Abstract
SARS-CoV-2 Nsp15 is a uridine-specific endoribonuclease with C-terminal catalytic domain belonging to the EndoU family that is highly conserved in coronaviruses. As endoribonuclease activity seems to be responsible for the interference with the innate immune response, Nsp15 emerges as an attractive target for therapeutic intervention. Here we report the first structures with bound nucleotides and show how the enzyme specifically recognizes uridine moiety. In addition to a uridine site we present evidence for a second base binding site that can accommodate any base. The structure with a transition state analog, uridine vanadate, confirms interactions key to catalytic mechanisms. In the presence of manganese ions, the enzyme cleaves unpaired RNAs. This acquired knowledge was instrumental in identifying Tipiracil, an FDA approved drug that is used in the treatment of colorectal cancer, as a potential anti-COVID-19 drug. Using crystallography, biochemical, and whole-cell assays, we demonstrate that Tipiracil inhibits SARS-CoV-2 Nsp15 by interacting with the uridine binding pocket in the enzyme’s active site. Our findings provide new insights for the development of uracil scaffold-based drugs.
Similar content being viewed by others
Introduction
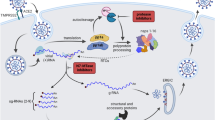
The current pandemic of COVID-19 is caused by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). At the time of writing, there is no vaccine or proven drug against SARS-CoV-2. Since February 2020, concerted efforts have focused on characterizing its biology and developing various detection and treatment options, ranging from vaccines through antibodies to antivirals.
As a typical member of the Coronaviridae family, it is spherical, enveloped, non-segmented, (+) sense RNA virus with a large ~30 kbs genome. These genomes are used as mRNA for translation of a replicase-transcriptase complex (RTC) constituents made of two large polyproteins, Pp1a and Pp1ab, and as a template for replication of its own copy1. These polypeptides are processed by two viral proteases: papain-like protease (PLpro, a domain within non-structural protein 3 (Nsp3)), and 3C-like protease (Nsp5 or 3CLpro or Mpro). For CoV-2 the cleavage yields 15 Nsps, (Nsp11 is just a 7-residues peptide) that assemble into a large membrane-bound RTC that exhibits multiple enzymatic and binding activities. In addition, several sub-genomic RNAs are generated from (-) sense RNA during virus proliferation, resulting in translation of 4 structural and 9-10 accessory proteins. Non-structural proteins are potential drug targets for therapies and, because of their essentiality, sequence, and function conservation, developed therapeutics might in principle inhibit all human coronaviruses.
Nsp15 is a uridine-specific endoribonuclease. Its catalytic C-terminal domain shows sequence similarity and functionality of the EndoU family enzymes. They are involved in RNA processing and broadly distributed in viruses, archaea, bacteria, plants, humans, and other animals. The viral EndoU subfamily has been named NendoU. The Nsp15 enzyme is active as a 234 kDa hexamer consisting of three dimers. It was reported that NendoU cleaves both single- and double-stranded RNA at uridine sites producing 2′,3′-cyclic phosphodiester and 5′-hydroxyl termini2. The 2′,3′-cyclic phosphodiester is then hydrolyzed to 3′-phosphomonoester. In coronaviruses, toroviruses, and arteriviruses, NendoU is mapping to Nsp15s and Nsp11s. Nsp15s are much more sequence conserved than Nsp11s. It was proposed that in coronaviruses Nsp15 affects viral replication by interfering with the host’s innate immune response3,4. To evade host pattern recognition receptor MDA5 responsible for activating the host defenses, the Nsp15 cleaves the 5′-polyuridine tracts in (-) sense viral RNAs, which are the product of polyA-templated RNA synthesis5. These polyU sequences correspond to MDA5-dependent pathogen-associated molecular patterns (PAMPs) and NendoU activity limits their accumulation. For SARS-CoV it was reported that Nsp15 cleaves highly conserved non-translated RNA on (+) sense strand showing that both RNA sequence and structure are important for cleavage6,7.
Recently we have determined the first two crystal structures of SARS-CoV-2 Nsp15 and demonstrated that it is similar to the SARS-CoV and MERS-CoV homologs8. We have also shown that purified recombinant SARS-CoV-2 Nsp15 can efficiently cleave synthetic oligonucleotide substrate containing single rU, 5′−6-FAM-dArUdAdA−6-TAMRA-3′ 8. Studies of NendoU subfamily members, Nsp15 and Nsp11, indicate that the enzymes vary in their requirement for metal ion, with Nsp15 variants displaying dependence on Mn2+. At the same time, the proteins are often compared to metal-independent eukaryotic RNase A, which has a completely different fold, but shares active site similarities and broad function. Specifically, it carries two catalytic histidine residues, His12 and His119, that correspond to indispensable His250 and His235 in SARS-CoV Nsp15. RNase A cleaves its substrates in a two-step reaction consisting of transphosphorylation that generates 2′,3′-cyclic phosphodiester followed by hydrolysis leading to 3′-phosphoryl terminus. The latter step has only been reported for SARS-CoV Nsp159. Here, we have expanded Nsp15 research to explore endoribonuclease sequence specificity, metal ion dependence, and catalytic mechanism. In addition to biochemical data, we report five new structures of the enzyme in complex with 5′UMP, 3′UMP, 5′GpU, uridine 2′3′-vanadate (UV)—a transition state analog, and Tipiracil—a uracil derivative. This compound is approved by FDA as a combination drug that is used with trifluridine in the treatment of colorectal cancer10. Tipiracil inhibits the enzyme thymidine phosphorylase which metabolizes trifluridine. We discovered that Tipiracil inhibits Nsp15 endoribonuclease activity, albeit not adequately to be an effective cure, but this scaffold can serve as a template for compounds that may block virus proliferation.
