Abstract
Rhodococcus opacus PD630 has considerable potential as a platform for valorizing lignin due to its innate “biological funneling” pathways. However, the transcriptional regulation of the aromatic catabolic pathways and the mechanisms controlling aromatic catabolic operons in response to different aromatic mixtures are still underexplored. Here, we identified and studied the transcription factors for aromatic degradation using GFP-based sensors and comprehensive deletion analyses. Our results demonstrate that the funneling pathways for phenol, guaiacol, 4-hydroxybenzoate, and vanillate are controlled by transcriptional activators. The two different branches of the β-ketoadipate pathway, however, are controlled by transcriptional repressors. Additionally, promoter activity assays revealed that the substrate hierarchy in R. opacus may be ascribed to the transcriptional cross-regulation of the individual aromatic funneling pathways. These results provide clues to clarify the molecule-level mechanisms underlying the complex regulation of aromatic catabolism, which facilitates the development of R. opacus as a promising chassis for valorizing lignin.
Similar content being viewed by others
Introduction
The serious environmental problems caused by society’s dependence on fossil fuels, including climate crisis and ocean pollution, make it important to seek alternative methods to produce chemicals and fuels, particularly from renewable non-food biomass1. Lignocellulose, which consists of carbohydrate polymers (e.g., cellulose and hemicellulose) and aromatic polymers (e.g., lignin), represents the most abundant raw material for the potential production of next-generation renewable biofuels and chemicals2. To date, the major components of lignocellulose—cellulose, and hemicellulose—have been efficiently converted into various chemicals through biochemical routes3. Lignin, which is the second most abundant polymer on earth, holds promise as a renewable feedstock for the production of fuels and platform chemicals, due to its high carbon-to-oxygen ratio (above 2:1) and great energy density4,5,6,7. Moreover, recent research indicates that converting lignin to high-value fuels and chemicals would improve the overall competitiveness of biorefineries8,9,10,11,12. However, due to the structural heterogeneity of lignin, the depolymerization process typically results in diverse aromatic products, which are challenging to valorize4,13; consequently, lignin is still under-utilized and treated primarily as waste14. To date, the most predominant strategies used for valorization—including depolymerization and fragmentation—require extensive separation and purification procedures, which are commercially non-profitable due to low yields and the low quality of their final products.
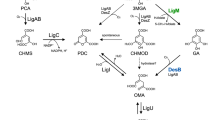
Beyond chemical processing, biological treatment is a promising choice for lignin valorization; particularly, bacterial systems are increasingly attracting attention due to their inherent “biological funneling” processes15,16. Diverse aromatic streams can be funneled into uniform compounds (catechol (CAT) and protocatechuic acid (PCA)) and then be degraded through the β-ketoadipate pathway17, a process that can potentially overcome the challenges associated with the heterogeneity of lignin breakdown products18. Rhodococcus opacus PD630 (hereafter, R. opacus), a Gram-positive soil bacterium that has the natural ability to tolerate and consume toxic aromatic compounds, has been considered a promising chassis for producing valuable products from aromatics19,20,21,22,23. Previous transcriptomic analysis of R. opacus cells grown on single aromatic compounds has identified the distinct funneling pathways for five lignin model compounds (benzoate, 4-hydroxybenzoate, phenol, vanillate, and guaiacol)18,24. Moreover, the roles of the two branches of the β-ketoadipate pathway in the degradation of those five compounds have been confirmed by gene knockout experiments18.
Constitutively maintaining this catabolic flexibility could impose a metabolic burden on the host microbe, but this handicap is typically overcome by arranging the genes of each degradative pathway as operons whose summed expression is controlled by specific regulators and inducers. Thus, the success of a particular catabolic pathway depends not only on the efficacy of the catabolic enzymes but also on the specific regulatory elements governing their expression25. As genomic, genetic, and biochemical data have been accumulated, various regulatory proteins that control the expression of the aromatic degradation pathways have been identified and classified into different protein families26. Of additional interest are the sensory mechanisms of the regulatory elements, because regulatory proteins and their cognate promoters have the potential to be developed into biosensors that can be used in many applications, including drug discovery, biomedicine, food safety, defense, security, and environmental monitoring27. In R. opacus, comparative transcriptomics has revealed that the five aromatic funneling pathways previously identified were significantly upregulated in response to a subset of the aromatic compounds tested, suggesting that the expression of those pathway genes is likely to be tightly controlled by specific regulatory mechanisms. Furthermore, via the application of whole genome sequencing and comparative genomics, we have identified different families of transcription factors (TFs) that are located adjacent to R. opacus’s proposed aromatic catabolic pathways18,24. However, the specific roles of these TFs in regulating the aromatic degradation pathways—such as the signals that trigger pathway expression and the exact mechanisms of activation and/or repression—are still unclear.
In natural environments, carbon sources are commonly found as heterogeneous mixtures. To handle these mixtures, including the portion of aromatic compounds that are toxic, most bacteria have evolved a hierarchy of substrate utilization that enables them to quickly adapt their intracellular metabolic network toward a preferred substrate, which is vital for competition in these environments. This phenomenon, termed carbon catabolite repression (CCR), has been extensively reported in the utilization of sugar mixtures and non-sugar substrates28,29,30, but the number of studies conducted on substrate combinations containing only aromatic compounds is limited. To date, most studies have focused on benzoate and 4-hydroxybenzoate, which are commonly metabolized via the two parallel branches of the β-ketoadipate pathway. For example, in the γ-proteobacteria Pseudomonas putida PRS2000 and Acinetobacter sp. Strain ADP1, benzoate has been found to be the preferred substrate of the two31,32. Similarly, in the β-proteobacterium Cupriavidus necator JMP134, the same utilization hierarchy between benzoate and 4-hydroxybenzoate has been observed33. In Rhodococcus sp. strain DK17, a catabolite repression-like response has been reported when cells are simultaneously provided with benzoate and phthalate34. Moreover, benzoate catabolite repression of phenol degradation has been observed in Acinetobacter calcoaceticus PHEA-235. Finally, in R. opacus, in a mixture of the five lignin model compounds previously described, benzoate was found to be consumed preferentially18, which suggests the existence of a substrate hierarchy. Although it is important for microbial lignin conversion strategies, this sequential consumption order is still underexplored.
In this study, to identify those TFs involved in regulating the degradation of lignin model compounds, we selected potential TFs in the genomic neighborhood of aromatic operons and knocked them out via homologous recombination. By using metabolite sensors derived from native R. opacus promoters that can detect aromatic compounds and by comparing the cell growth and aromatic consumptions of these TF deletion mutants to those of the wild type (WT), we evaluated the roles of the candidate TFs in regulating the degradation pathway of each lignin model compound. To establish the substrate hierarchy of the tested lignin model compounds, we performed time-course analyses of the consumption of individual aromatics in the mixture, revealing that these compounds were consumed in a distinct order. Moreover, by testing the responses of the funneling pathways in WT cells grown on both individual lignin model compounds and a mixture, we confirmed that R. opacus can differentially and specifically regulate the funneling pathways in response to specific compounds, which is critical for the utilization of the preferred aromatic substrate. Taken together, these results advance our understanding of the regulatory patterns of the aromatic degradation pathways, which is critical to constructing a more efficient bacterial chassis for comprehensively utilizing lignin. Moreover, these insights into the mechanism of hierarchical utilization of aromatics in R. opacus are of great significance for achieving rapid consumption of complex aromatic mixtures, enabling more cost-effective conversion of lignin into fuels, chemicals, and materials.
Results
Transcriptional regulation of the funneling pathways responsible for the degradation of aromatics via the CAT branch of the β-ketoadipate pathway
Previously, we found the ∆catB (cis,cis-muconate cycloisomerase knockout) strain was unable to grow using phenol, guaiacol, and benzoate as sole carbon sources, demonstrating that those three compounds are metabolized through the CAT branch of the β-ketoadipate pathway in R. opacus18. Additionally, by using RNA-Seq, several funneling pathway clusters, including those for phenol, guaiacol, and benzoate, were identified18. Because those genes were found to be upregulated by R. opacus in the presence of the relevant compounds, it was hypothesized that the induction of the funneling pathways is controlled by specific regulatory mechanisms. To identify the respective protein regulators, potential TFs, encoded by genes adjacent to the catabolic clusters, were selected (Supplementary Table 1).
