicates B. tectorum is armed with a complex array of genetic mechanisms to create adaptive variation underlying reproductive phenology and PH which has facilitated its invasion into N. America and suggests that it is likely to continue to spread north into western Canada as climate change facilitates range expansion79. Identifying candidate FHY3, FRS5, FRS6, FRS7, ABCB19, UVR8, and PHL genes that all act in response to light stimulus indicate that photoreceptor genes are critical for controlling variation in flowering time in B. tectorum. Photoreceptors controlling reproductive phenology will result in phenotypic plasticity because light signals of the environment will influence the underlying pathways80. Genetic control of traits was quite high, ranging from 0.99 to 0.94 for VPN and tiller number, respectively.
Furthermore, our results indicated the presence of small to moderate effect QTL controlling reproductive phenology traits, rather than singular large effect loci. The high heritability and moderate effect QTL detected for adaptive traits indicate that B. tectorum has already been adapting and will continue to adapt to a wide range of environments utilizing moderate-sized QTL—as genetic recombination is very rare. Further investigation into the plasticity and regulation of flowering time in B. tectorum is needed to understand how local populations respond to stress or climate variation, and how the genetic variation we have discovered confers success in the arid, dry southern reaches of the American southwest and northern Mexico north to western Canada.
Methods
Plant Material and DNA extraction for whole genome sequencing
For whole genome assembly, a single plant from the B. tectorum accession (FMH10) was grown hydroponically in an isolated, disease-free growth chamber under a 12-h photoperiod. Growing temperatures ranged from 18 °C (night) to 20 °C (day). The hydroponic growth solution was based on MaxiBloom® Hydroponics Plant Food (General Hydroponics, Sevastopol, CA, United States) at a concentration of 1.7 g/L. FMH10 is a common clade accession11. In preparation for PacBio CLR sequencing, high molecular weight DNA was extracted from 72-h dark-treated leaf samples using a CTAB-Qiagen Genomic-tip protocol as described by Vaillancourt and Buell81.
Whole genome sequencing
For whole-genome sequencing, large-insert SMRTBell libraries (>20 kb), selected using a SageElf (Sage Science, Inc., Beverly, MA, USA), were prepared according to standard manufacture protocols and sequenced at the BYU DNA Sequencing Center (Provo, UT, USA) using P6-C4 chemistry on a Sequel II instrument (Pacific BioSciences, Menlo Park, CA, USA). For whole genome polishing, DNA was sent to for Illumina HiSeq (2 × 150 bp) sequencing from standard 500‐bp insert libraries. Trimmomatic v0.3582 was used to remove adapter sequences and leading and trailing bases with a quality score < 20 or with an average per-base quality of 20 over a four-nucleotide sliding window. After trimming, any reads shorter than 75 nucleotides in length were removed. Raw PacBio and Illumina reads have been deposited in GenBank.
Genome assembly, polishing, and Hi-C scaffolding
A primary assembly of B. tectorum accession FMH10 was constructed using Canu v1.983 with default parameters (corMhapSensitivity = normal and corOutCoverage = 40). The primary assembly was polished twice with Illumina short reads using Arrow from the GenomicConsensus package in the Pacific BioSciences SMRT portal v5.1.0 followed by a single round of insertion/deletion correction using PILON v0.2284. The average read depth for the genome assembly was 68.6. Fresh leaf tissue from a single 3-week-old FMH10 plant was sent to Dovetail Genomics LLC (Santa Cruz, CA, USA) in preparation for construction of an Omni-C™ proximity-guided final chromosome-scale assembly. The Omni-C™ technology uses an approach to Hi-C library preparation via DNA digestion with a non-specific endonuclease to increase uniformity and genomic coverage (https://dovetailgenomics.com/omni-c/). The libraries were prepared using a standard Illumina library prep followed by sequencing on an Illumina HiSeq X in rapid run mode. The HiRiSE™ scaffolder and the Omni-C™ library-based read pairs were used to produce a likelihood model for genomic distance between read pairs, which was used to break putative miss-joins and to identify and make prospective joins in primary contig assembly to produce the final chromosome scale reference assembly.
GWAS plant materials collection
Bromus tectorum genotypes were obtained from Genome Resources Information Network (GRIN), field samples in eastern Washington collected by Jon Witkop and Amber Hauvermale in 2015 as part of the “Regional Approaches to Climate Change for Pacific Northwest Agriculture” a multi-disciplinary project that aims to mitigate the impacts climate change has on agriculture85, and samples from natural areas in Montana (contributed by Lisa Rew, Montana State University) with 11, 64, and 46 samples, respectively. Each genotype was grown for one generation in the greenhouse to increase seed and verify purity. Six replicates of each line were vernalized at 4 °C with 10 h light per day for 53 d, then planted in 1.4-liter square pots. Supplemental lighting was used to keep day lengths at least 15 h per day.
