Abstract
During ontogeny, macrophage populations emerge in the Yolk Sac (YS) via two distinct progenitor waves, prior to hematopoietic stem cell development. Macrophage progenitors from the primitive/”early EMP” and transient-definitive/”late EMP” waves both contribute to various resident primitive macrophage populations in the developing embryonic organs. Identifying factors that modulates early stages of macrophage progenitor development may lead to a better understanding of defective function of specific resident macrophage subsets. Here we show that YS primitive macrophage progenitors express Lyl-1, a bHLH transcription factor related to SCL/Tal-1. Transcriptomic analysis of YS macrophage progenitors indicate that primitive macrophage progenitors present at embryonic day 9 are clearly distinct from those present at later stages. Disruption of Lyl-1 basic helix-loop-helix domain leads initially to an increased emergence of primitive macrophage progenitors, and later to their defective differentiation. These defects are associated with a disrupted expression of gene sets related to embryonic patterning and neurodevelopment. Lyl-1-deficiency also induce a reduced production of mature macrophages/microglia in the early brain, as well as a transient reduction of the microglia pool at midgestation and in the newborn. We thus identify Lyl-1 as a critical regulator of primitive macrophages and microglia development, which disruption may impair resident-macrophage function during organogenesis.
Similar content being viewed by others
Introduction
Amongst the components of the transcription factor network that regulate hematopoietic cells features, Tal-1, Lmo2, Runx1, and Gata2 stand out as major regulators of hematopoietic progenitor development1,2. Tal-1, Lmo2, and Gata-2 belong to a transcriptional complex, which also includes the basic helix–loop–helix (bHLH) transcription factor lymphoblastic leukemia-derived sequence 1 (Lyl-1). Unlike its paralog Tal-1, which is mandatory for the specification of all hematopoietic progenitors3,4, Lyl-1 roles during developmental hematopoiesis remains poorly characterized. We analyzed these functions at the onset of YS hematopoietic development using Lyl-1LacZ/LacZ mutant mice5.
During ontogeny, hematopoietic progenitors are generated in three successive and overlapping waves6,7. The emergence of the Hematopoietic Stem Cells (HSC) that will maintain lifelong hematopoiesis in the adult occurs at mid-gestation in the third and definitive hematopoietic wave. HSC generated in the aorta region immediately migrate to the fetal liver (FL) where they mature and amplify before homing to the bone marrow before birth8,9. Prior to HSC generation, the production of blood cells relies on two hematopoietic waves provided by the YS. This HSC-independent hematopoiesis comprises first the primitive hematopoietic wave, with the transient production of progenitors with embryonic specific features: From Embryonic day (E) 7.00, the YS produces monopotent progenitors for erythrocytes, megakaryocytes and macrophages (MΦ)10,11, along with bipotent Erythro-Megakaryocytic progenitors12, in a Myb-independent pathway13,14. The second YS wave, called transient-definitive, provides for a limited duration progenitors (mostly erythro-myeloid) that seed the FL and produce a hematopoietic progeny that displays definitive/adult differentiation features. Erythro-myeloid cell production in this wave occurs in a Myb-dependent pathway, through the progressive differentiation of erythro-myeloid progenitors (EMP) in a pathway similar to the adult one6. As primitive and transient definitive YS waves both produce cells from erythro-myeloid lineages, they are also termed respectively “early EMP” and “late EMP”15,16.
Considering the MΦ lineage, fate-mapping approaches aimed at determining the embryonic origin of resident-MΦs indicated that most tissues harbor resident-MΦs of diverse origins (YS, FL and adult bone marrow)15,17,18, which complicates the characterization of wave-dependent functions of the various subsets. However, these fate-mapping analyses established that, contrary to others tissue, brain MΦs (microglia and Border Associated MΦ (BAM)) develop only from YS-derived MΦ-progenitors14,16,19,20,21, confirming a model we previously put forward22. Due to the coexistence of two waves in the YS, the origin of microglia has been debated (reviewed in refs. 6,15,18,23). An origin of microglia from MΦ-progenitors from the primitive/”early EMP” wave was supported by microglia labeling following an early (E7.0-E7.5) CRE-mediated induction of Runx119 and by the intact microglia pool in Myb-deficient mice14,21. The origin of microglia was also attributed to the primitive wave in zebrafish embryos, since primitive macrophage (MΦPrim) progenitors arise in this species from a location distinct from other hematopoietic progenitors24,25,26. Finally, the normal microglia development in mice lacking Kit ligand, leading to an impaired EMP development and the depletion of resident-MΦs in the skin, lung, and liver supports this model27.
We here show that, at the early stages of YS hematopoiesis, Lyl-1 expression characterizes primitive MΦ-progenitors. Through RNA-seq. analyses, it appears that these primitive MΦ-progenitors harbor an immune-modulatory phenotype, while those produce at a later stage favor the inflammatory signaling which promotes the emergence of HSC in the third and definitive hematopoietic wave28.
Our results also indicate that in the brain, Lyl-1 is expressed in the entire microglia/BAM cell population at the onset of brain colonization and appeared to regulate microglia/BAM development.
Altogether, these data point to Lyl-1 as a major regulator of early embryonic MΦ-progenitors development and advocate for further analyses to more precisely delineate Lyl-1 function during the development of resident-MΦs in homeostatic and pathological contexts.
Results and discussion
Lyl-1 expression marks MΦPrim progenitors from the early YS
Lyl-1 being expressed in the YS from the onset of YS hematopoiesis29, we first explored its function by characterizing the clonogenic potential of WT, Lyl-1WT/LacZ and Lyl-1LacZ/LacZ YS. E8-YS were maintained in organ culture for 1 day (E8 OrgD1-YS), allowing only the development of primitive and transient-definitive progenitors30,31. Compared to WT, the production of MΦ progenitors was increased in Lyl-1WT/LacZ and Lyl-1LacZ/LacZ OrgD1-YS. Otherwise, the clonogenic potential and progenitor distribution were unmodified (Fig. 1a).
a Lyl-1-deficiency leads to an increased production of MΦ-progenitors in the early YS: Clonogenic potential of E8 OrgD1-YS cells: production of MΦ-progenitors (MΦP) in WT, Lyl-1WT/LacZ and Lyl-1LacZ/LacZ OrgD1-YS. Distribution of other progenitors with a myeloid potential (EMP and GM) in WT, Lyl-1WT/LacZ and Lyl-1LacZ/LacZ E8 OrgD1-YS. (n = 3–5, 3–6 YS per sample; mean ± s.e.m.; Unpaired, two-tailed t-test). b Lyl-1 expression in MΦ-progenitors: FACS-Gal assay, using the β-Gal fluorescent substrate FDG, was used as a reporter for Lyl-1 expression. While all MΦ-progenitors in E9-YS expressed FDG/Lyl-1, E9.5 and E10-YS harbored two MΦ-progenitor subsets discriminated by their FDG/Lyl-1 expression. FDG+/Lyl-1+ and FDG-/Lyl-1- mature MΦs (CD11b+F4/80+) also coexisted in E10-YS. The contour plots in WT samples indicate the level of non-specific background β-Gal activity/FDG labeling in WT samples. Representative profiles of 3 independent samples, each consisting of 3–4 YS (see gating strategy in Supplementary Fig. 1a). c In clonogenic assays, the number of MΦ-progenitors obtained from E8-YS (0-3 Somites) was increased in Lyl-1WT/LacZ and Lyl-1LacZ/LacZ compared to WT. The majority of the 25–30 progenitors per YS were EryP (60 to 80% in each 3 genotypes). Other progenitors were occasionally and randomly detected in WT and mutant samples (less than one EMP (0.81% ± 0.66; n = 3) and/or GM progenitor per E8-YS), confirming that the assay was performed at a time when MΦT-Def progenitors were absent. (n = 3–4, 5–10 YS per sample; error bars show mean ± s.e.m.; Unpaired, two-tailed t-test). d MΦPrim progenitors express Lyl-1. Flow cytometry profiles of WT and Lyl-1WT/LacZ E8-YS (0-3 S). CD11b+CD31- MΦs correspond to maternal MΦs present in E8-YS11. All CD11b+CD31+ MΦ-progenitors displayed FDG/Lyl-1 expression (Red line). The contour plots in WT samples indicate the level of non-specific background β-Gal activity/FDG labeling in WT samples. Shown are representative profiles of 4 independent experiments. e RT-qPCR quantification of Myb expression levels: Kit+CD45+CD11b+ progenitors were sorted from WT E9-YS, WT and Lyl-1WT/LacZ E10-YS, and from FDG/Lyl-1 positive and negative fractions of MΦ-progenitors from Lyl-1WT/LacZ E10-YS. Lin-Sca+Kit+ (LSK) progenitors from WT E12-FL were used as positive control. FDG+/Lyl-1+ MΦ-progenitors from E10-YS expressed MybLow/Neg levels similar to MΦPrim progenitors from E9-YS. The FDG-/Lyl-1- fraction expressed significantly higher Myb levels, similar to LSK cells from E12-FL. Myb expression levels, shown on a Log2 scale, were normalized to the mean expression value obtained for WT E10-YS, considered as 1 (Each dot represent an independent experiment; Error bars show means ± s.e.m; Unpaired, two-tailed t-test). f FDG/Lyl-1 positive and negative myeloid progenitors produce a distinct progeny: Clonogenic assays characterization of the type of progenitors produced by myeloid progenitors (Ter119-Kit+CD45+CD11b+) sorted from WT and Lyl-1WT/LacZ E9-YS ( < 18 S; n = 7) and E10-YS (n = 15) in 3 independent experiments. At E10, myeloid progenitors from Lyl-1WT/LacZ YS were subdivided into FDG/Lyl-1 negative (n = 15) and positive (n = 12) fractions (5 independent experiments). Samples were biological replicates comprising 6–8 YS. 100 to 150 Kit+CD45+CD11b+ cells per condition were platted in triplicate. All samples produced few non-myeloid contaminants, such as Erythro-Megakaryocytic progenitors and EMP in similar, non-significant amounts. FDG+/Lyl-1+ progenitors essentially produced MΦ−progenitors (MΦP), while FDG-/Lyl-1- progenitors produced also granulo-monocytic- (GMP) and granulocyte- (GP) progenitors. Error bar shows means ± s.e.m.