For phenol degradation, a pair of similar phenol hydroxylase clusters (designated pheB1A1 and pheB2A2) were identified in the R. opacus genome, and these genes were significantly upregulated in the presence of phenol18,24. Furthermore, two AraC-type TFs, pheR1 (LPD06739) and pheR2 (LPD06574), which are located adjacent to these clusters but transcribed divergently (Fig. 1a, b), were selected for examination. To confirm the transcriptional regulation of these two clusters, transcriptional fusions of the respective promoters (PpheB1 and PpheB2) with the gfp+ reporter gene were constructed, and expressed in the WT strain (Fig. 1c, f). For PpheB1, the promoter activity was markedly increased with cultivation on phenol, but not on cis-cis muconate, 4-hydroxybenzoate, or PCA. In comparison, significant induction of PpheB2 was observed with cultivation on both phenol and cis-cis muconate (Fig. 1d, g and Supplementary Fig. 1).
The R. opacus PD630 genome contains two phenol degradation operons, depicted in a, b; the schematics include the promoters (small arrows) and proposed corresponding transcription factors (TFs), with operon genes indicated by LPD gene numbers corresponding to the NCBI database (Refseq, CP003949.1). For the full annotations of the pathway genes, see Supplementary Table 1. c, f To analyze the function of the proposed transcription factors pheR1 and pheR2 in regulating these operons, the transcriptional constructs PpheB1-GFP+ and PpheB2-GFP+ were expressed in WT and the corresponding TF deletion mutant strains. d, g The normalized fluorescence of the WT and ∆pheR1 and ∆pheR2 was measured with and without supplemental phenol (PHE) (****, P < 0.0001, unpaired two-tailed t test). e, h Comparisons of cell growth (OD) and phenol consumption (Con) between WT and the two TF deletion mutant strains ∆pheR1 and ∆pheR2 when fed 0.7 g/L PHE as the sole carbon source; cell growth and phenol consumption were significantly reduced in the ∆pheR2 strain (★, P < 0.05 for OD; ☆, P < 0.05 for Con; two-tailed mixed model ANOVA with Sidak’s multiple comparisons). i, j The transcriptional activity of the promoter PpheB1 in the mutant ∆pheR2 and the transcriptional activity of the promoter PpheB2 in the mutant ∆pheR1. Measurements were conducted with and without supplemental PHE (ns, not significant, unpaired two-tailed t test). k Comparison of the transcriptional activity of the promoter PpheB1 in the different knockout strains when treated with and without supplemental PHE (****, P < 0.0001, unpaired two-tailed t test). For all fluorescence assays, cell cultures contained 1 g/L glucose with (+) or without (−) 0.3 g/L PHE as the carbon source; all the fluorescence values were determined in the early stationary phase and normalized to an optical density at 600 nm (OD600). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
To elucidate the role of the two adjacent TFs in regulating the expression of these phenol funneling pathways, we generated two TF deletion mutants (∆pheR1 and ∆pheR2) and measured the promoter activity of PpheB1 and PpheB2, respectively, using the previously-described gfp+ constructs. In ∆pheR1, expression from promoter PpheB1 was severely inhibited but remained detectably ON (Fig. 1d). In ∆pheR2, however, the fluorescence output of PpheB2 was completely OFF (Fig. 1g). These results indicate that pheR1 and pheR2 act as activators of their respective phenol degradation clusters (Fig. 1c, f). Having established their roles, we analyzed cell growth and phenol consumption in the two mutants. When fed phenol as the sole carbon source, no cell growth or phenol consumption was observed in the ∆pheR2 strain; in contrast, cell growth and phenol consumption of ∆pheR1 strain were comparable to those of the WT strain (Fig. 1e, h). Cross-validation showed that knocking out either pheR1 or pheR2 had no effect on the induction of the non-cognate hydroxylase cluster (Fig. 1i, j). Additionally, we found that in a dual mutant (∆pheR1 ∆pheR2), the fluorescence output of the promoter PpheB1 was null, suggesting that this promoter may have crosstalk with TF pheR2 (Fig. 1k).
In R. opacus, although previous studies have shown that the genes encoding the phenol hydroxylase in these two clusters have a high identity to each other, and that this strain appears to utilize both copies of the two genes24, our results led us to hypothesize that only the pheB2A2 copy is essential for phenol degradation. To test this hypothesis, we generated a pheB1 deletion mutant. As expected, when fed phenol as the sole carbon source, the WT and ∆pheB1 strains showed statistically identical cell growth. Repeated attempts to disrupt the pheB2 failed, so a T7 RNAP-based CRISPRi platform36 was applied to test this putatively essential phenol degradation cluster (pheB2A2). In the strain expressing sgRNA PHE_1 (targeting pheB2), no significant increase in OD600 was observed under the induced condition, but growth was comparable to that of the control strain when uninduced (Supplementary Fig. 2). This result confirmed that pheB2A2 is the pivotal cluster for phenol degradation in R. opacus.
To identify the potential TF involved in regulating the guaiacol degradation pathway, a putative AraC-type TF guaR (LPD06577)—which is located adjacent to the deduced guaiacol funneling cluster—was selected as a knockout target (Fig. 2a)18. Specifically, the construct Pcyp255-GFP+ was expressed in both WT and ∆guaR strains (Fig. 2b). In the WT strain, the promoter activity was remarkably increased when cells were cultivated on guaiacol, but not on CAT, cis-cis muconate, 4-hydroxybenzoate, or PCA (Fig. 2c and Supplementary Fig. 1c). In the mutant ∆guaR, no expression from the promoter was observed with cultivation on guaiacol (Fig. 2c), indicating that guaR works as an activator with guaiacol as the inducer (Fig. 2b and Supplementary Fig. 1c). Furthermore, when the mutant was fed guaiacol as the sole carbon source, neither cell growth nor guaiacol consumption was detected (Fig. 2d), suggesting that guaR is the essential regulator for guaiacol degradation.
a, e The guaiacol (GUA) and benzoate (BEN) degradation operon with an annotated transcription factor (TF). Promoters are represented as small arrows. Genes are shown with LPD gene numbers from the NCBI database (Refseq, CP003949.1). For the full annotations of the pathway genes, see Supplementary Table 1. b The proposed model for the role of guaR. c, f To analyze the function of the proposed TF, the constructs Pcyp255-GFP+ and PbenA-GFP+ were expressed in WT and the corresponding TF deletion mutant strains; the normalized fluorescence of the WT and ∆guaR was measured with and without supplemental GUA (c) or BEN (f). d, g Comparisons of cell growth (OD) and aromatic consumption (Con) between the WT and ∆guaR strains when fed 0.6 g/L GUA (d) or 1.0 g/L BEN (g) as the sole carbon source. For all fluorescence assays, cell cultures contained 1 g/L glucose with (+) or without (−) 0.3 g/L GUA or 1.0 g/L BEN as the carbon source; all the fluorescence values were determined in the early stationary phase and normalized to OD600. Unpaired two-tailed t test was used to compare the variation in the change of fluorescence of the mutants treated with respective aromatic against that of the WT control (****, P < 0.0001); for cell growth and aromatic consumption, the variations in the changes of cell density (OD) and aromatic concentration (Con) of the mutant were compared against those of the WT control (★, P < 0.05 for OD; ☆, P < 0.05 for Con; two-tailed mixed model ANOVA with Sidak’s multiple comparisons). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
A putative funneling pathway for benzoate was proposed from the transcriptomic analysis18. By taking a close look at the genome architecture, we found that this cluster shares the AraC-type TF guaR (LPD06577) with the guaiacol funneling pathway (Fig. 2e). To analyze the potential role of guaR in regulating benzoate funneling pathway, the construct PbenA-GFP+ was expressed in both WT and ∆guaR strains, and the fluorescence output was measured in response to benzoate. The results showed that expression from this promoter decreased ~31% when knocking out guaR, but that the mutant maintained an identical benzoate consumption rate to that of the WT strain (Fig. 2f, g). These results indicate that guaR may not be the direct regulator of benzoate degradation.
