Abstract
How multipotential cells initiate distinct gene expression programs in response to external cues to instruct cell fate choice remains a fundamental question in biology. Establishment of CD4 and CD8 T cell fates during thymocyte development is critically regulated by T cell receptor (TCR) signals, which in turn control expression of the CD4-determining transcription factor ThPOK. However, the mechanism whereby differential TCR signals are molecularly interpreted to promote or antagonize ThPOK expression, and thereby CD4 versus CD8 lineage fates remains unknown. Here we show, using reverse genetic and molecular approaches that an autonomous, position-independent TCR-sensing switch is embedded within the ThPOK locus. Further, using an in vivo mutagenesis approach, we demonstrate that differential TCR signals are interpreted during lineage commitment by relative binding of EGR, NFAT and Ebox factors to this bistable switch. Collectively our study reveals the central molecular mechanism whereby TCR signaling influences differential lineage choice. Ultimately, these findings may provide an important new tool for skewing T cell fate to treat cancer and autoimmune diseases.
Similar content being viewed by others
Introduction
How bipotential cells use environmental cues to precisely orchestrate distinct gene expression programs in order to promote alternate developmental fates, remains a fundamental question in biology. CD4-CD8 T-cell lineage commitment in the thymus provides a valuable model system to understand this process1. T cells develop in the thymus from the early thymocyte progenitor stage to the mature SP CD4 and CD8 stages via a precisely ordered series of intermediate steps. The CD4+ CD8+ (double positive, or DP) stage is particularly important, as thymocytes first express a complete αβTCR at this stage, allowing them to engage MHC on antigen-presenting cells and undergo positive selection. Prior to positive selection DP thymocytes are believed to be lineage-uncommitted, i.e., not biased towards either CD4 or CD8 lineage choice. After positive selection, DP thymocytes split into alternate CD4 and CD8 lineages, by undergoing selective loss of one of the coreceptors. There is a tight correlation between a thymocyte’s TCR specificity for MHC class II or I, and differentiation to the CD4 or CD8 lineages, respectively.
The near-perfect correlation between lineage choice and MHC restriction can be explained by the kinetic-signaling model, which postulates that relatively long or short TCR signals promote CD4 versus CD8 commitment, respectively2,3,4. At the molecular level, it is established that the transcription factor (TF) ThPOK is necessary and sufficient to drive CD4 commitment, and to prevent CD8 commitment, of developing thymocytes5,6. Accordingly, ThPOK protein levels are higher in class II-restricted thymocytes than in class I-restricted thymocytes, suggesting a causal link between TCR engagement and ThPOK expression7. However, the fundamental question of how differential TCR signals control lineage-specific ThPOK expression, and thereby alternate lineage fate, remains to be resolved.
What is known so far is that ThPOK expression in thymocytes and mature T cells is controlled primarily at the transcriptional level via several stage- and lineage-specific cis elements8,9. Of particular importance is the 400 bp ThPOK silencer, SilThPOK, which is located 3 kb upstream of the distal ThPOK promoter8,9. Germline deletion of the SilThPOK in mice causes promiscuous expression of ThPOK and diverts all thymocytes towards the CD4 lineage, demonstrating that the SilThPOK is essential for repression of ThPOK transcription in cells that would normally adopt the CD8 lineage. Our understanding of how the SilThPOK is regulated, however, remains rudimentary. Deletion of 2 Runx consensus binding motifs severely impairs silencing function8,9, and mice lacking Runx1 and Runx3 or the obligate Runx-binding partner Cbfb, exhibit loss of the T-cytotoxic lineage. Interestingly, while constitutive expression of ThPOK causes redirection of class I-restricted thymocytes to the CD4 lineage, overexpression of Runx3 is not sufficient to redirect MHC II-restricted thymocytes to the CD8 + lineage10. Furthermore, Runx factors are bound to the SilThPOK at all stages of thymic development, indicating that differential binding by Runx factors is not responsible for differential silencer function in class I- versus II-restricted thymocytes. Hence, the molecular basis for how differential TCR signals regulate ThPOK expression in class I- versus class II-restricted thymocytes remains to be determined.
Regulation of the Cd4 gene during thymic development somewhat parallels that of ThPOK, in that it is also controlled by a stage-specific silencer element, SilCD4, which selectively represses Cd4 transcription in SP CD8 but not SP CD4 thymocytes11,12,13, and which contains functionally critical Runx-binding sites14,15. However, no evidence has emerged to date that Cd4 transcription and the SilCD4, in particular, are regulated by TCR signals. Indeed, this would seem intuitively unlikely given that Cd4 is transcribed in both unsignaled DP thymocytes and strongly signaled SP CD4 thymocytes.
As outlined above, the molecular genetic mechanisms by which TCR-dependent regulation of ThPOK transcription is controlled remain unknown. Here, we seek to resolve this critical issue using an in vivo gene targeting approach. First, through reciprocal swapping of the SilThPOK with the SilCD4, we provide molecular genetic proof that TCR signals directly target the SilThPOK. Second, using precise in vivo gene editing we identify an autonomous, position-independent TCR-sensing switch within the SilThPOK that controls ThPOK expression in developing thymocytes. Collectively, our study defines the central molecular genetic mechanism whereby TCR signaling influences lineage choice via regulation of ThPOK expression.
Results
In vivo silencer swap reveals the autonomous and position-independent function of SilThPOK
While we previously showed that strong TCR signals induce ThPOK transcription in thymocytes8, the molecular mechanisms that connect TCR signals with ThPOK induction are unknown. We reasoned that TCR signals may induce ThPOK expression either by (1) activation of positive regulatory elements (enhancers/promoters), or (2) inactivation of the SilThPOK silencer (Supplementary Fig. 1). To genetically test whether the SilThPOK encodes the autonomous and locus-independent capacity to sense differences in MHC class I- versus class II-restricted TCR signaling, we generated SilThPOK swap mice, in which the SilThPOK is inserted into the Cd4 gene in place of its own SilCD4 silencer element (CD4ThPOKsil mice). We used this approach because SilCD4 and SilThPOK share important functional attributes, i.e., both are active in developing class I-restricted thymocytes and both are dependent on binding of Runx factors14,16. The murine SilThPOK and SilCD4 elements are predicted to interact with 183 and 164 TFs, respectively, many of which are differentially regulated between mature CD4 and CD8 thymocytes and thus may contribute to lineage-specific regulation of the SilThPOK and SilCD4 elements. Interestingly, 129 of these TFs are predicted to bind both SilThPOK and SilCD4 elements (Supplementary Fig. 2a–c), suggesting substantial commonality in control of both elements. On the other hand, some TFs are predicted to show unique or highly preferred binding to the SilThPOK versus SilCD4 elements, suggesting a selective role in the control of the SilThPOK. Hence, we hypothesized that the SilThPOK might substitute in many respects for the SilCD4, but that some functions might be specialized, including the ability to respond to differential TCR signals.
