Abstract
Lung cancer is the main cause of cancer death worldwide, with lung squamous cell carcinoma (LUSC) being the second most frequent subtype. Preclinical LUSC models recapitulating human disease pathogenesis are key for the development of early intervention approaches and improved therapies. Here, we review advances and challenges in the generation of LUSC models, from 2D and 3D cultures, to murine models. We discuss how molecular profiling of premalignant lesions and invasive LUSC has contributed to the refinement of in vitro and in vivo models, and in turn, how these systems have increased our understanding of LUSC biology and therapeutic vulnerabilities.
Similar content being viewed by others
Introduction
Approximately 2.2 million lung cancer cases are diagnosed each year (https://gco.iarc.fr/today). As most patients present with a late-stage incurable disease, lung cancer is the major contributor to cancer mortality worldwide, accounting for 18.4% of deaths1. The design of more effective therapeutic strategies and the development of early detection and intervention approaches are therefore global health priorities.
Around 85% of lung cancer cases correspond to the non-small-cell lung cancer (NSCLC) subtype. Among these, one-third are lung squamous cell carcinoma (LUSC), and the rest are predominantly adenocarcinoma (LUAD)2. Established cancer cell lines have been instrumental in the investigation of the molecular mechanisms driving NSCLC and in drug discovery studies. However, in the absence of molecular fingerprinting and routine surveillance, intra- or inter-species contamination and cell line misidentification can severely compromise the potential of these models3. Currently, the availability of well characterised NSCLC cell lines is biased towards LUAD4.
Over the last decade, sequencing studies have not only identified recurrent genomic alterations across large cohorts of LUSC samples, but also highlighted the great inter- and intra-tumour heterogeneity and their complex evolutionary histories5,6, which cannot be recapitulated in traditional cancer cell line cultures. Additionally, it is clear that the tumour microenvironment plays an essential role in lung cancer progression7. Together, these observations stress the importance of developing alternative LUSC models that allow dissection of the molecular pathogenesis of LUSC, analyses of complex cell-cell and cell-microenvironment interactions, and assessment of tailored therapies at different stages of disease progression.
Here, we review advances in the development of in vitro and in vivo LUSC models and their implications in understanding the cellular and molecular processes driving LUSC development and responses to therapy.
In vitro systems
Modelling pre-invasive disease in air−liquid interface (ALI) culture
LUSC develops in a step-wise manner from increasingly disordered pre-invasive lesions in the bronchial epithelium. Pre- invasive lesions are associated with cigarette smoking, the main lung cancer risk factor8,9,10. The histology ranges from hyperplasia, metaplasia and dysplasia to carcinoma-in situ (CIS). Molecular studies have identified recurrent genomic, transcriptomic and epigenetic alterations in pre-invasive squamous lung lesions11,12,13,14. Both amplification of the distal region of chromosome 3q and mutations in TP53 are among the earliest changes commonly detected in pre-invasive lesions11,12,13,14,15.
A number of genes located within the 3q amplicon have been highlighted as potential LUSC drivers, including SRY-box 2 (SOX2), protein kinase C iota (PRKCI), epithelial cell transforming 2 (ECT2), phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA)15,16 and TRAF2 and NCK interacting kinase (TNIK)17. Gain- and loss-of-function studies in NSCLC cell lines have shown the transcription factor SOX2 to be a master regulator of squamous identity and a lineage-survival oncogene18. More recently, in vitro models have been used to investigate the effects of early SOX2 amplification and its potential interactions with other alterations occurring during the progression of pre-invasive lesions to invasive LUSC.
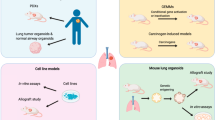
Since its development19, the ALI culture system has been widely used for functional in vitro analyses of the bronchial epithelium. Epithelial cells are seeded onto a biphasic cultivation chamber with a permeable support layered with an extracellular matrix gel substrate, with or without embedded stromal cells. Following initial growth in submerged culture, the apical surface of the epithelial cells is exposed to the air, while they keep receiving nutrients basally. After continuous culture in ALI conditions, ciliated and mucinous cell differentiation occurs, resulting in the formation of a polarised epithelial sheet (Fig. 1).
Establishment of air−liquid-interface (ALI) and organoid cultures from human or mouse airway epithelial cells and LUSC tissue. Following ALI or 3D culture, normal airway epithelial basal cells produce pseudostratified epithelial sheets or hollow organoids containing differentiated cells, respectively. In contrast, LUSC cells give rise to epithelial sheets with features of dysplasia and more solid, disorganised organoids. Cultured cells may be subjected to genetic and pharmacological manipulation to investigate the phenotypic consequences of molecular alterations recurrently identified in LUSC samples. Organoids can be used in in vitro drug screenings and may be implanted into mice to evaluate their ability to give rise to tumours in vivo and response to therapies. ECM extracellular matrix.
Enforced expression of SOX2 in human bronchial epithelial cells in ALI culture has been shown to induce the formation of squamous metaplasia20 and dysplasia21, enhance mucinous differentiation and reduce ciliated differentiation20. Silencing TP53 led to the formation of focal outgrowths within the epithelial cell sheet; when combined with SOX2 overexpression, the outgrowths were more diffuse21. This indicates that early dysregulation of SOX2 and TP53 in the bronchial epithelium jointly drives a dysplastic phenotype. The ability of SOX2 to induce squamous metaplasia and dysplasia was found to require phosphatidylinositol 3-kinase (PI3K) activity, as either pharmacological inhibition of the PI3K/AKT pathway20,21 or knockdown of PIK3CA20—encoding the p110α catalytic subunit of PI3K—suppressed squamous differentiation. The cooperative roles between SOX2 and PIK3CA provide mechanistic insights about the selective advantage of 3q amplification during pre-invasive LUSC evolution.
Human primary LUSC in 2D cell culture
Primary human epithelial cells from diverse tissues can be robustly expanded in vitro when cultured on a feeder cell layer of mitotically inactivated murine 3T3 fibroblasts and in the presence of Rho kinase (ROCK) inhibitor Y-2763222. Airway epithelial cells grown under these conditions, frequently referred to as ‘conditional reprogramming’ (CR), retain features of airway basal stem cells, including multi-lineage differentiation potential23. These methods have been applied in the establishment of primary NSCLC cell cultures24,25. However, reported success rates have been variable, with various studies indicating that CR conditions preferably support the growth of normal airway epithelial cells26,27,28.
In CR cultures established from resected primary NSCLC tumours, including several cases of LUSC, cancer cells were found to be outgrown by normal airway epithelial cells26,27. This resulted in the loss of both patient-specific cancer-associated genetic alterations26,27 and in vivo tumourigenic potential26. When cultures were initiated from lung cancer brain metastases or NSCLC patient-derived xenografts, both expected to lack contaminating human epithelial cells, they failed to expand26. The latter suggests that CR conditions cannot sustain the in vitro propagation of NSCLC cells.
A variation of the CR protocol that uses human instead of murine fibroblasts as a feeder layer, has been reported to support the establishment of NSCLC cell cultures from effusions and biopsies when used during the initial plating step24,29. Cultures are subsequently passaged off the feeder layer for further expansion. The resulting cell lines were found to retain key mutations identified in the corresponding patient tumour24,29. Yet, this method has been mainly used for LUAD. It is possible that CR conditions may be less permissive for the propagation of cells carrying LUSC-associated genetic and/or epigenetic changes. Side-by-side comparison of the two feeder cell types should help clarify the applicability of CR conditions both during the derivation and long-term expansion of primary LUSC cell cultures.
3D models
When embedded in basement membrane matrix hydrogels (e.g., Matrigel) and cultured in non-adherent conditions, either in the presence or absence of supporting stromal cells, adult airway stem and progenitor cells can give rise to self-organising 3D hollow structures called ‘organoids’. Lung organoid cultures may be established from primary murine or human cells and phenocopy basic cellular aspects of the airway epithelium, with polarised progenitor cells and their differentiated progeny arranged around a lumen30 (Fig. 1). This system has been used for the development of in vitro LUSC models.
Validating tumour suppressors and oncogenes in 3D culture
In addition to mutations in the tumour suppressor TP53 and amplification of chromosome 3q, molecular profiling studies of LUSC samples have identified significant dysregulation of a number of pathways, including cyclin-dependent kinase inhibitor 2A (CDKN2A)/retinoblastoma 1 (RB1), PI3K/AKT, squamous differentiation, and oxidative stress response6,18. The phenotypic consequences of several of these alterations, as well as potential synergistic effects, have been investigated using murine lung organoid models (Figs. 1 and 2).