Using FACS-Gal assay, we noticed that the entire MΦ-progenitor population (Kit+CD45+CD11b+) expressed Lyl-1 at E9. In contrast, two MΦ-progenitor subsets, discriminated by FDG/Lyl-1 expression, were present after E9.5 (Fig. 1b). Since after E9.5, the YS harbors both MΦPrim and transient-definitive (MΦT-Def) MΦ−progenitors, and as these progenitor subsets cannot be discriminated by phenotype11, we investigated the known features discriminating these two waves: the origin from monopotent progenitors for MΦPrim progenitors10,11 and the Myb-dependent14,32 differentiation of MΦT-Def progenitors from EMP, via the production of granulo-monocytic (GM-), then granulocyte (G-) and MΦ-progenitors6.
At E8 (0-5 somites), when the YS only harbors MΦPrim progenitors, clonogenic assays also pointed to an increased production of MΦ-progenitors in mutant E8-YS compared to WT (Fig. 1c). At this stage, all MΦ-progenitors, which harbor a CD11b+CD31+ phenotype11, expressed FDG/Lyl-1 (Fig. 1d). Most FDG+/Lyl-1+ CD11b+CD31+ cells (69.27%±0.33%) from Lyl-1WT/LacZ E8-YS reliably produced MΦ colonies (72.78 ± 9.65%; n = 3) in clonogenic assays, amounting 1-4 MΦ progenitors per YS, a value consistent with previously published data10,11.
Lyl-1 expression by MΦPrim progenitors was strengthened by RT-qPCR comparison of Myb expression: Both E9-YS MΦPrim progenitors and FDG+/Lyl-1+ progenitors from E10-YS expressed low Myb levels strengthening their primitive status, while FDG-/Lyl-1- progenitors from E10-YS progenitors displayed Myb levels similar to lineage-negative Sca1+Kit+ progenitors from E12-FL (Fig. 1e).
The differentiation potential of FDG+/Lyl-1+ and FDG-/Lyl-1- fractions of Ter119-Kit+CD45+CD11b+ myeloid progenitors isolated from E10-YS also pointed to Lyl-1 expression by MΦPrim progenitors (Fig. 1f): Similar to WT E9-YS MΦPrim progenitors, E10 FDG+/Lyl-1+ progenitors appeared monopotent, as they nearly exclusively produced MΦ colonies. In contrast, E10 FDG-/Lyl-1- myeloid progenitors produced GM, G and MΦ colonies, a feature typical of transient-definitive progenitors6. Overall, these data together suggested that Lyl-1 may mark MΦPrim progenitors from the earliest wave.
Distinct features of WT MΦ-progenitors at E9 and E10
The distinction between E9 and E10 MΦ-progenitors was confirmed in RNA-seq. analysis of CD45+CD11b+Kit+ MΦ-progenitors sorted at E9 (MΦPrim progenitors) and E10 (MΦPrim and MΦT-Def progenitors). Principal Component Analysis separated E9 and E10 MΦ-progenitors according to stage and genotypes (Fig. 2A). E9 and E10 WT MΦ-progenitors differed by the expression of 726 genes, 176 being up-regulated at E9 and 550 at E10. Considering the coexistence of MΦPrim and MΦT-Def progenitors in E10-YS, differentially expressed genes found at E10 may reflect wave-specific differences or stage-dependent changes related to MΦPrim progenitor maturation.
A Differentially expressed genes in MΦ-progenitors (Kit+CD45+CD11b+) sorted from WT and Lyl-1LacZ/LacZ YS at E9 and E10. Unsupervised principal component analysis plot positioned E9 and E10 MΦ-progenitors in two distinct groups, followed by segregation of WT and Lyl-1LacZ/LacZ samples. Volcano plot of E9 WT vs E10 WT MΦ-progenitors: Red and green dots indicate genes with statistically significant changes in expression level. (p-value < 0.05, absolute fold change≥2.0) (NDE: not deregulated genes; DE-Up: up-regulated genes; DE-Down: down-regulated genes). B Venn diagram comparing differentially expressed genes in E9 WT vs E10 WT MΦ-progenitors to the EMP or MΦ signatures defined by Mass et al.33 (GEO accession number GSE81774). The number and percentage of differentially expressed genes common to the EMP or MΦ signatures is shown. Expression profiles of the overlapping genes identified in the Venn diagram (Heatmap displays transformed log2-expression values; Unpaired t-test, two-tailed). Note the higher expression at E10 of genes involved in erythroid (Globins: Pink arrow; Transcription factors: green arrow), and megakaryocytic and granulocytic-related genes (blue arrow), and of Mrc1/CD206 (Asterisk). C Relative expression levels of Gata1 and Spi1/PU.1, indicated by their relative Transcripts per million kilo-bases. D Enriched pathways in E9 and E10 WT MΦ-progenitors with absolute z-score ≥2, from QIAGEN’s Ingenuity® Pathway Analysis. Bars: minus log of the p-value of each canonical pathway; Orange line: threshold p-value of 0.05. Ratio: genes detected/genes per pathway. E Expression profiles of differentially expressed genes related to IFNγ and IFNβ response, identified by g:Profiler (heatmap displays transformed log2-expression values; unpaired t-Test, two-tailed). F Expression profiles of differentially expressed genes related to MHC-II complex (Heatmap displays transformed log2-expression values; unpaired t-Test, two-tailed). G Expression profiles of differentially expressed genes related to cytokine signaling (Heatmap displays transformed log2-expression values; unpaired t-Test, two-tailed).
Overlapping the identified differentially expressed genes to the EMP and E10.25-E10.5 MΦs signatures obtained by Mass et al.33 confirmed that WT E9 MΦPrim progenitors were distinct from these two populations, since none of the 176 genes upregulated at E9 belonged to these signatures. Comparatively, about 5% of the genes up-regulated at E10 belonged to the EMP and MΦ signatures (Fig. 2B). A similar separation was observed in GSEA analyses (Supplementary Fig. 1b). These observations suggest that within E10 MΦ-progenitors some, likely the MΦT-Def ones, retain part of the EMPs signature.
In a WT context, E9 MΦPrim progenitors differed from E10 MΦ-progenitors by their transcription factors repertoire. Genes regulating erythroid development (Gata1, Gata2, Klf1) and globin genes, embryonic (Hbb-bh1, Hba-x, Hbb-y) and definitive (Hba-a2, Hba-a1, Hbb-bt), were enriched at E10 (Fig. 2B), while Spi1/PU.1 was highly expressed compared to Gata1 at both stages (Fig. 2C). The lower expression level at E9 of erythroid genes and of genes involved in granulo-monocytic (Mpo, Csf2r/GM-CSF receptors, Cebp, Jun) and megakaryocytic development (Pf4, TPO signaling) (Fig. 2B; Supplementary Table 1) sustains the monopotent/primitive status of E9 MΦ-progenitors, and suggests that MΦT-Def progenitors may retain the expression of genes that characterize their EMP ancestor.