Results and discussion
Biochemical assays
Nsp15 nuclease activity
We tested Nsp15 endoribonuclease activity using several assays and different substrates. We show that SARS-CoV-2 Nsp15 requires Mn2+ ions and retains only little activity in the presence of Mg2+ ions. The enzyme cleaves efficiently eicosamer 5′GAACU↓CAU↓GGACCU↓U↓GGCAG3’ at all four uridine sites (Fig. 1, Supplementary Figs. 1A and B), as well as synthetic EndoU substrate (5′−6-FAM-dArU↓dAdA−6-TAMRA-3′)8 in the presence of Mn2+, and the reaction rate increases upon rising metal ion concentration. The cleavage of the eicosamer seems to show no sequence preference as 5′CU↓C, 5′AU↓G, 5′CU↓U and 5′UU↓G are recognized and cut, especially at higher manganese concentration (Fig. 1). At 5 mM manganese, all four oligonucleotides accumulate, with the slowest cut sequence being the 5′AU↓G site. The eicosamer that does not have uridine in the sequence is not cleaved (Fig. 1, Supplementary Fig. 1C). The observed CoV-2 Nsp15 Mn2+ dependence is unlike its SARS-CoV homolog9. This is surprising as SARS-CoV-2 Nsp15 shares 88% sequence identity and 95% similarity with the SARS-CoV homolog and all active site residues are conserved. One explanation would be that the metal ion impacts the structure of RNA and the cleavage rate will depend on the sequence (and structure) of the substrate. Interestingly, by itself, the Nsp15 can cut RNAs at any uridine site, but as a component of RTC with Nsp8 and Nsp12, Nsp15 becomes a site-specific endonuclease which cuts RNA to leave short 5–10 uridine tails5. The enzyme also cuts the 3′ end of conserved non-coding region of RNA6,7 and the cleavage is RNA secondary structure-dependent.
Reaction products were separated in an 20% polyacrylamide gel containing 7 M urea. Digestion products of (A, B) GAACU5CAUGG10ACCUU15GGCAG20 and (C) GAACA5CAAGG10ACCAA15GGCAG20, respectively. U5-, U8-, and U14/15- uridine-specific ladder. XC and BB mark the final positions of the xylene cyanole and bromophenol blue dyes. The cleavage of the eicosamer is uridine-specific and shows no sequence preference as 5′CU↓C, 5′AU↓G, 5’CU↓U, and 5′UU↓G are recognized and cut, especially at higher manganese concentration (compare Figs. 1A and 1B). Eicosamer that does not contain uridine is not cut (Fig. 1C).
Our observations provide new insights that might be relevant to Nsp15 roles in the SARS-CoV-2 reproduction cycle. First of all, earlier studies have demonstrated that coronavirus NendoU endonucleases are uridine-specific and display differences in their requirement for Mn2+ ions. On one side of the spectrum are Nsp11s of Equine Arteritis Virus (EAV), Porcine Reproductive and Respiratory Syndrome Virus (PRRSV), and Turkey Coronavirus Nsp15 that do not need Mn2+ for their endoribonuclease activity11. SARS-CoV-2 Nsp15 might represent the other extreme end of the range. Differences in the requirement for Mn2+ ions may affect the outcomes of the endoribonuclease activity. For example, metal-independent Nsp11 from EAV and some Nsp15s generate oligonucleotides with 2′,3′-cyclic phosphodiesters at their 3′ ends9. In contrast, metal-dependent SARS-CoV Nsp15 seems to be able to hydrolyze 2’,3’-cyclic phosphodiesters and produce oligonucleotides with 3’-phosphoryl groups on their 3′-ends9. By analogy, the SARS-CoV-2 homolog should follow a two-step reaction and our structures provide strong backing, but this hypothesis is yet to be tested as our assays do not discriminate between the two.
The requirement for Mn2+ ions by SARS-CoV-2 Nsp15 may by itself enhance pathogen infectivity because Mn2+ is required for the host innate immune responses to viral infections. RNA binding of SARS-CoV Nsp15 was reported to be enhanced in the presence of Mn2+, whereas RNA binding by the cellular XendoU homolog from the African clawed frog was not affected by Mn2+ ions6,7.
Nevertheless, the role of metal ion is puzzling as it was proposed that Nsp15 may have similar activity to eukaryotic RNase A. However, the RNase A activity is metal independent, perhaps suggesting that metal ions may help to position RNA for binding or cleavage. No metal ions or strong metal-binding sites were found in any of the models of EndoU family members available in the Protein Data Bank (PDB). Our efforts to locate Mn2+ ion in the Nsp15 structures also failed, arguing against a direct role of metal ions in catalysis. However, we observe the increase of Nsp15 thermal stability in the presence of such ions (Supplementary Fig. 3), though the molecular basis of this behavior remains enigmatic. Previous data for SARS-CoV Nsp15, Mn2+ ions were found to change the intrinsic tryptophan fluorescence of SARS-CoV Nsp15, indicating conformational changes12,13.
Inhibition by Tipiracil
Since Tipiracil is an uracil derivative and Nsp15 is uridine-specific, we have speculated that the compound may inhibit the enzyme. This hypothesis was tested by using octamer RNA with single uridine site 5′AGGAAGU32pC, which under experimental conditions remains single-stranded and does not form duplexes. Upon endonuclease reaction, 3′-32P-labeled heptamer and unlabeled 3’CMP are produced. The transfer of the 32P from the pCp to the 3′ terminus of the heptamer is consistent with transphosphorylation mechanism. In the presence of 5 mM MnCl2, only partial cleavage is observed. This reaction is decreased by 50% in the presence of 7.5 μM Tipiracil (Fig. 2). At the higher Mn2+ ion concentration, the cleavage reaction proceeds to completion but no inhibition by Tipiracil can be measured.
Lane 10: minus enzyme control. Reaction products were separated in a 20% polyacrylamide gel containing 7 M urea. 32P -labeled phosphates are marked with an asterisk. Raw image of the gel is shown in Supplementary Fig. 2. The x- and y-axes are plotted on a linear scale. Points represent two independent assays (black and open circles). Raw data for Fig. 2 are included in Supplementary Data file. The continuous line represents the best fit for the Morrison equation46.