Phenotyping
Plants in the greenhouse were observed daily to record reproductive phenology-associated phenotypes. The phenotypes measured included days until first panicle visible (VPN, Feekes 10.1), days to first joint (J1, Feekes 6), days until first mature seed (FRS), days until 50% of seeds dry with awns angled outward (AWN50), number of tillers, and height of the tallest panicle (PH). Number of tillers were counted for each plant at the end of the experiment before harvest. The height of the tallest panicle was measured in centimeters, measuring from the base of the plant to the tip of the longest panicle.
GWAS DNA extraction and resequencing
DNA was extracted using a bromide CTAB protocol as previously described86. Samples were diluted to a 50 ng/µl concentration. Genotyping by sequencing libraries were prepared for each sample by LGC Genomics (Berlin, Germany) following Elshire et al.87 using the Msl1 restriction enzyme. Barcode adapters were ligated to each sample and the samples were put into 48-plex library plates. The polymerase chain reaction was used to amplify samples on the plates which were then sequenced using a single lane of Illumina NextSeq 500 V2. Approximately 1.5 million (2 × 150 bp) reads were generated per sample.
After sequencing, all the library groups were de-multiplexed with bcl2fastq v2.17.1.14 (https://support.illumina.com/sequencing/sequencing_software/bcl2fastq-conversion-software.html) software allowing for up to two mismatches on the barcodes. Library groups were de-multiplexed further into separate samples according to the inline barcodes, where no mismatches were allowed. The adapter barcodes were clipped and reads <20 bases in length were discarded as were any reads where the 5′ end did not match the restriction enzyme cutting motif. Reads were quality trimmed from the 3′ end so that the average Phred quality score across ten neighboring bases >20.
GWAS SNP calling
De-multiplexed filtered reads for each sample were aligned to the de novo reference genome using BWA mem88 with default settings for paired end reads. SAM files generated from the alignments were converted to BAM files and sorted using SAMTOOLS89. The mpileup and call functions from Bcftools90 were used to call SNPs. The vcf file generated from Bcftools was filtered so only SNP variants were kept, minor allele frequency (MAF) > 0.05, Missing Alleles <25% and a QUAL score of at least 30 for each SNP, using Bcftools89. Although the vcf file was not explicitly filtered for read depth, read depths were adequate with a minimum read depth of 54 and a median read depth of 808. Scripts in R statistical programming language were used to read the vcf file into an allelic dosage table and filter out markers with more than two alleles or more than 5 heterozygous calls (B. tectorum is an autogamous species). Missing calls were imputed with a kth nearest neighbor imputation using the “impute” package91 from the Bioconductor project92 in the R statistical programming language93.
Linkage-disequilibrium
Linkage-disequilibrium (LD) was calculated on a pairwise basis between SNP on the same chromosome within 3 Mb using 1500 randomly sampled SNP from each chromosome. LD, measured as r2sv, was calculated using the method described by Mangin et al.94, that calculates the Pearson correlation between SNP but corrects for population structure and kinship in LD calculations implemented in the “LDcorSV” R package95. A genomic kinship matrix was estimated from the SNP data using the method from VanRaden96 implemented in GAPIT97. Landscape and Ecological Analysis (LEA)98 was used in R to determine the optimal number of ancestral groups, and then calculate the admixture of each of these ancestral groups. Values of K, ranging from 1 to 20, were evaluated using the snmf function in LEA for cross entropy in ten replications and the lowest K near the lowest cross entropy was selected as the optimal K. The kinship matrix and population components from snmf described above were used with LDcorSV to correct for population structure, with distances above 3 Mb being discarded. The data from all the chromosomes was pooled together after filtering SNP by distance. The nlrq function of the quantreg R package99 was used estimate an asymptotic decay by physical genetic distance for the 90th percent quantile of LD values. The LD decay was defined as the distance required for the LD (90th percent quantile of corrected r2) to drop from its initial starting point to half-way between the starting point and the lower asymptotic limit (LD90,1/2). LD90,1/2 as a measure was found by simulation to be a more accurate estimate then r2 = 0.1 as a measure of LD decay100.
GWAS analysis
GWAS was performed using the Bayesian-information and Linkage-disequilibrium Iteratively Nested Keyway (BLINK) algorithm53 with the BLUEs for each phenotype as the response variables and the numeric SNP matrix as the genetic data. BLINK was ran using the GAPIT package97 in R. Three principal components were chosen to be included as a covariate in the BLINK which represented the smallest number of principal components that controlled inflation on the P-P diagnostic plots generated by GAPIT. Potential candidate genes within a 1 Mb window, based on LD (Fig. S2) for each of the genomic regions identified in the GWAS analysis were manually identified from the previously described annotated gene set with functional annotations indicating association with maturity traits.