Transcriptional regulation of the funneling pathways responsible for the degradation of aromatics via the PCA branch of the β-ketoadipate pathway
In addition to the CAT branch of the β-ketoadipate pathway, the PCA branch has been identified as playing a role in the degradation of aromatics, including 4-hydroxybenzoate and vanillate. Similar to phenol, guaiacol, and benzoate in the CAT branch, the funneling pathways for these two compounds have been proposed based on transcriptomic analysis18. For 4-hydroxybenzoate, a putative 4-hydroxybenzoate monooxygenase pobA (LPD06764) was identified, which is responsible for the conversion of 4-hydroxybenzoate to protocatechuate. Adjacent to this gene are two proposed TFs, IclR-type regulator hbaR1 (LPD06765) and Tet/AcrR-type regulator hbaR2 (LPD06763) (Fig. 3a, e). To study the roles of both TFs in regulating the 4-hydroxybenzoate degradation pathway, the construct PpobA-GFP+ was expressed in both WT and the TF deletion mutant strains (∆hbaR1 and ∆ hbaR2). In the WT strain, the promoter activity was markedly increased during cultivation on 4-hydroxybenzoate, but not on PCA or on CAT (Supplementary Fig. 1e). In the mutant ∆hbaR1, no transcription from the promoter was observed, but in ∆hbaR2, the fluorescence output of the promoter was identical to that of the WT strain (Fig. 3c, f). These results demonstrate that hbaR1 acts as an activator in regulating the 4-hydroxybenzoate funneling pathway with 4-hydroxybenzoate as the inducer (Fig. 3b). We also tested the two knockout mutants by feeding them with 4-hydroxybenzoate as the sole carbon source. No cell growth or 4-hydroxybenzoate consumption was observed in ∆hbaR1, but in ∆hbaR2, cell growth and 4-hydroxybenzoate consumption were comparable to those of the WT strain (Fig. 3d, g). These results indicate that hbaR1 is necessary for the regulation of 4-hydroxybenzoate funneling pathway in R. opacus.
a, e Two potential transcription factors, hbaR1 (LPD06765) and hbaR2 (LPD06763), are located near 4-hydroxybenzoate hydroxylase pobA (LPD06764). Promoters are represented as small arrows, and genes are shown with LPD gene numbers from the NCBI database (Refseq, CP003949.1). For the full annotations of the pathway genes, see Supplementary Table 1. b The proposed working model for hbaR1. c, f Analysis of the function of the proposed transcription factors. The construct PpobA-GFP+ was expressed in the WT and the corresponding TF deletion mutant strains. The normalized fluorescence of the WT and ∆hbaR1 and ∆hbaR2 was measured with and without supplemental 4-hydroxybenzoate (HBA). d, g Comparisons of cell growth (OD) and HBA consumption (Con) between WT and the two TF deletion mutant strains ∆hbaR1 and ∆hbaR2 when fed 1.0 g/L HBA as the sole carbon source. For all fluorescence assays, cell cultures contained 1 g/L glucose with (+) or without (−) 1.0 g/L HBA as the carbon source; all the fluorescence values were determined in the early stationary phase and normalized to OD600. Unpaired two-tailed t test was used to compare the variation in the change of fluorescence of the mutants treated with HBA against that of the WT control (****, P < 0.0001; ns, not significant). To assess cell growth and aromatic consumption, the variations in the changes of cell density (OD) and aromatic concentration (Con) of the mutants were compared against those of the WT control (★, P < 0.05 for OD; ☆, P < 0.05 for Con; two-tailed mixed model ANOVA with Sidak’s multiple comparisons). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
Similar to the 4-hydroxybenzoate funneling pathway, two putative TFs, IclR-type regulator vanR1 (LPD00562) and RrfR-type regulator vanR2 (LPD00561), are found adjacent to the vanillate funneling pathway (Fig. 4a, e). To elucidate the roles of these TFs in regulating the vanillate degradation pathway, the construct PvanA-GFP+ was expressed in both WT and the corresponding TF deletion mutants (Fig. 4b). In the WT strain, the promoter demonstrated a remarkably strong response to vanillate, but not to PCA or CAT (Supplementary Fig. 1f). In ∆vanR1, the fluorescence output of PvanA was completely OFF and no cell growth or vanillate consumption was observed when cells were fed with vanillate as the sole carbon source (Fig. 4c, d). Knocking out vanR2 had the unexpected effect of significantly decreasing the promoter activity of PvanA, as well as limiting VAN consumption (Fig. 4f, g). When examining the genome architecture more closely, we noted that these two TFs have a short overlap region in the C terminus. The phenotype of ∆vanR2, which was created with vanR1 intact, is characterized in part by the negative effect on the expression of vanR1, an effect which may be explained by the gene deletion causing unexpected context effects on neighboring genes36. Taken together, these findings suggest that vanR1 is a key activator that regulates the vanillate funneling pathway in R. opacus.
a, e The vanillate degradation operon in R. opacus is adjacent to two transcription factors, vanR1 (LPD00562) and vanR2 (LPD00561). Promoters are represented as small arrows, and genes are shown with LPD gene numbers from the NCBI database (Refseq, CP003949.1). For the full annotations of the pathway genes, see Supplementary Table 1. b The proposed working model for vanR1. c, f Analysis of the function of the proposed transcription factors. The construct PvanA-GFP+ was expressed in WT and in the corresponding TF deletion mutant strains. The normalized fluorescence of the WT, ∆vanR1, and ∆vanR2 was measured with and without supplemental vanillate (VAN). d, g Comparison of the cell growth (OD) and vanillate consumption (Con) between the WT and the two TF deletion mutant strains ∆vanR1 and ∆vanR2 when fed 0.6 g/L VAN as the sole carbon source. For all fluorescence assays, cell cultures contained 1 g/L glucose with (+) or without (−) 0.3 g/L VAN as the carbon source; all the fluorescence values were determined in the early stationary phase and normalized to OD600. Unpaired two-tailed t test was used to compare the variation in the change of fluorescence of the mutants treated with VAN against that of the WT control (**, P < 0.0021; ****, P < 0.0001). To assess cell growth and aromatic consumption, the variations in the changes of cell density (OD) and aromatic concentration (Con) of the mutants were compared against those of the WT control (★, P < 0.05 for OD; ☆, P < 0.05 for Con; two-tailed mixed model ANOVA with Sidak’s multiple comparisons). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
Transcriptional regulation of the β-ketoadipate pathway
The funneling pathways described above convert aromatic compounds into intermediate compounds, CAT, and protocatechuate for subsequent degradation by the β-ketoadipate pathway. Based on the previous transcriptomic analysis, three potential β-ketoadipate gene clusters are upregulated in WT when treated with aromatic compounds; furthermore, the fundamental roles of the CAT and PCA branches of the β-ketoadipate pathway have been confirmed by gene knockout experiments18. For the CAT branch, the critical genes and an IclR-type TF catR (LPD06569) are clustered but transcribed divergently (Fig. 5a). To analyze the transcriptional regulation of this cluster, we first localized the promoters PcatA and PcatR by comparative analysis37. Proposed transcription start sites (TSSs) were found 80 nt and 127 nt upstream of the translation start sites of catA and catR, respectively. Additionally, the −35 and -10-like elements of the promoter regions were found with appropriate spacing within the sequences upstream of the TSSs (Fig. 5g). Moreover, for the promoter PcatR, the supposed -35 and -10 elements overlap with the putative IclR-type regulator binding site, indicating a self-regulating mechanism for the expression of this TF.
a A catechol (CAT) degradation operon with an annotated transcriptional factor (TF). Promoters are represented as small arrows. Genes are shown with LPD gene numbers from the NCBI database (Refseq, CP003949.1). For the full annotations of the pathway genes, see Supplementary Table 1. b, c The proposed models for the role of catR. d, e To analyze the function of catR in regulating the CAT degradation cluster, the fusion construct PcatA-GFP+ was expressed in both the WT and the TF deletion mutant strains. Normalized fluorescence of the WT and ∆catR strains is shown in response to CAT and cis-cis muconate (CCMA). f The fusion construct PcatR-GFP+ was used to analyze the self-regulation pattern of catR. Normalized fluorescence of the WT and ∆catR strains is shown in response to CCMA. All cultures contained 1 g/L glucose with (+) or without (−) the respective aromatic carbon source (CAT 0.3 g/L and CCMA 5 g/L). All the fluorescence values were determined in the early stationary phase and normalized to OD600. Unpaired two-tailed t test was used to compare the variations in the changes of fluorescence of the strains under different conditions (****, P < 0.0001). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean. g The potential regulatory sequences from the intergenic region between catR and catA. All the regulatory features and sequence elements were identified by comparison with the position identified in Rhodococcus erythropolis37. Transcriptional start sites (TSS, +1) are underlined, and transcriptional initiation is indicated by bent arrows. The proposed −35 and −10 regions are in bold, and the putative IclR-type regulator binding site is boxed. h The deduced schematic of the regulation model of the CAT degradation pathway in R. opacus.