We performed the SilThPOK > SilCD4 swap in 2 steps: first we generated mice in which the SilCD4 is deleted (CD4Δsil mice) (Fig. 1a). As previously reported, CD4Δsil mice exhibit gain of CD4 expression by mature CD8 T lymphocytes both in heterozygous and homozygous condition, consistent with the fact that the SilCD4 is required to suppress Cd4 expression after CD8 commitment9 (Fig. 1b–d). Next, we inserted the SilThPOK into the CD4Δsil allele (at the site of the SilCD4 deletion) to test whether it was able to restore normal regulation of Cd4 expression (CD4ThPOKsil mice), as assessed at the single-cell level by FACS (Fig. 1a). Expression of the CD4ThPOK.Sil allele was first assessed in heterozygous CD4ThPOKsil/+ mice, in which the CD4ThPOK.Sil allele is expressed in combination with a normal Cd4 allele regulated by the endogenous SilCD4. T cells from these mice showed no change in coreceptor expression pattern in thymocytes or peripheral T cells, including no gain of CD4 expression on CD8 T cells, suggesting that the SilThPOK substituted fully for the function of the endogenous SilCD4 in the CD8 lineage and/or is functionally complemented by the wt Cd4 allele. We next generated hemizygous CD4ThPOKsil/o mice, in which only the CD4ThPOKsil allele is expressed, by crossing CD4ThPOKsil mice to Cd4-deficient (CD4o/o) mice17. Importantly, these mice show severe alteration of CD4 expression in the thymus, i.e., downregulation of CD4 on most immature (TCRlo/−) thymocytes. As a result, normal immature DP (CD4 + CD8 + TCRlo) thymocytes are largely replaced by aberrant SP CD8 TCRlo cells (distinguishable from mature SP CD8 cells by the absence of surface TCR; Supplementary Fig. 3) (Fig. 1b–d). Thus, insertion of SilThPOK into the Cd4 locus represses transcription of Cd4 in most DP thymocytes, revealing an inherent capacity of the SilThPOK to repress transcription at the DP stage.
a Schematic of Cd4 gene organization in wt mice (top row), CD4ΔSil (second row) or CD4ThPOK.Sil knock-in mice (bottom row). Black boxes indicate exons. Enhancers are shown as white boxes, the Cd4 silencer as a red circle, and the ThPOK silencer as a red triangle. b FACS analysis of CD4, and CD8a expression of total thymocytes (top row), or indicated gated thymocyte subsets (bottom 2 rows) of CD4Δsil/Δsil, CD4O/ThPOK.Sil, CD4O/O, and wt mice. Note that CD4 expression is severely reduced at immature DP-like stage, but is substantially upmodulated in many CD69+ TCRβ+ thymocytes that have received a recent TCR signal. c FACS analysis of CD4 and CD8a expression of total mesenteric lymph node (LN) cells (top row), or gated TCRβ+ LN cells subsets (bottom row) of same strains of mice as above. Results are representative of three independent experiments (n = 3, for each strain). d Plots showing % of DP, SP CD4, SP CD8, and DN thymocytes for mice of indicated genotypes. N = 3 independent animals. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Significant differences were determined by one-way ANOVA with post hoc Tukey HSD (honest significant difference), and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to wt mice.
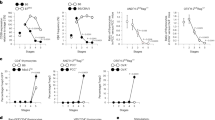
Downmodulation of CD4 at the DP stage is expected to impair TCR signaling by MHC class II-restricted thymocytes leading to a defect in their differentiation, similar to CD4 knockout mice in which most class II-restricted thymocytes are redirected to the CD8 lineage18. Indeed, CD4ThPOKsil/CD4o mice show a reduction in the frequency of SP CD4 thymocytes and peripheral CD4 T cells compared to wt mice (~50% of normal) (Fig. 1b–d). To distinguish whether this reflects a block in the development of class II-restricted cells, or redirection to the CD8 lineage, we backcrossed CD4ThPOK.Sil mice to a β2m-deficient background, in which only MHC class II-restricted cells can develop19. While, the frequency of SP CD4 cells was reduced, particularly in the periphery (spleen and lymph node), the CD8 compartment was not substantially increased (Fig. 2a, b), implying that the defect in CD4 generation reflects a partial block in development rather than redirection to the CD8 lineage. We speculated that remaining CD4 cells in CD4ThPOK.Sil/ThPOK.Sil mice might be strongly skewed toward higher affinity TCRs that are relatively less dependent on CD4 coengagement for efficient TCR signaling. To test this hypothesis, we introduced the AND TCR transgene, which recognizes the MHC II I-Ak allele with high affinity, onto the CD4ThPOK.Sil background. Notably, AND TCR+ I-Ak+ CD4ThPOK.Sil/ThPOK.Sil mice showed normal frequencies of mature CD4 thymocytes, supporting the view that high-affinity TCRs are unaffected on the CD4ThPOK.Sil background (Supplementary Fig. 4a; note that SP CD8 thymocytes are immature DP-like cells, not mature CD8 thymocytes). In contrast to most MHC class II-restricted cells, the development of class I-restricted T cells appears normal. Thus, on the MHC class II-deficient background, in which only class I-restricted cells can develop, all cells mature appropriately to the CD8 lineage and give rise to normal frequencies of CD8 cells (Supplementary Fig. 4b).
a FACS analysis of CD4 and CD8a expression by indicated thymic or peripheral lymphocyte populations of wt, CD4ThPOK.Sil/+ or CD4ThPOK.Sil/ThPOK.Sil mice crossed to the β2 m-deficient background. b Plots showing % of DP, SP CD4, and SP CD8 thymocytes and LN for mice of indicated strains on β2m−/− background. N = 3 independent animals, for each strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Significant differences between indicated mutant mice and WT mice were determined by one-way ANOVA with post hoc Tukey HSD (honest significant difference), and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to β2m−/− mice.
SilThPOK encodes inherent capacity to respond to TCR signal
To directly test whether the SilThPOK encodes the inherent capacity for TCR responsiveness in the context of CD4ThPOKsil mice, we treated CD4ThPOKsil/ThPOKsil MHC II deficient mice in vivo with anti-TCR antibody, as an inducer of strong TCR signaling, and used Cd4 transcription from the knock-in locus as a readout of TCR responsiveness. We focused on CD69+ CD4lo/− CD8+ thymocytes, because they are the functional equivalent of CD69+ DP thymocytes (since SilThPOK is repressing Cd4 expression at the immature DP equivalent stage). In normal thymic development the first signaled thymocytes appear as CD69+ CD4+ CD8+20. Importantly, we detected a strong increase of Cd4 mRNA levels in activated (CD69+) thymocytes derived from treated versus untreated mice, providing direct biochemical evidence that the SilThPOK encodes the inherent capacity for inducibility in an autonomous and position-independent manner in response to strong TCR signals (Supplementary Fig. 5a, b). Since TCR signaling increases Cd4 expression from the CD4ThPOK.Sil knock-in locus, TCR signaling appears to counteract SilThPOK function. Altogether our data indicate that the SilThPOK represses the Cd4 locus at the DP stage via a cell-autonomous and position-independent mechanism, and confers susceptibility to inactivation by strong TCR signals in activated (CD69+) thymocytes.
SilCD4 element cannot substitute for the TCR-sensing capacity of the SilThPOK
Next, we tested the proposition whether the capacity to respond to TCR signals is unique to the SilThPOK or might be shared with other regulatory elements at the ThPOK locus. For this purpose, we carried out a converse silencer swap approach in which the SilThPOK was replaced by an exogenous silencer element. We first used the human adult-specific γ-globin silencer21. However, this silencer failed to repress ThPOK expression in DP thymocytes or in any other thymocyte stage (Supplementary Fig. 6), suggesting that the γ-globin gene does not work in the context of T lymphocytes. To overcome this limitation, we instead used the SilCD4, which is known to be active in the T-cell lineage. Accordingly, we replaced the 418 bp SilThPOK by the 434 bp SilCD4 in the context of the endogenous ThPOK locus, to generate ThPOKCD4Sil mice (Fig. 3a). As mentioned above, the SilCD4 is selectively active in SP CD8 cells, but unlike the SilThPOK is not active in DP thymocytes. FACS analysis of homozygous ThPOKCD4Sil knock-in mice showed essentially normal CD4:CD8 T-lymphocyte ratios in mature thymocytes and peripheral T cells, consistent with the efficient functional substitution for the SilThPOK by the SilCD4 during T-cell development (Fig. 3b, c). Furthermore, RT-PCR analysis showed normal repression of ThPOK transcription in sorted mature SP CD8 thymocytes, However, expression levels of ThPOK at other stages of T-lymphocyte development were severely altered, i.e., upmodulated in DP T-lymphocyte precursors, and downmodulated in CD4+ 8lo (eightfold) and SP CD4 lymphocytes in thymus and peripheral immune sites (five- to sixfold) (Fig. 3d).
a Schematic of ThPOK gene organization in ThPOKCD4.Sil knock-in mice. Black boxes indicate exons. Enhancers are shown as white boxes, and the Cd4 silencer as a red circle. b FACS analysis of CD4 and CD8α expression by indicated thymic populations of wt or ThPOKCD4-Sil/CD4-Sil mice. c FACS analysis of CD4 and CD8a expression by indicated splenic populations of same mice as in panel b. Graphic comparison of proportions of DN, SP CD4, and SP CD8 subsets within gated TCRβ + spleen lymphocytes of wt or ThPOKCD4-Sil/CD4-Sil mice (n = 7) (right panel). There were no statistically significant differences between mice of ThPOKCD4.Sil/CD4.Sil and ThPOK+/+ genotypes by one-way ANOVA with post hoc Tukey HSD. d RT-PCR analysis showing relative expression of ThPOK mRNA in indicated sorted thymocyte subsets of wt or ThPOKCD4-Sil/CD4-Sil mice. Results are a combination of three replicates per strain. RT-PCR data represent four technical replicates, each derived from pooled RNA of three animals. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. All FACS results are representative of at least three experiments. Statistical significance was determined between mice of ThPOKCD4.Sil/CD4.Sil and ThPOK+/+ genotypes by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001).