Transcriptome assembly and genome annotation
RNA-Seq data (2 × 150 bp Illumina reads), derived from a bulk tissue sample consisting of 7-d old seedlings and leaf, roots, and stems from hydroponically grown B. tectorum (FMH10) plants, was trimmed using Trimmomatic72 and aligned to Omni-C reference assembly using HiSat2 v2.0.4101 with default parameters and max intron length set to 50,000 bp. The resulting SAM file was sorted and indexed using SAMtools v1.689 and assembled into putative transcripts using StringTie v1.3.4102. The quality of the assembled transcriptome was assessed relative to completeness using BLAST comparisons to the reference Brachypodium distachyon L. (ftp://ftp.ensemblgenomes.org/pub/plants/release-37/fasta/brachypodium_distachyon/pep/).
Prior to annotation with MAKER2 v2.31.10103, RepeatModeler v1.0.11104 and RepeatMasker v4.0.7105 were used to identify repetitive elements in the final reference assembly, relative to RepBase libraries v20181026; www.girinst.org. Transcriptome evidence for the annotation included the de novo transcriptome for B. tectorum as well as the cDNA models from Brachypodium distachyon (v 1.0; Ensembl genomes). Protein evidence included the uniprot-sprot database (downloaded September 25, 2018) as well as the peptide models from B. distachyon (v 1.0; Ensembl genomes). Repeats within the reference assembly were masked based on the species-specific sequences produced by RepeatModeler. For ab initio gene prediction, B. tectorum-specific AUGUSTUS gene prediction models were provided to MAKER as well as rice (Oryza sativa L.)-based SNAP models. Benchmarking Universal Single-Copy Orthologs (BUSCO) v3.0.2106 was used to assess the completeness of the final assembly using the Embryophyta odb10 dataset, with the “–long” argument.
Syntenny analysis was performed using McScanX27 with results of a pBlast with default settings between the predicted genes of our OMNI-C B. tectorum and the barley reference genome. Gene density was calculated for each 1 Mb window of the B. tectorum reference genome. Telomere density was also calculated for 1 Mb windows using the Blast results between the nucleotide sequence of the B. tectorum reference genome and “TTAGGG” the telomeric repeat of most plant species107 repeated four times. Custom Python code was used to create simplified GFF files to run McScanX and format the results for creating a circle plot. Circos108 was used to create a circle plot using the results from the protein blast between barley and B. tectorum.
Statistics and reproducibility
Three replicates, or blocks, were placed in one of two greenhouses with the same N = 121 genotypes in each. The plants were arranged in a completely randomized block design, with each block grown on a separate greenhouse bench. linear mixed effect (LME) and generalized linear mixed effect (GLME) models were fit using the “lme4” R package109 using lmer and glmer functions, respectively. LMEs were used for traits that are continuous measurements, GLMEs with a sqrt link function were used for count data. The linear model for the LME is:
Genotype ∼ N (0, σ2g)
Block within Greenhouse ~ N (0, σ2β(E))
Greenhouse ~ N (0, σ2E)
Genotype × Greenhouse ~ N (0, σ2 (gE))
Error ~ N (0, σ2ε)
Where yijk the phenotype, µ is the mean, gi is a random effect due to genotype, β(E)jk is a random effect of the block within greenhouse, Ek is the random effect of greenhouse, and ∈ijk is the error term. Broad Sense Heritability for each trait was estimated according to Cullis et al.110.
where \(\bar{v}Blup\) is the mean variance of a difference of two blups and \({{{{{{\rm{\sigma }}}}}}}_{g}^{2}\) is the variation of the random effect for genotype. For count data where a generalized mixed linear model was used heritability was calculated in the latent (transformed) distribution. BLUEs were calculated for the GWAS using the model for heritability modified to have genotype set as a fixed effect instead of a random effect, and a log link function instead of square root link function.