To study the roles of catR in regulating this cluster, we expressed transcriptional constructs PcatA-GFP+ and PcatR-GFP+ in both the WT and corresponding TF deletion mutant strains (Fig. 5b, c). In the WT strain, the promoter PcatA was responsive to benzoate, phenol, guaiacol, and CAT, but not to PCA (Supplementary Fig. 1g and 3b–d). In ∆catR, the activity of PcatA was strongly increased with the cultivation of both glucose and the respective aromatic compounds (Supplementary Fig. 3b–d), indicating that catR acts as a repressor (Fig. 5b). Moreover, as expected, knocking out catR modestly enhances the consumption of all three aromatic compounds (Supplementary Fig. 3e–g). This promoter is responsive when cultivated on phenol, benzoate, or guaiacol, so we hypothesized that the real effector compound might be one of the intermediates of the CAT degradation pathway. To confirm this hypothesis, we first tested the WT and TF deletion mutant strains with cultivation on CAT. Analysis showed that PcatA displayed only low basal activity in the glucose condition, but that the promoter activity was markedly enhanced in CAT (Fig. 5d). Surprisingly, the same trend of promoter activity was observed when cells were grown on cis-cis muconate (Fig. 5e). As it is known that CAT can be spontaneously converted into cis-cis muconate by the basal expression of the CAT 1,2-dioxygenase (catA), this finding suggests that the real effector compound may be cis-cis muconate.
A previous study has revealed that in Rhodococcus erythropolis, expression of the TF responsible for regulating the CAT degradation cluster is regulated by auto-regulatory repression37. To examine whether the regulation pattern of catR (LPD06569) in R. opacus is comparable, the promoter activity of PcatR was examined with cultivation on various carbon sources. In the WT strain, promoter PcatR showed constitutive activity when glucose was present, but in ∆catR, only basal expression from this promoter was observed under the same condition. Positing that ligand binding could change the conformation of catR, leading to the detachment of protein from the cognate promoter, we hypothesized that with ligand present in the media, the promoter activity of PcatR would be decreased. As expected, with the addition of cis-cis muconate, the output of this promoter was significantly decreased (Fig. 5f). Taken together, these results indicate that when regulating its own expression, catR acts as an activator in the absence of the ligand cis-cis muconate (Fig. 5c, h). The same assay was repeated with an array of aromatic compounds; promoter PcatR displayed reduced activity in all cases except when cultivated on PCA, which is not converted into cis-cis muconate (Supplementary Fig. 1h). Moreover, to test whether catR displays crosstalk between the benzoate, phenol, and guaiacol funneling pathways, the constructs PbenA-GFP+ (benzoate responsive), PpheB2-GFP + (phenol responsive), and Pcyp255-GFP+ (guaiacol responsive) were expressed in WT and the mutant ∆catR. In the absence of catR, the promoter activities of PbenA and PpheB2 were identical to that of the WT strain; however, the output of the promoter Pcyp255 was significantly decreased in ΔcatR, implying that catR may work as a weak co-activator in regulating the guaiacol funneling pathway (Supplementary Fig. 4). More intriguingly, no putative binding sites for catR were identified in Pcyp255, suggesting catR may play an indirect role in regulating this funneling pathway.
In addition to the role of the CAT branch of the β-ketoadipate pathway in degrading phenol, guaiacol, and benzoate, the PCA branch has been identified as the key pathway for the degradation of 4-hydroxybenzoate and vanillate in R. opacus. Moreover, the putative gene cluster has been confirmed by gene knockout experiments18. In contrast to the related mycobacterium Corynebacterium glutamicum38, only one IclR-type TF pcaR (LPD05454) is located in the PCA operon. To investigate how this single TF regulates the entire PCA operon, PpcaH-GFP+ and PpcaI-GFP+ constructs were expressed in both WT and the corresponding TF deletion mutant strains. For the orientation corresponding to coding region pcaH-pcaF (LPD05450-05455), promoter PpcaH was responsive to all the compounds we tested (Supplementary Fig. 1i), suggesting that—similar to observations in the CAT degradation pathway—the real inducer is also one of the intermediate compounds in the PCA degradation pathway. In the ∆pcaR strain, the promoter activity of PpcaH was significantly higher during cultivation on glucose, suggesting that pcaR may work as a transcriptional repressor (Fig. 6a-c). For the pcaI-pcaJ coding region (LPD05448-05449), in the presence of pcaR, carbon source did not make a significant difference in the promoter activity of PpcaI. However, in the mutant ∆pcaR, the promoter activity was dramatically enhanced on glucose, compared to that of WT, indicating that in regulating this coding region, pcaR works ligand-free as a repressor (Fig. 6d, e).
a A protocatechuic acid (PCA) degradation operon with an annotated transcription factor (TF). Promoters are represented as small arrows. Genes are shown with LPD gene numbers from the NCBI database (Refseq, CP003949.1). For the full annotations of the pathway genes, see Supplementary Table 1. b, d The proposed models for the role of pcaR (LPD05454). c, e To analyze the function of pcaR, the fusion construct PpcaH-GFP+ and PpcaI-GFP+ were expressed in both the WT and the corresponding TF deletion mutant strains. The normalized fluorescence of the WT and ∆pcaR strains was measured with and without supplemental PCA. All cultures contained 1 g/L glucose with (+) or without (−) 0.3 g/L PCA. All the fluorescence values were determined in the early stationary phase and normalized to OD600. Unpaired two-tailed t test was used to compare the variations in the changes of fluorescence of the strains under different conditions (**, P < 0.0021; ****, P < 0.0001; ns, not significant). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
Transcriptional cross-regulation of the funneling pathways during growth on alternative substrates
Bacteria often degrade mixed compounds sequentially; this type of ordered catabolism, termed CCR, has been noted in many different strains32,39,40,41. In R. opacus specifically, a targeted metabolomic approach was applied to establish the time-resolved concentrations of each individual aromatic compound in a defined mixture, showing that this strain preferred benzoate and 4-hydroxybenzoate over phenol, guaiacol, and vanillate. In the same work, a transcriptomic study revealed that R. opacus can differentially and specifically regulate aromatic utilization pathways in response to aromatic mixtures18, which led us to investigate the adaptability of R. opacus to the presence of multiple aromatic hydrocarbons as carbon sources.
As an initial step in studying the response of R. opacus to simultaneously available aromatic mixtures, the effects of 4-hydroxybenzoate and benzoate on the other three model compounds’ catabolism were examined. Time-course analyses of cell growth and the aromatic concentrations were conducted, and the results showed that the presence of 4-hydroxybenzoate inhibited the catabolism of phenol and guaiacol, but not vanillate (Fig. 7a-c). Even though the utilization of 4-hydroxybenzoate was preferential, diauxic growth was not observed (Fig. 7a, c). Because the funneling pathways play important roles in the degradation of aromatics, we hypothesized that 4-hydroxybenzoate might mediate a certain form of transcriptional repression over the funneling pathway of phenol or guaiacol, in particular by inhibiting the activity of the cognate promoters. To test our hypothesis, different strains harboring the constructs PpheB2-GFP+ (phenol responsive), PvanA-GFP+ (vanillate responsive), or Pcyp255-GFP+ (guaiacol responsive) were selected to test the activities of each promoter when treated with different combinations of aromatic mixtures (Fig. 7d–f). For phenol, transcription from PpheB2 was significantly decreased by the presence of 4-hydroxybenzoate for the first 16 h of cultivation (Fig. 7g). Promoter PpheB1, which drives the expression of non-essential phenol degradation cluster was also selected and tested. Similar to the findings for PpheB2, the data showed that the presence of 4-hydroxybenzoate precipitates a significant delay in the induction of this promoter by phenol, and that expression increased in the latter stages of cultivation, suggesting that the transcriptional repression observed earlier may be eased as 4-hydroxybenzoate is preferentially consumed (Supplementary Fig. 5a, b). For the guaiacol-reactive promoter, Pcyp255, a similar trend was observed (Fig. 7i). In contrast, the addition of 4-hydroxybenzoate did not cause a significant delay in vanillate induction, except at the 16 and 20 h time points (Fig. 7h).