Given the severe misregulation of ThPOK levels in ThPOKCD4Sil mice, we questioned whether the coreceptor expression pattern of their SP thymocytes really correlates appropriately with MHC restriction, despite the superficially normal CD4:CD8 ratio. To definitively address this issue, we crossed ThPOKCD4Sil mice with MHC class I- or II-deficient mice (or TCR transgenic mice expressing class II- or I-restricted TCRs) to restrict development exclusively to class II- or I-restricted cells, respectively. ThPOKCD4Sil MHC I−/− mice developed a considerable proportion of misdirected class II-restricted CD8 T lymphocytes in the thymus compared to control MHC I−/− mice, of which a few are also detected in the periphery, although less than in the thymus, presumably reflecting homeostatic mechanisms that favor class II-restricted T lymphocytes expressing the appropriate CD4 coreceptor (Fig. 4a). Interestingly, redirected CD8 SP thymocytes were evident only at the mature CD69- stage, not among signaled (TCRβ + CD69+) thymocytes, which are thought to contain lineage-committed semi-mature CD4 and CD8 SP cells in WT mice. Hence, derepression of ThPOK appears to be influencing post-commitment events in CD8 SP differentiation. Even more strikingly, ThPOKCD4Sil MHC II−/− mice develop almost equal frequencies of mature CD4 and CD8 T lymphocytes in the thymus, in contrast to MHC II−/− control mice (Fig. 4b). The proportion of SP CD4 cells was also elevated in the periphery compared to control MHC II−/− mice, although to a lesser degree (Fig. 4b). Conversely, there is substantial misdirection of class II-restricted thymocytes to the CD8 lineage in ThPOKCD4Sil MHC I−/− mice compared to control MHC I−/− mice (Fig. 4b). A few misdirected MHC II-restricted CD8 T cells are also detected in the periphery, although proportionally less than in the thymus. These data demonstrate that the SilCD4 cannot substitute functionally for the SilThPOK, especially failing to repress ThPOK transcription in DP thymocytes and causing inappropriate reduction of ThPOK transcription in CD4+ 8lo and SP CD4 thymocytes. Most importantly, other endogenous ThPOK regulatory elements cannot confer TCR sensitivity in the absence of the SilThPOK. As a consequence, thymocytes from ThPOKCD4Sil mice exhibit random lineage commitment independent of MHC specificity (Fig. 4a, b). Altogether these results suggest that ThPOK promoters and enhancers are insufficient to support normal levels of ThPOK transcription in response to TCR signal when linked to the SilCD4. Thus TCR sensitivity is exclusive to the SilThPOK.
a FACS analysis of CD4 and CD8a expression by indicated thymic or peripheral lymphocyte populations of wt or ThPOKCD4-Sil/CD4-Sil mice crossed to the β2 m-deficient background. b FACS analysis of CD4 and CD8α expression by indicated thymic or peripheral lymphocyte populations of wt or ThPOKCD4-Sil/CD4-Sil mice crossed to the MHC II-deficient background. N = 3 independent animals per strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Statistical significance was determined between mice of ThPOKCD4.Sil/CD4.Sil and ThPOK+/+ genotypes on β2m−/− or MHC II−/− background by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to β2m−/− mice (a), or MHC II−/− mice (b).
TCR responsiveness of the SilThPOK may be encoded by an evolutionarily conserved TF consensus site signature
To identify TFs that could be responsible for TCR responsiveness of the murine SilThPOK during thymic development, we first assessed interspecies conservation of the ThPOK and Cd4 silencers. This revealed that the SilThPOK but not the SilCD4 showed extensive sequence homology between all mammalian species examined22 (Supplementary Fig. 7a). Further, organization of TF consensus sites (as predicted by JASPAR algorithm) was well conserved between species for the SilThPOK, but not the SilCD4 element. Thus 524 of 1315 (40%) predicted TF sites for the mouse SilThPOK element were also conserved in relative position/orientation in placental and marsupial mammals (human versus opossum), compared to only 106 of 1071 (10%) for the mouse SilCD4 element (Supplementary Fig. 7b)22. Given that precise motif grammar may not be critical for function of some cis elements (e.g., billboard enhancers), we compared just the relative number of sites for each TF within each element, regardless of position. We excluded TFs not expressed at the DP > SP transition (according to IMMGEN Skyline RNA-seq database), and not evolutionarily conserved (i.e., not predicted to bind both human and mouse homologs of either SilThPOK or SilCD4). For the remaining 170 TFs, the number of binding sites within SilThPOK and SilCD4 elements for three mammalian species (human, mouse, opossum) was averaged across all three species, and the ratio between SilThPOK and SilCD4 calculated for each TF, to reveal candidate TFs that may selectively bind SilThPOK or SilCD4. Importantly, TF sites that are overrepresented in the SilThPOK versus the SilCD4 include many TFs implicated in TCR signaling, such as Ebox factors (HEB, E2A)23, Egr1, NFAT, and NfkB (Supplementary Fig. 8a). Conversely, the SilCD4 is enriched for different TF consensus sites that do not include sites for TFs implicated in TCR signaling (Supplementary Fig. 8b). The most enriched sites were mapped onto the respective sequence coordinates of the SilThPOK or SilCD4 to define distinct, evolutionarily conserved TF site signatures. Strikingly, the distribution of preferentially represented TF sites is closely conserved across species for the SilThPOK, but not for the SilCD4 (Supplementary Fig. 8c). Of note, this does not imply that the latter are functionally unimportant, but rather that motif grammar (relative order and spacing of TF sites) may be more important for the function of the SilThPOK, than the SilCD4.
SilThPOK regions responsible for lineage-specific silencing bind multiple effectors of TCR signaling
Before determining the role of specific TF binding to the SilThPOK for its lineage-specific function, we first determined the essential region of the SilThPOK required for its lineage and stage-specific function by creating three different SilThPOK deletion mutant alleles: (A) Δ1–150 bp (line NR82); (B) Δ1–261 bp (line QK27); and (C) Δ357–418 bp (line QC48) (Fig. 5a). All three mutant mouse lines were bred to homozygosity. Strikingly, both QC48 and QK27 deletions resulted in a severe reduction in SP CD8 cells in both the thymus and periphery, suggesting the failure of silencing activity during thymic development (Fig. 5b–e). In contrast, homozygous mutant NR82 mice developed normal proportions of CD8 T cells in the thymus and periphery24. These results indicate that the regions deleted in lines QC48 and QK27, but not NR82, encode non-redundant functions essential for silencing. Next, we tested whether the absence of CD8 SP thymocytes in QC48−/− and QK27−/− mice reflected a block in the development of MHC class I-restricted thymocytes or redirection to the CD4 lineage by crossing each mutant onto a MHC class II-deficient background, which restricts development to class I-restricted cells. Strikingly, even on the MHC class II-deficient background QC48 and QK27 mice still generate CD4 T cells, in contrast to MHC II−/− control mice which develop only CD8 cells, suggesting redirection of class I-restricted T cells to the CD4 lineage (Fig. 5f–i). Collectively, nonoverlapping regions of the SilThPOK, i.e., 150–260 bp and 358–418 bp, are individually essential for lineage-specific SilThPOK function. Notably, both of these regions are enriched for EGR consensus motifs, while the 358–418 bp region also includes two NFAT motifs.