The statistical testing for the GWAS analyses and ad-hoc analyses for the GWAS all used BLUEs generated from [1] with N = 121 genotypes. Interactions were tested for when protein products of two genes near two QTL are known to interact through previous studies are found near significant (q < 0.05) QTL for the same trait. The following models for each trait where QTL associated with the trait were fit using the lme4qtl111 R package. For kinship matrix required we used the “Van-Raden” method implemented in GAPIT R package on our full SNP matrix to generate a kinship matrix. Models were solved using restricted maximum likelihood (REML) for calculating percent variation explained by SNPs while maximum likelihood (ML) was used to solve the models when the models were used for likelihood ratio test of an interaction112.
where ynx1 is a vector of the standardized BLUEs, Xnx3 is a matrix of fixed effects which in our case is the first three principal components of the full SNP matrix with the corresponding vector of coefficients b3x1. Mnxp is a numeric SNP matrix comprising of the significant (q < 0.05) SNP found by BLINK in the GWAS, vpxn is the vector of corresponding fixed-effect coefficients for the SNP found by Blink. (mi⊙mj) is a nx1 vector is the product of component wise vector multiplication of the SNPs being tested for an interaction between SNPi and SNPj with vij a scalar as its fixed effect coefficient. Znxn is an incidence matrix representing the genetic relationship between genotypes and unx1 is a vector of random polygenic effects that follows the distribution \(N(0,{\sigma }_{g}^{2}{A}_{nxn})\)where Anxn is the additive relationship matrix. enx1 is a vector of residual error that follows the distribution \(N(0,{\sigma }_{e}^{2}{I}_{nxn})\).
The likelihood ratio tests were implemented using the anova function in r on the models fit for [3] and [4]. Because only one interaction was tested at a time the [3] was nested in the more general model [4]. The anova function was used with both fit models are the input. The anova function calculates the log-likelihood of each model to calculate the likelihood ratio that is tested using a chi-sq test with one degree of freedom ([4] has one more parameter then [3]). Significant results (p < 0.05) indicate the more general model (model with SNP interaction [2]) is more likely to be explained by the data thus there is likely an interaction between those SNP.
Percent variation explained by QTL was calculated for each QTL identified by BLINK for each trait using the following formula for each QTL in model [1]. For each trait the markers were fit to a model simultaneously to estimate coefficients.
where \({h}_{qtl}^{2}\)is the percent of phenotypic variation explained by a given QTL, \({\sigma }_{y}^{2}\) is total phenotypic variation, \({\sigma }_{qtl}^{2}\) is the phenotypic variation explained by the QTL, \(\hat{B}\) is the estimated effect of a marker (vi) from equation [3], P is the allele frequency which follows a binomial distribution with 2 trials and probability p, f is the minor allele frequency (1-p). By standardizing the BLUEs, \({\sigma }_{y}^{2}\)is set to 1 thus simplifying the equation to:
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The raw sequences used for the B. tectorum genome assembly are deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive database under the BioProject PRJNA728981 with the following accession numbers: SRR14498212–SRR14498217 (PacBio reads), SRR14578284–SRR14578290 (Hi-C reads), SRR14578282 (Transcriptome) and SRR14498209–SRR14498211 (Polishing short reads). The raw reads for the resequencing panel are found in BioProject PRJNA728981 with the following NCBI accession numbers: SRR15308470–SRR15308851 (resequencing panel). Genome browsing and bulk data downloads, including annotations and BLAST analysis of the final proximity-guided assemblies are available at CoGe (https://genomevolution.org/coge/) with genome ID: id64356. Source data pertaining to general information on the genotypes is available in Supplementary Data 1 and source data of the BLUEs used to produce Fig. 2 and perform GWAS analyses is available in Supplementary Data 2. Genotype files (VCF, numeric SNP matrix), GWAS summary statistics are all freely available for download in a figshare repository associated with this manuscript. (https://doi.org/10.6084/m9.figshare.c.6419786.v1).
Materials availability
All the genotypes that were phenotyped and genotyped in this manuscript are available upon request.
Code availability
The custom scripts in R and Python for the phenotypic analysis, automating bioinformatics, and performing GWAS analyses are freely available without restriction on the figshare repository associated with this manuscript (https://doi.org/10.6084/m9.figshare.c.6419786.v1).
References
Bradley, B. A. et al. Cheatgrass (Bromus tectorum) distribution in the intermountain western United States and its relationship to fire frequency, seasonality, and ignitions. Biol. Invasions 20, 1493–1506 (2018).
Balch, J. K., Bradley, B. A., D’Antonio, C. M. & Gomez-Dans, J. Introduced annual grass increases regional fire activity across the arid western USA (1980–2009). Glob. Change Biol. 19, 173–183 (2012).
Pilliod, D. S., Welty, J. L. & Arkle, R. S. Refining the cheatgrass–fire cycle in the Great Basin: precipitation timing and fine fuel composition predict wildfire trends. Ecol. Evol. 7, 8126–8151 (2017).
Zimmer, S. N., Grosklos, G. J., Belmont, P. & Adler, P. B. Agreement and uncertainty among climate change impact models: a synthesis of sagebrush steppe vegetation projections. Rangel. Ecol. Manag. 75, 119–129 (2021).
Blackshaw, R. E. Downy brome (Bromus tectorum) density and relative time of emergence affects interference in winter wheat (Triticum aestivum). Weed Sci. 41, 551–556 (1993).