SOX2, ECT2, PKCι (encoded by PRKCI), and PI3K signalling cooperate to promote a neoplastic cell fate in LUSC models. AKT is a downstream effector of stimulated p110α. Full AKT activation is achieved when phosphorylated at both positions S473 and T308127. High levels of SOX2 have been correlated with upregulated phospho-AKT21. PKCι phosphorylates and directly interacts with ECT2 to promote anchorage-independent growth and invasion through downstream targets. PKCι phosphorylates SOX2 favouring squamous cell fate and decreased differentiation16. Loss of p53, PTEN, and KEAP1 have been used to model LUSC phenotypes both in vitro and in vivo. Simultaneous loss of p53 and KEAP1 has shown synergistic effects, inducing increased proliferation, metastatic potential, and resistance to oxidative stress31. p53 activity can inhibit PI3K signalling through PTEN-dependent and potentially-independent mechanisms in squamous cell carcinomas128. Additional interactions between depicted proteins have been described in other cellular contexts.
Inactivation of Trp53 (the mouse orthologue of TP53) in bulk mouse tracheal epithelial cells has been shown to enhance organoid formation efficiency31. When 3D cultures were initiated with purified basal cells, p53 deficiency increased both organoid number and size and enabled longer-term passaging, indicative of aberrant stem/progenitor cell self-renewal32. Depletion of Kelch-like ECH associated protein (KEAP1), a negative regulator of the oxidative stress response, also augmented organoid forming capacity of tracheal epithelial cells31. Concurrent inactivation of Keap1 and Trp53 resulted not only in greater increase of organoid number than individual gene deletion, but also in the formation of solid spheres, suggesting synergistic effects of the two genetic alterations. The loss of Keap1 in Trp53-deficient organoids lowered intracellular reactive oxygen species (ROS) levels. This was found to mediate the increase in organoid forming ability, to enhance the metastatic potential of organoid-derived cells upon implantation in mice, and to confer resistance to oxidative stress and radiotherapy31.
Using lentiviral vectors to model amplification of different genes located within the 3q26 chromosomal region, it has been shown that SOX2, ECT2 and PRKCI exert cooperative interactions to induce neoplastic transformation of Trp53-depleted murine basal cells32. In organoid cultures, SOX2, ECT2, and PRKCI were found to each have a dominant role in the control of basal cell self-renewal, cell proliferation and epithelial polarity, respectively. Simultaneous overexpression of the three genes in p53-deficient basal cells led to increased number and size of organoids and acquisition of disorganised solid morphology. When grafted into mice, these cells gave rise to tumours with features of LUSC32. As both SOX2 and ECT2 are phosphorylation targets of PRKCI33,34, phosphorylation-resistant mutant forms of each gene were used in organoid cultures to demonstrate that the effects of PRKCI on airway epithelial cell polarity and proliferation are mediated by SOX2 and ECT2, respectively32. The latter emphasises the multifunctional role of SOX2 in early LUSC development.
In a different study, 3D cultures were established from bulk bronchial cells isolated from mice with Cre-inducible expression of Sox2 and Cas9. The resulting organoids were genetically manipulated in vitro to both induce Sox2 overexpression and have CRISPR/Cas9-mediated loss-of-function mutations in Trp53, Cdkn2a, and phosphate and tensin homologue (Pten), a negative regulator of PI3K signalling. Longitudinal analysis of organoid diameters showed that double- and triple-mutant organoids displayed a growth advantage. Upon engraftment into mice, triple-mutant organoids formed tumours reminiscent of well-differentiated LUSC35.
Overall, organoid cultures, together with the diverse array of currently available genetic manipulation techniques, constitute a powerful tractable platform to dissect molecular drivers and vulnerabilities of LUSC in vitro. Furthermore, the ability to assess organoid tumour-forming potential in vivo following transplantation into murine hosts makes murine organoid cultures a valuable tool for the generation of immunocompetent LUSC mouse models (Fig. 1), as we discuss later.
Patient-derived lung cancer organoids
Organoid culture methods for tissue stem cells have been adapted for the 3D expansion of patient-derived tumour cells from different epithelial cancers (Fig. 1). To prevent normal cells from outgrowing cancer cells, culture conditions need to be selective or organoids initiated exclusively with tumour cells36. In contrast to organoids derived from normal bronchial epithelial cells, NSCLC-derived 3D cultures produce more solid spheres lacking apico-basal polarity37.
Different strategies have been used for the derivation of patient-derived lung cancer organoids. One approach exploited the high prevalence of TP53 mutations in NSCLC6,38 and used the TP53 stabilising drug Nutlin-3a to selectively inhibit the growth of cells with wild-type TP53, thereby enriching for mutant tumour cells39. In an alternative approach, tumour-derived organoids were established from five different lung cancer types, including LUSC, by using a minimum basal medium (MBM) that does not support the expansion of normal airway epithelial cells40. To generate patient-matched normal control lung organoids, the MBM was supplemented with Wnt family member 3A (WNT3A) and inhibitors of the transforming growth factor-beta (TGFβ) and bone morphogenic protein (BMP) signalling pathways. With both methodologies, LUSC-derived organoids retained the histology and overall pattern of genomic alterations of the original tumour39,40. Yet, a subset of LUAD-derived organoids exhibited higher variant allele frequencies than the original tumours40. While this was suggested to be the result of in vitro tumour cell enrichment, rather than of the acquisition of novel mutations ex vivo, the possibility of ongoing tumour evolution in organoid cultures cannot be ruled out.
Following cryopreservation, around 75% of LUSC-derived organoids can be successfully reconstituted, enabling the generation of biobanks40. In drug screening trials, the response of individual lung cancer organoids varied according to their mutational profile, highlighting their potential as tools for predicting patient-specific therapeutic responses, as well as for drug development studies39,40. However, given the propensity of NSCLC-derived organoid cultures to be outgrown by normal cells, tumour purity should be verified regularly through a combination of strategies, including immunostaining and genetic profiling41.
Autochthonous murine models of LUSC
Chemical carcinogenesis
In vivo lung cancer modelling was at first limited to the study of spontaneously developing lung cancers in susceptible mouse strains such as A/J and SWR42. These strains have since been widely exploited in chemical carcinogenesis models43. Most chemical models have focused on the application of cigarette smoke44,45 and its toxic components to induce LUSC46. Exposing female B6C3F1 mice to cigarette smoke was shown to induce broad airway carcinogenesis, predominantly adenoma, and adenocarcinoma, but very few incidences (4.2%) of LUSC47. Individual toxic cigarette smoke components have shown more specific induction of LUSC development.
Benzo(a)pyrene, a polycyclic aromatic hydrocarbon commonly found in the environment, cigarette smoke, and coal tar, has shown an almost exclusive induction of LUSC in ~80% of C3H/He mice when administered intratracheally alongside charcoal powder. Yet, this selective induction of LUSC is highly dose- and regimen-dependent, with 74% of incidences characterised as either LUAD or papillary-type adenomas when the dose of Benzo(a)pyrene was halved. Doubling the dosing regimen length restored almost exclusive induction of LUSC (87% of mice) at the lower dose of Benzo(a)pyrene. Similar results were observed in the C57BL/6 strain48. 3-methylcholanthrene (MCA), another polycyclic aromatic hydrocarbon has been shown to induce LUSC when intratracheally injected into BC3F1 mice with 86% incidence, 2.5−6.5 months following treatment. However, induction is strain- and regimen-dependent with LUSC observed only in 6% of DBA/2 mice 7 months after treatment49. Ultimately, it is likely that the necessity of intratracheal administration of polycyclic hydrocarbons has favoured the use of topically applied agents such as nitrosourea derivatives to study LUSC progression.
Since the discovery of the toxicity of nitrosamine compounds, commonly found in processed meats and cigarette smoke, they have been used in models of several types of cancer50,51,52. N-nitroso-tris-chloroethylurea (NTCU), a triple chlorinated nitrosamine derivative, has an almost exclusive propensity to induce the development of LUSC when topically applied to female Swiss mice compared to other nitrosoalkylureas, which widely trigger the development of skin carcinomas, LUAD and leukaemia50. Despite this, dosing regimen, treatment length (2−32 weeks), mouse strain and sex all affect NTCU-induced carcinogenesis53,54,55. Overall, SWR/J, NIH Swiss, A/J, BALB/cJ, and FVB/NJ have been shown to be sensitive to NTCU, with females displaying higher susceptibility than males. Resistant strains included AKR/J, 129/svJ, and C57BL/6J56.
The epithelial histological changes induced by NTCU application are representative of the step-wise evolution of human LUSC52,54,57. Early NTCU-induced changes have been shown to begin in the trachea, with Keratin 5 (KRT5)+ and KRT14+ tracheal basal cell dysplasia and loss of Club and ciliated cells prior to bronchial dysplasia development57. NTCU-induced pre-invasive lesions express oxidative phosphorylation proteins at higher levels than control-treated lungs, akin to human bronchial pre-invasive lesions, indicating increased cellular respiration58.