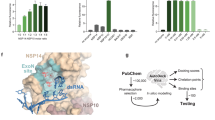
We have also performed S antigen ELISA assay in A549 cells and CoV-2 virus replication assay using qRT-PCR. Tipiracil was not affecting viability of cells but the inhibition of virus was found to be modest in the concentration range 1–50 μM (Fig. 3). This preliminary data suggests that the affinity of the compound may need to be improved in order to serve as an antiviral drug. Structural information is essential for this process (see below).
A549-hACE2 cells were pre-treated with Tipiracil or carrier (0 mM) for 2 h and infected with CoV-2 at MOI 1. After 48 h, cells were harvested to check (A) spike protein and (B) RNA expression, *P < 0.05. Raw data for Fig. 3 are included in Supplementary Data file.
Structure determination and binding of ligands
SARS-CoV-2 Nsp15 protein was crystallized with 5′UMP, 3′UMP, 5′GpU, uridine 2′,3′-vanadate (UV), and Tipiracil using methods described previously8 and the structures were determined at 1.82 Å, 1.85 Å, 1.97 Å, 2.25 Å, and 1.85 Å resolution, respectively. With exception of the complex with UV, crystals with ligands diffract to a higher resolution than the apoprotein (2.20 Å). Of note, we were unable to obtain co-crystals of Nsp15 with 5′AMP, 3′,5′-cyclic AMP, 5′GMP, 5′TMP, and 5′CMP using the same set of conditions as tested for 5′UMP. All structures were solved by molecular replacement using Nsp15 (PDB id: 6WVV) and refined as explained in Materials and Methods and Table 1. The majority of the uncleaved His-tag residues are not ordered and hence not visible but for all residues from Met1 to Gln347 and for all bound ligands the electron density is excellent (Fig. 4). In all five structures, ligands bind to the C-terminal catalytic domain active site (Fig. 4), though the exact positions vary, as described below. In 5′GpU and Tipiracil complexes, there is also a phosphate ion bound in the catalytic pocket. The compound binding is facilitated by side chains of seven conserved residues His235, His250, Lys290, Trp333, Thr341, Tyr343, and Ser294 and the main chain of Gly248, Lys345, and Val292, as well as water molecules (Fig. 4). One non-conserved residue, Gln245, is also involved through a water-mediated interaction. The interactions do not trigger any major protein conformational changes either globally or locally. In fact, the catalytic residues (His235, His250) and other active site residues discussed below maintain very similar conformations in all complexes (RMSD 0. 29 Å) over Cα atoms for residues His235, Gly248, His250, Lys290, Trp333, Thr341, Tyr343, and Ser294 in the pairwise superposition of complexes with the apo-structure (the highest is 0.39 Å for B chain of the Tipiracil complex and the lowest is 0.29 Å for the chain B of the 5′UMP complex). The position of phosphate ion or 3′-phosphoryl group of 3′UMP is also preserved. The vanadate moiety of UV overlaps with the bound phosphate. Specific interactions of ligands with the protein are described below.
5′UMP Binding
The model of uracil binding by Nsp15 was proposed based on the EndoU and RNase A structures2, speculating that Ser294 might be responsible for base discrimination. Here we describe experimental details of pyrimidine recognition. The base of 5′UMP forms van der Waals contacts with Tyr343 and several hydrogen bonds with active site residues, including sidechain OG and main-chain nitrogen atom of Ser294 (Fig. 4A). These interactions define O2 and N3 specificity. O4 interacts with the main chain nitrogen atom of Leu346 defining the uracil specificity. Potentially, cytosine and thymine pyrimidines may also bind—an amino group in position 4 of C should be compatible with the recognition pattern and a methyl group in position 5 should not interfere with binding either as uracil position 5 is solvent accessible. In fact, recognition of C has been reported for distantly related, though with similar active site, bacterial EndoU anticodon tRNase14. The ribose ring makes several hydrogen bonds with protein residues: (i) 2′OH interacts with Lys290 (NZ) and with NE2 of the catalytic His250, (ii) 3′OH makes water-mediated hydrogen bonds with catalytic His235 (NE2), His250 (NE2), Thr341 (OG1) and Gly248 (main chain nitrogen atom), and (iii) O4′ is hydrogen-bonded to main chain of Val292 via a water molecule. Interestingly, the 5′-phosphoryl group projects into solvent with no interaction with protein atoms. Its only ordered interaction is with 3′OH through a water molecule. This 5′-phosphoryl group location overlaps with that of Nsp15/5′GpU complex (see below). Also worth mentioning is that the ribose in the Nsp15/5′UMP together with the phosphate ion in the Nsp15/Tipiracil mimic 2′3′-cyclic phosphodiester. The structure with 5′UMP shows how the enzyme discriminates between uracil and purine bases with Ser294 serving as the key discriminatory residue, as has been hypothesized before6,12,15,16.
5′GpU Binding
In RNA sequence containing uracil, such as 5′NpGpUpN3′, where N corresponds to any base, the Nsp15 cleavage would produce 5′NpGpU3′p, if transphosphorylation is followed by hydrolysis of 5′NpGpU2′3′p. In the crystal structure of Nsp15/5′GpU, the dinucleoside monophosphate binds to the active site with uracil interacting with Tyr343 and Ser294 (Fig. 4B), as seen in the Nsp15/5′UMP complex. However, the distance between O4 and the main chain nitrogen of Leu346 is too long to make a hydrogen bond. This implicates some flexibility at the protein C-terminus and suggests that the amino group in position 4 of cytosine can be accommodated, as we suggested above and reported previously8. The guanine ring is stacking against Trp333 and makes two hydrogen bonds with water molecules. The absence of defined base-side chain interactions suggests a lack of specificity for this site in the substrate sequence. The Nsp15/5′GpU complex binds also a phosphate ion in the active site, most likely from the crystallization buffer. The ion interacts with the protein side chains (His235, His250, Thr341, and Lys290) and uridine ribose 2′OH and 3′OH groups. It most likely mimics the binding of scissile phosphoryl group of the substrate. The backbone phosphoryl group (5′ of U) faces solvent as in the Nsp15/5′UMP complex and makes a hydrogen bond to a water molecule. This structure and the Nsp15/5′UMP complex illustrate location and specificity determinants of the uridine with a 5′-phosphoryl group. The binding of guanine in the structure identifies a strong base binding site at Trp333, which, however, may not necessarily be dedicated to 5′end of the oligonucleotide (see discussion below).