Novak, S. J. & Mack, R. N. Genetic variation in Bromus tectorum (Poacea): comparison between native and introduced populations. Heredity 71, 167–176 (1993).
Mack, R. N. Invasion of Bromus tectorum L. into western North America: an ecological chronicle. Agro-Ecosyst. 7, 145–165 (1981).
Merrill, K. R., Meyer, S. E. & Coleman, C. E. Population genetic analysis of Bromus tectorum (Poaceae) indicates recent range expansion may be facilitated by specialist genotypes. Am. J. Bot. 99, 529–537 (2012).
Wolkovich, E. M. & Cleland, E. E. The phenology of plant invasions: a community ecology perspective. Front. Ecol. Environ. 9, 287–294 (2010).
Arnesen, S., Coleman, C. E. & Meyer, S. E. Population genetic structure of Bromus tectorum in the mountains of western North America. Am. J. Bot. 104, 879–890 (2017).
Merrill, K. R., Coleman, C. E., Meyer, S. E., Leger, A. L. & Collins, K. A. Development of single-nucleotide polymorphism markers for Bromus tectorum (Poaceae) from a partially sequenced transcriptome. Appl. Plant Sci. 4, 1600068 (2016).
Meyer, S. E., Leger, E. A., Eldon, D. R. & Coleman, C. E. Strong genetic differentiation in the invasive annual grass Bromus tectorum across the Mojave– Great Basin ecological transition zone. Biol. Invasions 18, 1611–1628 (2016).
Lawrence, N. C., Hauvermale, A. L. & Burke, I. C. Downy brome (Bromus tectorum) vernalization: variation and genetic controls. Weed Sci. 66, 310–316 (2018).
Mack, R. N. & Pyke, D. A. The demography of Bromus tectorum: variation in time and space. J. Ecol. 71, 69–93 (1983).
Rice, K. J. & Mack, R. N. Ecological genetics of Bromus tectorum. I. A hierarchical analysis of phenotypic variation. Oecologia 88, 77–83 (1991).
Meyer, S. E. Ecological genetics of seed germination regulation in Bromus tectorum L. I. phenotypic variance among and within populations. Oecologia 120, 27–34 (1999).
Meyer, S. E., Nelson, D. L. & Carlson, S. L. Ecological genetics of vernalization response in Bromus tectorum L. (Poaceae). Ann. Bot. 2004 93, 653–663 (2003).
Fernández-Calleja, M., Casas, A. M. & Igartua, E. Major flowering time genes of barley: allelic diversity, effects, and comparison with wheat. Theor. Appl. Genet. 134, 1867–1897 (2021).
Mathews, S., Tsai, R. C. & Kellogg, E. A. Phylogenetic structure in the grass family (Poaceae): evidence from the nuclear gene phytochrome b. Am. J. Bot. 87, 96–107 (2000).
Fortune, P. M., Pourtau, N., Viron, N. & Ainouche, M. L. Molecular phylogeny and reticulate origins of the polyploid Bromus species from Section Genea (Poaceae). Am. J. Bot. 95, 454–464 (2008).
Pais, A. L., Whetten, R. W. & Xiang, Q. Y. Population structure, landscape genomics, and genetic signatures of adaptation to exotic disease pressure in Cornus florida L.— Insights from GWAS and GBS data. J. Syst. Evol. 58, 546–570 (2020).
Kosch, T. A. et al. Genetic potential for disease resistance in critically endangered amphibians decimated by chytridiomycosis. Anim. Conserv. 22, 238–250 (2019).
Nichols, K. M., Kozfkay, C. C. & Narum, S. R. Genomic signatures among Oncorhynchus nerka ecotypes to inform conservation and management of endangered Sockeye Salmon. Evol. Appl. 9, 1285–1300 (2016).
Wright, B. R. et al. A demonstration of conservation genomics for threatened species management. Mol. Ecol. Resour. 20, 1526–1541 (2020).
Duntsch, L. et al. Polygenic basis for adaptive morphological variation in a threatened Aotearoa New Zealand bird, the hihi (Notiomystis cincta). Proc. R. Soc. Lond. B Biol. Sci. 287, 20200948 (2020).
Hsiao, C., Chatterton, N. J., Asay, K. H. & Jensen, K. B. Molecular phylogeny of the Pooideae (Poaceae) based on nuclear RDNA (ITS) sequences. Theor. Appl. Genet. 90, 389–398 (1995).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Han, R. et al. Enhancing xanthine dehydrogenase activity is an effective way to delay leaf senescence and increase rice yield. Rice 13, 16 (2020).