Linkage disequilibrium genetic mapping identified three genetic loci significantly associated with NTCU susceptibility: D1Mit169, D3Mit178, and D18Mit91, which are syntenic to the human chromosome loci 6q12−14, 3q26.2−26.31, and 5q23−31, respectively56. This suggests that NTCU-induced LUSC may resemble human LUSC genetically, as 3q26 amplification is the most frequent human LUSC genetic alteration16 and losses in chromosome 5q have been described as common early clonal events, potentially key for LUSC induction5. Whole-exome sequencing of NTCU-induced lung tumours in NIH-Swiss mice revealed that 80% (47/59) of recurrently mutated genes were also altered in human LUSC tumours, these included Kmt2d, Zeb2, Braf, and Igf2r59. Analysis of transcriptional profiles of laser capture microdissected NTCU-induced LUSC in NIH-Swiss mice has shown that the top 150 dysregulated genes in human LUSC are also altered in NTCU-induced LUSC53. RNA sequencing studies of airway brushings from NTCU-treated mice have linked the PI3K pathway and activation of nuclear factor kappa B (NFκB) with NTCU-induced LUSC60. The transcriptomic similarity with human LUSC has allowed NTCU-induced LUSC gene expression to be included in a dataset alongside human bronchial biopsies, brushes, and TCGA LUSC tumours, to identify four molecular subtypes of human premalignant lesions61.
Although further understanding of the genetic triggers and evolution in chemical models is still required, it is clear that NTCU-induced LUSC has the potential to recapitulate some of the heterogeneity, mutational burden, and tumour−microenvironment interactions occurring during different stages of human LUSC development, making it an ideal substrate for intervention trials. Chemopreventative studies thus far have focused on evaluating the potential of natural herbs62, vitamin levels63, antioxidants64, and anti-inflammatory agents65 to protect against NTCU-induced LUSC development.
Genetically-engineered mouse models
The high genomic complexity of LUSC and the extensive inter- and intra-tumour heterogeneity5,6 have hindered the development of genetically-engineered mouse models (GEMMs) that accurately recapitulate the molecular alterations and histopathology of the human disease. Yet, continuous efforts are being made to generate clinically relevant transgenic LUSC models (Table 1).
Expression of Kirsten rat sarcoma viral oncogene (Kras) mutant KrasG12D in the murine airway epithelium is sufficient to drive LUAD formation66. However, when combined with additional somatic alterations, mice may develop tumours with features of LUSC. Deletion of the tumour suppressor serine/threonine kinase 11 (Stk11, also known as Lkb1) in the context of oncogenic KRAS leads to the formation of LUAD, lung adenosquamous tumours (LUASC), and LUSC67. Longitudinal analyses of this model revealed that LUSC arises from squamous differentiation of LUAD lesions, with LUASC constituting an intermediate state68. This transition, which is accompanied by epigenetic de-repression of Sox2, ∆Np63, nerve growth factor receptor (Ngfr), and Krt5, as well as decreased collagen deposition-associated extracellular matrix (ECM) remodelling, also occurred following Stk11 ablation in established KrasG12D-induced LUAD68,69. However, mutations or downregulation of STK11 are rare in human LUSC (Fig. 3) and more prevalent in LUAD38.
Oncoprint showing frequency of genetic and transcriptional changes in the indicated genes across 469 lung squamous cell carcinoma samples from human donors included in The Cancer Genome Atlas (TCGA) PanCancer Atlas dataset. Normal adjacent tissue samples in the cohort were used as a reference for gene expression changes (z-score threshold ± 2.0) (downloaded from https://www.cbioportal.org/).
Depletion of TGFβ receptor 2 (Tgfbr2) concurrently to KrasG12D activation substantially increased LUSC incidence, without eliminating LUAD formation70. LUSC development was associated with dampened extracellular signal-regulated protein kinase 1 and 2 (ERK1/2) activity and upregulation of Sox2. While LUSC appeared predominantly along the bronchioles, LUAD tumours were located in the alveolar space, resembling human pathology. Similar spatial segregation of the two tumour types was seen when lung tumourigenesis was induced by the concomitant activation of KrasG12D and ablation of F-box/WD repeat-containing protein 7 (Fbxw7)71. In this model, LUSC and LUAD arose at 1:1 ratio, enabling side-by-side comparison of cisplatin chemotherapy response in both tumour types. There are, however, caveats to the use of KrasG12D in GEMMs of LUSC. Activating mutations of KRAS are around ten times more prevalent in LUAD than in LUSC6,38. Additionally, multi-region whole-exome sequencing analyses indicate that, when present in LUSC, these tend to occur late during tumour evolution5.
Although rarely affected by genetic alterations, the serine/threonine kinase component of inhibitor of nuclear factor kappa B kinase complex (CHUK, also known as IKKA) is downregulated in a fraction of human LUSC cases. Knock-in mice expressing a kinase-dead form of CHUK (ChukK44A) develop severe skin lesions and spontaneous LUSC72. Re-introduction of wild-type Chuk under the control of the Loricrin promoter rescued the skin phenotype, while LUSC developed with 100% penetrance. Tumours exhibited elevated ERK activity and decreased p53 levels and their formation was preceded by increased expression of ∆Np63 and tripartite motif-containing 29 (TRIM29), inflammation, and lung enlargement.
Additional GEMMs have been developed by targeting components of core pathways significantly altered across large cohorts of LUSC patients. Alterations in the PI3K/AKT pathway are seen in around half of LUSC cases6. PTEN is mutated or deleted in ~21% of LUSC cases (Fig. 3), and these alterations tend to occur during the early stages of tumour evolution5. Ablation of both Pten and Stk11 in the airway epithelium induced LUSC formation with high penetrance, although some phenotypic variations were seen depending on the induction method73,74. Pten- and Stk11-deficient LUSC formed both near the main bronchi and at peripheral locations and exhibited high levels of phospho-AKT and low ERK activity73,74. A subpopulation of tumour cells, identified by co-expression of epithelial cell adhesion molecule (EPCAM), the basal cell marker NGFR, and the bronchioalveolar stem cell (BASC) marker stem cell antigen-1 (SCA1, also known as LY6A), was found to have the ability to propagate the tumour upon implantation into immunocompromised mice73. However, Sca1 does not have a human orthologue, so the relevance of this cellular population for human cancer has not to be established.
SOX2 gains and upregulation occur in ~40% and ~77% of LUSC cases, respectively (Fig. 3). Enforced expression of Sox2 in the murine airway epithelium, however, is not enough to drive LUSC formation75,76. Instead, it can result in bronchial hyperplasia and eventually LUAD with aberrant TRP63 expression75. Yet, a few genetic alterations have been found to cooperate with Sox2 to promote LUSC formation. Overexpression of Sox2 coupled with the loss of Stk11 induces predominantly the formation of peripheral LUSC, with a fraction of tumours expressing low levels of NK2 homeobox 1 (NKX2-1)77,78. Further co-deletion of Nkx2-1 in this model, significantly reduced tumour latency78. Longitudinal analyses suggested that the combination of enforced Sox2 expression with co-ablation of Stk11 and Nkx2-1 caused the formation of early mucinous LUAD lesions that differentiated into LUSC over time78. This is reminiscent of what occurs in the KrasG12D; Stk11–/– model68, and supports a previously identified role for NKX2-1 in suppressing squamous differentiation76. Indeed, ablation of Nkx2-1 with concurrent Sox2 overexpression has been shown to be sufficient to drive LUSC formation in mice76.
The great majority of LUSC cases display cell-cycle dysregulation through alterations in the tumour suppressors genes CDKN2A and RB16. Deletions or mutations in CDKN2A and/or its neighbouring gene CDKN2B occur in approximately 42% of LUSC tumours (Fig. 3). Concurrent ablation of Cdkn2ab and Pten in the murine airway induced the formation of a range of lung neoplasias, but no instances of LUSC were observed79. In contrast, co-deletion of Cdkn2ab and Pten concomitant with enforced expression of Sox2 resulted solely in pre-invasive squamous lesions and LUSC79, highlighting the key role of SOX2 in promoting a squamous fate. In an alternative model where PI3K dysregulation is driven by expression of a constitutively active form of p110α (Pik3caH1047R), concurrent overexpression of Sox2 and loss of Cdkn2ab also led to the formation of LUSC with high penetrance80. Ablation of nuclear receptor binding SET domain protein 3 (NSD3), frequently upregulated in human LUSC, significantly extended animal survival and reduced tumour size in this model, indicating an oncogenic role for this methyltransferase in LUSC80.
Transplantation models
Allogenic and syngeneic models
Murine airway epithelial cells may be genetically modified in vitro and transplanted either as allografts into immunocompromised mice or into syngeneic immunocompetent mouse hosts to evaluate their ability to form tumours in vivo. Immunocompromised host strains used in transplantation models include athymic nude (nu/nu), non-obese diabetic/severe combined immunodeficiency (NOD/scid), and NOD/scid/Il2rgnull (NSG). These systems have been used as an alternative to GEMMs for the phenotypic characterisation of candidate LUSC driver genes, assessment of genetic interactions, investigation of tumour cell-of-origin, and evaluation of therapeutic candidates (Table 2). Syngeneic models have also been established using tumours or cell lines derived from chemically-induced or transgenic murine LUSC models32,35,81,82,83 (Table 2). The derivation of cell lines from murine LUSC normally involves serial passaging in vivo and in vitro to select for cell populations capable of engraftment.