3′UMP Binding
We co-crystallized Nsp15 with 3′UMP, assuming that the enzyme would dock it in a manner expected for uridine monophosphate nucleotide in the contexts of larger RNA substrate, preserving the uracil specific interactions described above. Surprisingly, the uracil base is anchored by Trp333 in the guanine site observed in the Nsp15/5′GpU complex (Fig. 4C), confirming that this site can accommodate purine and pyrimidine bases. The 3′-phosphoryl group occupies the phosphate ion site created by His235, His250, and Thr341, as observed in Nsp15/5′GpU. It is surprising that uracil does not go into its dedicated site, but the result demonstrates higher affinity for the base in the Trp333 site than in uracil-recognition site, potentially governed by the strong stacking interactions with the aromatic side chain that take precedence over hydrogen bonds observed in the uracil binding of 5′UMP. The identity of the Trp333-interacting base is irrelevant, especially given that the enzyme’s substrate is most likely a larger RNA molecule.
Tipiracil binding – a uracil derivative and Nsp15 inhibitor
The Tipiracil molecule binds to the uracil site as observed in 5′UMP and 5′GpU (Figs. 4D and 5). The molecule makes several substrate analog-like interactions. The uracil ring stacks against Tyr341 and makes hydrogen bonds with Ser294 (interacting with O2 and N3), Lys345 (O4), and His250. The carbonyl group of Leu346 makes a water-mediated hydrogen bond in the structure with Tipiracil (Fig. 4D). The N1 atom makes a hydrogen bond with a phosphate ion and through it connects to Lys290. There are also two water-mediated interactions to Ser294 and the main-chain carbonyl oxygen atom of Val292. Iminopyrrolidin nitrogen atom binds to Gln245, representing the only interaction unique to the ligand. Nsp15 binds Tipiracil in its active site in a manner compatible with competitive inhibition and the compound and its derivatives may serve as inhibitors of the enzyme. This structure suggests that uracil alone may have similar inhibitory properties and provides basis for the uracil scaffold-based drug development.
Binding of uridine 2′,3′-vanadate – SN2 transition state analog
The structure of the complex with UV was determined at 2.25 Å resolution. UV binds to the uridine binding site as observed in the structures with 5′UMP, GpU, and Tipiracil. The uracil moiety is recognized by Ser294 and Leu346 defining the uracil recognition and interacts also with Tyr343 (Fig. 4E). The pentavalent vanadate is covalently bound to 2′ and 3′ ribose oxygen atoms and it corresponds to the SN2 transition state17,18,19,20,21. The metal ion, O3′, and two vanadate oxygen atoms are in a plane. One oxygen (O2′) atom below the plane is interacting with His250 and the other oxygen above the plane is in contact with His235 (Figs. 4E and 6B). The vanadate position is consistent with 3′-phosphoryl group that is cleaved in the RNA substrate and observed in the GpU and Tipiracil complexes. The vanadate interacts with key catalytic site residues including His250, His235, Lys290, Thr341, Gly247 and has several water-mediated interactions (Fig. 4E). Particularly, Lys290 makes hydrogen bonds to O2′ and one of the vanadate oxygen atoms in the plane reminiscent of Lys41 in RNase A. These interactions confirm previous experimental observations for Nsp15 and are consistent with the proposed SN2 mechanism. The structure also provides insights into the structure of 2′,3′-cyclic phosphodiester, which is the product of transphosphorylation reaction. This structure suggests that Nsp15 should follow a two-step reaction mechanism with the final product being 3′UMP.
A, B Schematic drawings of reaction mechanism of Nsp15. A Transphosphorylation reaction that generates 2′,3′-cyclic phosphodiester. B Hydrolysis reaction from transition state. Residues shown in red are for RNase A and in black for Nsp15. Arrows depict electron transfer. The active site of RNase A (C) is compared with that of Nsp15 in the complex with transition state analog uridine 2′,3′-vanadate (D), and with Tipiracil (E). The transition state of the phosphate is present in both reactions.
Compounds influence on Nsp15 thermal stability
The differential scanning fluorimetry experiments (DSF) showed that melting temperature (Tm) of the Nsp15 in a presence of Tipiracil, 5′GpU, 5′UMP, 3′UMP, and 3′TMP is approximately 60.5 °C under-investigated conditions with buffer that contains 10 mM MnCl2 (Supplementary Fig 1. A, B). Depicted local minima of the Nsp15 first derivative of fluorescence signals have the same Tm values as a control. The denaturation profile of the Nsp15 in the presence of Tipiracil is broader and consistently shifted (0.5 °C) to higher temperatures (Supplementary Fig 1. A). Therefore, DSF results indicate the small increase of stability of the Nsp15/ Tipiracil complex in comparison to the control sample and other tested Nsp15 complexes with 5′GpU, 5′UMP, 3′UMP, and TMP. Additional change in the Nsp15 Tm is observed at 83 °C (Supplementary Fig. 1B). This is caused by all ligands and may be related to increased stability of the hexamer or EndoU catalytic domain. In the presence of Mn2+ ions the main Tm of Nsp15 is increased from 56.5 °C to 60.5 °C and is Mn2+ concentration-dependent. Interestingly, at 5 mM and 20 mM concentrations of Mn2+ a new local Tm minimum is observed at 83 °C potentially suggesting increased stability of the EndoU domain (Supplementary Fig. 1C).