Kim, J. I. et al. Overexpression of Arabidopsis YUCCA6 in potato results in high- auxin developmental phenotypes and enhanced resistance to water deficit. Mol. Plant 6, 337–349 (2013).
Rohilla, M. et al. Genome-wide association studies using 50 K rice genic SNP chip unveil genetic architecture for anaerobic germination of deep-water rice population of Assam, India. Mol. Genet. Genome 295, 1211–1226 (2020).
Brookbank, B. P., Patel, J., Gazzarrini, S. & Nambara, E. Role of basal ABA in plant growth and development. Genes 12, 1936 (2021).
Kim, J. I. et al. yucca6, a dominant mutation in Arabidopsis, affects auxin accumulation and auxin-related phenotypes. Plant Physiol. 45, 722–735 (2007).
Jia, P., Bayaerta, T., Li, X. & Du, G. Relationships between flowering phenology and functional traits in eastern Tibet alpine meadow. Arct. Antarct. Alp. Res. 43, 585–592 (2011).
Bolmgren, K. & Cowan, P. D. Time - size tradeoffs: a phylogenetic comparative study of flowering time, plant height and seed mass in a north-temperate flora. Oikos 117, 424–429 (2008).
Pham, A. T. et al. Identification of wild barley derived alleles associated with plant development in an Australian environment. Euphytica 216, 148 (2020).
Hazard, L., Betin, M. & Molinari, N. Correlated response in plant height and heading date to selection in perennial ryegrass populations. J. Agron. 98, 1384–1391 (2006).
Brown, R. F. Tiller development as a possible factor in the survival of the two grasses. Aristida armata Thyridolepis mitchelliana. Rangel. 4, 34–38 (1982).
Williamson, M. A. et al. Fire, livestock grazing, topography, and precipitation affect occurrence and prevalence of cheatgrass (Bromus tectorum) in the central Great Basin, USA. Biol. Invasions 22, 663–680 (2020).
Manolio, T. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Barson, N. et al. Sex-dependent dominance at a single locus maintains variation in age at maturity in salmon. Nature 528, 405–408 (2015).
Sinclair-Waters, M. et al. Beyond large-effect loci: large-scale GWAS reveals a mixed large-effect and polygenic architecture for age at maturity of Atlantic salmon. Genet. Sel. Evol. 52, 9 (2020).
Totland, O. Effects of temperature and date of snowmelt on growth, reproduction, and flowering phenology in the arctic/alpine herb, Ranunculus glacialis. Oecologia 133, 168–175 (2002).
Hall, M. C. & Willis, J. H. Divergent selection on flowering time contributes to local adaptation in mimulus guttatus populations. Evolution 60, 2466–2477 (2007).
Ågren, J. & Schemske, D. W. Reciprocal transplants demonstrate strong adaptive differentiation of the model organism Arabidopsis thaliana in its native range. N. Phytol. 194, 1112–1122 (2012).
Salomé, P. A. et al. Genetic architecture of flowering-time variation in Arabidopsis thaliana. Genetics 188, 421–433 (2011).
Xu, Y. et al. Quantitative trait locus mapping and identification of candidate genes controlling flowering time in Brassica napus L. Front. Plant Sci. 11, 626205 (2021).
Buckler, E. S. et al. The genetic architecture of maize flowering time. Science 325, 5941 (2009).
Hill, B. H. et al. Hybridisation-based target enrichment of phenology genes to dissect the genetic basis of yield and adaptation in barley. Plant Biotechnol. J. 17, 932–944 (2018).
Lowry, D. B. et al. The genetics of divergence and reproductive isolation between ecotypes of Panicum hallii. N. Phytol. 205, 402–414 (2014).
Leinonen, P. H., Salmela, M. J., Greenham, K., McClung, C. R. & Willis, J. H. populations are differentiated in biological rhythms without explicit elevational clines in the plant Mimulus laciniatus. J. Biol. Rhythms 35, 452–464 (2020).
Friedman, J. & Willis, J. H. Major QTLs for critical photoperiod and vernalization underlie extensive variation in flowering in the Mimulus guttatus species complex. N. Phytol. 199, 571–583 (2013).
Rice, B. R., Fernandes, S. B. & Lipka, A. E. Multi-trait genome-wide association studies reveal loci associated with maize inflorescence and leaf architecture. Plant Cell Physiol. 61, 1427–1437 (2020).
Huang, M., Liu, X., Zhou, Y., Summers, R. M. & Zhang, Z. BLINK: A Package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience 8, giy154 (2019).
Sun, X. et al. The Oryza sativa regulator HDR1 associates with the kinase OsK4 to control photoperiodic flowering. PLoS Genet. 12, e1005927 (2016).
Dieterle, M. et al. Molecular and functional characterization of Arabidopsis Cullin 3A. Plant J. 41, 386–399 (2005).