Before implantation, grafts can be modified to express a luciferase reporter, allowing for non-invasive monitoring of tumour growth and metastasis via bioluminescence imaging32,81,84,85. Implantation site may vary, but common establishment routes include subcutaneous graft, intravenous inoculation, and orthotopic injection in the lungs. LUSC allogenic and syngeneic models have been successfully used for the evaluation of therapeutic response to conventional, targeted, and combination therapies31,35,85,86 (Table 3).
Xenograft models
Xenograft models involve the injection of either human cancer cell lines (CDXs) or patient-derived tissue (PDXs) into immunocompromised mice. After implantation, often in combination with a basement membrane mix, cells are left for tumour engraftment and growth. Xenografts have been applied across many types of cancer87,88. Serial passaging of orthotopic CDXs derived from the LUSC cell lines H520 and SKMES-1 has produced subclones that metastasise to LUSC characteristic sites (i.e., adrenal glands and chest wall)89 and could be used as a xenograft models of LUSC metastasis. However, CDXs are likely to be less reflective of human lung cancers due to inherent in vitro biases on top of xenograft selection.
To increase engraftment rates of injected cells or tumours, host-cell rejections are minimised by using immune-deficient mouse strains. NSG mice are reported to have higher engraftment rates, across cell types, than NOD/scid mice. NOD/scid mice have a complete loss of both T and B cells and impaired functioning of NK cells, whereas NSG mice are more immune-deficient with additional impaired cytokine signalling, requiring costly higher biosecurity measures (www.jax.org). In an effort to reduce the cost and time spent generating PDXs for preclinical cancer modelling the EurOPDX consortium initiative is establishing a large collection of PDX models across 30 pathologies, including NSCLC (www.europdx.eu).
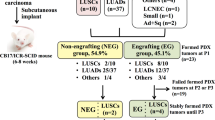
Subcutaneous implantation rates for LUSC PDXs have been recorded as high as ~45% in NOD/scid mice90, whereas renal capsule implantation of patient NSCLC tumours have shown engraftment rates of up to 90% in the same murine strain91. Engraftment rates are dependent on primary tumour and microenvironment characteristics, with poorly differentiated, larger, higher-grade tumours more likely to engraft92. Choice of xenograft implantation site is often a trade-off between tumour engraftment rate and ease of tumour development tracking. Subcutaneous implantation is most common due to the ease of tumour growth tracking. However, this limits the recapitulation of normal tumour−microenvironment interaction and hence is unlikely to directly follow normal LUSC development. Orthotopic xenograft models offer the potential of modelling the tumour microenvironment93. Yet, orthotopic endobronchial implantation of lung cancer cell lines has been shown to have much higher tumour-related mortality rates than subcutaneous implantation94.
NSCLC PDXs have been shown to be representative of the original patient tumour histologically over serial passaging95. Patient tumour copy number alterations are well conserved throughout PDX engraftment and passaging. With copy number variations likely to originate from patient intratumoural heterogeneity96. Ninety-three percent of mutations found in the original tumours were maintained in NSCLC PDXs. However, allele frequencies varied (>1.5 fold), and PDXs harboured a significant amount of additional mutations not found in the original patient tumour with differences attributed to either tumour heterogeneity or PDX-induced tumour evolution90. Transcriptomic comparison between NSCLC patient tumours and tumour-derived PDXs have demonstrated a high degree of correlation97,98. The immune compartment represents a substantial difference between PDXs and their corresponding patient tumours, with immune-response and extracellular matrix gene signatures being the top differentially expressed97,99. The ability of PDXs to broadly recapitulate both the genome and transcriptome of original patient tumours presents an opportunity for individualised therapy screening. This has motivated the use of PDXs for the derivation of tumour cell lines and long-term organoid cultures from patient samples100,101.
NSCLC PDXs have been found to be good predictors of therapeutic outcome and metastases, with 6/7 patients that experienced recurrence or metastases shown to have PDXs non-responsive to corresponding therapies91. LUSC PDXs have been used to improve the stratification of patients likely to respond to fibroblast growth factor receptor (FGFR) inhibitors. Contrary to previous clinical trials that used FGFR1 amplification, a genetic change that occurs in ~17% of human LUSC (cBioPortal, TCGA PanCancer Atlas), as a predictor of inhibitor response, PDX responses demonstrated that high FGFR1 RNA levels instead were indicative of FGFR inhibitor-induced tumour cell differentiation and enhanced tumour cell death, particularly when combined with cisplatin99.
Comparative studies of LUSC PDXs carrying different PI3K pathway alterations indicate that PDXs expressing mutant PIK3CA are more likely to respond to the PI3K inhibitor BKM120 than those with PIK3CA amplification or PTEN loss. BKM120-responsive mutant PIK3CA PDXs were further found to display loss of CDKN2A, which encodes the cell-cycle regulator p16, an inhibitor of cyclin-dependent kinases 4 and 6 (CDK4/6) activity. When compared to single therapies, combined PI3K and CDK4/6 inhibition, resulted in enhanced anti-tumour effects, including tumour regression102.
LUSC cell-of-origin
The resemblance of LUSC to airway basal epithelial cells highlighted them as the suspected cellular origin of this tumour type. However, this view has been challenged by studies in murine LUSC models.
In the great majority of inducible GEMMs of LUSC, driver mutations are activated via intranasal or intratracheal delivery of virus expressing Cre under the control of a constitutive promoter (e.g., cytomegalovirus, CMV) (Table 1). When used on the naive airways, these methods preferentially lead to infection of secretoglobin family 1A member 1 (SCGB1A1)+ Club cells in the distal airways31,70,79, introducing bias for cell-of-origin investigation. The use of transgenic murine lines driving basal cell-specific tamoxifen-inducible Cre expression is frequently unfeasible for lung carcinogenesis studies, as it may result in faster cancer formation in other epithelia31,71. In autochthonous LUSC models, these limitations have been overcome by using cell-type-restricted Cre-expressing viruses and/or by depleting the airways from luminal cells (using naphthalene or SO2) to allow infection of the underlying basal cells31,71,79. This has allowed assessment of the ability of distinct airway epithelial subpopulations to initiate LUSC in response to defined genetic drivers.
Lineage tracing analyses following Ad-CMV-Cre administration have tracked the origin of murine LUSC tumours induced by activation of KrasG12D and Tgfbr2 ablation to SCGB1A1+ Club cells70. Targeting the same hits to Krt5-expressing basal cells resulted in substantially lower LUSC incidence103. Similarly, when cell-type-restricted Cre expressing viruses were used to simultaneously activate KrasG12D and delete Fbxw7 in distinct subpopulations of airway epithelial cells, LUSC only developed when targeting SCGB1A1+ Club cells. KRT5+ basal cells did not produce any tumours, whereas ciliated (Forkhead box J1, FOXJ1+) and alveolar Type II (Surfactant protein C, SFTPC+) gave rise to LUAD. Lineage tracing analyses demonstrated that although the targeted KrasG12D; Fbxw7∆/∆ Club cells showed enhanced proliferation shortly after recombination, activation of squamous markers only occurred after the establishment of SCGB1A1+ hyperplastic lesions71.
In the LUAD-LUSC transition model driven by expression of KRASG12D along with STK11 depletion, tumours have been found to originate from SCGB1A1+ Club cells and SFTPC+SCGB1A1+ BASCs, as assessed by allograft assays following 3D organoid culture. Basal cells carrying the same mutations were unable to produce any organoids69. It has been suggested that since basal cells express high levels of epidermal growth factor receptor (EGFR), and co-expression of mutant forms of KRAS and EGFR is deleterious104, activation of oncogenic KRAS in basal cells may lead to oncogene-induced senescence, potentially explaining their inability to propagate either in vitro or in vivo69.
The GEMM driven by co-ablation of Pten and Cdkn2ab, along with Sox2 overexpression, combines molecular alterations occurring early during LUSC evolution and is among the few murine models that have been reported to result solely in the formation of squamous lung tumours. Remarkably, these three hits could lead to LUSC development when targeted to KRT14+ basal cells, SCGB1A1+ Club cells, or SFTPC+ Type II cells and/or BASCs. While tumours mainly arose in the bronchi and bronchioles after targeting KRT14+ cells, they were restricted to peripheral locations when SCGB1A1+ or SFTPC+ cells were targeted. In peripheral pre-invasive lesions, replacement of SCGB1A1 expression by KRT5 occurred as lesions progressed to LUSC, with the two markers showing mutually exclusive patterns79.
Transplantation studies have shown that following loss of Keap1 and Tp53 in airway epithelial cells, tumours with features of LUSC only arise from Integrin Alpha 6 (ITGA6)+ basal cells. In contrast to other models where mutant Club cells could give rise to LUSC, tracheal luminal cells carrying the same alterations did not generate any tumours, while peripheral luminal cells gave rise to LUAD31. This suggests differential susceptibility among distinct subpopulations of airway epithelial progenitor cells to specific genetic hits.
Tumour microenvironment
Reciprocal interactions between lung cancer cells and both the stromal and immune components of their microenvironment influence not only the evolution of the tumour105,106, but also its response to immune-based therapies7.