a Schematic of positions of transcription factor (TF) consensus binding sites within murine SilThOPK (top row). Different TF motifs are color-coded according to the legend at the left. b FACS analysis of CD4, CD8a, TCRβ, and CD69 expression of total thymocytes (top and second rows), or CD4 and CD8a expression of indicated gated thymocyte subsets (bottom 2 rows) of ThPOKsil.ΔQC48, ThPOKsil.ΔQK27 and wt mice, as indicated. Note that mature CD8 population is absent in both mutant lines. c FACS analysis of TCR expression of total mesenteric lymph node (LN) cells (top row), or gated TCRβ + LN cells subsets (bottom row) of same strains of mice as above. Results are representative of multiple experiments. N = 3 independent animals per strain. d Plots showing % of DP, SP CD4, SP CD8, and DN thymocytes, or (e) SP CD4 and SP CD8 T cells from LN for mice of indicated genotypes. N = 5 independent animals per strain. Data are presented as mean values +/− SEM. A P value <0.05 was considered significant. Significant differences were determined by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). f FACS analysis of TCRβ, CD69, CD4, and CD8α expression by indicated thymic and (g) peripheral lymphocyte populations of ThPOKsil.ΔQC48, ThPOKsil.ΔQK27 and wt mice, as indicated, crossed to the MHC II-deficient background. h, i Plots showing % of DP, SP CD4, SP CD8, and DN thymocytes (h) or SP CD4 and SP CD8 T cells (i) for mice of indicated genotypes. N = 6 independent animals per strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Significant differences were determined between indicated mutant mice and WT control mice by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to wt mice (d, e), or MHC II−/− mice (h, i).
Of note, a 34-bp region containing conserved Runx-binding motifs, which have previously been shown to be essential for SilThPOK function8,9,25, was left intact in both QC48 and QK27 mutants, so that their phenotypes are not attributable to disruption of Runx binding. Nevertheless, we noticed that this 34-bp region also encodes several predicted EGR, Ebox, and NFAT-binding sites (Fig. 6a). In order to evaluate the importance of these motifs, we generated further knock-in mice in which this 34-bp segment of the murine SilThPOK was replaced with a 34-bp segment of the murine SilCD4 that also contains two closely spaced Runx sites, but lacks any of the predicted Egr, Ebox or NFAT motifs (ThPOKSIL.CD4Rx mice) (Fig. 6a). Importantly, ThPOKSIL.CD4Rx mice showed a striking reduction in SP CD8 thymocytes and CD8 peripheral T cells, albeit less severe than control ThPOKSIL.ΔRUNX mice in which both Runx sites are deleted (Fig. 6b). Crossing ThPOKSIL.CD4Rx mice to OT-1 TCR transgenic mice reveal substantial redirection of class I-restricted thymocytes to the CD4 lineage, indicating aberrant ThPOK upmodulation as a result of impaired silencer function (Fig. 6c). Hence, the presence of Runx motifs at this location within the SilThPOK is not sufficient to confer normal regulation of silencer activity. Rather other motifs surrounding the Runx sites, including predicted Egr, Ebox, and NFAT sites, are also critical, suggesting functional synergy between these factors and Runx factors in the control of SilThPOK activity.
a Schematic of ThPOKSIL.CD4.Rx and ThPOKSIL.ΔRUNX knock-in alleles, indicating the location of conserved Runx sites (red bars). Blue color indicates swapped region from the SilCD4. Bottom panels show the position/orientation of indicated TF consensus sites for the swapped regions. Note that the occurrence of Ebox consensus motifs at indicated positions within the SilThPOK but not SilCD4. b FACS analysis of CD4, and CD8 expression of total thymocytes, or indicated gated thymocyte and peripheral T-cell subsets of wt, CD4 SIL.CD4Rx/SIL.CD4Rx and CD4SIL.ΔRUNX/SIL.ΔRUNX mice. Results are representative of multiple experiments. N = 3 independent animals per strain. Plots showing % of DP, SP CD4, SP CD8, and DN thymocytes for mice of indicated genotypes are shown at right. N = 3 independent animals per strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. c FACS analysis of CD4 and CD8 expression by indicated thymic or peripheral lymphocyte populations of wt or CD4 SIL.CD4Rx/SIL.CD4Rx mice expressing the MHC class I-restricted OT-1 TCR transgene on the selecting H-2b/b background. Results are representative of multiple experiments. Note that the proportion of SP CD4 mature thymocytes is strongly or moderately increased for mature thymocytes a or gated peripheral T cells, respectively, from CD4 SIL.CD4Rx/SIL.CD4Rx mice. N = 3 independent animals per strain. Statistical significance was determined between indicated mutant mice and ThPOK+/+ mice by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to wt mice.
Given that regions required for SilThPOK function are notably enriched for EGR and NFAT consensus motifs, we directly tested whether EGR and NFAT can bind to the SilThPOK by EMSA. We used two different labeled oligos corresponding to 161–268 bp (probe 1) and 169–371 bp (probe 2) regions of the SilThPOK. For each region, we generated alternate probes encoding either the endogenous SilThPOK sequence (wt probes), or mutant sequences in which consensus Egr or NFAT motifs (as determined by JASPAR algorithm) are disrupted (mutant probes). We observed that (1) both wt probes bind to EGR1 and NFAT2 factors, as evidenced by band shift upon incubation with cell lysates overexpressing either factor, while (2) mutant probes failed to undergo such band shift, or to a much lesser extent, demonstrating site-specificity of this TF binding (Fig. 7a).
a EMSA analysis showing EGR1 and NFAT2 binding to oligos comprising wt silencer sequence (region surrounding Runx sites), or mutated at consensus EGR or NFAT motifs, as indicated. b Position of TF consensus binding sites for wt silencer, or indicated variant alleles. FACS analysis of TCRβ, CD69, CD4, and CD8 expression of total thymocytes, gated thymocytes (c), and peripheral T-cell subsets (d) of wt mice or mutant lines, as indicated. Results are representative of multiple experiments. N = 6 independent animals per strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Statistical significance was determined between indicated mutant mice and ThPOK+/+ mice by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to wt mice.
Effect of combinatorial disruption of EGR, NFAT, and Ebox motifs on SilThPOK function
Next, to test the in vivo relevance of EGR, Ebox (TCF12/3), and NFAT motifs for SilThPOK function, we generated a series of mutant mouse lines in which we mutated different combinations of these sites (numbered as shown in Fig. 7b) within the context of the endogenous full-length SilThPOK element: (1) the QK9 allele, containing mutations of three EGR (E6, E7, E9), one Ebox (T6), and one NFAT (N5) site, (2) the QW82 allele, containing mutations of two EGR (E6, E7), and one Ebox (T6) site, and (3) The QY53 allele, containing mutations of three EGR (E6, E7, E9), and one NFAT (N5) (Fig. 7b).