Rieu, I. et al. The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J. 53, 488–504 (2008).
Zhao, H. et al. The ATP-binding cassette transporter ABCB19 regulates postembryonic organ separation in Arabidopsis. PLoS ONE 8, e60809 (2013).
Lin, R. & Wang, H. Arabidopsis FHY3/FAR1 Gene family and distinct roles of its members in light control of Arabidopsis development. Plant Physiol. 136, 4010–4022 (2004).
McKown, A. D. et al. Geographical and environmental gradients shape phenotypic trait variation and genetic structure in Populus trichocarpa. N. Phytol. 201, 1263–1276 (2014).
Magome, H., Yamaguchi, S., Hanada, A., Kamiya, Y. & Oda, K. dwarf and delayed- flowering 1, a novel Arabidopsis mutant deficient in gibberellin biosynthesis because of overexpression of a putative AP2 transcription factor. Plant J. 37, 720–729 (2004).
Endo, M., Tanigawa, Y., Murakami, T., Araki, T. & Nagatani, A. PHYTOCHROME- DEPENDENT LATE-FLOWERING accelerates flowering through physical interactions with phytochrome B and CONSTANS. Proc. Natl Acad. Sci. USA 110, 18017–18022 (2013).
Škiljaica, A. et al. The protein turnover of Arabidopsis BPM1 is involved in regulation of flowering time and abiotic stress response. Plant. Mol. Biol. 102, 359–372 (2020).
Dotto, M., Gómez, M. S., Soto, M. S. & Casati, P. UV-B radiation delays flowering time through changes in the PRC2 complex activity and miR156 levels in Arabidopsis thaliana. Plant Cell Environ. 41, 1394–1406 (2018).
He, Y., Doyle, M. R. & Amasino, R. M. PAF1-complex-mediated histone methylation of FLOWERING LOCUS C chromatin is required for the vernalization-responsive, winter-annual habit in Arabidopsis. Genes Dev. 18, 2774–2784 (2004).
Finnegan, E. J., Sheldon, C. C., Jardinaud, F., Peacock, W. J. & Dennis, E. S. A cluster of Arabidopsis genes with a coordinate response to an environmental stimulus. Curr. Biol. 14, 911–916 (2004).
Ritter, A. et al. The transcriptional repressor complex FRS7-FRS12 regulates flowering time and growth in Arabidopsis. Nat. Commun. 8, 15235 (2017).
Xie, Y. et al. FHY3 and FAR1 Integrate light signals with the miR156-SPL module- mediated aging pathway to regulate Arabidopsis flowering. Mole. Plant 13, 483–498 (2020).
Werner, T. et al. Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15, 2532–2550 (2003).
D’Aloai, M. et al. Cytokinin promotes flowering of Arabidopsis via transcriptional activation of the FT paralogue. Tsf. Plant J. 65, 972–979 (2011).
Wiegmann, M. et al. Barley yield formation under abiotic stress depends on the interplay between flowering time genes and environmental cues. Sci. Rep. 9, 6397 (2019).
Yan, H. et al. Position validation of the dwarfing gene Dw6 in oat (Avena sativa L.) and its correlated effects on agronomic traits. Front. Plant Sci. 12, 668847 (2021).
Nadolska-Orczyk, A., Rajchel, I. K., Orczyk, W. & Gasparis, S. Major genes determining yield-related traits in wheat and barley. Theor. Appl. Genet. 130, 1081–1098 (2017).
Hedden, P. The genes of the Green Revolution. Trends Genet. 19, 5–9 (2003).
Lin, Y. R., Schertz, K. F. & Paterson, A. H. Comparative analysis of QTLs affecting plant height and maturity across the Poaceae, in reference to an interspecific sorghum population. Genetics 141, 391–411 (1995).
Tessmann, E. W. & Sanford, D. A. V. GWAS for fusarium head blight related traits in winter wheat (Triticum Aestivum L.) in an artificially warmed treatment. Agronomy 8, 68 (2018).
Biscarini, F. et al. Genome-wide association study for traits related to plant and grain morphology, and root architecture in temperate rice accessions. PLoS ONE 11, e0155425 (2016).
Pasam, R. K. et al. Genome-wide association studies for agronomical traits in a world wide spring barley collection. BMC Plant Biol. 12, 16 (2012).
Hodgins, K. A. & Yeaman, S. Mating system impacts the genetic architecture of adaptation to heterogeneous environments. N. Phytol. 224, 1201–1214 (2019).
DeBeer, C. M., Wheater, H. S., Carey, S. K. & Chun, K. P. Recent climatic, cryospheric, and hydrological changes over the interior of western Canada: a review and synthesis. Hydrol. Earth Syst. Sci. 20, 1573–1598 (2016).