Comparison of CoV-2 Nsp15 active site with RNase A – mechanistic implications
Our structures of complexes with nucleotides can inform catalytic mechanism of Nsp15 endoribonuclease. We compared our structures with eukaryotic RNase A, a very well-studied model system22 (Fig. 7). RNase A recognizes pyrimidine nucleotides in RNA, preferring C over U, and catalyzes a two-step reaction, the transphosphorylation of RNA to form a 2′,3′-cyclic CMP intermediate followed by its hydrolysis to 3′CMP. Nsp15 also recognizes pyrimidines, preferring U over C, and was proposed to catalyze an analogous reaction9. In RNase A the base selectivity is achieved by Thr45 that forms specific hydrogen bonds similar to those created by Ser294 in the Nsp15/5′UMP, Nsp15/Tipiracil, and Nsp15/5′GpU complexes (Fig. 7). In RNase A the transphosphorylation reaction proceeds via an asynchronous concerted general acid/base mechanism involving His12, His119, and Lys4123. In this mechanism the 2′OH proton is transferred to the deprotonated form of His12 to activate the 2′O nucleophile. Then, the protonated His119 donates a proton to the departing 5′OH group. Lys41 function is to stabilize the negative charge that accumulates on the nonbridging phosphoryl oxygen atoms in the transition state. In the hydrolysis step the role of two histidine residues are inverted. The Nsp15 mechanism of catalysis is consistent with that of RNase A as observed in the structure of the complex with UV transition state (Figs. 4E and 7B). The 2′OH proton is transferred to the deprotonated form of His250 to activate the 2′O nucleophile, the protonated His235 donates a proton to the departing 5′OH group and Lys290 stabilizes the negative charge of the vanadate ion.
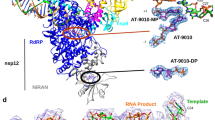
A Nsp15/5’GpU complex compared with RNase A/5’dApdTpdApdA complex (PDB id 1RCN). The least square superposition is calculated with the Nsp15/RNase A residues H250/H12, S294/T35, K290/K41, and U/T bases using COOT (RMSD 0.6 Å on Cα atoms). Nsp15 is shown in dark blue, RNase A in aqua, 5’GpU in yellow and 5’dApdTpdApdA oligonucleotide in pink. Base positions B0-B3 and phosphoryl groups positions P0 and P1 are indicated in red (bold). B Model of 5'GpUpU oligonucleotide binding to the Nsp15 active site. Red arrows show proton flows (extraction and donation) in A & B.
The RNase A active site is well organized and has several distinct pockets for binding RNA substrates (e.g., bases B1, B2, B3, and phosphoryl groups P0, P1, and P2) (for review see24,25). The B1 site provides base specificity and P1 site binds the scissile phosphoester bond. B0 represent the binding site for the base upstream of the B1-P1 cleavage site. In the RNase A complex with the 5′dApdTp↓dApdA deoxyoligonucleotide (PDB id: 1RCN,26), B0 interacts with adenine, B1 binds thymine, etc. The P1 site represents the above-mentioned catalytic machinery consisting of His12, His119, and Lys41, and Lys41 assisted by Gln11 in phosphoryl binding.
Structural alignment of the Nsp15 and RNase A catalytic site residues and RNA ligands shows that they adopt similar positions in the two enzymes, despite sequence and structure dissimilarity. Specifically, His250, Ser294, and Lys290 virtually overlap with the His12, Thr45, and Lys41. His250, by analogy His12 in RNase A, is in position to serve as the key residue in deprotonation of 2′OH. It is close to 2′OH in both 5′GpU and 5′UMP structures (3.2 and 3.8 Å respectively) and is in similar orientation in the complex of RNase A with the DNA substrate analog (Fig. 6). Therefore, His250 is very likely to directly activate 2′OH. Lys290 seems to play a role of Lys41 in RNase A. Main chain amide of Gly248 may provide function of Gln11 in binding to the substrate phosphoryl group. If His250 is a base in Nsp15 then the His235 must be a proton donor for the departing 5′OH group and equivalent of His119 of RNase A. However, these residues are ~8 Å apart and they approach the phosphoryl group from different directions (Fig. 6A). In RNase A, His119 forms a hydrogen bond with Asp121 which may provide proton for the reaction. The structural environment for His235 is different in Nsp15. Thr341 forms a hydrogen bond with His235 and there is also Asp240 further away that makes a water-mediated hydrogen bond with His235. In the hydrolysis step of converting the 2′3′-cyclic phosphate back to 2′OH and 3′-phosphoryl group the roles of histidine residues are reversed and now His235 must be a base deprotonating a water molecule and His250 serves as a proton donor for the 2′OH leaving group. A different set of interactions involving Nsp15 and RNase A catalytic residues may explain why Nsp15 activity is more sensitive to low pH and it is expected that kinetics of the reactions will be different.
Besides similarities in P1, the two enzymes share the organization of B1 pocket. Here, Nsp15 has a very well-defined uracil recognition site made of Ser294, Tyr343, and Leu346 that are equivalent to Thr45, Phe120, and Ser123 in RNase A. Further extrapolation from the RNase A model of the deoxyoligonucleotide binding allows us to hypothesize that B2 site, dedicated to the base on the 3′ end of scissile bond, in Nsp15 is created by Trp333. Its sidechain provides stacking option for a base with no base selectivity function. Yet in our Nsp15/5′GpU structure this site is occupied by the G base located on the 5′end of U. We speculate, that for oligonucleotides that are flanked on both sides of U, mimicking the RNase A ligand, 5′ guanine (or a different base) may adopt position of 5′dA that locates to the B0 site owing to the rotation of the P-5′O bond. Then, the available Trp333 base binding site can accept base on the 3′ end of the uridine moiety, somewhat resembling our Nsp15/3′UMP structure. When we combine oligonucleotides from our structures with RNase A/DNA complex a plausible model can be built of 5’GpUp↓U nucleotide bound to the Nsp15 active site (Fig. 6B). This model underscores importance of conserved Trp333 in anchoring RNA in the active site. While Ser294 is the key residue in discriminating base, the hydrophobic interaction with Trp333 may be a significant force for ligand docking. The B3 site is not easily identifiable in the available Nsp15 structures. In addition, unlike in RNase A, where all P0-P2 subpockets contribute to the backbone binding, in Nsp15 only P1 site is currently well-defined. 5′-phosphoryl groups in position P0 of the ligands do not form any direct contacts with the protein, while RNase A P0 site has Lys66 participating in RNA binding. P2 site of RNase A is created by Lys7 and it appears that His243 may fulfill such a role in Nsp15.