All mutants showed major defects in thymic development: (1) Homozygous QK9−/− mice completely lack CD8 T cells in thymus and periphery, suggesting loss of SilThPOK function (Fig. 7c, d). Furthermore, crossing to MHC class II-deficient background to limit thymocyte development to class I-restricted cells, showed the preferential generation of CD4 rather than CD8 T cells, compared to MHC II−/− control mice, indicating redirection of class I-restricted thymocytes to the CD4 lineage, presumably consequent to aberrant ThPOK upmodulation (Fig. 8a, b). (2) Homozygous QW82 mice display a similar phenotype to QK9 mice, i.e., lack of CD8 T-cell development, and substantial redirection of class I-restricted cells to the CD4 lineage, as evidenced by the presence of a large proportion of CD4 T cells in class II- deficient QW82−/− mice (Fig. 8a–d), suggesting a severe defect of SilThPOK repression. (3) Homozygous QY53−/− mouse also displays a striking block in CD8 development and redirection of a large proportion of class I-restricted thymocytes to the CD4 lineage as QK9 and QW82 mice (Figs. 7c, d and 8a–d). Collectively, our mutational analysis indicates that (a) two EGR consensus motifs that are mutated in all three mutant lines (E6, E7 sites; Fig. 7a) are indispensable for CD8 lineage-specific ThPOK silencing, while (b) NFAT, Ebox, and EGR sites (N3, E9, and T6 sites) that are mutated in only some lines are unnecessary or redundant for this process. Of note, while the QC48 deletion showed a severe developmental phenotype (Fig. 5), this may not be attributable to Egr or Nfat sites contained within this region, as mutating all Egr sites (line TO61) or all Nfat sites (line RQ17) within this region had no effect on CD4/CD8 ratio. It remains possible that Egr and NFAT sites within the QC48 region are collectively required for correct lineage choice.
a, b FACS analysis of TCRβ, CD69, CD4, and CD8 expression of total thymocytes, gated thymocytes (a), and peripheral T-cell subsets (b) of wt, or mutant mouse lines crossed to MHC II−/− background. c, d Plots showing % of DP, SP CD4, SP CD8, and DN thymocytes for mice of indicated genotypes. N = 6 independent animals per strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Statistical significance was determined between mutant mice and ThPOK+/+ mice on MHC II−/− background by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to wt mice.
Effect of disruption of specific NFAT and Ebox motifs on SilThPOK function
To separately evaluate the contribution of these NFAT and Ebox factor binding sites to silencer activity independent of EGR site mutations, we generated two additional mutant alleles, with mutations of (A) Ebox site T6 (line RJ59;), or (B) NFAT site N3, which also affects adjoining Stat and EGR motifs (line RS59) (Fig. 9a). As expected from our previous mutants, neither homozygous line induced skewing toward the CD4 lineage. Surprisingly, however, they displayed substantial skewing toward the CD8 lineage in the thymus, resulting in CD4:CD8 ratios of 1.3:1 and 1.8:1 for homozygous RJ59−/− and RS59−/− mice, respectively (versus 3:1 in wt mice) (Fig. 9b–d). In addition, there was a notable increase in atypical mature DN T cells in the thymus (Fig. 9b). Similar CD8 skewing was evident in peripheral T cells of Ebox mutant RJ59−/− mice, but not NFAT mutant RS59−/− mice (Fig. 9c, e). We did not observe substantial misdirection of either class I- or class II-restricted cells in RJ59−/− or RS59−/− mice crossed to MHC II and MHC I-deficient backgrounds, suggesting that reduction in ThPOK expression in these lines may occur too late in development to allow CD8 commitment. Altogether, these data suggest that Ebox and NFAT binding to specific sites near conserved Runx-binding sites of the SilThPOK is necessary to oppose silencer function during the development of class II-restricted thymocytes to the CD4 lineage.
a Position of TF consensus binding sites for wt silencer, or indicated mutant RJ59 and RS59 alleles. FACS analysis of TCRβ, CD69, CD4, and CD8 expression of (b) total thymocytes, gated thymocytes and (c) peripheral T-cell subsets of wt mice or mutant lines, as indicated. d, e Plots showing % of DP, SP CD4, SP CD8, and DN thymocytes for mice of indicated genotypes. N = 6 independent animals per strain. Data are presented as mean values +/− SEM. A P value < 0.05 was considered significant. Statistical significance was determined between mice of ThPOK SIL.CD4Rx/SIL.CD4Rx and ThPOK+/+ genotypes by one-way ANOVA with post hoc Tukey HSD, and indicated by asterisks (*P < 0.01; **P < 0.005; ***P < 0.001). Statistical significance was calculated for each indicated mutant line relative to wt mice.
CD4 commitment involves intra-locus chromatin looping between the SilThPOK and other cis-regulatory elements
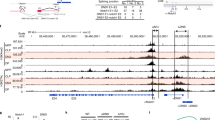
Given that enhancers and silencers function in part by controlling chromatin topology26,27, we asked whether topological assembly of ThPOK regulatory regions may be regulated in a stage- and/or lineage-specific manner. Accordingly, we performed 3C analysis on different thymocyte populations, i.e., predominantly class I-restricted thymocytes from OT-1 transgenic mice, predominantly class II-restricted thymocytes from AND transgenic mice, unsignaled DP thymocytes from CD3δ−/− thymocytes, and ThPOKΔsil thymocytes which express ThPOK constitutively due to absence of the SilThPOK (Fig. 10a, b). Interestingly, we observed a selective interaction between SilThPOK with the lymphoid enhancer (GTE) in OT-1 transgenic thymocytes, correlating with transcriptional repression of ThPOK. Conversely, interaction between the lymphoid enhancer (GTE) and the distal promoter was only observed in AND transgenic and ThPOKΔsil thymocytes, correlating with transcriptional activation of ThPOK. These observations suggest that the SilThPOK may specifically sequester the GTE enhancer away from the distal promoter in class I-restricted thymocytes (Fig. 10c). Analysis of publicly available databases indicates striking stage- and lineage-specific changes in chromatin accessibility and histone modification of the SilThPOK, GTE enhancer, and distal promoter during thymic development (Supplementary Fig. 9a, b).
a Schematic of ThPOK gene organization showing exons (dark blue boxes), promoters (light blue boxes), enhancers (green boxes), and silencer (red box). Arrowheads at bottom of the panel indicate the position/orientation of PCR primers, while connected arrows at top of the panel show different primer combinations that were utilized. b 3C analysis of physical interactions of SilThPOK with other ThPOK cis elements in total thymocytes from AND Tg+ (class II-restricted), OT-1 Tg+ (class I-restricted), CD3δ−/− (uncommitted; blocked at DP stage), and ThPOKΔSil (DRE knockout) mice, respectively. N = 3 independent animals per strain. c Model of proposed ThPOK cis element interactions in class I- versus class II-restricted thymocytes.
Discussion
CD4 versus CD8 lineage commitment is controlled by TCR specificity, such that long and/or strong TCR signals elicited by MHC class II lead to CD4 commitment, whereas weak and/or transient signals elicited by MHC class I lead to CD8 commitment. But how differential TCR signals culminate in alternate transcriptional programs driving CD4 versus CD8 choice has remained an open question. In this study, using a regulatory element swap approach, we provide compelling evidence that the SilThPOK encodes the intrinsic capacity to respond to TCR signaling leading to MHC restricted lineage choice, and show how this occurs at the molecular level.
Our comparison of the SilThPOK and SilCD4 elements shows that while they share the ability to suppress gene transcription in committed CD8 cells, they function very differently in early thymocyte developmental stages, particularly at the DP and CD4 + 8lo stages. Thus, these silencers possess intrinsically distinct functional capacities regardless of the genomic context, and dominantly control other cis elements in both endogenous and exogenous gene contexts. For example, the SilThPOK suppresses the transcription of Cd4 in DP thymocytes of CD4ThPOK.Sil mice similar to the suppression of endogenous ThPOK in DP thymocytes of wt mice. Conversely, derepression of ThPOK transcription in DP thymocytes of ThPOKCD4.Sil mice mirrors expression of endogenous Cd4 in DP thymocytes of wt mice. Hence, differential stage-specific regulatory control of the ThPOK and Cd4 loci appears to be autonomous and encoded primarily within their respective silencers.