PDX-derived cell lines have been applied to 3D co-culture systems with cancer-associated fibroblasts (CAFs) to model interactions with the tumour microenvironment101. It was found that forced-expression of SOX2 stimulates transition from hyperplasia to dysplasia and CAFs promote the acquisition of an invasive phenotype, while unexpectedly, suppressing the SOX2-induced dysplastic phenotype.
The angiogenic landscape of LUSC displays intratumour and interpatient heterogeneity. In-depth multi-omic phenotyping of tumour endothelial cells has highlighted the possibility of targeting endothelial subtypes as a potential therapeutic target107. In H520 CDXs, the vascular disrupting agent CKD-516 showed enhanced anti-tumour efficacy when administered in combination with radiotherapy compared to single treatments108.
Tumour immune microenvironment
In comparison with LUAD, human LUSC tumours have been found to be enriched in neutrophils, diverse subsets of CD4+ cells—including immunosuppressive regulatory T (Treg) cells—and CD8+ cells expressing programmed cell death (PD1, encoded by PDCD1), while containing lower numbers of macrophages109. Analyses of the tumour immune microenvironment (TIME) in different GEMMs of LUSC have revealed, in most instances, several similarities with their human counterparts (Table 1).
The presence of large neutrophilic infiltrates has been linked to increased levels of neutrophil recruitment-associated molecules, including CXC ligands, in distinct LUSC models70,73,78. Chromatin immunoprecipitation-RNA sequencing analyses on a set of murine LUSC and LUAD tumours, suggested that SOX2 and NKX2-1 exert antagonistic effects on the regulation of Cxcl5 expression. Indeed, either overexpression of Sox2 or ablation of Nkx2-1 in a KrasG12D; Tp53∆/∆ LUAD model—which normally does not contain neutrophilic infiltrates—resulted in both CXCL5 expression and neutrophil recruitment. The ability of SOX2 to induce expression of CXCL6, the human orthologue of Cxcl5, was conserved in human lung cancer cells in vitro. However, altering the levels of NKX2-1 had no effect on CXCL6 expression in this setting78.
In the LUAD-to-LUSC transdifferentiation model driven by Sox2 overexpression alongside co-deletion of Stk11 and Nkx2-1, it was found that, in contrast to peripheral blood neutrophils, tumour-associated neutrophils (TANs) produce increased levels of ROS and include a subpopulation of cells expressing high levels of sialic acid-binding immunoglobulin-like lectin F (SiglecF)67. Both features are suggestive of a tumour-supportive role78,110,111. It would be important to determine whether TANs present in tumours from LUSC patients and in murine models exhibiting squamous features throughout LUSC development display similar properties.
High infiltration of CD11b+ myeloid populations has been detected in orthotopic syngeneic LUSC models35. The ChukK44A/K44A; Tg(Loricrin-CHUK) GEMM exhibits extensive infiltration of F4/80+ macrophages within tumours. Macrophage depletion or haematopoietic reconstitution with wild-type bone marrow following irradiation of mutant mice both prevented LUSC development. Macrophage depletion was associated with decreased oxidative stress-induced DNA damage, suggesting that Chuk-mutant macrophages promote tumour development through enhanced ROS production72.
Immune suppression appears to play a role in murine LUSC progression. Increased levels of several immunosuppressive molecules, including programmed cell death ligand 1 (PDL1, also known as CD274), have been observed in various transgenic LUSC models71,73,78,79. Analyses across three different mouse strains demonstrated expression of PDL1 in two-thirds of NTCU-induced LUSC cases and an overall frequency of less than 5% of CD4+ and CD8+ tumour infiltrating lymphocytes; while lymphoid aggregates were evident close to the main bronchi and blood vessels53. Additionally, forkhead box P3 (FOXP3)+ Treg cells have been found to be enriched in some murine LUSC tumours73,78. In the Pten∆/∆; Stk11∆/∆ mouse LUSC model, the ratio of CD8+ T cells to Tregs showed an inverse correlation with tumour burden73. These attributes emphasise the potential of these autochthonous LUSC models for the evaluation of immune-based therapies.
Bioinformatic studies have identified a subset of human LUSC tumours enriched in inflammatory monocytes (IMs) expressing CD1481. In an immune-competent orthotopic murine LUSC model, it was shown that in response to tumour necrosis factor alpha (TNAα)/NFκB pathway activation, LUSC cells secrete C−C motif chemokine ligand 2 (CCL2), leading to IM recruitment. IMs in turn, were found to express high levels of factor XIIIA (FXIIIA, encoded by F13a1) and thereby to promote fibrin cross-linking, which favoured invadopodia formation in LUSC cells and metastasis. In human LUSC, increased F13A1 expression and high levels of intra-tumoural fibrin cross-linking were associated with worse overall and recurrence-free patient survival, respectively81.
Application of LUSC models for immunotherapy studies
Tumour establishment and growth are reliant on the ability to evade immune-mediated clearance. One such way immune evasion can be achieved is tumour-expression of PDL1. PD1 is expressed on T cells and binds to its ligand PDL1 on tumour cells and tumour-associated stromal cells. This results in inhibition of T cell activation and T cell exhaustion, preventing immune-mediated clearance. PD1/PDL1 monoclonal antibodies used effectively in the treatment of NSCLC include: nivolumab, pembrolizumab, atezolizumab, and durvalumab112. However, these achieve a response in only ~15−25% of patients with high incidence of immune-mediated side-effects113. A better understanding of the mechanisms through which LUSC cells evade immune response is key for the identification of biomarkers for patient stratification and the development of combination therapies that can improve response to current treatments.
Tumours with high mutational load have shown improved responses to immune checkpoint inhibitors in a variety of cancers, including NSCLC114. High non-synonymous mutational tumour burden in NSCLC has been associated with better patient response to pembrolizumab115. Similarly, the mutational burden of circulating tumour cells in blood has been identified as a potential biomarker for immunotherapy in NSCLC116. These observations emphasise the importance for preclinical immunotherapy models to represent a high mutational/neo-antigen load, characteristic of LUSC. LUAD GEMMs have shown a lower non-synonymous mutational load than human adenocarcinomas (0.7 mutations per Mb compared to 1.97 mutations per Mb)117, highlighting potential limitations of GEMMs for immunotherapy studies.
Both the presence and localisation of T cell populations affect immunotherapy treatment efficacy and patient outcomes. Increased tumour infiltration of FOXP3+ Tregs is associated with a decrease in recurrence free survival in NSCLC patients118. Whereas high diversity of PD1+CD8+ T cell repertoires in peripheral blood has been associated with improved response to anti-PD1/PDL1 therapies and increased progression-free survival119.
High levels of neutrophil infiltration predict poor response to immunotherapy in a subset of myeloid-rich NSCLC tumours. Using multiplexed-IHC, it was found that the ratio of CD8+ T cells to neutrophils within the tumour mass was able to distinguish between patients who were responsive to immune checkpoint inhibitors and those with stable or progressive disease120. In the Pten- and Stk11-deficient GEMM LUSC model, inhibition of neutrophil’s CXC chemokine receptors 1 and 2 (CXCR1/2)—which regulate their recruitment to the TIME110—combined with anti-PD1 treatment, decreased tumour masses, unlike single therapy treatment120. This was associated with a reduction of Lymphocyte antigen 6 complex locus G (Ly6G)+ neutrophils in the tumour mass and increased relocation of CD8+ T cells from the tumour periphery into the tumour mass, suggesting that TANs impede CD8+ T cell infiltration120.
The combination of targeted inhibitors alongside anti-PD1 treatment has shown increased tumour responses, thought to be mediated by increased immune system activation. Inhibition of WEE1 G2 checkpoint kinase (WEE1), a CDK1 negative-regulator, alongside PD1 blockade resulted in diminished tumour growth in two LUSC subcutaneous syngeneic models. The associated increase in DNA damage triggered by WEE1 inhibition and consequent CDK1 activation stimulated expression of type 1 interferon (IFN) signalling and major histocompatibility complex (MHC) class I antigen presentation genes in LUSC cells, increased tumour-infiltrating CD8+ T cells, and decreased TANs35. This demonstrates the enhanced efficacy of combination therapies in the treatment of LUSC by further unleashing the immune system. In renal, colon, and prostate syngeneic cancer models, orthotopic implanted grafts had a lesser response to immunotherapeutic agents when compared to subcutaneous implants121. It would be important to address whether syngeneic LUSC models display a similar behaviour, so that the implantation site can be considered during experimental design.
The creation of humanised PDXs allows modelling of human immune and stromal interactions in mice122, enabling the application of xenograft models to lung immunotherapy studies. Humanised murine models have been generated by reconstituting the immune system of mice with either human peripheral blood mononucleated cells (PBMCs) or haematopoietic stem and progenitor cells (HSPCs). PBMC implantation has been shown to reconstitute the immune system faster than HSPC implantation (4 weeks as opposed to 10−14 weeks). Studies utilising humanised NSCLC CDX/PDX models have demonstrated the ability of anti-PD1 and anti-PDL1 to slow tumour growth in vivo123,124 (Table 3), presenting new opportunities for combination trials.