Conclusions
The active site of Nsp15 can accommodate all five ligands, three nucleotides (5′UMP, 3′UMP, 5′GpU), one synthetic analog (Tipiracil), and uridine 2′,3′-vanadate—transition state analog. Interactions with these small molecules seem to stabilize the active site residues and entire catalytic EndoU domain resulting in a better-ordered protein. Previous mutational studies demonstrated that two histidine (His235 and His250) and one serine (Ser294) residues are essential for SARS-CoV Nsp15 activity. We, for the first time, illustrated how Ser294 participates in uracil (or pyrimidine) recognition and we concur with previous suggestions that His250 may be key to deprotonate the 2′OH to allow nucleophilic attack on the phosphoryl group. Comparisons of Nsp15 with RNase A active site display some similarities (active site residues conservation) and indicate a common catalytic mechanism with a two-step reaction releasing 3′UMP, but organization of the RNA binding site is distinct, especially at sites more distant from the elements crucial for chemistry.
Our structures are consistent with the binding of single-stranded nucleic acids, such as loops or bulges, as was shown for Nsp15s and other EndoUs. Both uracil and preceding base must be in an unpaired region in order to bind to Nsp15, which is consistent with degradation of polyU tracks as reported recently5. Our structures show that the preceding base can be guanine or uracil, or other bases as well (see below). Accommodation of 5′GpU and 5′GpUpU is in agreement with previous reports of guanine preference in anterior position with respect to U and efficient degradation of polyU tracks. However, SARS-CoV-2 Nsp15 can cleave RNA, at high Mn2+ concentrations, at uridine sites connected to any base in anterior position; for example, the synthetic EndoU substrate used for the nuclease assay has A.
Nsp15 binds nucleotides to the catalytic domain of each monomer independently, therefore it is not clear why the hexamer is required for the EndoU activity of the enzyme. We show that Nsp15 can bind and hydrolyze 4, 7, and 20 nucleotide long RNA. It is possible that the hexamer is needed to bind longer RNA substrates or is involved in interactions with other proteins (for example Nsp8 and Nsp12) within RTC27.
The role for metal ion requirement remains a puzzle. Although the Mn2+ dependence has been reported for some EndoU members and appears to be a common feature of NendoU subfamily, the metal-binding site was never located. Past studies indicated that single strand polyU RNA is relatively unstructured under most conditions. It was showed that Nsp15 cleavage is RNA sequence and structure dependent6. It is possible that the metal is required for maintaining conformation of the RNA substrate during catalysis. For example, metal ions like Mg2+, Mn2+ and Zn2+ form complexes with purine nucleotides to affect outcome of many enzymatic reactions28. Binding of Mn2+ ions to RNA molecules may dramatically transform their structure, as it was shown for riboswitch29 and the Bacillus subtilis M-box aptamer that sequence contains U track (PDB id: 3PDR,30). As manganese can exist in several oxidation states, it is also possible that metal redox properties can affect protein and RNA interactions and chemistry.
Tipiracil, an uracil derivative, binds to Nsp15 uracil site in a manner consistent with competitive inhibition. In vitro it inhibits Nsp15 RNA nuclease activity and shows modest inhibition of CoV-2 virus replication in the whole cell assay. While the compound itself is not optimal for the therapeutic applications, our work shows that uracil and its derivatives may represent a plausible starting point for nucleotide-like drug development. Moreover, interaction of Trp333 with bases may provide additional site to build much higher affinity inhibitors.
Materials and methods
3′-CMP was obtained from Sigma. Crude [gamma-32P]ATP was purchased from PerkinElmer. T4 Polynucleotide Kinase (3’ phosphatase minus) was from New England BioLabs. All other chemicals were reagent grade.
Cytidine and uridine 3,5′-bisphophates synthesis
[5′-32P]pCp was prepared by phosphorylation of 3′-CMP as described by England et al.31.
RNA synthesis
RNA eicosamers were synthetized by runoff transcription of synthetic double-stranded DNA templates published by Sherlin et al.32. 5′-32P-labeled RNA eicosamers and 3′-32P-labeled RNA octamers were prepared according to Zwieb et al.33 and England et al.31, respectively.
Protein purification
Protein was expressed and purified using protocol developed by Kim et al.8. Briefly, a 4 l culture of LB Lennox medium was grown at 37 °C (190 rpm) in presence of ampicillin 150 mg/ml. Once the culture reached OD600 ~1.0, the temperature setting was changed to 4 °C. When bacterial suspension cooled down to 18 °C it was supplemented with the following components to indicated concentration: 0.2 mM IPTG, 0.1% glucose, 40 mM K2HPO4. The temperature was set to 18 °C for 20 h incubation. Bacterial cells were harvested by centrifugation at 7,000 g and cell pellets were resuspended in a 12.5 ml lysis buffer (500 mM NaCl, 5% (v/v) glycerol, 50 mM HEPES pH 8.0, 20 mM imidazole, and 1 mM TCEP) per liter culture and sonicated at 120 W for 5 min (4 s ON, 20 s OFF). The cellular debris was removed by centrifugation at 30,000 g for 1 h at 4 °C. Supernatant was mixed with 4 ml of Ni2+ Sepharose (GE Healthcare Life Sciences) equilibrated with lysis buffer supplemented to 50 mM imidazole pH 8.0 and suspension was applied on Flex-Column (420400-2510) connected to Vac-Man vacuum manifold (Promega). Unbound proteins were washed out via controlled suction with 160 ml of lysis buffer (50 mM imidazole). Bound proteins were eluted with 20 ml of lysis buffer supplemented imidazole to 500 mM pH 8.0. Then 1 mM TCEP was added followed by Tobacco Etch Virus (TEV) protease treatment at 1:20 protease:protein ratio. The solution was left at 4 °C overnight. For this particular construct, TEV protease was not able to cleave off the His tag. Nsp15 was successfully separated from TEV protease on Superdex 200 column equilibrated in lysis buffer where 10 mM β-mercaptoethanol was replaced by 1 mM TCEP. Fractions containing Nsp15 were collected. Lysis buffer was replaced on 30 kDa MWCO filter (Amicon-Millipore) via 10× concentration/dilution repeated 3 times to crystallization buffer (150 mM NaCl, 20 mM HEPES pH 7.5, 1 mM TCEP). Final concentration of Nsp15 was 19 mg/ml.