a–c Utilization profiles of the aromatic mixtures. A glucose-grown WT cell culture was used as the inoculum. Cells were cultured in minimal medium with different aromatic mixtures as carbon sources: a 0.75 g/L HBA + 0.75 g/L PHE; b 0.5 g/L HBA + 0.5 g/L VAN; c 0.75 g/L HBA + 0.75 g/L GUA. At each time point, a 200 μL cell suspension was taken to measure the cell density (OD), and concentrations of the aromatics (Con) in the supernatant were determined by HPLC. d–f The proposed working models for the interference between HBA and PHE (d), HBA and VAN (e), and HBA and GUA (f). The different transcriptional constructs were transformed into WT R. opacus strain and used for fluorescence assays. g–i The normalized fluorescence in response to the aromatic mixtures was measured for each mixture: g HBA + PHE, h HBA + VAN, and i HBA + GUA. All cultures (g–i) contained 1 g/L glucose, with either 0.3 g/L PHE, 0.3 g/L VAN, or 0.3 g/L GUA. The concentrations of HBA were set at 0, 0.6, and 0.9 g/L. All of the fluorescence values were determined and normalized to OD600. Two-tailed mixed model ANOVA with Sidak’s multiple comparisons was used to compare the variations in the changes of concentration (Con) of the two aromatic compounds (☆, P < 0.05); for fluorescence assays, the variations in the changes of fluorescence of the two-compound treatment were compared against that of the one-compound treatment only (★, P < 0.05 for 0.6 g/L HBA; ☆, P < 0.05 for 0.9 g/L HBA; two-tailed mixed model ANOVA with Sidak’s multiple comparisons). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
The effect of benzoate on the catabolism of phenol, vanillate, and guaiacol was also analyzed; the results indicate that while R. opacus preferentially consumes benzoate over phenol and vanillate (Fig. 8a, b), no such preference was observed between benzoate and guaiacol (Fig. 8c). To test whether the similarities between 4-hydroxybenzoate and benzoate preferential consumption extended to transcriptional repression of parallel funneling pathways, the activities of the cognate promoters were examined by feeding cultures with different combinations of aromatic monomers (Fig. 8d–f). For phenol and vanillate, transcription from the cognate promoter decreased with the addition of benzoate (Fig. 8g, h and Supplementary Fig. 5c), which is consistent with the consumption profile data. Intriguingly, even though R. opacus could simultaneously consume benzoate and guaiacol, significant transcriptional repression of promoter Pcyp255 was observed when benzoate was available during the cultivation (Fig. 8i). Similarly, we compared the consumption profiles of benzoate and 4-hydroxybenzoate. We observed that in the WT strain, the consumption of benzoate is faster than that of 4-hydroxybenzoate (Supplementary Fig. 6a), and further observed that, in the presence of benzoate, the induction of the 4-hydroxybenzoate funneling pathway was delayed (Supplementary Fig. 6b, c).
a–c Utilization profiles of the aromatic mixtures. A glucose-grown WT cell culture was used as the inoculum. Cells were cultured in minimal medium with different aromatic mixtures as carbon sources: a 0.75 g/L BEN + 0.75 g/L PHE, b 0.75 g/L BEN + 0.75 g/L VAN, and c 0.75 g/L BEN + 0.75 g/L guaiacol (GUA). At each time point, a 200 μL cell suspension was taken to measure the cell density (OD), and concentrations of the aromatics (Con) in the supernatant were determined by HPLC. d–f The proposed working models for the interference between BEN and PHE (d), BEN and VAN (e), and BEN and GUA (f). The different transcriptional constructs were transformed into the WT R. opacus strain and used for the fluorescence assays. g–i Normalized fluorescence was measured in response to the aromatic mixtures: g BEN + PHE, h BEN + VAN, and i BEN + GUA. All cultures (g–i) contained 1 g/L glucose with either 0.3 g/L PHE, 0.3 g/L VAN, or 0.3 g/L GUA. The concentrations of BEN were set at 0, 0.9, and 1.2 g/L. All of the fluorescence values were determined and normalized to OD600. Two-tailed mixed model ANOVA with Sidak’s multiple comparisons was used to compare the variations in the changes of concentration (Con) of the two aromatic compounds (☆, P < 0.05); for fluorescence assays, the variations in the changes of fluorescence of the two-compound treatment were compared against that of the one-compound treatment (★, P < 0.05 for 0.9 g/L BEN; ☆, P < 0.05 for 1.2 g/L BEN; two-tailed mixed model ANOVA with Sidak’s multiple comparisons). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
To further specify the sequential prioritization of vanillate, phenol, and guaiacol, we conducted a combinatorial examination of the consumption profiles. For phenol+vanillate, the consumption data showed that R. opacus could simultaneously consume vanillate and phenol, and that VAN induction was not repressed by the addition of phenol (Supplementary Fig. 7a, b, d). In contrast, the activity of the phenol-responsive promoter PpheB1 was significantly decreased when vanillate was available (Supplementary Fig. 7b, c). For the combinations of phenol+guaiacol and vanillate+guaiacol, we observed that R. opacus consumed phenol and vanillate prior to guaiacol (Supplementary Figs. 8a and 9a). As this delay corresponded to a delay in guaiacol induction in the presence of vanillate or phenol (Supplementary Figs. 8d and 9d), we posited that vanillate and phenol may inhibit the consumption of guaiacol.
The above results suggest that the sequential consumption of aromatics in R. opacus may be mediated through a certain form of transcriptional repression over the respective funneling pathways. To confirm this possibility, we examined the promoter activity of Pcyp255 in different mutants with cultivation on guaiacol plus 4-hydroxybenzoate or the intermediate compound PCA (4-hydroxybenzoate degradation through the PCA branch and the corresponding transcriptional regulation pattern being clearly demonstrated; Fig. 9a). In the WT, guaiacol induction was significantly inhibited when 4-hydroxybenzoate was available as additional carbon source, but with the addition of an equal amount of PCA, the induction pattern of the promoter Pcyp255 reverted to that of the guaiacol-only treatment (Fig. 9b). Similar repression was observed in the PCA scenario when additional 4-hydroxybenzoate was added (Fig. 9b), suggesting that, rather than the intermediates of 4-hydroxybenzoate degradation, 4-hydroxybenzoate itself precipitates the observed transcriptional repression. Because the TF-based “induction prevention” mechanism has been identified to describe the preferential consumption of aromatics in bacteria31,33, to better address the possibility of 4-hydroxybenzoate repression on guaiacol utilization, we generated mutant strains defective in the catabolism of 4-hydroxybenzoate. First, the TF hbaR1 (LPD06765)—a transcriptional activator responsible for regulating the expression of 4-hydroxybenzoate monooxygenase pobA (LPD06764)—was knocked out. The fusion construct Pcyp255-GFP+ was expressed in this ∆hbaR1 mutant, and the resulting fluorescence data indicated a lack of transcriptional repression, regardless of whether cells were cultivated with or without 4-hydroxybenzoate (Fig. 9c). Next, we deleted pobA (LPD06764). In this mutant, the conversion of 4-hydroxybenzoate to PCA is abolished, whereas TF hbaR1 is still active. Monitoring the promoter activity when cells were cultivated on different aromatic combinations showed no transcriptional repression in ∆pobA strain (Fig. 9d), suggesting that the transcriptional repression cannot attribute to induction prevention. This finding indicates that diverse CCR mechanisms have evolved in R. opacus to deal with mixtures of several substrates, an idea that needs to be further investigated.
a Schematics depicting the funneling pathway for 4-hydroxybenzoate (HBA) and the annotated transcription factor. b–d Proposed working models for the promoter Pcyp255 during growth in minimal medium with guaiacol (GUA) and HBA (upper) and fluorescence of Pcyp255-GFP+ in response to GUA and HBA in WT and knockout strain backgrounds (lower). All cultures contained 1 g/L glucose with either 0.3 g/L GUA, or 0.3 g/L GUA plus 0.9 g/L protocatechuic acid (PCA) or 0.9 g/L HBA. The concentrations of the PCA and HBA mixtures were set at 0.45 g/L. All the fluorescence values were determined in the early stationary phase and normalized to OD600. Unpaired two-tailed t test was used to compare variations in the changes of fluorescence of the two aromatic mixtures against that of the GUA-only control (****, P < 0.0001; ns, not significant). All values represent the mean of triplicate cultures, with error bars depicting the standard deviation from that mean.