Due to the simultaneous presence of MΦPrim and MΦT-Def progenitors in E10-YS, the differential expressions of markers that are wave-specific at these stages (Primitive: Lyl-1; transient-definitive: Myb and Tlr234) were not significant despite a tendency to decrease for Lyl-1 and increase for Myb and Tlr2 (Supplementary Fig. 1c).
IPA and GSEA analyses indicated that E9 MΦPrim progenitors were more active in Eicosanoid signaling than E10 progenitors (Fig. 2D). They were also enriched in type I interferon (IFN) β and type II IFNγ signaling (Fig. 2E) and in MHC-II related genes, especially Cd74 (top 1 IPA network) (Fig. 2F; Supplementary Fig. 1d). Cytometry analyses confirmed a low, but significant, enrichment of MHC-II expression at E9, compared to E10 (Supplementary Fig. 1e). Comparatively, E10 MΦ-progenitors were more active in inflammatory signaling (Fig. 2D, E; Supplementary Fig. 1d, f–h; Supplementary Table 1), and metabolically active (Supplementary Table 1). The complement cascade and phagocytosis also prevailed at E10 (Supplementary Fig. 1i, j).
Altogether, the signature for WT E9 MΦPrim progenitors points to an immuno-modulatory function, while E10 MΦ-progenitors appear involved in phagocytosis and inflammatory signaling. Interestingly, inflammatory signaling has been revealed as a key factor favoring embryonic HSC emergence (reviewed in ref. 35). The source of inflammatory signals was further identified as MΦ-progenitors expressing Mrc1/CD20636, a marker up-regulated in E10 WT MΦ-progenitors compared to E9 (Fig. 2B).
Lyl-1 deficiency impacts embryonic development
When evaluating the effect of Lyl-1-deficiency at the earliest stage of MΦPrim development, clonogenic assays pointed to an increased production of MΦ-progenitors in Lyl-1LacZ/LacZ MΦPrim E8-YS compared to WT (Fig. 1c). The analysis of the RNA-seq. comparison of E9 WT and Lyl-1LacZ/LacZ MΦPrim suggests that the increased size of the initial MΦ-progenitor pool could results from an elevated commitment of mesodermal/pre-hematopoietic cells to a MΦ fate, rather than from a defective differentiation (Supplementary Fig. 2a–d and Supplementary note 1). The high increase of Itga2b/CD41 expression level in E9 Lyl-1LacZ/LacZ MΦPrim progenitors (Fig. 3a) may reflect this elevated commitment. Lyl-1 expression in YS mesoderm, where it cannot substitute for Tal-1 mandatory function for the generation of hematopoietic progenitors4, was already established3,29. Recently, Lyl-1 was identified as a regulator of mesoderm cell fate37 and of the maintenance of primitive erythroid progenitors38.
a Relative expression levels of the CD41 coding gene Itg2b in WT and Lyl-1LacZ/LacZ MΦ-progenitors at E9 (error bars show mean ± s.e.m.; unpaired t-Test, two-tailed). b Relative expression levels of transcription factors regulating hematopoietic progenitor emergence in Lyl-1LacZ/LacZ MΦ-progenitors compared to WT at E9 (Error bars show mean ± s.e.m.; unpaired, two-tailed t-Test). c GSEA pathways (FDR q-value <0.29) and GO terms (FDR q-value < 0.01) enriched in E9 Lyl-1LacZ/LacZ compared to E9 WT MΦ-progenitors. Highlighted are the pathways specifically related to embryo patterning (blue) and to the development of skeletal (green) and nervous systems (yellow). Pink arrows point to changes related to metabolic pathways.
The transcription factors network that controls developmental hematopoiesis2 was also modified (Fig. 3b): beside the expected reduction of Lyl-1 expression, the expression of Lmo2, a Lyl-1 target39, was down-regulated, while Tal-1 up-regulation might reflect some compensatory function3. The consequences were apparent in GSEA analyses: both pathways and GO terms uncovered an up-regulation of signaling pathways involved in embryo patterning (Wnt, Hox and Smad) in Lyl-1LacZ/LacZ MΦ-progenitors, as well as a highly modified collagen, integrin, and cadherin usage (Supplementary Table 2). Accordingly, developmental trajectories were affected (Fig. 3c), with the up-regulation in E9 Lyl-1LacZ/LacZ MΦ-progenitors of gene sets related to “anterior-posterior pattern specification” and “anatomical structure formation involved in morphogenesis”, notably skeletal and nervous system development.
GSEA and KEGG comparison of Lyl-1LacZ/LacZ and WT MΦ-progenitors at E10 highlighted another patterning modification, namely the down-regulation of gene sets involved in heart development (Supplementary Fig. 2e; Supplementary Table 3), which might stem from a defective MΦ development. The heart harbors three resident-MΦ subsets, two of which originate from the YS40. Amongst the features that distinguish WT E9 MΦPrim progenitors from E10 MΦ-progenitors, the enriched expression of MHC-II (Fig. 2F) and poor expression of phagocytosis-related genes (Supplementary Fig. 1i) at E9 also characterize one of the two YS-derived CCR2- resident-MΦ subsets in the heart40,41. Therefore, a function for Lyl-1+ MΦPrim progenitors in heart development may be considered. This observation reinforces the need to better characterize the contributions of MΦ-progenitors from both primitive and transient-definitive waves to tissues harboring YS-derived resident-MΦs.
The patterning defects highlighted in defective MΦPrim, might be responsible, at least in part, for the significantly decreased litter size and increased perinatal lethality observed in Lyl-1LacZ/LacZ mice compared to WT (Supplementary Fig. 2F), which indicates the requirement for Lyl-1 during various developmental processes. Whether this function is specific to mesodermal stage or related to a patterning function for MΦPrim progenitors remains to be determined. Other Lyl-1-expressing lineages29,42 might also participate to impaired development processes in Lyl-1LacZ/LacZ embryos, calling for the development of new mouse models that will allow lineage specific reports of Lyl-1 expression and function in endothelial or hematopoietic lineages during discrete steps of early YS development and beyond.
Defective MΦPrim development in Lyl-1 mutant YS
The analysis of Lyl-1 expression in A1-A2-A3 subsets from Cx3cr1WT/GFP YS indicated that Lyl-1 is expressed throughout MΦ-progenitor differentiation, with levels decreasing from A1 to A3 (Supplementary Fig. 3a). We monitored the distribution of A1-A2 and A3 MΦ subsets (Supplementary Fig. 2a) at E10-YS, when all three subsets are present, using the Cx3cr1WT/GFP:Lyl-1LacZ/LacZ strain. While the size of the whole MΦ population was not overtly modified, Lyl-1-deficiency impacted the subset distribution, with increased A1 and reduced A2 and A3 pool sizes (Fig. 4A). Lyl-1 thus appears to regulate MΦ differentiation towards mature MΦs. A limited/delayed differentiation of E9 Lyl-1LacZ/LacZ MΦPrim progenitors could result from an altered cytokine signaling: Signaling through Jak-Stat pathway (Jak1-Stat1/6), which underlays cytokine signaling in hematopoietic cells, was down-regulated In E9 mutant progenitors, (Fig. 4B; Supplementary Fig. 3b), with profoundly decreased expression of interleukins and their receptors (Il1a-b/Il1r1-2; Il4i1/Il4Ra; Il6st/Il6ra; Il10/ Il10ra). It was also supported by a down-regulated Spi1/PU.1 signaling pathway (Supplementary Fig. 3c; Supplementary Tables 4–5) and by the decreased expression of Ptprc/CD45, Csf1r, Itgam/CD11b and CD33 (Fig. 4C).