The shared ability of SilThPOK and SilCD4 elements to repress transcription following CD8 lineage commitment may largely reflect the presence of evolutionarily conserved Runx-binding motifs in both silencers, which have previously been shown to be critical for the activity of both the SilCD4 and SilThPOK elements9,14,16,28,29. Nevertheless, the mechanism by which Runx binding supports silencing is not well understood. ChIP analysis shows binding of Runx complexes (Runx1 or 3) to the SilThPOK at all stages of thymic development, regardless of whether ThPOK is expressed, suggesting that the TCR responsiveness of the ThPOK silencer requires Runx factors to synergize with other factors bound to the SilThPOK. Our finding that knocking the Runx motifs from the SilCD4 into the SilThPOK (ThPOKSIL.CD4.Rx mice) causes aberrant redirection of class I-restricted thymocytes to the CD4 lineage, indicates that precise TF-binding site syntax surrounding the Runx sites is critical for SilThPOK function, and that the mere presence of Runx sites even at the same relative position of the silencer is not sufficient. Interestingly, some thymocytes still develop to the CD8 lineage in ThPOKSIL.CD4.Rx mice, compared to mice which lack Runx sites, indicating that silencing still occurs for some cells, possibly those expressing particularly low-affinity TCRs. We conclude that the regions surrounding the Runx sites of the SilThPOK and SilCD4 elements recruit different TFs that critically control stage-specific activity of these elements. Our analysis reveals that motifs for TFs implicated in TCR signaling are preferentially represented in the SilThPOK versus SilCD4, and that disrupting these motifs perturbs normal regulation of silencing, leading to abnormal CD4-8 development. In particular, specific EGR sites nearby to conserved Runx-binding sites are necessary to promote silencer function and oppose CD4 development, while certain Ebox and NFAT sites are necessary to oppose silencer function and permit CD4 development.
There is substantial data to support an important role of TCR signaling in the induction of EGR30, NFAT31, and Ebox factors32. EGR factors have in turn been implicated in CD4 development33,34,35 and regulation of Th differentiation and function36. NFATc proteins are induced strongly in CD4 compared to DP and CD8 thymocytes37, and have been previously implicated in the control of positive selection38,39. Finally, we previously showed that conditional deletion of HEB/E2A at the DP stage leads to a severe defect in CD4 T-cell development, while deletion of the E protein antagonists Id2 and Id3 at the DP stage favors CD4 over CD8 lineage development40. Our current data provide a partial molecular genetic mechanism for some of these effects. Targeting specific Ebox and NFAT sites in silencer (T6/ N5) causes loss of mature CD4 thymocytes, and gain of mature DN and CD8 thymocytes, indicating increased silencing activity, which results in diminished ThPOK expression and consequent loss of CD4 expression. So HEB and NFAT factors appear to play a role in tuning SilThPOK activity in response to different TCR signals, such that strong/long TCR signals elicited in MHC class II-restricted thymocytes cause preferential association of HEB and NFAT factors to the SilThPOK leading to its loss of activity. Cooperative interaction of Ebox and NFAT factors has been established in other cellular contexts41. On the other hand, targeting specific EGR sites in the silencer (E6 / E7) drives the development of all thymocytes to the CD4 lineage, indicating blockade of silencer function and constitutive ThPOK expression. Hence, it appears that even weak class I-restricted signals are sufficient to trigger EGR binding to these sites and EGR binding to SilThPOK is obligatory for silencing function. The effect of mutation of these EGR sites appears to be dominant over the effect of mutation of Ebox and NFAT-binding sites (since QK9, QW82, and QY53 mice have both E6/E7 sites and one or more T6/N5 sites mutated). It’s been shown that CD4-inducing TCR signals result in increased Egr1 and Egr2 induction versus CD8-inducing signals42. So, EGR factors induced by TCR signals are necessary to actively turn on silencer activity but may not be involved in tuning silencer activity in response to differential TCR signaling.
Analysis of available ATAC-seq data from purified thymocyte subsets indicates that the SilThPOK region is accessible in DN1-DP stages, suggesting that it is poised for activity, but that this accessibility is substantially lost in both CD4 and CD8 thymocytes (Supplementary Fig. 9a). We suggest that this loss of accessibility is due to dense binding by TFs that are induced during positive selection and lineage commitment, including NFAT, Ebox, and EGR factors. Based on ATAC data, it further appears that the ThPOK distal promoter and GTE enhancer become markedly more accessible during the transition from the DP to SP CD4 stage, and that the GTE simultaneously comes into close proximity with the distal ThPOK promoter according to 3C analysis. These changes do not occur in class I-restricted cells, where the GTE instead loops to the SilThPOK. SilThPOK-deficient cells which instead exhibit constitutive interaction of distal promoter and lymphoid enhancer (GTE), suggesting that weak TCR signals promote differential silencer-dependent chromatin organization at the ThPOK locus. In the context of our mutational analysis, one might hypothesize that EGR factors promote looping of the silencer to the GTE, while Ebox/NFAT factors may oppose it.
CD4 versus CD8 lineage commitment is controlled by TCR signals of different strength and duration, which in turn promote different downstream patterns of TF expression. It appears that initially many/most of the TFs involved in CD4 vs CD8 commitment are induced by all TCR signals, e.g., Egr, NFAT, etc., but that their kinetics of expression may differ in response to TCR signals of different strength/duration. As a consequence, there will be differences in the level and/or duration of binding of these shared TFs to the ThPOK silencer. By mutating particular consensus motifs within the silencer, we have prevented the binding of cognate TFs to these sites, and potentially affected the recruitment of other TFs to nearby sites. The fact that some silencer mutations result in intermediate phenotypes is consistent with a multilayered and redundant control mechanism. For instance, in QW82 mutants some class I-restricted thymocytes differentiate to the CD8 lineage while others progress aberrantly to the CD4 lineage. Given that thymocytes are heterogeneous in terms of their surface TCR affinity/avidity to the MHC I and MHC II encoded by their clonotypic TCRs, it’s reasonable to speculate that those class I-restricted thymocytes which are still able to differentiate to the CD8 lineage in QW82 mice express particularly low-affinity TCRs, resulting in distinct TF-binding patterns at the ThPOK silencer that are able to maintain its activity. Finally, it’s entirely possible that other non-TCR-dependent signal pathways also impinge on the ThPOK silencer and contribute to control of its activity, including the IL-7 signal pathway which has been implicated in CD8 commitment43. Indeed, we have identified consensus Stat factor binding sites in the silencer, and in the future will test how TCR signals may cooperate with or antagonize IL-7 signal to control silencer activity and lineage choice.
In summary, we postulate that stage-specific SilThPOK activity is controlled by TFs that are transcribed/activated downstream of TCR signals, including EGR, NFAT, and Ebox factors, which in turn control topological organization of in the ThPOK regulatory region particularly the apposition of the SilThPOK with other cis elements.
Methods
Mice
RAG1−/− (stock # 034159), β2m−/− (stock # 002087), and AND TCR transgenic (stock # 002761) mice have been procured from Jackson Laboratory. OT-1 TCR transgenic RAG2−/− (stock # 2334) and MHC II−/− (stock # ABBN12) mice were obtained from Taconic. CD3δ−/− mice are from our own colony44. All other mouse lines described in this paper have been generated by the FCCC Transgenic Facility on the C57BL/6J strain of M. musculus. Animals used in all experiments were 6–10 weeks of age, and males and females were used in equal proportions (no difference was noted between males and females in any experiment). Animal care was in accordance with NIH guidelines. Mice were maintained on a 12 h light/dark cycle, at 75 °F and 50% humidity. All experimentation involving animals was approved by the Institutional Animal Care and Use Committee (IACUC) of Fox Chase Cancer Center.
In vivo treatment with anti-TCRβ mAb
CD4ThPOKsil/ThPOKsil were crossed to MHC II −/− mice to generate compound mutant CD4ThPOKsil/ThPOKsil MHC II −/− mice. Four to six-week-old animals were injected i.p. with 30 μg of azide-free anti-TCRβ mAb H57-597 and thymocytes isolated 15 h later.
ZFN-mediated gene targeting in mouse embryos
Site-specific mutagenesis was carried out, according to established procedures22. Briefly, a pair of ZFN RNAs that recognize a specific target site near the SilThPOK was designed and generated by Millipore-Sigma (Genome Editing division). The ZFN target sequence is ACCGCTACCCTAACCcataaCTGGAAGGGGTTTAG (capital letters denotes nucleotides actually bound by right and left ZFN proteins). mRNAs encoding the two site-specific ZFNs (50 ng/μl) were introduced together with double-stranded DNA-targeting constructs bearing the desired mutations/deletions into 1-cell C57BL/6J mouse oocytes by pronuclear injection, and injected oocytes were transferred to a pseudopregnant surrogate mother. Targeting constructs contained 1.5 and 0.8 kb arms of homology on either side of the desired mutations/deletion. Positive founder pups were identified based on the reduced size of PCR product using primers F1 (5′-ATCCCTACGAAGAAGCCTCT-3′) and R1 (5′-AGGCTTTCCATGTCAGGGTC-3′), and mated to C57BL/6 mice for seven generations to produce stable heritable knock-in lines.