Discussion
Unraveling the complex transcriptional regulation of the catabolism of aromatics in R. opacus is a prerequisite for engineering this promising chassis for many biotechnological applications. It was discovered that many related catabolic pathways did not carry the same regulatory system, suggesting that the regulatory systems and their target operons seem to become associated independently, which makes the regulatory system varied and complex26. In this study, the AraC-type regulators we found for controlling the expression of the phenol and the guaiacol funneling pathways are transcriptional activators (Figs. 1 and 2). Our results also demonstrated that phenol is the effector compound for inducing expression of the pheR2-pheB2A2 cluster, but intriguingly, there is evidence to suggest that cis-cis muconate is also a ligand for TF pheR2 (Supplementary Fig. 1b). While unexpected, this behavior is not unprecedented: our multi-omics data have revealed that the corresponding phenol degradation cluster (pheB2A2) is also highly upregulated when treated with guaiacol (121-477-fold)18, another compound which is degraded through the CAT branch of the β-ketoadipate pathway.
IclR-type TFs are generally recognized as transcriptional repressors26; however, assays of promoter activity in both the WT and the mutant strains indicated that the IclR-type TF hbaR1 and vanR1 both work as transcriptional activators in regulating the expression of 4-hydroxybenzoate and vanillate funneling pathways, respectively (Figs. 3 and 4). This is not inconsistent with previous data, as IclR-type TFs have also been found to work as activators in regulating the catabolic pathways42. In addition, our findings also demonstrated that the transcriptional regulation of the two different branches of the β-ketoadipate pathway is controlled by separate IclR-type transcriptional repressors (Figs. 5 and 6). For the CAT branch, two regulatory modes have been reported: when CAT-degradation genes are controlled by LysR-type activators, the effector compound is usually the intermediate cis-cis muconate, whereas in operons under the regulation of IclR-type repressors, this role is mostly fulfilled by aromatic substrates37. Although the IclR-type transcriptional regulators have a similar structure as the LysR-type regulators43, the rather dissimilar amino acid sequences distinguish these two families. In this study, our results confirmed that the effector compound for induction of the CAT degradation pathway is cis-cis muconate, which is dramatically different from that of closely-related strain R. erythropolis CCM2595, where the expression of CAT-degradation genes is induced by phenol, rather than CAT or cis-cis muconate37. Although, the entire PCA branch of β-ketoadipate pathway consists of 8 individual genes, our analysis indicated a single, IclR-type repressor, pcaR (LPD05454), is responsible for regulating the entire PCA operon (Fig. 6). This is also a departure from precedent, as for example, the mycobacterium C. glutamicum places the PCA degradation pathway under the control of two different regulators: pcaIJ and pcaFDO are regulated by the IclR-type repressor pcaR, whereas the expression of pcaHG is controlled by an atypically large ATP-binding LuxR family (LAL)-type activator pcaO38,44.
In general, the coding gene for an IclR-type TF lies upstream of its target gene cluster and is transcribed in the opposite direction26. In this study, however, we found that the location and transcription orientation of IclR-type regulators varied. For instance, the catR regulator is located upstream of the CAT degradation operon, and is transcribed divergently (Fig. 5a); however, the upstream-located hbaR1 is transcribed in the same orientation as the critical gene in the 4-hydroxybenzoate funneling pathway (Fig. 3a). Notably, the vanR1 and pcaR are located within their target operons (Figs. 4a and 6a), arranged in their genomic neighborhoods in a way which may provide extra regulatory functions. vanR1 is located immediately downstream of the vanillate monooxygenase reductase vanB (LPD00563) and shares a promoter with this operon (Fig. 4a), an arrangement which suggests that the vanillate degradation process may be regulated through a positive feedback loop (PFL)45. While studies have demonstrated that the positive-feedback response to an environmental signal is slower than in those systems that produce the regulatory protein constitutively46, this moderate delay could be beneficial in R. opacus as a means of ordering the action of cellular response mechanisms in time, e.g., by upregulating the gene clusters involved in one-carbon compound metabolism to prepare for the harmful formaldehyde released during a demethylation step of vanillate catabolism. For pcaR, however, the regulator lies in the middle of the pcaH-pcaF (LPD05450-05455) coding region and thus shares the promoter, resulting in a negative feedback loop (NFL). Because some of the pca genes (e.g., pcaIJ and pcaFD) are also involved in the degradation of CAT, this NFL allows the enhanced PcaR to downregulate the expression of those genes, which consequently serves as a metabolic node for controlling the carbon flux of the β-ketoadipate pathway towards the TCA cycle to reduce succinate overflow47.
In nature, carbon and energy resources are often limited. Thus, specific bacteria that are more efficient or more selective in utilizing the carbon sources in their habitat will have significantly higher growth rates and therefore greater competitive success than other microorganisms48. In this work, we found that benzoate is the most-preferred substrate among the model compounds we tested. Additionally, we were able to firmly establish the substrate hierarchy of all tested compounds in R. opacus (Figs. 7–8 and Supplementary Figs. 5–9). The preference for benzoate can presumably be traced to the different energetic demands of the funneling pathways—the conversion of benzoate to CAT consumes no net reducing equivalents because the NADH oxidized in the first step is recovered during the next dehydrogenation reaction by an NAD+-dependent dehydrogenase. In contrast, the conversions of 4-hydroxybenzoate or vanillate to PCA and phenol or guaiacol to CAT require the oxidation of NAD(P)H18,34. Energetic considerations cannot, however, explain the preference for 4-hydroxybenzoate, since the initial steps in the metabolisms of 4-hydroxybenzoate, phenol, and guaiacol have similar requirements for reducing equivalents. To facilitate understanding the complex intracellular dynamics, a cybernetic model has been previously developed to describe the resource allocation and microbial kinetics that influence the hierarchical utilization of carbon sources49; different variants of this model have been able to account for a variety of instances of preferential carbon uptake in E. coli50. From this resource-allocation point of view, it is reasonable to find that R. opacus prefers 4-hydroxybenzoate over phenol and guaiacol. Compared to phenol, which has redundant funneling pathways, and guaiacol, which requires an accessory demethylation pathway to detoxify formaldehyde released from the funneling process, the 4-hydroxybenzoate funneling pathway’s “single regulator-single enzyme” unit is relatively simple and thus less resource-intensive.
Intriguingly, although the vanillate and guaiacol funneling pathways both require reducing equivalents and the accessory demethylation pathway, our results indicated preferential utilization of vanillate over guaiacol (Supplementary Fig. 9). This preference cannot be explained by the two theories discussed above, but the difference in the structures of the two funneling pathway operons may indicate that the preference is attributable to the mechanistic model51. Specifically, in a mixture of vanillate and guaiacol, the proposed PFL regulation pattern for vanillate keeps the TF vanR1 preferentially expressed, which in turn could further promote the induction of the vanillate funneling pathway, supporting fast cell growth. Fast growth on vanillate may result in dilution of guaR, thus preventing effective induction of the guaiacol funneling pathway. While these models developed in E. coli can be used to provide a coarse-grained description of the preferential utilization of aromatics, we believe that a new generation of models, specifically and precisely tuned for R. opacus, is still needed.
Our results also provide some insights into the mechanism of the substrate hierarchy in R. opacus, or more specifically, the transcriptional cross-regulation of the funneling pathways. Induction prevention has been used to describe the preferential utilization of aromatics in other strains31,32,33. However, our results showed that in the mutants ∆hbaR1 and ∆pobA (for which degradation of 4-hydroxybenzoate was abolished), no transcriptional repression of the phenol or guaiacol funneling pathway was observed (Fig. 9 and Supplementary Fig. 10), suggesting that the preference for 4-hydroxybenzoate cannot be explained by induction prevention alone. Inspired by the resource allocation view, we speculate that this preference might be controlled by a novel global regulation mechanism. More specifically, due to the relative simplicity of the 4-hydroxybenzoate funneling pathway, the investment towards the utilization of 4-hydroxybenzoate is much smaller, which allows cells to maximize profit. In this scenario, once cells detect 4-hydroxybenzoate in the environment, more resources are allocated to the 4-hydroxybenzoate degradation pathway, with transcription and translation of the operons which enable the degradation of phenol or guaiacol consequently decreased, resulting in the sequential utilization of the three compounds. Transporter-mediated inducer exclusion has been used to describe the molecular mechanism of the substrate hierarchy in Bacillus species29. In this study, we also studied the transcriptional regulation of the annotated shikimate transporter LPD06699 and found that the expression of this transporter could be induced by all the model compounds we tested; furthermore, this upregulation was enhanced in the TF deletion mutant (∆LPD06698) (Supplementary Figs. 11–12). Interestingly, in the vanillate scenario alone, the upregulation of this transporter improved carbon utilization (Supplementary Fig. 12e), supporting the hypothesis that inducer exclusion might be one of the mechanisms responsible for the substrate hierarchy in R. opacus that still needs to be evaluated.