Conclusions and perspectives
Molecular profiling analysis of patient samples corresponding to different developmental stages of LUSC has provided insights into the alterations underlying cancer initiation and progression5,6,14,105,106. This has led to the identification of candidate driver genes and biomarkers, and fuelled the refinement of LUSC models to more closely mimic human pathology. Advances in cell culture methods for airway epithelial cells, such as 3D organoid cultures, and in genetic manipulation technologies, including CRISPR/Cas9 gene editing, have further contributed to streamlining lung cancer modelling. An increasingly growing array of in vitro and in vivo systems, each with their own strengths and limitations, are now used for both fundamental and translational LUSC research.
A major area of the current investigation is the interactions between LUSC cells and their microenvironment, particularly the TIME7. Due to their immune proficiency, the great majority of studies have thus far utilised GEMMs and murine syngeneic models. Yet, the development of humanised PDXs and co-culture systems is opening new avenues for research using human material. For example, autologous co-cultures of tumour-derived organoids and PBMCs from NSCLC patients, including one case of LUSC, have been used to assess the ability of T cells to recognise and eliminate tumour cells125. In LUAD, the high relative expression of receptor tyrosine kinase-like orphan receptor-1 (ROR1) in lung tumours to normal tissue has been exploited by engineering ROR1-specific chimeric antigen receptor (CAR) T cells, showing promising results in both static and dynamic co-culture systems with A549 LUAD cells126.
With programmes for early cancer detection and intervention becoming top priorities, the availability of models recapitulating the pre-invasive stages of LUSC is essential for the identification of relevant biomarkers and therapeutic targets. NTCU-induced murine LUSC currently is one of the best characterised models in this sense. A better understanding of the genetic events underlying NTCU-induced carcinogenesis will be key for its optimal use in preclinical studies. While the combination of genetic manipulation techniques, pharmacological modulators, and 2D and 3D co-culture systems is expected to support the creation of novel human premalignant LUSC in vitro models.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J. Clin. 68, 394–424 (2018).
Herbst, R. S., Morgensztern, D. & Boshoff, C. The biology and management of non-small cell lung cancer. Nature 553, 446–454 (2018).
Gazdar, A. F., Girard, L., Lockwood, W. W., Lam, W. L. & Minna, J. D. Lung cancer cell lines as tools for biomedical discovery and research. J. Natl Cancer Inst. 102, 1310–1321 (2010).
Gazdar, A. F., Hirsch, F. R. & Minna, J. D. From mice to men and back: an assessment of preclinical model systems for the study of lung cancers. J. Thorac. Oncol.: Off. Publ. Int. Assoc. Study Lung Cancer 11, 287–299 (2016).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Altorki, N. K. et al. The lung microenvironment: an important regulator of tumour growth and metastasis. Nat. Rev. Cancer 19, 9–31 (2019).
Auerbach, O., Stout, A. P., Hammond, E. C. & Garfinkel, L. Changes in bronchial epithelium in relation to cigarette smoking and in relation to lung cancer. N. Engl. J. Med. 265, 253–267 (1961).
Alberg, A. J., Brock, M. V., Ford, J. G., Samet, J. M. & Spivack, S. D. Epidemiology of lung cancer: diagnosis and management of lung cancer, 3rd ed: American College of Chest. Physicians evidence-based clinical practice guidelines. Chest 143, e1S−e29S (2013).
Yoshida, K. et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266–272 (2020).
Sundaresan, V. et al. Somatic genetic changes in lung cancer and precancerous lesions. Ann. Oncol.: Off. J. Eur. Soc. Med. Oncol. 6(Suppl. 1), 27–31 (1995).
Ooi, A. T. et al. Molecular profiling of premalignant lesions in lung squamous cell carcinomas identifies mechanisms involved in stepwise carcinogenesis. Cancer Prev. Res. 7, 487–495 (2014).
van Boerdonk, R. A. et al. DNA copy number aberrations in endobronchial lesions: a validated predictor for cancer. Thorax 69, 451–457 (2014).
Teixeira, V. H. et al. Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions. Nat. Med. 25, 517–525 (2019).
McCaughan, F. et al. Progressive 3q amplification consistently targets SOX2 in preinvasive squamous lung cancer. Am. J. Respir. Crit. Care Med. 182, 83–91 (2010).
Fields, A. P., Justilien, V. & Murray, N. R. The chromosome 3q26 OncCassette: a multigenic driver of human cancer. Adv. Biol. Regul. 60, 47–63 (2016).
Torres-Ayuso, P. et al. TNIK is a therapeutic target in lung squamous cell carcinoma and regulates FAK activation through Merlin. Cancer Discov. https://doi.org/10.1158/2159-8290.Cd-20-0797 (2021).
Bass, A. J. et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat. Genet. 41, 1238–1242 (2009).
Whitcutt, M. J., Adler, K. B. & Wu, R. A biphasic chamber system for maintaining polarity of differentiation of cultured respiratory tract epithelial cells. In Vitro Cell. Dev. Biol.: J. Tissue Cult. Assoc. 24, 420–428 (1988).
Kim, B. R. et al. SOX2 and PI3K cooperate to induce and stabilize a squamous-committed stem cell injury state during lung squamous cell carcinoma pathogenesis. PLoS Biol. 14, e1002581 (2016).
Correia, L. L. et al. SOX2 drives bronchial dysplasia in a novel organotypic model of early human squamous lung cancer. Am. J. Respir. Crit. Care Med. 195, 1494–1508 (2017).
Liu, X. et al. ROCK inhibitor and feeder cells induce the conditional reprogramming of epithelial cells. Am. J. Pathol. 180, 599–607 (2012).
Butler, C. R. et al. Rapid expansion of human epithelial stem cells suitable for airway tissue engineering. Am. J. Respir. Crit. Care Med. 194, 156–168 (2016).
Crystal, A. S. et al. Patient-derived models of acquired resistance can identify effective drug combinations for cancer. Science 346, 1480–1486 (2014).
Park, K. S. et al. CRIPTO1 expression in EGFR-mutant NSCLC elicits intrinsic EGFR-inhibitor resistance. J. Clin. Invest. 124, 3003–3015 (2014).
Sette, G. et al. Conditionally reprogrammed cells (CRC) methodology does not allow the in vitro expansion of patient-derived primary and metastatic lung cancer cells. Int J. Cancer 143, 88–99 (2018).
Gao, B. et al. Non-malignant respiratory epithelial cells preferentially proliferate from resected non-small cell lung cancer specimens cultured under conditionally reprogrammed conditions. Oncotarget 8, 11114–11126 (2017).
Hynds, R. E. et al. Expansion of airway basal epithelial cells from primary human non-small cell lung cancer tumors. Int J. Cancer 143, 160–166 (2018).
Piotrowska, Z. et al. Heterogeneity underlies the emergence of EGFRT790 wild-type clones following treatment of T790M-positive cancers with a third-generation EGFR inhibitor. Cancer Discov. 5, 713–722 (2015).
Barkauskas, C. E. et al. Lung organoids: current uses and future promise. Development 144, 986–997 (2017).
Jeong, Y. et al. Role of KEAP1/NRF2 and TP53 mutations in lung squamous cell carcinoma development and radiation resistance. Cancer Discov. 7, 86–101 (2017).
Liu, Y. et al. Chromosome 3q26 gain is an early event driving coordinated overexpression of the PRKCI, SOX2, and ECT2 oncogenes in lung squamous cell carcinoma. Cell Rep. 30, 771–782.e776 (2020).
Justilien, V., Jameison, L., Der, C. J., Rossman, K. L. & Fields, A. P. Oncogenic activity of Ect2 is regulated through protein kinase C iota-mediated phosphorylation. J. Biol. Chem. 286, 8149–8157 (2011).
Justilien, V. et al. The PRKCI and SOX2 oncogenes are coamplified and cooperate to activate Hedgehog signaling in lung squamous cell carcinoma. Cancer Cell 25, 139–151 (2014).
Hai, J. et al. Generation of genetically engineered mouse lung organoid models for squamous cell lung cancers allows for the study of combinatorial immunotherapy. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. https://doi.org/10.1158/1078-0432.Ccr-19-1627 (2020).
Drost, J. & Clevers, H. Organoids in cancer research. Nat. Rev. Cancer 18, 407–418 (2018).
Fessart, D., Begueret, H. & Delom, F. Three-dimensional culture model to distinguish normal from malignant human bronchial epithelial cells. Eur. Respir. J. 42, 1345–1356 (2013).
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Sachs, N. et al. Long-term expanding human airway organoids for disease modeling. EMBO J. https://doi.org/10.15252/embj.2018100300 (2019).
Kim, M. et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 10, 3991 (2019).
Dijkstra, K. K. et al. Challenges in establishing pure lung cancer organoids limit their utility for personalized medicine. Cell Rep. 31, 107588 (2020).