Flow cytometry
All fluorescently labeled antibodies used were obtained from commercial sources (eBioscience, Biolegend, BD, or Invitrogen), including TCRβ-PE/Cy5 (H57-597 Cat # 109210; Lot # B170070), CD4-BV421 (clone RM4-5; BioLeg Cat # 100544; Lot # B293278), CD8a-APC/Cy7 (clone 53-6.7; BioLeg Cat # 100714; Lot # B276265), CD69-PE/Cy7 (clone H1.2F3; eBio Cat # 25-0691-82; Lot # E07583-1635), CD24-FITC (clone M1-69; BioLeg Cat 101806; Lot # B184710), CD62L-PerCP/Cy5.5 (clone MEL-14; BioLeg Cat 104432; Lot # B272105). 1 × 106 cells were stained in 100 μl of PBS, 5% FCS at 4 °C for 30 min with 0.5 μg/ml of each antibody in 96-well round-bottomed microtiter plates, cells washed three times by centrifugation at 1500 rpm for 5 minutes, and then resuspended in 200 μl of PBS, 5% FCS. In total, 5 µL of PI solution (10 μg/mL PI in PBS) were added to each sample just prior to analysis. Dead cells, doublets, and debris were excluded in all analyses. Flow cytometry analyses were conducted on a FACS LSRII. Cell sorting was performed on a FACSAria II (Becton, Dickinson, and Company). FACS data were collected using FACS Diva version 7.0 or 9.0, and data were analyzed using FlowJo software (versions 9.3.3, 10.1, or 10.2, FlowJo, Ashland, OR, USA). In contour plots, expression levels were shown at 5% probability, unless indicated otherwise in the figure legend. Total thymus cell counts were performed for each animal in presence of trypan blue, showing that none of the silencer mutations alter the absolute number of total thymocytes relative to WT mice (1.6 × 108 +/− 0.3 for 6–10-week-old mice). Furthermore, the frequency of pre-selection (TCRlo CD69−) DP thymocytes is unaffected in any mutant (except for CD4ThPOK.Sil mice), excluding the possibility that mutations of ThPOK silencer cause reduction of these precursors. Immature signaled (TCRβ+ CD69+) and mature (TCRβ+ CD69−) thymocytes are defined according to the gating strategy shown in Supplementary Fig. 10.
EMSA
Nuclear extracts were prepared from human embryonic kidney (HEK) 293T cells cultured on flat-bottom six-well cell culture plates were transfected with Flag-tagged murine Nfat2 or Egr1 constructs (cloned into the pcDNA3 expression vector) in the presence of 10 µg/ml polybrene. Negative controls included nuclear extracts from cells transfected with vector alone. TF expression was verified by immunoblot analysis and used as a protein source for binding assay. DNA-binding probes were generated by annealing of synthetic double-stranded oligonucleotides corresponding to the target region and end-labeling with polynucleotide kinase and digoxigenin-11-ddUTP using EMSA Kit (Sigma). The anti–Flag Ab (Sigma) was used for ‘supershifting’ of TF protein–DNA complexes.
Quantitative RT-PCR
Cell sorting was carried out using a BD FACSAria and FACS Diva software. 100,000 cells were sorted directly into RNA lysis buffer (4 M guanidinium thiocyanate, 25 mM sodium citrate, pH 7.0, 0.5% (wt/vol) N- laurosylsarcosine (Sarkosyl) and 0.1 M 2-mercaptoethanol)45. cDNA was synthesized using the High Capacity RNA-to-cDNA kit (ThermoFisher). For mouse Cd4, we used commercial Taqman (probe-based) assay Mm00442754_m1 (Life Technologies), with the QuantStudio6 thermocycler (Life Technologies) and QuantStudio Realtime PCR Software.
For mThPOK, we performed quantitative RT-qPCR using SYBR Green Master Mix (ThermoFisher) and our own forward and reverse primers8:
ThPOK For 5′-ACCCAACGGCTGAAAGGA-3′
ThPOK Rev 5′-GCTGCTGTGGTCTGGCAAT-3′
Transcript levels are expressed as 2^(-J), where J refers to the difference between Ct (transcript of interest) and Ct (Rps6).
3C analysis
Chromatin crosslinking was performed by adding 9.5 ml of 2% formaldehyde/10% FCS/PBS per 1 × 107 thymocytes from AND Tg+ (class II-restricted), OT-1 Tg+ (class I-restricted), CD3δ−/− (uncommitted; blocked at DP stage), and ThPOKΔSil (DRE knockout) mice, followed by incubation at room temperature for 10 min. The crosslinking reaction was quenched by the addition of 1.425 ml of 1 M glycine (ice cold). Cells were isolated by spinning for 8 min at 225 × g at 4 °C, and supernatants were carefully removed. Cell pellets were resuspended in 5 ml cold lysis buffer (10 mM Tris-HCl, pH 7.5; 10 mM NaCl; 5 mM MgCl2; 0.1 mM EGTA; 1 × complete protease inhibitor; 11836145001 Roche) and incubated for 10 min on ice, and nuclei isolated by centrifugation at 400 × g for 5 min. Quantitative analysis of chromosome conformation capture assays was performed46 using 4 BP cutter NlaIII and ThPOK BAC plasmid was used as the positive control.
Statistics and reproducibility
No statistical method was used to determine the sample size. Instead, sample sizes were rationalized by weighing sufficient replication (to determine the extent of biological variation) with reduction of total animals used. Data were excluded only for technical reasons, such as low cell viability. Regarding replication, all in vivo analyses were performed on a total of three to six animals per genotype (across at least three separate experiments). All attempts at replication were successful. Randomization was not used; assignment to experimental groups was based on genotype. To exclude physiological and environmental covariates, mice of different genotypes were derived from the same litters as control mice (as much as possible), or cohoused prior to analysis. Statistical analysis for nonsequencing data was performed using GraphPad Prism software. Data were analyzed by applying one-way ANOVA with post hoc Tukey HSD (Honest Significant Difference) method. A P value of less than 0.05 was considered significant. *P < 0.05, **P < 0.01, ***P < 0.001.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
References
Jameson, S. C., Hogquist, K. A. & Bevan, M. J. Positive selection of thymocytes. Annu. Rev. Immunol. 13, 93–126 (1995).
Katz, G. et al. T cell receptor stimulation impairs IL-7 receptor signaling by inducing expression of the microRNA miR-17 to target Janus kinase 1. Sci. Signal 7, ra83 (2014).
Park, J. H. et al. Signaling by intrathymic cytokines, not T cell antigen receptors, specifies CD8 lineage choice and promotes the differentiation of cytotoxic-lineage T cells. Nat. Immunol. 11, 257–264 (2010).
Singer, A. & Bosselut, R. CD4/CD8 coreceptors in thymocyte development, selection, and lineage commitment: analysis of the CD4/CD8 lineage decision. Adv. Immunol. 83, 91–131 (2004).
He, X. et al. The zinc finger transcription factor Th-POK regulates CD4 versus CD8 T-cell lineage commitment. Nature 433, 826–833 (2005).
Keefe, R., Dave, V., Allman, D., Wiest, D. & Kappes, D. J. Regulation of lineage commitment distinct from positive selection. Science 286, 1149–1153 (1999).
Park, K. et al. TCR-mediated ThPOK induction promotes development of mature (CD24−) gammadelta thymocytes. EMBOJ 29, 2329–2341 (2010).
He, X. et al. CD4-CD8 lineage commitment is regulated by a silencer element at the ThPOK transcription-factor locus. Immunity 28, 346–358 (2008).
Setoguchi, R. et al. Repression of the transcription factor Th-POK by Runx complexes in cytotoxic T cell development. Science 319, 822–825 (2008).