A Distribution of A1-A2 and A3 MΦ subsets in E10-YS from Cx3cr1WT/GFP:Lyl-1WT/WT, Cx3cr1WT/GFP:Lyl-1WT/LacZ and Cx3cr1WT/GFP: Lyl-1LacZ/LacZ embryos. While the size of the whole MΦ population is similar in the three genotypes, Lyl-1-deficiency leads to a modified distribution of the MΦ subsets with an increased size of the A1 subset and a reduced A3 pool (5–12 independent analyses, 6-8 YS per sample; Plots show mean ± s.e.m.; Unpaired, two-tailed t-Test). B GSEA pathway indicates a deficit in Jak1-Stat signaling in Lyl-1LacZ/LacZ MΦ-progenitors compared to WT at E9 (NES: normalized enrichment score; FDR: false discovery rate). C Relative expression levels (read counts) of hematopoietic markers in WT and Lyl-1LacZ/LacZ MΦ-progenitors at E9 (Error bars show mean ± s.e.m.; unpaired t-Test, two-tailed). D Top 1 GSEA pathway indicates that the IFN signaling pathway, which characterize E9 MΦPrim progenitors, is defective in Lyl-1LacZ/LacZ MΦ-progenitors (NES: normalized enrichment score; FDR: false discovery rate). The expression of Irf8 is particularly affected (error bars show mean ± s.e.m.; unpaired t-test, two-tailed). E From the 53 canonical pathways identified by Ingenuity® Pathway Analysis as differentially expressed, 9 were enriched with an absolute Z score ≥ 1. Bars: minus log of the p-value of each canonical pathway; Orange line: threshold p-value of 0.05. Ratio: genes detected/genes per pathway. F Venn diagram comparing the differentially expressed genes in E9 Lyl-1LacZ/LacZ vs E9 WT to those in E10 Lyl-1LacZ/LacZ vs E10 WT MΦ-progenitors. Expression profiles of the differentially expressed genes common to both stages identified by the Venn comparison (Heatmap displays transformed log2-expression values; unpaired t-Test, two-tailed). G Relative expression levels (read counts) of phagocytosis genes in WT and Lyl-1LacZ/LacZ MΦ-progenitors at E10 (Box and whisker plots, min to max; median is shown; two tailed, unpaired Mann–Whitney test).
Within the MΦ lineage, Lyl-1 function during normal development would initially consist to restrict the size of the MΦPrim progenitor pool and/or the duration of its production, which is transient6, as indicated by the maintenance of the intermediate mesoderm to MΦ-progenitor pool observed in Lyl-1LacZ/LacZ E8-YS. Indeed, the increased size of the MΦ-progenitor pool in E8-E9 YS appears independent from the defective/delayed differentiation of MΦ-progenitors observed at E10, since this process starts after E9.510,11,15. Subsequently, the increased size of MΦPrim progenitor pool in E10 Lyl-1LacZ/LacZ YS likely results from a defective/delayed differentiation mediated by a defective cytokine signaling, implying that during normal development, Lyl-1 would promote the differentiation of MΦPrim progenitors.
Lyl-1LacZ/LacZ MΦ-progenitors were also deficient in the IFN signaling that characterize E9 MΦPrim progenitors, notably Irf8, a factor involved in YS-MΦ and microglia development21,43 (Fig. 4D). Compared to WT, MΦ-progenitors from Lyl-1LacZ/LacZ E9-YS up-regulated the LXR/RXR activation pathway (Fig. 4E) and metabolic pathways, some enriched WT MΦ-progenitors at E10 (Butanoate, steroid) (Supplementary Table 1), and other not (Fructose/mannose, fatty acid) (Fig. 3c). They were also less active in inflammatory signaling pathways, particularly through NFκb, a factor known to interact with Lyl-144, and in TLR signaling (Fig. 4D; Supplementary Fig. 3b, d–e; Supplementary Table 5).
Overlapping the differentially expressed genes identified in Lyl-1LacZ/LacZ MΦ-progenitors at E9 and E10 identified the core signature of Lyl-1-deficiency, independent of the maturation occurring between these stages (Fig. 4F). Unfortunately, the co-existence of MΦPrim and MΦT-Def progenitors in E10-YS complicates the attribution of gene expression changes to a stage-dependent maturation of MΦPrim progenitors or to a signature specific to MΦT-Def progenitors. However, most pathways favored by E10 progenitors were insensitive to Lyl-1-deficiency, except for a decreased expression of phagocytosis genes (Axl1, Mertk, Trem2, P2y6r/P2ry6) (Fig. 4G). Conversely, the TLR signaling pathway was increased in Lyl-1LacZ/LacZ MΦ-progenitors at E10, compared to E9.
Lyl-1-expressing MΦ-progenitors contribute to the fetal liver and brain
The FL9 and brain19,22 are colonized as early as E9 by YS-derived resident-MΦ progenitors. We evaluated the contribution of Lyl-1-expressing MΦPrim progenitors to these rudiments at E10 (Fig. 5A). While E10-YS comprised FDG+/Lyl-1+ and FDG-/Lyl-1- MΦ-progenitors and mature (F4/80+) MΦ subsets (Fig. 1b), the brain from the same embryos essentially harbored FDG+/Lyl-1+ MΦ-progenitors and MΦs. In contrast, both FDG+/Lyl-1+ and FDG-/Lyl-1- MΦ-progenitors were present in E10-FL, as in E10-YS, and MΦ-progenitors were more abundant in mutant FL than in WT (Fig. 5B).
A All MΦ-progenitors from E10-Brain expressed Lyl-1, contrary to the corresponding YS (Fig. 1b) which harbor both FDG+/Lyl-1+ and FDG-/Lyl-1- subsets. MΦ-progenitors from E10 Lyl-1LacZ/LacZ FL harbored both FDG+/Lyl-1+ and FDG-/Lyl-1 MΦ-progenitor subsets. in E10-brain, mature MΦs (CD11b+F4/80+ gate) were all FDG+/Lyl-1+. The contour plots in WT samples indicate the level of non-specific background β-Gal activity/FDG labeling in WT samples. Representative profiles of 3 independent samples, each consisting of 3-4 E10-Brain or 8-12 E10-FL. B Quantification of Kit+CD45+CD11b+ MΦ-progenitors in E10-FL (plots show mean ± s.e.m.; Unpaired, two-tailed t-Test). C Lyl-1 marks the entire F4/80 + microglia/BAM population from the onset of brain colonization. FDG/Lyl-1 expression in F4/80+ microglia/BAM from the brain of Lyl-1WT/LacZ and Lyl-1LacZ/LacZ embryos at E9 to E11. The rare CD11b+ F4/80low-neg cells present in the brain at E9 are FDG+/Lyl-1+. Gray histograms indicate non-specific background β-Gal activity/FDG levels in WT samples. Representative profiles of 3–5 independent samples, each consisting of 3-5 brains. D MΦ-progenitors from E10-brain express Myb levels similar to E9-YS MΦPrim progenitors. RT-qPCR quantification of Myb expression levels in Kit+CD45+CD11b+ MΦ-progenitors sorted from WT E9-YS and from WT and Lyl-1WT/LacZ brain at E10. Lin-Sca+cKit+ (LSK) progenitors from WT E12-FL were used as positive control. Myb expression levels, shown on a Log2 scale, were normalized to the mean expression value obtained for WT E10-YS, considered as 1 (error bars show mean ± s.e.m.; Unpaired, two-tailed t-Test). E RT-qPCR analyses of Lyl-1 expression in A1 to A3 MΦ subsets isolated from Cx3cr1WT/GFP brain at E10. Lyl-1 is expressed by the 3 subsets, with levels decreasing with differentiation. Expression levels were normalized to the mean value obtained for Cx3cr1WT/GFP YS A1 progenitors (n = 2; Error bars show mean ± s.e.m). F Defective differentiation of brain MΦ-progenitor in Lyl-1 mutant embryos. Distribution of A1-A2 and A3 MΦ subsets in E10 brain from Cx3cr1WT/GFP:Lyl-1WT/WT, Cx3cr1WT/GFP:Lyl-1WT/LacZ and Cx3cr1WT/GFP:Lyl-1LacZ/LacZ embryos. The size of the whole MΦ population was similar in the three genotypes, but Lyl-1 deficiency modified the distribution of the MΦ subsets with an increased size of the A1 subset and a reduced A3 pool (5-12 independent analyses, 6–8 brains per sample. Error bars show mean ± s.e.m.; Unpaired, two-tailed t Test). G Heatmap expression profile of the genes that mark the development of tissue resident-MΦs in WT E9 and E10 MΦ-progenitors (Heatmap displays transformed log2-expression values; Unpaired, two-tailed t-test).
We next focused on brain MΦs during the colonization stage, which lasts until E1145. At this stage, microglia and perivascular, meningeal and choroid plexus MΦs, collectively referred to as BAMs, are all located in the brain mesenchyme and therefore undistinguishable16,46. FACS-Gal assay demonstrated that the whole F4/80+ microglia/BAM expressed Lyl-1 throughout the settlement period (Fig. 5C). The presence of FDG+/Lyl-1+F4/80+ microglia/BAM at early stage of brain colonization suggests that MΦs could participate to this step.