Wakamatsu, N., Devereux, T. R., Hong, H. H. & Sills, R. C. Overview of the molecular carcinogenesis of mouse lung tumor models of human lung cancer. Toxicol. Pathol. 35, 75–80 (2007).
Shimkin, M. B. & Stoner, G. D. Lung tumors in mice: application to carcinogenesis bioassay. Adv. Cancer Res. 21, 1–58 (1975).
Witschi, H. A/J mouse as a model for lung tumorigenesis caused by tobacco smoke: strengths and weaknesses. Exp. Lung Res. 31, 3–18 (2005).
Witschi, H., Espiritu, I., Dance, S. T. & Miller, M. S. A mouse lung tumor model of tobacco smoke carcinogenesis. Toxicol. Sci. 68, 322–330 (2002).
Dwyer-Nield, L. D., McArthur, D. G., Tennis, M. A., Merrick, D. T. & Keith, R. L. An improved murine pre-malignant squamous cell model: tobacco smoke exposure augments NTCU-induced murine airway dysplasia. Cancer Prev. Res. https://doi.org/10.1158/1940-6207.CAPR-20-0332 (2020).
Hutt, J. A. et al. Life-span inhalation exposure to mainstream cigarette smoke induces lung cancer in B6C3F1 mice through genetic and epigenetic pathways. Carcinogenesis 26, 1999–2009 (2005).
Yoshimoto, T. et al. Differential induction of squamous cell carcinomas and adenocarcinomas in mouse lung by intratracheal instillation of benzo(a)pyrene and charcoal powder. Cancer Res. 40, 4301–4307 (1980).
Nettesheim, P. & Hammons, A. S. Induction of squamous cell carcinoma in the respiratory tract of mice. J. Natl Cancer Inst. 47, 697–701 (1971).
Lijinsky, W. & Reuber, M. D. Neoplasms of the skin and other organs observed in Swiss mice treated with nitrosoalkylureas. J. Cancer Res. Clin. Oncol. 114, 245–249 (1988).
Magee, P. N. & Farber, E. Toxic liver injury and carcinogenesis. methylation of rat-liver nucleic acids by dimethylnitrosamine in vivo. Biochemical J. 83, 114–124 (1962).
Rehm, S., Lijinsky, W., Singh, G. & Katyal, S. L. Mouse bronchiolar cell carcinogenesis. Histologic characterization and expression of Clara cell antigen in lesions induced by N-nitrosobis-(2-chloroethyl) ureas. Am. J. Pathol. 139, 413–422 (1991).
Riolobos, L. et al. The effect of mouse strain, sex, and carcinogen dose on toxicity and the development of lung dysplasia and squamous cell carcinomas in mice. Cancer Prev. Res. 12, 507–516 (2019).
Tago, Y. et al. Novel medium-term carcinogenesis model for lung squamous cell carcinoma induced by N-nitroso-tris-chloroethylurea in mice. Cancer Sci. 104, 1560–1566 (2013).
Hudish, T. M. et al. N-nitroso-tris-chloroethylurea induces premalignant squamous dysplasia in mice. Cancer Prev. Res. 5, 283–289 (2012).
Wang, Y. et al. A chemically induced model for squamous cell carcinoma of the lung in mice: histopathology and strain susceptibility. Cancer Res. 64, 1647–1654 (2004).
Ghosh, M. et al. Tracheal dysplasia precedes bronchial dysplasia in mouse model of N-nitroso trischloroethylurea induced squamous cell lung cancer. PLoS One 10, e0122823 (2015).
Beane, J. et al. Detecting the presence and progression of premalignant lung lesions via airway gene expression. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 23, 5091–5100 (2017).
Xiong, D. et al. Novel mutational landscapes and expression signatures of lung squamous cell carcinoma. Oncotarget 9, 7424–7441 (2018).
Pan, J. et al. Airway brushing as a new experimental methodology to detect airway gene expression signatures in mouse lung squamous cell carcinoma. Sci. Rep. 8, 8895 (2018).
Beane, J. E. et al. Molecular subtyping reveals immune alterations associated with progression of bronchial premalignant lesions. Nat. Commun. 10, 1856 (2019).
Wang, Y. et al. Chemoprevention of lung squamous cell carcinoma in mice by a mixture of Chinese herbs. Cancer Prev. Res. 2, 634–640 (2009).
Mazzilli, S. A. et al. Vitamin D repletion reduces the progression of premalignant squamous lesions in the NTCU lung squamous cell carcinoma mouse model. Cancer Prev. Res. 8, 895–904 (2015).
Surien, O., Ghazali, A. R. & Masre, S. F. Histopathological effect of pterostilbene as chemoprevention in N-nitroso-tri-chloroethylurea (NTCU)-induced lung squamous cell carcinoma (SCC) mouse model. Histol. Histopathol. https://doi.org/10.14670/hh-18-247 (2020).
Song, J. M. et al. Dietary diindolylmethane suppresses inflammation-driven lung squamous cell carcinoma in mice. Cancer Prev. Res. 8, 77–85 (2015).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007).
Han, X. et al. Transdifferentiation of lung adenocarcinoma in mice with Lkb1 deficiency to squamous cell carcinoma. Nat. Commun. 5, 3261 (2014).
Zhang, H. et al. Lkb1 inactivation drives lung cancer lineage switching governed by polycomb repressive complex 2. Nat. Commun. 8, 14922 (2017).
Wang, Y. et al. Dysregulated Tgfbr2/ERK-Smad4/SOX2 signaling promotes lung squamous cell carcinoma formation. Cancer Res. 79, 4466–4479 (2019).
Ruiz, E. J. et al. LUBAC determines chemotherapy resistance in squamous cell lung cancer. J. Exp. Med. 216, 450–465 (2019).
Xiao, Z. et al. The pivotal role of IKKalpha in the development of spontaneous lung squamous cell carcinomas. Cancer Cell 23, 527–540 (2013).
Xu, C. et al. Loss of Lkb1 and Pten leads to lung squamous cell carcinoma with elevated PD-L1 expression. Cancer Cell 25, 590–604 (2014).
Liu, J. et al. JNK(1/2) represses Lkb(1)-deficiency-induced lung squamous cell carcinoma progression. Nat. Commun. 10, 2148 (2019).
Lu, Y. et al. Evidence that SOX2 overexpression is oncogenic in the lung. PLoS One 5, e11022 (2010).
Tata, P. R. et al. Developmental history provides a roadmap for the emergence of tumor plasticity. Dev. Cell 44, 679–693.e675 (2018).
Mukhopadhyay, A. et al. Sox2 cooperates with Lkb1 loss in a mouse model of squamous cell lung cancer. Cell Rep. 8, 40–49 (2014).
Mollaoglu, G. et al. The lineage-defining transcription factors SOX2 and NKX2-1 determine lung cancer cell fate and shape the tumor immune microenvironment. Immunity 49, 764–779.e769 (2018).
Ferone, G. et al. SOX2 Is the determining oncogenic switch in promoting lung squamous cell carcinoma from different cells of origin. Cancer Cell 30, 519–532 (2016).
Yuan, G. et al. Elevated NSD3 histone methylation activity drives squamous cell lung cancer. Nature 590, 504–508 (2021).
Porrello, A. et al. Factor XIIIA-expressing inflammatory monocytes promote lung squamous cancer through fibrin cross-linking. Nat. Commun. 9, 1988 (2018).
Kaneko, T. & LePage, G. A. Growth characteristics and drug responses of a murine lung carcinoma in vitro and in vivo. Cancer Res. 38, 2084–2090 (1978).
Azpilikueta, A. et al. Successful immunotherapy against a transplantable mouse squamous lung carcinoma with anti-PD-1 and anti-CD137 monoclonal antibodies. J. Thorac. Oncol.: Off. Publ. Int. Assoc. Study Lung Cancer 11, 524–536 (2016).
Marien, E., Hillen, A., Vanderhoydonc, F., Swinnen, J. V. & Vande Velde, G. Longitudinal microcomputed tomography-derived biomarkers for lung metastasis detection in a syngeneic mouse model: added value to bioluminescence imaging. Lab. Investig. 97, 24–33 (2017).
He, X. et al. Upfront dose-reduced chemotherapy synergizes with immunotherapy to optimize chemoimmunotherapy in squamous cell lung carcinoma. J. Immunother. Cancer https://doi.org/10.1136/jitc-2020-000807 (2020).
Mender, I. et al. Telomerase-mediated strategy for overcoming non-small cell lung cancer targeted therapy and chemotherapy resistance. Neoplasia 20, 826–837 (2018).
Rosfjord, E., Lucas, J., Li, G. & Gerber, H. P. Advances in patient-derived tumor xenografts: from target identification to predicting clinical response rates in oncology. Biochem. Pharm. 91, 135–143 (2014).
Yoshida, G. J. Applications of patient-derived tumor xenograft models and tumor organoids. J. Hematol. Oncol. 13, 4 (2020).
Harrison, E. B. et al. A Circle RNA regulatory axis promotes lung squamous metastasis via CDR1-mediated regulation of Golgi trafficking. Cancer Res. 80, 4972–4985 (2020).