Crystallization of complexes
Crystallization experiments were conducted using previously published protocols8. Except for Nsp15/UV, all complexes Nsp15/5′UMP, Nsp15/3′UMP, Nsp15/5′GpU, and Nsp15/Tipiracil were prepared by mixing 5–15 mM of each ligand with 0.2 mM Nsp15 and incubate for at least 30 min before crystallization. Crystals of Nsp15/3′UMP, Nsp15/5′GpU, and Nsp15/Tipiracil, the hexagonal crystal form, in P63 space group, were obtained from MCSG1 screen A3 condition containing 0.2 M sodium chloride, 0.1 M sodium/potassium phosphate pH 6.2, 10 %(w/v) PEG8000. These crystals diffracted x-rays to 1.85, 1.93, and 1.85 Å for Nsp15/3′UMP, Nsp15/5′GpU, and Nsp15/Tipiracil, respectively. The crystals of Nsp15/5’UMP and Nsp15/uridine vanadate complexes grew from the Crystal Screen Classic HTP (Jena Bioscience) C2 condition containing 16 %(w/v) PEG4000, 0.1 M Tris pH 8.5, 0.2 M sodium acetate and diffracted to 1.82 Å and 2.25 Å respectively. To achieve higher occupancies of the ligands in the structures, except for uridine vanadate, each ligand compound was added to the cryo-solution and the co-crystal was soaked for 2–3 min before frozen in the liquid nitrogen.
Endoribonuclease assays
Typical reaction contained 5 × 105 CPM of 5′-32P-labeled RNA eicosamers or 3′-32P-labeled RNA octamers (0.5 µM final RNA concentration) and 10 nM SARS-CoV-2 Nsp15 in 20 mM HEPES-KOH (pH 7.5), 50 mM KCl, 1 mM DTT and either 5 mM or 11 mM MnCl2. To inhibit endonuclease, Tipiracil dissolved in water was added. Following incubation at 30 °C for up to 60 min, the reactions were stopped by adding an equal volume of a gel-loading buffer containing 95% formamide, 10 mM EDTA, and 0.025% SDS. The products were analyzed in 20% polyacrylamide gels (acrylamide/bisacrylamide ratio, 19:1) buffered with 0.5x Tris-borate-EDTA containing 7 M urea.
Synthesis of uridine 2′,3′-vanadate
The protocol was modified from the references34,35 to yield higher concentration of uridine vanadate. 1 mg of ammonium meta vanadate is dissolved in 200 μl of 60 °C water and 2.93 mg of uridine is added then incubated for 10 min at the same temperature to obtain ~ 40 mM uridine vanadate solution. Without further purification, this solution and protein were mixed in a 10:1 molar ratio to set up crystals.
Cells and virus
A549 cells expressing human ACE2 (kind gift of Dr. Benjamin R. tenOever, Mt. Sinai School of Medicine) were infected under biosafety level 3 conditions with SARS-CoV-2 (nCoV/Washington/1/2020, kindly provided by the National Biocontainment Laboratory, Galveston, TX).
Cell viability
Cell viability was measured by staining with CellTracker™ Red CMTPX (ThermoFisher Scientific). SARS-CoV-2 infected cell was stained with 2 µM of CellTracker™ Red CMTPX for 30 min and then detected by Tecan Infinite m200 (Tecan) at ex577/em602 nm. After reading, cells were fixed by 10% neutral buffered formalin (NBF) for immunohistochemistry assay.
Immunostaining against Spike protein
Immunostaining was performed on 10% NBF fixed SARS-CoV-2 infected cells in 96-well plate. After fixation, 10% NBF was removed, and cells were washed with PBS, followed by washing with PBS-T (0.1% Tween 20 in PBS), and then blocked for 30 min with PBS containing 1% BSA at room temperature. After blocking, endogenous peroxidases were quenched by 3% hydrogen peroxide for 5 min. Then, cells were washed with PBS and PBS-T and incubated with a monoclonal mouse-anti- SARS-CoV-2 spike antibody (GeneTex, 1:1000) in PBS containing 1% BSA overnight at 4 °C. Primary antibody was washed with PBS and PBS-T and then cells were incubated in secondary antibody (ImmPRESS Horse Anti-Mouse IgG Polymer Reagent, Peroxidase; Vector Laboratories) for 60 min at room temperature. After washing with PBS for 10 min, color development was achieved by applying diaminobenzidine tetrahydrochloride (DAB) solution (Metal Enhanced DAB Substrate Kit; ThermoFisher Scientific) for 30 min and detected by Tecan Infinite m200 (Tecan) at 492 nm after replacing DAB solution to PBS. The result was normalized by cell viability.
RNA Extraction and qRT-PCR
Total RNA from SARS-CoV-2 infected cells was isolated using a NucleoSpin 96 RNA kit following the manufacturer’s instructions (Macherey-Nagel). SARS-CoV-2 RNA was quantified by qRT-PCR using SuperScript™ III Platinum™ One-Step qRT-PCR Kit w/ROX (ThermoFisher Scientific) and normalized using Eukaryotic 18 S rRNA Endogenous Control (VIC™/MGB probe, Applied Biosystems) via a StepOnePlus real-time PCR system (Applied Biosystems). All reactions performed in a dual-plex qRT-PCR using the CDC-recommended primers for N1. Primer and probe sequences are as follows: 2019-nCoV_N1 Forward Primer (2019-nCoV_N1-F), GACCCCAAAATCAGCGAAAT; 2019-nCoV_N1 Reverse Primer (2019-nCoV_N1-R), TCTGGTTACTGCCAGTTGAATCTG; Probe (2019-nCoV_N1-P), FAM-ACCCCGCATTACGTTTGGTGGACC-BHQ1.