In conclusion, we identified and investigated the TFs involved in regulating several aromatic degradation pathways in R. opacus by combining gene knockouts with aromatic sensors. We also observed that individual lignin model compounds in an aromatic mixture are consumed by R. opacus in sequential order and that this preferential utilization pattern can be ascribed to the transcriptional cross-regulation of the funneling pathways. While we have been able to describe many mechanisms of individual pathway control, aspects of how these pathways interact have yet to be fully explained. Nonetheless, our results can inform the development of strain-specific models of R. opacus metabolism for industrial applications.
Methods
Strains and plasmids
R. opacus PD630 (DSMZ 44193) was used as the strain (WT) to construct mutants. Unless otherwise indicated, cells were grown at 30 °C and 250 rpm in a previously described minimal salts medium B52 with different carbon sources. All the plasmids constructed in this study were confirmed by DNA sequencing and are summarized in Supplementary Table 2.
Plasmid construction
For constructing the TF deletion strains, the deletion construct was assembled as described previously53. ~600 bp regions up- and downstream of the selected transcriptional regulator were amplified from the genome. These two pieces of DNA fragments were assembled into a plasmid containing an E. coli origin of replication (p15a) and a chloramphenicol selection cassette. To analyze the potential function of the selected TF, the upstream region containing the promoter and ribosome-binding site (RBS) of the proposed target genes was cloned in front of gfp+. The resulting transcriptional construct was then expressed in both WT and the corresponding TF deletion mutant strains. Unless otherwise described, all plasmids were assembled in E. coli DH10B using Gibson Assembly54.
T7 RNAP-based CRISPRi
The T7 RNAP expression platform was developed in our previous study36. Specifically, the T7 RNAP gene was integrated into the R. opacus chromosomal neutral site (ROCI3) under the control of the phenol inducible promoter (PpheB2). The codon-optimized dcas9 from Streptococcus thermophilus (dcas9Sth1) was placed under the control of the pBAD promoter. The expression of the guide RNA was driven by T7 promoter. One sgRNA (sgRNA PHE_1) was designed to target pheB2 (LPD06575), and a strain with dcas9Sth1 and T7 RNAP but no sgRNA was used as a control. The uninduced condition represented 0 mM arabinose and 0.3 g/L PHE, while the induced condition represented 50 mM arabinose and 0.3 g/L PHE.
Transformation of R. opacus
Competent cells were made as described previously52. Cells were transformed with ~500 ng plasmid DNA by electroporation. To generate R. opacus knockout mutants, a previously developed homologous recombination method was applied with modifications53. Briefly, a helper plasmid expressing modified viral recombinases was introduced into the strain via electroporation, and electrocompetent cells were made from this strain. The cells were transformed with ~2 μg suicide plasmid DNA. Positive colonies were verified by colony PCR. All the strains used in this study are summarized in Supplementary Table 3.
Cell growth and fluorescence measurements
The optical density at 600 nm (OD600), absorbance at 600 nm (Abs600), and fluorescence were measured using a Tecan Infinite M200 Pro plate reader. For the measurements of GFP+ fluorescence, the excitation and emission wavelengths were 488 and 530 nm, respectively52. Fluorescence values were normalized using Eq. (1):
where Fluorescencenorm is the normalized fluorescence, A600sample is the absorbance (at 600 nm) of the test strain, Fluorescencesample is the fluorescence value of the test strain, A600control is the absorbance of the empty vector control strain, and Fluorescencecontrol is the fluorescence value of the empty vector control strain.
Aromatic consumption profiling
To measure the aromatics, 200 μL samples from each cell culture were centrifuged at 3500 rcf for 5 min, and the cell culture supernatant was subsequently analyzed. For cell cultures using a single aromatic compound as the carbon source, the concentration of each lignin model compound was measured by comparing UV absorbance values to a standard curve for each lignin model compound. For cultures using the aromatic mixture, overnight cultures of WT R. opacus cells grown on 1 g/L glucose were harvested, washed, and then resuspended in fresh minimal medium containing different combinations of aromatic compounds at an initial OD600 of ~0.05. Aromatics in the culture supernatant were detected using an Agilent 1260 Infinity II HPLC system equipped with the Agilent Poroshell 120 EC-C18 column (4.6 × 100 mm, 2.7 μm) and a UV detector (at 280 nm). The temperature was set at 60 °C, and the flow rate was 1 mL/min. Mobile phase A (water with 0.1% formic acid) and mobile phase B (acetonitrile with 0.1% formic acid) were used as follows (A%/B% with a gradient elution): 92/8 at 0 min, 74/26 at 5 min, 50/50 at 8 min, and 92/8 at 10 min. Concentrations were determined by comparing UV absorbance values to a standard curve for each lignin model compound.
Statistics and reproducibility
All experiments were conducted with at least three biological replicates. Differences between the control and engineered strains were analyzed using GraphPad Prism 9.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The main data and raw data supporting the results described in this study are available within the main paper, its Supplementary Information, Supplementary Data 1, and Supplementary Data 2. Additional data generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Stephanopoulos, G. Challenges in engineering microbes for biofuels production. Science 315, 801–804 (2007).
Ragauskas, A. J. et al. The path forward for biofuels and biomaterials. Science 311, 484–489 (2006).
Maity, S. K. Opportunities, recent trends and challenges of integrated biorefinery: part II. Renew. Sust. Energy Rev. 43, 1446–1466 (2015).
Xu, Z., Lei, P., Zhai, R., Wen, Z. & Jin, M. Recent advances in lignin valorization with bacterial cultures: microorganisms, metabolic pathways, and bio-products. Biotechnol. Biofuels 12, 32 (2019).
Becker, J. & Wittmann, C. A field of dreams: lignin valorization into chemicals, materials, fuels, and health-care products. Biotechnol. Adv. 37, 107360 (2019).
Brink, D. P., Ravi, K., Liden, G. & Gorwa-Grauslund, M. F. Mapping the diversity of microbial lignin catabolism: experiences from the eLignin database. Appl. Microbiol. Biotechnol. 103, 3979–4002 (2019).
Kumar, M. et al. Lignin valorization by bacterial genus Pseudomonas: state-of-the-art review and prospects. Bioresour. Technol. 320, 124412 (2021).
Wang, H., Pu, Y., Ragauskas, A. & Yang, B. From lignin to valuable products-strategies, challenges, and prospects. Bioresour. Technol. 271, 449–461 (2019).
Eng, T. et al. Engineering Pseudomonas putida for efficient aromatic conversion to bioproduct using high throughput screening in a bioreactor. Metab. Eng. 66, 229–238 (2021).
Yaguchi, A. L., Lee, S. J. & Blenner, M. A. Synthetic biology towards engineering microbial lignin biotransformation. Trends Biotechnol. 39, 1037–1064 (2021).
Varman, A. M. et al. Decoding how a soil bacterium extracts building blocks and metabolic energy from ligninolysis provides road map for lignin valorization. Proc. Natl Acad. Sci. 113, E5802–E5811 (2016).
Salvachúa, D., Karp, E. M., Nimlos, C. T., Vardon, D. R. & Beckham, G. T. Towards lignin consolidated bioprocessing: simultaneous lignin depolymerization and product generation by bacteria. Green. Chem. 17, 4951–4967 (2015).
Arvay, E., Biggs, B. W., Guerrero, L., Jiang, V. & Tyo, K. Engineering acinetobacter baylyi ADP1 for mevalonate production from lignin-derived aromatic compounds. Metab. Eng. Commun. 13, e00173 (2021).
Gellerstedt, G., Sjöholm, E. & Brodin, I. The wood-based biorefinery: a source of carbon fiber? Open Agric. J. 4, 119 (2010).
Linger, J. G. et al. Lignin valorization through integrated biological funneling and chemical catalysis. Proc. Natl. Acad. Sci. USA 111, 12013–12018 (2014).
Beckham, G. T., Johnson, C. W., Karp, E. M., Salvachúa, D. & Vardon, D. R. Opportunities and challenges in biological lignin valorization. Curr. Opin. Biotechnol. 42, 40–53 (2016).
McLeod, M. P. et al. The complete genome of <em>Rhodococcus</em> sp. RHA1 provides insights into a catabolic powerhouse. Proc. Natl. Acad. Sci. 103, 15582–15587 (2006).
Henson, W. R. et al. Multi-omic elucidation of aromatic catabolism in adaptively evolved Rhodococcus opacus. Metab. Eng. 49, 69–83 (2018).