Casal, J. J., Fankhauser, C., Coupland, G. & Blázquez, M. A. Signalling for developmental plasticity. Trends Plant Sci. 9, 309–314 (2004).
Vaillancourt, B. & Buell, C. R. High molecular weight DNA isolation method from diverse plant species for use with Oxford Nanopore sequencing. Preprint at https://www.biorxiv.org/content/10.1101/783159v2.full.pdf (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Eigenbrode, S. D., Binns, W. P. & Huggins, D. R. Confronting climate change challenges to dryland cereal production: a call for collaborative, transdisciplinary research, and producer engagement. Front. Ecol. Evol. 5, 164 (2018).
Porebski, S. L., Bailey, G. & Baum, B. R. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol Components. Plant Mol. Biol. Rep. 15, 8–15 (1997).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379 (2011).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA- MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Hastie, T., Tibshirani, R., Narasimhan, B. & Chu, G. Impute: imputation for microarray data. R package version 1.58.0. (2019).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/ (2019).
Mangin, B. et al. Novel measures of linkage disequilibrium that correct the bias due to population structure and relatedness. Heredity 108, 285–291 (2012).
Desrousseaux, D., Sandron, F., Siberchicot, A., Cierco‐Ayrolles, C. & Mangin, B. LDcorSV: linkage disequilibrium corrected by the structure and the relatedness. R package version 1.3.2 (2017).
VanRaden, P. M. Efficient methods to compute genomic predictions. J. Dairy Sci. 91, 4414–4423 (2008).
Lipka et al. Genome association and prediction integrated tool. Bioinformatics 28, 2397–2399 (2012).
Frichot, E. & Francois, O. LEA: an R package for landscape and ecological association studies. Methods Ecol. Evol. 6, 925–929 (2015).
Koenker, R. Qauntreg: quantile regression. R package version 5.83. (2021).
Vos, P. G. et al. Evaluation of LD decay and various LD-decay estimators in simulated and SNP-array data of tetraploid potato. Theor. Appl. Genet. 130, 123–135 (2017).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinform. 12, 491 (2011).
Smit, A. F. A. & Arian, F. A. RepeatModeler Open-1.0. (2008).
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker Open-4.0. (2015).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Cox, A. V. et al. Comparison of plant telomere locations using a PCR-generated synthetic probe. Ann. Bot. 72, 239–247 (1993).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Bates, D., Mächler, M., Bolker B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Cullis, B. R. & Smith, A. B. On the design of early generation variety trials with correlated data. J. Agric. Biol. Environ. Stat. 11, 381 (2006).
Ziyatdinov, A. et al. lme4qtl: linear mixed models with flexible covariance structure for genetic studies of related individuals. BMC Bioinform. 19, 68 (2018).
Revolinski, S., Coleman, C. E., Maughan, P. J. & Burke, I. C. Supplementary Code and Data: preadapted to adapt: underpinnings of adaptive plasticity revealed by the downy brome genome. figshare. Collect. https://doi.org/10.6084/m9.figshare.c.6419786.v1 (2023).
Acknowledgements
We would like to acknowledge Amber Hauvermale for providing sequence data on B. tectorum that was used to conceive the experiment but was not included in analyses present in the manuscript. We would also like acknowledge Lisa Rew for providing seed for the B. tectorum genotypes from Montana. This work was funded in part by the Washington Grain Commission and by U.S. Department of Agriculture National Institute of Food and Agriculture grant No. 2017-68002-26819, the Washington Grain Commission R. J. Cook Chair Endowment, and the USDA National Institute of Food and Agriculture, Hatch project 1017286.
Author information
Authors and Affiliations
Contributions
S.R.R. wrote the manuscript, designed experiments and analyzed data. P.J.M. assembled and annotated the reference genome, analyzed data and edited the manuscript. C.E.C. contributed to the assembly and annotation of the reference genome and edited the manuscript. I.C.B. contributed to writing the manuscript, designed experiments and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All relevant permissions were obtained for the collection of B. tectorum genotypes. The methods carried out in our manuscript were in accordance with the local, national, and international guidelines and regulations.
Peer review
Peer review information
Communications Biology thanks Anh Pham and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: David Favero. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Revolinski, S.R., Maughan, P.J., Coleman, C.E. et al. Preadapted to adapt: underpinnings of adaptive plasticity revealed by the downy brome genome. Commun Biol 6, 326 (2023). https://doi.org/10.1038/s42003-023-04620-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04620-9
This article is cited by
-
Trait plasticity: a key attribute in the invasion success of Ageratina adenophora in different forest types of Kumaun Himalaya, India
Environment, Development and Sustainability (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.