Lyl-1+ MΦPrim progenitors and early microglia/BAM shared similar features, such as an early appearance timing and low level of Myb expression (Fig. 5D), concordant with a Myb-independent development of microglia14,21. Lyl-1 was also similarly expressed in A1-A2 and A3 MΦ subsets from the YS and brain (Fig. 5E; Supplementary Fig. 3a). Lyl-1-deficiency impacted the distribution of MΦs subsets in E10 Cx3cr1WT/GFP:Lyl-1LacZ/LacZ brain: an increased A1 and a reduced A3 pool size indicated that Lyl-1 regulates MΦ-progenitor differentiation in both YS and brain (Fig. 5F).
The proximity between YS MΦPrim progenitors and microglia was also apparent in RNA-seq. data: E9 WT MΦPrim progenitors expressed significantly lower Mrc1/CD206 and higher Sall3 levels than E10 MΦ-progenitors, and a slightly increased Sall1 level (Fig. 5G), a transcriptomic pattern that characterizes microglia33,45,47. This partial bias toward a microglia signature suggests that the first stage of microglia development program is already initiated in MΦPrim /“early EMP”progenitors in E9-YS.
Lyl-1 inactivation impairs microglia development at two development stages
Having defined Lyl-1 implication during microglia/BAM settlement in the brain, we turned to later development stages. Cytometry and database analyses45 confirmed the continuous expression of Lyl-1 in CD45low microglia until adulthood (Supplementary Fig. 4a). LYL-1 expression was also reported in microglia from healthy murine and human adults48,49,50,51. We examined the impact of Lyl-1-deficiency on microglia pool size during development. Microglia quantification pointed to E12 as the first step impacted. The arrested increase of microglia pool in Lyl-1LacZ/LacZ brain at E12 (Fig. 6a) resulted from a reduced proliferation (Fig. 6b) rather than an increased apoptosis (Supplementary Fig. 4b). Moreover, Lyl-1-deficiency provoked morphological changes in E12 Cx3cr1WT/GFP:Lyl-1LacZ/LacZ, compared to Cx3cr1WT/GFP microglia, namely a reduced number and extent of ramifications (Fig. 6c; Supplementary Fig. 4c, d). From E14, the microglia pool size returned to levels similar to WT (Fig. 6a), probably due to the highly reduced apoptosis level in Lyl-1LacZ/LacZ microglia at E14 (Supplementary Fig. 4b).
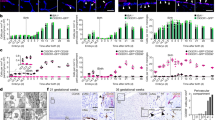
a Quantification of the microglia population in E12 and E14 brain showing the decreased size of the microglia pool at E12 and its recovery to a normal pool size at E14. (Each dot represents an independent experiment. Error bars show mean ± s.e.m.; two tailed, unpaired t-test.) b The decreased microglia pool at E12 may result from a reduced proliferation, as shown by the two folds decrease of BrdU-labeled cells in Lyl-1LacZ/LacZ compared to WT brains. (4 independent experiments; error bars show mean ± s.e.m.; two tailed, unpaired Mann–Whitney test). c At E12, Cx3cr1WT/GFP:Lyl-1LacZ/LacZ microglia displayed a reduced number and extent of ramifications compared to their Cx3cr1WT/GFP counterpart. Microglia morphology was classified into subtypes depending on the number of main ramifications (A: none, B: 2, C: 3 and D: > 3). Microglia deprived of ramifications predominated in Lyl-1-deficient microglia. 65 and 61 cells were respectively acquired from the midbrain of E12 Cx3cr1WT/GFP and Cx3cr1WT/GFP:Lyl-1LacZ/LacZ embryos (for each genotype, brains from 12 embryos were acquired in 3 independent experiments). Microglia were identified by Cx3cr1-driven GFP expression and F4/80-APC immuno-staining. Bar = 10 μm. (Error bars show mean ± s.e.m.; two tailed, unpaired t-test). d In Lyl-1LacZ/LacZ newborns, the cellularity of the brain was consistently lower than in WT, and so was the estimated microglia number. (3 independent experiments. Error bars show mean ± s.e.m.; two tailed, unpaired t-test). e Kinetic evolution of Lyl-1 expression levels in WT microglia from embryonic stages to adulthood. Expression levels were normalized to the mean value obtained for E12 microglia (n = 3; error bars show mean ± s.e.m). An increased expression of Lyl-1 from embryonic stages to adulthood was also inferred from timeline RNA-seq. data45. (GEO accession number GSE79812). f Quantitative RT-PCR analyses also point to E12 and P0 as key development stages regulated by Lyl-1. CD11b+F4/80+CD45low microglia were isolated at sequential development stages. Bar graphs show the kinetic of expression of genes modified in Lyl-1LacZ/LacZ microglia (arrowheads), normalized to the mean expression value in WT E12 microglia (n = 3). Floating bars indicate mean ± s.e.m. Two tailed, unpaired t-test. g Cx3Cr1 and Lyl-1 expression in mutant microglia. The expression level of Cx3CR1, analyzed as in f, was decreased in Lyl-1 mutants at E12, while Lyl-1 expression levels were unmodified in CX3CR1GFP/GFP microglia at E12 and in newborns. h Mafb expression in mutant microglia. Mafb expression level, analyzed as in f, was reduced in the microglia of Lyl-1LacZ/LacZ newborns. i The expression of genes enriched in microglia and/or essential for their function are deregulated in Lyl-1LacZ/LacZ MΦ-progenitors at E9. Relative expression levels (read counts) in WT and Lyl-1lacZ/lacZ MΦ-progenitors from E9-YS. (3 independent experiments. Box and whisker plots, min to max; median is shown; Unpaired, two-tailed t-Test). j Relative expression levels (read counts) of phagocytosis genes in sorted microglia from WT and Lyl-1LacZ/LacZ newborn (Error bars show mean ± s.e.m.; one tailed unpaired t-test).
P0-P3 was identified as a second stage altered in Lyl-1LacZ/LacZ microglia. At birth, the cellularity of Lyl-1LacZ/LacZ brain was significantly decreased compared to WT (Fig. 6d), which was not the case at earlier stages (Supplementary Fig. 4e). CD11b+ cells recovery was also reduced (WT: 140.96 ± 0.91 × 103, n = 9; Lyl-1LacZ/LacZ: 87.18 ± 0.37 × 103, n = 9). Consequently, Lyl-1-deficiency triggered a nearly 2-fold reduction of the microglia population (Fig. 6d). This perinatal reduction of microglia appeared transient, since no difference with WT brain was observed in the adult (Supplementary Fig. 4f). Transient decreases of microglia pool size, such as those we observed at E12 and P0-P3 in Lyl-1LacZ/LacZ mutant, have been reported to occur during normal development in postnatal weeks 2–352, but also in Cx3cr1 mutant mice during the 1st postnatal week53. This indicates a highly dynamic control of the microglia pool size during key steps of neural development that seems preserved in Lyl-1 mutant, with the exception of the E12 and P0-P3 time-points. At this later stage, the reduction of brain cellularity in Lyl-1LacZ/LacZ mice points to Lyl-1 as a possible regulator of the trophic function of microglia on brain cells54,55.
The identification of E12 and P0-P3 as key stages for Lyl-1 function in microglia development was confirmed by RT-qPCR analyses of the expression of genes essential for MΦs (Spi1/PU.1, Csf1r, Mafb) and/or microglia (Runx1, Cx3cr1, Irf8) development and function, of known regulators of developmental hematopoiesis (Tal-1, Lmo2, Runx1) and related factors (Tcf3/E2A, Tcf4/E2.2) (Fig. 6e, f; Supplementary Fig. 4g). Time-course analyses highlighted the down-regulation of Csf1r, Irf8 and Lmo2 in Lyl-1LacZ/LacZ microglia at E12 and P0-P3, while Cx3cr1 was only decreased at E12 (Fig. 6g). Note that Lyl-1 expression was unmodified in Cx3cr1GFP/GFP mutants (Fig. 6g). Interestingly, Cx3cr1, as well as Irf8 and Lmo2, belong to potential Lyl-1 target genes2.