Grueter, B. et al. Runx3 regulates integrin alpha E/CD103 and CD4 expression during development of CD4−/CD8+ T cells. J. Immunol. 175, 1694–1705 (2005).
Donda, A., Schulz, M., Bürki, K., De Libero, G. & Uematsu, Y. Identification and characterization of a human CD4 silencer. Eur. J. Immunol. 26, 493–500 (1996).
Sawada, S., Scarborough, J. D., Killeen, N. & Littman, D. R. A lineage-specific transcriptional silencer regulates CD4 gene expression during T lymphocyte development. Cell 77, 917–929 (1994).
Siu, G., Wurster, A. L., Duncan, D. D., Soliman, T. M. & Hedrick, S. M. A transcriptional silencer controls the developmental expression of the CD4 gene. EMBOJ 13, 3570–3579 (1994).
Taniuchi, I., Sunshine, M. J., Festenstein, R. & Littman, D. R. Evidence for distinct CD4 silencer functions at different stages of thymocyte differentiation. Mol. Cell 10, 1083–1096 (2002).
Woolf, E. et al. Runx3 and Runx1 are required for CD8 T cell development during thymopoiesis. Proc. Natl Acad. Sci. USA 100, 7731–7736 (2003).
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002).
Rahemtulla, A. et al. Normal development and function of CD8+ cells but markedly decreased helper cell activity in mice lacking CD4. Nature 353, 180–184 (1991).
Matechak, E. O., Killeen, N., Hedrick, S. M. & Fowlkes, B. J. MHC class II-specific T cells can develop in the CD8 lineage when CD4 is absent. Immunity 4, 337–347 (1996).
Koller, B. H., Marrack, P., Kappler, J. W. & Smithies, O. Normal development of mice deficient in beta 2M, MHC class I proteins, and CD8+ T cells. Science 248, 1227–1230 (1990).
Karimi, M. M. et al. The order and logic of CD4 versus CD8 lineage choice and differentiation in mouse thymus. Nat. Commun. 12, 99 (2021).
Stamatoyannopoulos, G., Josephson, B., Zhang, J. W. & Li, Q. Developmental regulation of human gamma-globin genes in transgenic mice. Mol. Cell. Biol. 13, 7636–7644 (1993).
Mookerjee-Basu, J. et al. Functional conservation of a developmental switch in mammals since the Jurassic Age. Mol. Biol. Evol. 36, 39–53 (2019).
Murre, C. Helix-loop-helix proteins and lymphocyte development. Nat. Immunol. 6, 1079–1086 (2005).
Basu, J. et al. Essential role of a ThPOK autoregulatory loop in the maintenance of mature CD4(+) T cell identity and function. Nat. Immunol. 22, 969–982 (2021).
Issuree, P. D., Ng, C. P. & Littman, D. R. Heritable gene regulation in the CD4:CD8 T cell lineage choice. Front. Immunol. 8, 291 (2017).
Jiang, H. & Peterlin, B. M. Differential chromatin looping regulates CD4 expression in immature thymocytes. Mol. Cell. Biol. 28, 907–912 (2008).
Kolovos, P., Knoch, T. A., Grosveld, F. G., Cook, P. R. & Papantonis, A. Enhancers and silencers: an integrated and simple model for their function. Epigenetics Chromatin 5, 1 (2012).
Egawa, T., Tillman, R. E., Naoe, Y., Taniuchi, I. & Littman, D. R. The role of the Runx transcription factors in thymocyte differentiation and in homeostasis of naive T cells. J. Exp. Med. 204, 1945–1957 (2007).
Sato, T. et al. Dual functions of Runx proteins for reactivating CD8 and silencing CD4 at the commitment process into CD8 thymocytes. Immunity 22, 317–328 (2005).
Carleton, M. et al. Early growth response transcription factors are required for development of CD4(-)CD8(-) thymocytes to the CD4(+)CD8(+) stage. J. Immunol. 168, 1649–1658 (2002).
Shaw, J. P. et al. Identification of a putative regulator of early T cell activation genes. Science 241, 202–205 (1988).
Barndt, R., Dai, M. F. & Zhuang, Y. A novel role for HEB downstream or parallel to the pre-TCR signaling pathway during alpha beta thymopoiesis. J. Immunol. 163, 3331–3343 (1999).
Bettini, M., Xi, H., Milbrandt, J. & Kersh, G. J. Thymocyte development in early growth response gene 1-deficient mice. J. Immunol. 169, 1713–1720 (2002).
Lauritsen, J. P. et al. Egr2 is required for Bcl-2 induction during positive selection. J. Immunol. 181, 7778–7785 (2008).
Miyazaki, T. & Lemonnier, F. A. Modulation of thymic selection by expression of an immediate-early gene, early growth response 1 (Egr-1). J. Exp. Med. 188, 715–723 (1998).
Mookerjee-Basu, J. et al. Suppression of Ca(2+) signals by EGR4 controls Th1 differentiation and anti-cancer immunity in vivo. EMBO Rep. 21, e48904 (2020).
Rincon, M. & Flavell, R. A. Regulation of AP-1 and NFAT transcription factors during thymic selection of T cells. Mol. Cell. Biol. 16, 1074–1084 (1996).
Bueno, O. F., Brandt, E. B., Rothenberg, M. E. & Molkentin, J. D. Defective T cell development and function in calcineurin A beta -deficient mice. Proc. Natl Acad. Sci. USA 99, 9398–9403 (2002).
Canté-Barrett, K., Winslow, M. M. & Crabtree, G. R. Selective role of NFATc3 in positive selection of thymocytes. J. Immunol. 179, 103–110 (2007).
Jones-Mason, M. E. et al. E protein transcription factors are required for the development of CD4(+) lineage T cells. Immunity 36, 348–361 (2012).
Alfonso-Jaume, M. A., Mahimkar, R. & Lovett, D. H. Co-operative interactions between NFAT (nuclear factor of activated T cells) c1 and the zinc finger transcription factors Sp1/Sp3 and Egr-1 regulate MT1-MMP (membrane type 1 matrix metalloproteinase) transcription by glomerular mesangial cells. Biochem. J. 380, 735–747 (2004).
Basson, M. A. et al. Early growth response (Egr)-1 gene induction in the thymus in response to TCR ligation during early steps in positive selection is not required for CD8 lineage commitment. J. Immunol. 165, 2444–2450 (2000).
Brugnera, E. et al. Coreceptor reversal in the thymus: signaled CD4+8+ thymocytes initially terminate CD8 transcription even when differentiating into CD8+ T cells. Immunity 13, 59–71 (2000).
Dave, V. P. et al. CD3 delta deficiency arrests development of the alpha beta but not the gamma delta T cell lineage. EMBOJ 16, 1360–1370 (1997).
Chomczynski, P. & Sacchi, N. The single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction: twenty-something years on. Nat. Protoc. 1, 581–585 (2006).
Hagège, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nat. Protoc. 2, 1722–1733 (2007).
Acknowledgements
This work was supported by NIH grants R01 AI068907 (D.J.K.), R01 GM107179 (D.J.K.), and P30 CA006927 (FCCC Comprehensive Cancer Center Core Grant). J.B. received a fellowship from W. J. Avery foundation. We acknowledge the assistance of the following core facilities of the Fox Chase Cancer Center: Flow Cytometry, Cell Culture, DNA Sequencing, and Laboratory Animal. We thank D. Wiest and K. Hogquist for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.B. and D.J.K.; methodology: J.B., D.J.K., and X.H.; investigation, J.B., J.Z., E.N., M.C., P.C., X.H., L.G., and D.J.K.; writing—original draft: J.B. and D.J.K.; writing—review & editing: J.B. and D.J.K.; funding acquisition: D.J.K.; supervision: D.J.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: George Inglis.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Basu, J., Zha, J., Nicolas, E. et al. An autonomous TCR signal-sensing switch influences CD4/CD8 lineage choice in mice. Commun Biol 5, 84 (2022). https://doi.org/10.1038/s42003-022-02999-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-02999-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.