Alvarez, H. M., Mayer, F., Fabritius, D. & Steinbuchel, A. Formation of intracytoplasmic lipid inclusions by Rhodococcus opacus strain PD630. Arch. Microbiol. 165, 377–386 (1996).
Tsitko, I. V., Zaitsev, G. M., Lobanok, A. G. & Salkinoja-Salonen, M. S. Effect of aromatic compounds on cellular fatty acid composition of Rhodococcus opacus. Appl. Environ. Microbiol. 65, 853–855 (1999).
Navas, L. E. et al. Bacterial transformation of aromatic monomers in softwood black liquor. Front. Microbiol. 12, 735000 (2021).
Li, X. et al. Discovery of potential pathways for biological conversion of poplar wood into lipids by co-fermentation of Rhodococci strains. Biotechnol. Biofuels 12, 60 (2019).
Xie, S. X. et al. Mechanism-guided design of highly efficient protein secretion and lipid conversion for biomanufacturing and biorefining. Adv. Sci. 6, 1801980 (2019).
Yoneda, A. et al. Comparative transcriptomics elucidates adaptive phenol tolerance and utilization in lipid-accumulating Rhodococcus opacus PD630. Nucleic Acids Res. 44, 2240–2254 (2016).
deLorenzo, V. & PerezMartin, J. Regulatory noise in prokaryotic promoters: How bacteria learn to respond to novel environmental signals. Mol. Microbiol. 19, 1177–1184 (1996).
Tropel, D. & van der Meer, J. R. Bacterial transcriptional regulators for degradation pathways of aromatic compounds. Microbiol. Mol. Biol. Rev. 68, 474–500 (2004).
Vigneshvar, S., Sudhakumari, C. C., Senthilkumaran, B. & Prakash, H. Recent advances in biosensor technology for potential applications - an overview. Front. Bioeng. Biotechnol. 4, 11 (2016).
Deutscher, J. The mechanisms of carbon catabolite repression in bacteria. Curr. Opin. Microbiol. 11, 87–93 (2008).
Stulke, J. & Hillen, W. Regulation of carbon catabolism in Bacillus species. Annu. Rev. Microbiol. 54, 849–880 (2000).
Lane, S. et al. Glucose repression can be alleviated by reducing glucose phosphorylation rate in Saccharomyces cerevisiae. Sci. Rep. 8, 2613 (2018).
Brzostowicz, P. C., Reams, A. B., Clark, T. J. & Neidle, E. L. Transcriptional cross-regulation of the catechol and protocatechuate branches of the beta-ketoadipate pathway contributes to carbon source-dependent expression of the Acinetobacter sp. strain ADP1 pobA gene. Appl. Environ. Microbiol. 69, 1598–1606 (2003).
Nichols, N. N. & Harwood, C. S. Repression of 4-hydroxybenzoate transport and degradation by benzoate - a new layer of regulatory control in the pseudomonas-putida beta-ketoadipate pathway. J. Bacteriol. 177, 7033–7040 (1995).
Donoso, R. A., Perez-Pantoja, D. & Gonzalez, B. Strict and direct transcriptional repression of the pobA gene by benzoate avoids 4-hydroxybenzoate degradation in the pollutant degrader bacterium Cupriavidus necator JMP134. Environ. Microbiol. 13, 1590–1600 (2011).
Choi, K. Y., Zylstra, G. J. & Kim, E. Benzoate catabolite repression of the phthalate degradation pathway in Rhodococcus sp. strain DK17. Appl. Environ. Microbiol. 73, 1370–1374 (2007).
Zhan, Y. et al. Benzoate catabolite repression of the phenol degradation in Acinetobacter calcoaceticus PHEA-2. Curr. Microbiol. 59, 368–373 (2009).
DeLorenzo, D. M., Diao, J., Carr, R., Hu, Y. & Moon, T. S. An improved CRISPR interference tool to engineer Rhodococcus opacus. ACS Synth. Biol. 10, 786–798 (2021).
Vesely, M., Knoppova, M., Nesvera, J. & Patek, M. Analysis of catRABC operon for catechol degradation from phenol-degrading Rhodococcus erythropolis. Appl. Microbiol. Biotechnol. 76, 159–168 (2007).
Brinkrolf, K., Brune, I. & Tauch, A. Transcriptional regulation of catabolic pathways for aromatic compounds in Corynebacterium glutamicum. Genet. Mol. Res. 5, 773–789 (2006).
Ampe, F., Leonard, D. & Lindley, N. D. Repression of phenol catabolism by organic acids in Ralstonia eutropha. Appl. Environ. Microbiol. 64, 1–6 (1998).
Gaines, G. L. 3rd, Smith, L. & Neidle, E. L. Novel nuclear magnetic resonance spectroscopy methods demonstrate preferential carbon source utilization by Acinetobacter calcoaceticus. J. Bacteriol. 178, 6833–6841 (1996).
Heinaru, E. et al. Reversible accumulation of p-hydroxybenzoate and catechol determines the sequential decomposition of phenolic compounds in mixed substrate cultivations in pseudomonads. FEMS Microbiol. Ecol. 37, 79–89 (2001).
Romero-Steiner, S., Parales, R. E., Harwood, C. S. & Houghton, J. E. Characterization of the pcaR regulatory gene from Pseudomonas putida, which is required for the complete degradation of p-hydroxybenzoate. J. Bacteriol. 176, 5771–5779 (1994).
Schell, M. A. Molecular biology of the LysR family of transcriptional regulators. Annu. Rev. Microbiol. 47, 597–626 (1993).
Zhao, K. X., Huang, Y., Chen, X., Wang, N. X. & Liu, S. J. PcaO positively regulates pcaHG of the beta-ketoadipate pathway in Corynebacterium glutamicum. J. Bacteriol. 192, 1565–1572 (2010).
Mitrophanov, A. Y. & Groisman, E. A. Positive feedback in cellular control systems. Bioessays 30, 542–555 (2008).
Alon, U. Network motifs: theory and experimental approaches. Nat. Rev. Genet. 8, 450–461 (2007).
Roell, G. W. et al. A concerted systems biology analysis of phenol metabolism in Rhodococcus opacus PD630. Metab. Eng. 55, 120–130 (2019).
Gorke, B. & Stulke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Ramkrishna, D. & Song, H. S. Dynamic models of metabolism: review of the cybernetic approach. Aiche J. 58, 986–997 (2012).
Ramakrishna, R., Ramkrishna, D. & Konopka, A. E. Cybernetic modeling of growth in mixed, substitutable substrate environments: preferential and simultaneous utilization. Biotechnol. Bioeng. 52, 141–151 (1996).
Narang, A. & Pilyugin, S. S. Bacterial gene regulation in diauxic and non-diauxic growth. J. Theor. Biol. 244, 326–348 (2007).
DeLorenzo, D. M., Henson, W. R. & Moon, T. S. Development of chemical and metabolite sensors for Rhodococcus opacus PD630. ACS Synth. Biol. 6, 1973–1978 (2017).
DeLorenzo, D. M., Rottinghaus, A. G., Henson, W. R. & Moon, T. S. Molecular toolkit for gene expression control and genome modification in Rhodococcus opacus PD630. ACS Synth. Biol. 7, 727–738 (2018).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Acknowledgements
The authors thank Dr. Shulin Chen for pAL5000 (S) and KanR, Dr. Anthony Sinskey for GentR, Dr. Graham Hatfull for the Che9c recombinases, and Dr. Sarah Fortune for the dCas9Sth1 gene. The authors also thank James Ballard for his comments on the manuscript. This work was supported by the United States Department of Energy (DE-SC0018324 to T.S.M.) and the United States Department of Agriculture (2020-33522-32319 to T.S.M.).
Author information
Authors and Affiliations
Contributions
T.S.M. conceived the project. J.D. and T.S.M. designed experiments and analyzed the data. J.D. performed the experiments. J.D., R.C., and T.S.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Johann Heider and Seetharaman Vaidyanathan for their contribution to the peer review of this work. Primary Handling Editors: Shahid Mukhtar and Gene Chong. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diao, J., Carr, R. & Moon, T.S. Deciphering the transcriptional regulation of the catabolism of lignin-derived aromatics in Rhodococcus opacus PD630. Commun Biol 5, 1109 (2022). https://doi.org/10.1038/s42003-022-04069-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-04069-2
This article is cited by
-
Mining microbial metabolism
Nature Chemical Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.