Mafb expression levels in Lyl-1LacZ/LacZ microglia transiently decreased at P0-P3 and later returned back to WT expression levels (Fig. 6h). As Mafb represses resident-MΦ self-renewal56, the recovery of a normal amount of microglia after birth may stem from this transient decrease. Spi1/PU.1, Tcf3/E2A and Tcf4/E2.2 expression levels were unmodified in Lyl-1LacZ/LacZ microglia, while Runx1 expression was only affected after birth. Tal-1 expression was decreased at E14 and increased after birth, suggesting that this Lyl-1 paralog3 does not compensate Lyl-1-deficiency during embryonic stages, but may do so at postnatal stages (Supplementary Fig. 4g). Remarkably, RNA-seq. results indicated that some genes deregulated in Lyl-1LacZ/LacZ microglia at E12 and P0-P3 were also down-regulated in Lyl-1LacZ/LacZ MΦ-progenitors at E9 (Csf1r: Supplementary Fig. 3c, Lmo2: Fig. 3b; Irf8: Fig. 4D; Cx3cr1: Fig. 6i). These deregulations were transient, however in both locations and stages they coincided with a defective MΦ/microglia differentiation.
Other genes enriched in microglia (Fcrls, Mef2c, Maf)57 or involved in the maintenance of microglia homeostasis (P2ry12)58 were also expressed in E9 Lyl-1LacZ/LacZ MΦ-progenitors at a lower level than in the WT, except Lpr8 and Aif-1/Iba1 (Fig. 6i). Interestingly, the lower expression of phagocytosis genes (Axl1, Mertk, Trem2, P2y6r/P2ry6) uncovered in Lyl-1LacZ/LacZ MΦ-progenitors from E10-YS (Fig. 4G), was present and stronger in the microglia from Lyl-1LacZ/LacZ newborn, except for Trem2 which expression was unmodified (Fig. 6j).
These deregulations highlight again shared features between MΦPrim progenitors and microglia/neural development which became apparent upon Lyl-1 inactivation considering the large number of neural signaling pathways up-regulated in E9 Lyl-1LacZ/LacZ MΦ-progenitors (Fig. 3c) and the relationship of the differentially expressed genes enriched in E9 MΦPrim progenitors with brain formation and neuro-development uncovered in IPA analysis (Supplementary Fig. 4h).
Based on the gene expression pattern of Lyl-1-deficient microglia and the signature of MΦPrim progenitors in the early YS, a contribution of Lyl-1-deficiency to neurodevelopmental disorders may be considered. Synaptic pruning and neural maturation, which characterize the perinatal phase of microglia development45, might be impaired in Lyl-1LacZ/LacZ embryos considering the defects observed at P0-P3, the later key developmental stage regulated by Lyl-1. Indeed, Lyl-1 deregulation has been observed in datasets reporting pathological models of brain myeloid cells (http://research-pub.gene.com/BrainMyeloidLandscape/#Mouse-gene/Mouse-gene/17095/geneReport.html)59, as well as in human neuropathies60,61, including the 19p13.13 micro-deletion neuro-developmental disabilities62. However, since Lyl-1 is expressed in endothelial cells, including in the brain42, a contribution of LYL-1-deficient endothelial cells to these diseases must be considered.
Conclusions
Altogether, our findings reveal Lyl-1 as a key factor regulating the production and differentiation of YS MΦ-progenitors and the development of microglia. Lyl-1 is the least studied partners of the transcription factor complex that regulates developmental hematopoiesis. The development of more appropriate models is required to precise Lyl-1 functions in microglia and determine its role in the development of other resident-MΦs populations.
Methods
Mice
The following mouse strains, housed in Gustave Roussy Institute animal facility (License #H94-076-11) were used (See Table 1 for a summary of the breeding schemes): 1- C57BL/6 mice (Harlan or Charles Rivers Laboratories), referred to as wild type (WT); 2- Lyl-1LacZ mice5. To avoid the possible detection of FDG/Lyl-1 expression from maternally-derived MΦ11 in heterozygous embryos, Lyl-1LacZ/LacZ males were crossed with WT females; 3- Cx3cr1GFP mice63. Cx3cr1GFP/GFP males were crossed with WT females to generate Cx3cr1WT/GFP mice/embryos or with Lyl-1LacZ/LacZ females to generate Cx3cr1WT/GFP:Lyl-1WT/LacZ mice/embryos. 4-Cx3cr1GFP/GFP:Lyl-1LacZ/LacZ double mutant strain developed from Cx3cr1WT/GFP:Lyl-1WT/LacZ crosses. Cx3cr1WT/GFP:Lyl-1WT/LacZ and Cx3cr1WT/GFP:Lyl-1LacZ/LacZ mice/embryos were obtained by crossing Cx3cr1GFP/GFP:Lyl-1LacZ/LacZ males respectively to C57BL/6 or Lyl-1LacZ/LacZ females. Experiments were conducted in compliance with French/European laws, under authorized project #2016-030-5798, approved by officially accredited local institutional animal (committee n°26) and French “Ministère de la Recherche” ethics boards..
The day of vaginal plug observation was considered as E0.5. Pregnant females were sacrificed by cervical dislocation. Pre-somite embryos were staged according to Downs et al.64, by somite counting from E8 to E10.5 and thereafter by morphological landmarks.
Tissues
Cells from the yolk sac (E7.5-E10.5), from E10 FL or whole brain (E9-E14) were obtained after mechanical disruption9,65. For cytometry analyses, the dissected region of the brain comprises the di-, mes- and metencephalon at E9-E10, the ectoderm was carefully removed22. After E12, microglia were recovered from the midbrain following Percoll (P1644, Sigma) separation66. Cells were filtered through a 70 μm cell strainer and centrifuged at 300 g for 10 min. Cells were resuspended in Fc block BD solution and incubated at room temperature for 10 min. After washing in PBS + 10% FCS and centrifugation, cells were resuspended in 50 μl + 10% FCS containing the fluorochrome coupled antibodies and incubated at 4 °C for 20 min. After washing in PBS + 10%FCS and centrifugation, cells were resuspended in 50ul + 10% FCS. Dead cells were excluded by adding 1 μg/mL DAPI (Sigma) before acquisition.
The number of microglia per brain was estimated by reporting the percentage of CD11b+CD45lowF4/80+ microglia to the cellularity of the corresponding sample.
Tissue culture
YS explants, maintained for 1 day in organ culture in plates containing OptiMEM+Glutamax, 1% Penicillin-streptomycin, 0.1% β-mercaptoethanol (ThermoFisher) and 10% fetal calf serum (Hyclone), are referred to as OrgD1-YS.
In clonogenic assays, YS suspension or sorted cells were plated in triplicate (3 × 103 or 100-150 cells/mL) in Methocult® M3234 (StemCell Technologies) always supplemented the cytokines listed in Table 2. Colonies were scored at day 5 for primitive erythroblasts and day 7 for other progenitors.
Flow cytometry
Cells, stained with antibodies listed in Table 3, for 30 min. on ice, were acquired (Canto II) or sorted (FACS-Aria III or Influx, BD Biosciences) and analyzed using FlowJo (Treestar) software.
The β-Galactosidase substrate fluorescein di-β-galactopyranoside (FDG; Molecular probe), was used as reporter for Lyl-1 expression in FACS-Gal assay67,68. For apoptosis analysis, microglia were immune-stained and incubated with Annexin V-FITC. 7AAD was added before acquisition. For proliferation assays, pregnant females (12 gestational days) were injected with BrdU (10μM) and sacrificed 2 h later. Microglia were isolated, immune-stained and prepared according to kit instruction (BD Pharmingen 552598). BrdU incorporation was revealed using anti-BrdU-APC.
Brain imaging
To assess microglia morphology in E12 embryos, the midbrain was dissected from Cx3cr1WT/GFP:Lyl-1WT/WT and Cx3cr1WT/GFP:Lyl-1LacZ/LacZ embryos and sectioned through the midline. After fixation (4% paraformaldehyde) overnight at 4 °C, whole midbrains were washed in phosphate-buffered saline (PBS)/0.1 M glycine and incubated overnight in PBS/15% sucrose at 4 °C. Midbrains were washed with PBS + 0.1% Tween and incubated 90 min. in blocking buffer (PBS + 10% FCS) at room temperature (RT). Midbrains were subsequently immune-labeled with F4/80-APC overnight at 4 °C. After washing, they were incubated 3 min. in PBS + DAPI (1 μg/mL) at RT and washed. Finally, midbrains were placed in the central well of glass-bottom culture dishes (P35G-1.5-10-C; MatTek, USA) filled with PBS + 10% FCS. After appropriate orientation of the sample, the well was covered with a 12 μm ∅ glass coverslip. Image stacks were collected using a Leica SP8 confocal microscope. To unsure an unbiased choice of the cells imaged, taking into account possible changes in cell distribution induced by Lyl-1 deficiency, we always acquired cells in similar positions regarding the landmark set in the midbrain flat mount, as shown in Supplementary data and Fig. 4C. Images were processed using Imaris x64 (version 7.7.2; Bitplane) and Photoshop 8.0 (Adobe Systems, San Jose, CA) softwares.