Hao, C. et al. Gene mutations in primary tumors and corresponding patient-derived xenografts derived from non-small cell lung cancer. Cancer Lett. 357, 179–185 (2015).
Dong, X. et al. Patient-derived first generation xenografts of non-small cell lung cancers: promising tools for predicting drug responses for personalized chemotherapy. Clin. Cancer Res. 16, 1442–1451 (2010).
Chen, Y. et al. Tumor characteristics associated with engraftment of patient-derived non-small cell lung cancer xenografts immunocompromised mice. Cancer 125, 3738–3748 (2019).
Kellar, A., Egan, C. & Morris, D. Preclinical murine models for lung cancer: clinical trial applications. BioMed. Res. Int. 2015, 621324 (2015).
McLemore, T. L. et al. Novel intrapulmonary model for orthotopic propagation of human lung cancers in athymic nude mice. Cancer Res. 47, 5132–5140 (1987).
Kita, K. et al. Patient-derived xenograft models of non-small cell lung cancer for evaluating targeted drug sensitivity and resistance. Cancer Sci. 110, 3215–3224 (2019).
Woo, X. Y. et al. Conservation of copy number profiles during engraftment and passaging of patient-derived cancer xenografts. Nat. Genet. 53, 86–99 (2021).
Li, H. et al. Integrated analysis of transcriptome in cancer patient-derived xenografts. PLoS One 10, e0124780 (2015).
Fichtner, I. et al. Establishment of patient-derived non-small cell lung cancer xenografts as models for the identification of predictive biomarkers. Clin. Cancer Res. 14, 6456–6468 (2008).
Weeden, C. E. et al. Cisplatin increases sensitivity to FGFR inhibition in patient-derived xenograft models of lung squamous cell carcinoma. Mol. Cancer Ther. 16, 1610–1622 (2017).
Shi, R. et al. Organoid cultures as preclinical models of non-small cell lung cancer. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 26, 1162–1174 (2020).
Chen, S. et al. Cancer-associated fibroblasts suppress SOX2-induced dysplasia in a lung squamous cancer coculture. Proc. Natl Acad. Sci. USA 115, E11671–e11680 (2018).
Shi, R. et al. Targeting the CDK4/6-Rb pathway enhances response to PI3K inhibition in PIK3CA-mutant lung squamous cell carcinoma. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 24, 5990–6000 (2018).
Malkoski, S. P. et al. Loss of transforming growth factor beta type II receptor increases aggressive tumor behavior and reduces survival in lung adenocarcinoma and squamous cell carcinoma. Clin. Cancer Res.: Off. J. Am. Assoc. Cancer Res. 18, 2173–2183 (2012).
Unni, A. M., Lockwood, W. W., Zejnullahu, K., Lee-Lin, S. Q. & Varmus, H. Evidence that synthetic lethality underlies the mutual exclusivity of oncogenic KRAS and EGFR mutations in lung adenocarcinoma. Elife 4, e06907 (2015).
Pennycuick, A. et al. Immune surveillance in clinical regression of preinvasive squamous cell lung cancer. Cancer Discov. 10, 1489–1499 (2020).
Mascaux, C. et al. Immune evasion before tumour invasion in early lung squamous carcinogenesis. Nature https://doi.org/10.1038/s41586-019-1330-0 (2019).
Goveia, J. et al. An integrated gene expression landscape profiling approach to identify lung tumor endothelial cell heterogeneity and angiogenic candidates. Cancer Cell 37, 421 (2020).
Kim, M. Y. et al. Anti-tumor efficacy of CKD-516 in combination with radiation in xenograft mouse model of lung squamous cell carcinoma. BMC Cancer 20, 1057 (2020).
Kargl, J. et al. Neutrophils dominate the immune cell composition in non-small cell lung cancer. Nat. Commun. 8, 14381 (2017).
Jaillon, S. et al. Neutrophil diversity and plasticity in tumour progression and therapy. Nat. Rev. Cancer 20, 485–503 (2020).
Engblom, C. et al. Osteoblasts remotely supply lung tumors with cancer-promoting SiglecF(high) neutrophils. Science https://doi.org/10.1126/science.aal5081 (2017).
Onoi, K. et al. Immune checkpoint inhibitors for lung cancer treatment: a review. J. Clin. Med. https://doi.org/10.3390/jcm9051362 (2020).
Johnson, D. B., Rioth, M. J. & Horn, L. Immune checkpoint inhibitors in NSCLC. Curr. Treat. Options Oncol. 15, 658–669 (2014).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Wang, Z. et al. Assessment of blood tumor mutational burden as a potential biomarker for immunotherapy in patients with non-small cell lung cancer with use of a next-generation sequencing cancer gene panel. JAMA Oncol. 5, 696–702 (2019).
McFadden, D. G. et al. Mutational landscape of EGFR-, MYC-, and Kras-driven genetically engineered mouse models of lung adenocarcinoma. Proc. Natl Acad. Sci. USA 113, E6409–E6417 (2016).
Shimizu, K. et al. Tumor-infiltrating Foxp3+ regulatory T cells are correlated with cyclooxygenase-2 expression and are associated with recurrence in resected non-small cell lung cancer. J. Thorac. Oncol. 5, 585–590 (2010).
Han, J. et al. TCR repertoire diversity of peripheral PD-1. Cancer Immunol. Res. 8, 146–154 (2020).
Kargl, J. et al. Neutrophil content predicts lymphocyte depletion and anti-PD1 treatment failure in NSCLC. JCI insight https://doi.org/10.1172/jci.insight.130850 (2019).
Devaud, C. et al. Tissues in different anatomical sites can sculpt and vary the tumor microenvironment to affect responses to therapy. Mol. Ther. 22, 18–27 (2014).
Morton, J. J. et al. XactMice: humanizing mouse bone marrow enables microenvironment reconstitution in a patient-derived xenograft model of head and neck cancer. Oncogene 35, 290–300 (2016).
Pyo, K. H. et al. Promising preclinical platform for evaluation of immuno-oncology drugs using Hu-PBL-NSG lung cancer models. Lung Cancer 127, 112–121 (2019).
Lin, S. et al. Establishment of peripheral blood mononuclear cell-derived humanized lung cancer mouse models for studying efficacy of PD-L1/PD-1 targeted immunotherapy. MAbs 10, 1301–1311 (2018).
Dijkstra, K. K. et al. Generation of tumor-reactive T cells by co-culture of peripheral blood lymphocytes and tumor organoids. Cell 174, 1586–1598.e1512 (2018).
Wallstabe, L. et al. ROR1-CAR T cells are effective against lung and breast cancer in advanced microphysiologic 3D tumor models. JCI Insight https://doi.org/10.1172/jci.insight.126345 (2019).
Vanhaesebroeck, B., Stephens, L. & Hawkins, P. PI3K signalling: the path to discovery and understanding. Nat. Rev. Mol. Cell Biol. 13, 195–203 (2012).
Singh, B. et al. p53 regulates cell survival by inhibiting PIK3CA in squamous cell carcinomas. Genes Dev. 16, 984–993 (2002).
Acknowledgements
We want to thank Marie-Belle El Mdawar, Lukas Kalinke, Kyren Lazarus (all from UCL Respiratory), and Rob Hynds (UCL Cancer Institute) for helpful comments during the preparation of this manuscript, and Rebecca Graham (UCL Respiratory) for proofreading. This work was funded by the Wellcome Trust [WT107963AIA] and a CRUK programme award to S.M.J. S.M.J. is further supported by the Roy Castle Lung Cancer Foundation, the Rosetrees Trust, UCLH Charitable Foundation, the Longfonds BREATH consortium, and the MRC UKRMP2 cell therapy platform. Z.E.W. is funded by a CRUK-UCL Centre PhD studentship [A27437].
Author information
Authors and Affiliations
Contributions
S.G.-L. and Z.E.W wrote the paper with input from S.M.J.
Corresponding author
Ethics declarations
Competing interests
Paid advisory boards: 2018 and 2020 Astra-Zeneca; 2018 and 2019 BARD1 Life Sciences; 2018 Achilles Therapeutics; 2019 Johnson and Johnson (S.M.J.). Assistance or travel to meetings: Astra-Zeneca, ATS 2018 San Diego; Takeda, WCLC 2019 Barcelona (S.M.J.). Grant Investigator lead 2018-2028 GRAIL Inc; 2018 Owlstone (S.M.J.). Personal: Spouse works for and owns shares in Astra-Zeneca (S.M.J.). S.G.-L. and Z.E.W. declare no competing interests relating to this work.
Additional information
Peer review information Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Christina Karlsson Rosenthal. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gómez-López, S., Whiteman, Z.E. & Janes, S.M. Mapping lung squamous cell carcinoma pathogenesis through in vitro and in vivo models. Commun Biol 4, 937 (2021). https://doi.org/10.1038/s42003-021-02470-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-02470-x
This article is cited by
-
Micro-CT acquisition and image processing to track and characterize pulmonary nodules in mice
Nature Protocols (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.