RT-qPCR analyses
RNA was extracted using Trizol. After cDNA synthesis (SuperScript™ VILO™ Master-Mix reverse transcriptase, ThermoFisher), Real Time (RT)-PCR was performed (SYBR Premix Ex TaqII, Takara Bio). Reference genes were Actin, Hprt and Tubulin. Gene expressions were normalized to the values obtained for E10-YS MΦ-progenitors, E12 WT Lin-Sca+cKit+ (LSK) progenitors or E12 WT microglia. Primers are listed in Table 4.
RNA-seq
For samples preparation, MΦ-progenitors (Kit+CD45+CD11b+) were sorted from E9 (MΦPrim progenitor) and E10 (MΦPrim + MΦT-Def progenitors) YS pools from WT or Lyl-1LacZ/LacZ embryos (Four biological replicates). RNA was extracted as described above.
The RNA integrity (RNA Integrity Score ≥ 7.0) was checked on the Agilent 2100 Bioanalyzer (Agilent) and quantity was determined using Qubit (Invitrogen). SureSelect Automated Strand Specific RNA Library Preparation Kit was used according to manufacturer’s instructions with the Bravo Platform. Briefly, 50 ng of total RNA sample was used for poly-A mRNA selection using oligo(dT) beads and subjected to thermal mRNA fragmentation. Fragmented mRNA samples were subjected to cDNA synthesis and converted into double stranded DNA using reagents supplied in the kit. The resulting dsDNA was used for library preparation. The final libraries were bar-coded, purified, pooled together in equal concentrations and subjected to paired-end sequencing (2 × 100) on Novaseq-6000 sequencer (Illumina) at Gustave Roussy genomic facility.
The quality of RNA-seq. reads were assessed with FastQC 0.11.7 and MultiQC 1.569. Low quality reads were trimmed with Trimmomatic 0.3370. Salmon 0.9.0 tool71 was used for quantifying the expression of transcripts using geneset annotation from Gencode project release M17 for mouse72. The version of transcriptome reference sequences used was GRCm38.p6.
Statistical analysis was performed using R with the method proposed by Anders and Huber implemented in the DESeq2 Bioconductor package73. The differential expression analysis in DESeq2 uses a generalized linear model (GLM) where counts are modeled using a negative binomial distribution. Counts were normalized from the estimated size factors using the median ratio method and a Wald test was used to test the significance of GLM coefficients. Genes were considered differentially expressed when adjusted p-value < 0.05 and fold-change > 2.
Data were analyzed through the use of Ingenuity® Pathway Analysis (QIAGEN Inc., https://www.qiagenbioinformatics.com/products/ingenuity-pathway-analysis)74, Gene set enrichment analysis (GSEA; https://www.gsea-msigdb.org/gsea/index.jsp)75,76, Morpheus (https://software.broadinstitute.org/morpheus/) and Venny (https://bioinfogp.cnb.csic.es/tools/venny/) softwares.
Statistics and reproducibility
Statistical analyses including mean, SEM and p values were performed using using Prism 7 software (GraphPad). Statistical significance between two groups was assessed by unpaired two-tailed Student’s t tests or Mann–Whitney tests. Data are presented as mean ± SEM or by Box and whisker plots, min to max, with the median shown. Statistical significances were indicated by p-values and/or as *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001, ns: not significant). P-values below 0.05 were considered as significantly different. The number of biological replicates (n) and independent experiments is reported in the corresponding figure legend.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its supplementary information files or from the corresponding author upon reasonable request. RNAseq data have been deposited in the ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/arrayexpress) under accession code E-MTAB-9618. The source data underlying figures and supplementary figures are provided as a Supplementary Data 1.
References
Pina, C. & Enver, T. Differential contributions of haematopoietic stem cells to foetal and adult haematopoiesis: insights from functional analysis of transcriptional regulators. Oncogene 26, 6750–6765 (2007).
Wilson, N. K. et al. Combinatorial transcriptional control in blood stem/progenitor cells: genome-wide analysis of ten major transcriptional regulators. Cell Stem Cell 7, 532–544 (2010).
Curtis, D. J., Salmon, J. M. & Pimanda, J. E. Concise review: blood relatives: formation and regulation of hematopoietic stem cells by the basic helix-loop-helix transcription factors stem cell leukemia and lymphoblastic leukemia-derived sequence 1. Stem Cells 30, 1053–1058 (2012).
Porcher, C., Chagraoui, H. & Kristiansen, M. S. SCL/TAL1: a multifaceted regulator from blood development to disease. Blood 129, 2051–2060 (2017).
Capron, C. et al. The SCL relative LYL-1 is required for fetal and adult hematopoietic stem cell function and B-cell differentiation. Blood 107, 4678–4686 (2006).
McGrath, K. E., Frame, J. M. & Palis, J. Early hematopoiesis and macrophage development. Semin. Immunol. 27, 379–387 (2015).
Palis, J. Hematopoietic stem cell-independent hematopoiesis: emergence of erythroid, megakaryocyte, and myeloid potential in the mammalian embryo. FEBS Lett. 590, 3965–3974 (2016).
Cumano, A. & Godin, I. Ontogeny of the hematopoietic system. Annu Rev. Immunol. 25, 745–785 (2007).
Kieusseian, A., Brunet de la Grange, P., Burlen-Defranoux, O., Godin, I. & Cumano, A. Immature hematopoietic stem cells undergo maturation in the fetal liver. Development 139, 3521–3530 (2012).
Palis, J., Robertson, S., Kennedy, M., Wall, C. & Keller, G. Development of erythroid and myeloid progenitors in the yolk sac and embryo proper of the mouse. Development 126, 5073–5084 (1999).
Bertrand, J. Y. et al. Three pathways to mature macrophages in the early mouse yolk sac. Blood 106, 3004–3011 (2005).
Tober, J. et al. The megakaryocyte lineage originates from hemangioblast precursors and is an integral component both of primitive and of definitive hematopoiesis. Blood 109, 1433–1441 (2007).
Sumner, R., Crawford, A., Mucenski, M. & Frampton, J. Initiation of adult myelopoiesis can occur in the absence of c-Myb whereas subsequent development is strictly dependent on the transcription factor. Oncogene 19, 3335–3342 (2000).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Hoeffel, G. & Ginhoux, F. Fetal monocytes and the origins of tissue-resident macrophages. Cell Immunol. 330, 5–15 (2018).
Utz, S. G. et al. Early fate defines microglia and non-parenchymal brain macrophage development. Cell 181, 557–573 e518 (2020).
Ginhoux, F. & Guilliams, M. Tissue-resident macrophage ontogeny and homeostasis. Immunity 44, 439–449 (2016).
Mass, E. Delineating the origins, developmental programs and homeostatic functions of tissue-resident macrophages. Int Immunol. 30, 493–501 (2018).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Gomez Perdiguero, E. et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature 518, 547–551 (2015).
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013).
Alliot, F., Godin, I. & Pessac, B. Microglia derive from progenitors, originating from the yolk sac, and which proliferate in the brain. Brain Res. Dev. Brain Res. 117, 145–152 (1999).
Wu, Y. & Hirschi, K. K. Tissue-resident macrophage development and function. Front. Cell Dev. Biol. 8, 617879 (2021).
Herbomel, P., Thisse, B. & Thisse, C. Zebrafish early macrophages colonize cephalic mesenchyme and developing brain, retina, and epidermis through a M-CSF receptor-dependent invasive process. Dev. Biol. 238, 274–288 (2001).
Ferrero, G. et al. Embryonic microglia derive from primitive macrophages and are replaced by cmyb-dependent definitive microglia in zebrafish. Cell Rep. 24, 130–141 (2018).
Wittamer, V. & Bertrand, J. Y. Yolk sac hematopoiesis: does it contribute to the adult hematopoietic system? Cell Mol. Life Sci. 77, 4081–4091 (2020).
Azzoni, E. et al. Kit ligand has a critical role in mouse yolk sac and aorta-gonad-mesonephros hematopoiesis. EMBO Rep. 19, e45477 (2018).
Espin-Palazon, R. et al. Proinflammatory signaling regulates hematopoietic stem cell emergence. Cell 159, 1070–1085 (2014).
Giroux, S. et al. lyl-1 and tal-1/scl, two genes encoding closely related bHLH transcription factors, display highly overlapping expression patterns during cardiovascular and hematopoietic ontogeny. Gene Expr. Patterns 7, 215–226 (2007).
Cumano, A., Dieterlen-Lievre, F. & Godin, I. Lymphoid potential, probed before circulation in mouse, is restricted to caudal intraembryonic splanchnopleura. Cell 86, 907–916 (1996).
Cumano, A., Ferraz, J. C., Klaine, M., Di Santo, J. P. & Godin, I. Intraembryonic, but not yolk sac hematopoietic precursors, isolated before circulation, provide long-term multilineage reconstitution. Immunity 15, 477–485 (2001).
Hoeffel, G. et al. C-Myb+ erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678 (2015).
Mass, E. et al. Specification of tissue-resident macrophages during organogenesis. Science 353, aaf4238 (2016).
Balounova, J. et al. Toll-like receptor 2 expression on c-kit(+) cells tracks the emergence of embryonic definitive hematopoietic progenitors. Nat. Commun. 10, 5176 (2019).
Espin-Palazon, R., Weijts, B., Mulero, V. & Traver, D. Proinflammatory signals as fuel for the fire of hematopoietic stem cell emergence. Trends Cell Biol. 28, 58–66 (2018).
Mariani, S. A. et al. Pro-inflammatory aorta-associated macrophages are involved in embryonic development of hematopoietic stem cells. Immunity 50, 1439–1452.e5 (2019).
Palpant, N. J. et al. Chromatin and transcriptional analysis of mesoderm progenitor cells identifies HOPX as a regulator of primitive hematopoiesis. Cell Rep. 20, 1597–1608 (2017).
Chiu, S. K. et al. Shared roles for Scl and Lyl1 in murine platelet production and function. Blood 134, 826–835 (2019).
McCormack, M. P. et al. Requirement for Lyl1 in a model of Lmo2-driven early T-cell precursor ALL. Blood 122, 2093–2103 (2013).
Epelman, S. et al. Embryonic and adult-derived resident cardiac macrophages are maintained through distinct mechanisms at steady state and during inflammation. Immunity 40, 91–104 (2014).
Leid, J. et al. Primitive embryonic macrophages are required for coronary development and maturation. Circulation Res. 118, 1498 (2016).
Pirot, N. et al. LYL1 activity is required for the maturation of newly formed blood vessels in adulthood. Blood 115, 5270–5279 (2010).
Hagemeyer, N. et al. Transcriptome-based profiling of yolk sac-derived macrophages reveals a role for Irf8 in macrophage maturation. EMBO J. 35, 1730–1744 (2016).
Ferrier, R. et al. Physical interaction of the bHLH LYL1 protein and NF-kappaB1 p105. Oncogene 18, 995–1005 (1999).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE79812 (2016). Data sets.
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. 17, 797–805 (2016).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Wehrspaun, C. C., Haerty, W. & Ponting, C. P. Microglia recapitulate a hematopoietic master regulator network in the aging human frontal cortex. Neurobiol. Aging 36, 2443 e2449–2443 e2420 (2015).
Bennett, M. L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl Acad. Sci. 113, E1738–E1746 (2016).
Zhan, Y. et al. Deficient neuron-microglia signaling results in impaired functional brain connectivity and social behavior. Nat. Neurosci. 17, 400–406 (2014).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Antony, J. M., Paquin, A., Nutt, S. L., Kaplan, D. R. & Miller, F. D. Endogenous microglia regulate development of embryonic cortical precursor cells. J. Neurosci. Res. 89, 286–298 (2011).
Ueno, M. et al. Layer V cortical neurons require microglial support for survival during postnatal development. Nat. Neurosci. 16, 543–551 (2013).
Soucie, E. L. et al. Lineage-specific enhancers activate self-renewal genes in macrophages and embryonic stem cells. Science 351, aad5510 (2016).
Crotti, A. & Ransohoff, R. M. Microglial physiology and pathophysiology: insights from genome-wide transcriptional profiling. Immunity 44, 505–515 (2016).
Bisht, K., Sharma, K. & Tremblay, M.-È. Chronic stress as a risk factor for Alzheimer’s disease: Roles of microglia-mediated synaptic remodeling, inflammation, and oxidative stress. Neurobiol. Stress 9, 9–21 (2018).
Friedman, B. A. et al. Diverse brain myeloid expression profiles reveal distinct microglial activation states and aspects of Alzheimer’s disease not evident in mouse models. Cell Rep. 22, 832–847 (2018).
Colangelo, V. et al. Gene expression profiling of 12633 genes in Alzheimer hippocampal CA1: transcription and neurotrophic factor down-regulation and up-regulation of apoptotic and pro-inflammatory signaling. J. Neurosci. Res. 70, 462–473 (2002).
Thomas, D. M., Francescutti-Verbeem, D. M. & Kuhn, D. M. Gene expression profile of activated microglia under conditions associated with dopamine neuronal damage. Faseb J. 20, 515–517 (2006).
Nimmakayalu, M. et al. Apparent germline mosaicism for a novel 19p13.13 deletion disrupting NFIX and CACNA1A. Am. J. Med. Genet. A 161A, 1105–1109 (2013).
Jung, S. et al. Analysis of fractalkine receptor CX(3)CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell Biol. 20, 4106–4114 (2000).
Downs, K. M. & Davies, T. Staging of gastrulating mouse embryos by morphological landmarks in the dissecting microscope. Development 118, 1255–1266 (1993).
Bertrand, J. Y., Giroux, S., Cumano, A. & Godin, I. Hematopoietic stem cell development during mouse embryogenesis. Methods Mol. Med. 105, 273–288 (2005).
Mildner, A. et al. Microglia in the adult brain arise from Ly-6ChiCCR2+ monocytes only under defined host conditions. Nat. Neurosci. 10, 1544–1553 (2007).
Fiering, S. N. et al. Improved FACS-Gal: flow cytometric analysis and sorting of viable eukaryotic cells expressing reporter gene constructs. Cytometry 12, 291–301 (1991).
Guo, W. & Wu, H. Detection of LacZ expression by FACS-Gal analysis. Nat. Protoc. Exch. (2008).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–d773 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Krämer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
The authors thank Julien Bertrand for critical reading of the manuscript. We are grateful to the staff of the facilities at Gustave Roussy, the animal facility (PFEP, UMS AMMICa UMS 3655/US23, directed by P. Gonin), the imaging facility (PFIC, UMS AMMICa UMS 3655/US23, directed by C. Laplace-Builhe), the genomic facility directed by N. Droin, the bioinformatics facility (G. Meurice), directed by M. Deloger. This work was supported by fundings from Institut National de la Santé et de la Recherche Médicale to W. Vainchenker, I. Plo and H. Raslova, from Centre National de la Recherche Scientifique and Université de Paris-Saclay to I. Godin, from grants INCA PLBio to I. Plo, “Ligue Nationale contre le Cancer” Certified Team to H. Raslova, “Association pour la Recherche sur le Cancer” (n°4878) to I. Godin, Gustave Roussy (TA DERE 17) and National Natural Science foundation of China (N°32000669) to D. Ren, Grant Agency of the Czech Republic (GACR n°19–23154 S) to D. Filipp and from fellowships from “Association pour la Recherche sur le Cancer” to A.-L. Kaushik; “Société Française d’Hématologie” to S. Wang and Chinese Scholarship Council fellowships to S. Wang and D. Ren.
Author information
Authors and Affiliations
Contributions
SW: Conceptualization; Methodology; Validation; Formal analysis; Investigation; Data curation; Writing – original draft preparation; Visualization; DR: Formal: analysis; Investigation; Data curation; Writing – original draft preparation; Visualization; A-LK; BA: Investigation, methodology; Formal analysis; Visualization; GM: Formal analysis; Investigation; methodology; YL: Investigation; methodology; DF: Writing—review & editing: critical review, commentary and revision; WV, HR, IP: Funding acquisition; Writing—review & editing: critical review, commentary and revision; IG: Conceptualization; Project administration; Supervision; Methodology; Validation; Formal analysis; Resources; Data curation; Writing – original draft preparation; Writing—review & editing; Visualization; Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Communications Biology thanks Ki-Wook Kim, Noelia Alonso Gonzalez and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editors: Eirini Trompouki and George Inglis. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, S., Ren, D., Arkoun, B. et al. Lyl-1 regulates primitive macrophages and microglia development. Commun Biol 4, 1382 (2021). https://doi.org/10.1038/s42003-021-02886-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-02886-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.