Protein differential scanning fluorimetry
The Nsp15 differential scanning fluorimetry assays were done in a buffer 20 mM Tris pH 7.5, 100 mM NaCl, 1 mM TCEP that was supplemented with SYPRO Orange (Invitrogen) dye36 to 5x final concentration. We used 10 µM of the Nsp15 with addition of 1 mM of Tipiracil, 5′GpU, 5′UMP, 3′UMP, TMP, respectively to get the final molar ratio of protein to ligand 1:100. Additionally, the samples with ligands were supplemented with 10 mM MnCl2. After 30 min of incubation of samples at room-temperature fluorescence measurements were done using CFX Connect Real-Time System (BioRad). Fluorescent signal was detected with temperature rate 1 °C per 60 s. Graphs represents the first derivative of observed fluorescence signal. Samples were measured in triplicates and the representative standard deviation of measurements are depicted for Nsp15 control sample and complex of the Nsp15 with the addition of Tipiracil.
Data collection, structure determination, and refinement
The x-ray diffraction experiments were carried out at the Structural Biology Center 19-ID beamline at the Advanced Photon Source, Argonne National Laboratory. The diffraction images were recorded at 100 K at the wavelength of 0.9793 Å from all crystal forms on the PILATUS3 X 6 M detector using 0.5° rotation and 0.5 s exposure for 140°, 180°, 110°, 210°, and 125° for Nsp15/5′UMP, Nsp15/3′UMP, Nsp15/5′GpU, Nsp15/Tipiracil, and Nsp15/UV, respectively. The data were integrated and scaled with the HKL3000 suite37. Intensities were converted to structure factor amplitudes in the Ctruncate program38,39 from the CCP4 package40 and using the apo-form SARS-CoV-2 Nsp15 structure (PDB id: 6VWW) as a search model, the structures were determined using MOLREP41, all implemented in the HKL3000 software package. The initial solutions were refined, both rigid-body refinement and regular restrained refinement by REFMAC40,42 as a part of HKL3000. The models including the ligands were manually adjusted using COOT43 and then iteratively refined using COOT and PHENIX44. Throughout the refinement, the same 5% of reflections were kept out throughout from the refinement (in both REFMAC and PHENIX refinement). The stereochemistry of the structure was checked with PROCHECK45 and the Ramachandran plot and validated with the PDB validation server. For the structures Nsp15/5′UMP, Nsp15/3′UMP, Nsp15/5′GpU, Nsp15/Tipiracil, Nsp15/UV, Ramachandran favored/allowed/outliers are 98.12/1.88/0.0, 97.38/2.47/0.15, 97.39/2.61/0.0, 97.97/2.03/0.0, and 96.96/3.04/0.0%, respectively. The data collection and processing statistics are given in Table 1.
Statistics and reproducibility
The results included in this manuscript can be reproduced by following protocols and using materials described in Materials and Methods.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The structural datasets generated during the current study are available in the Protein Data Bank repository (https://www.rcsb.org/) under accession codes: 6WLC, 6X4I, 6X1B, 6WXC, and 7K1L. Diffraction images are available on the server in Dr. W. Minor laboratory https://proteindiffraction.org.
Plasmid for expression Nsp15 (NR-52426 Vector pMCSG53 Containing the SARS-Related Coronavirus 2, Wuhan-Hu-1 Non-Structural Protein 15) is available in the NIH the BEI Resources Repository (https://www.niaid.nih.gov/research/bei-resources-repository). All other data generated during the current study including the raw kinetic and biophysical data are available upon request.
References
Kim, D. et al. The architecture of SARS-CoV-2 transcriptome. Cell 181, 914–921 (2020).
Ulferts, R. & Ziebuhr, J. Nidovirus ribonucleases: structures and functions in viral replication. RNA Biol. 8, 295–304 (2011).
Deng, X. et al. Coronavirus nonstructural protein 15 mediates evasion of dsRNA sensors and limits apoptosis in macrophages. Proc. Natl Acad. Sci. USA 114, E4251–E4260 (2017).
Liu, X. et al. Porcine deltacoronavirus nsp15 antagonizes interferon-beta production independently of its endoribonuclease activity. Mol. Immunol. 114, 100–107 (2019).
Hackbart, M., Deng, X. & Baker, S. C. Coronavirus endoribonuclease targets viral polyuridine sequences to evade activating host sensors. Proc. Natl Acad. Sci. USA 117, 8094–8103 (2020).
Bhardwaj, K., Sun, J., Holzenburg, A., Guarino, L. A. & Kao, C. C. RNA recognition and cleavage by the SARS coronavirus endoribonuclease. J. Mol. Biol. 361, 243–256 (2006).
Gioia, U. et al. Functional characterization of XendoU, the endoribonuclease involved in small nucleolar RNA biosynthesis. J. Biol. Chem. 280, 18996–19002 (2005).
Kim, Y. et al. Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2. Protein science: a publication of the Protein Society, https://doi.org/10.1002/pro.3873 (2020).
Nedialkova, D. D. et al. Biochemical characterization of arterivirus nonstructural protein 11 reveals the nidovirus-wide conservation of a replicative endoribonuclease. J. Virol. 83, 5671–5682 (2009).
Kish, T. & Uppal, P. Trifluridine/tipiracil (Lonsurf) for the treatment of metastatic colorectal cancer. Drugs 41, 314–325 (2016).
Cao, J., Wu, C. C. & Lin, T. L. Turkey coronavirus non-structure protein NSP15–an endoribonuclease. Intervirology 51, 342–351 (2008).
Bhardwaj, K., Guarino, L. & Kao, C. C. The severe acute respiratory syndrome coronavirus Nsp15 protein is an endoribonuclease that prefers manganese as a cofactor. J. Virol. 78, 12218–12224 (2004).
Guarino, L. A. et al. Mutational analysis of the SARS virus Nsp15 endoribonuclease: identification of residues affecting hexamer formation. J. Mol. Biol. 353, 1106–1117 (2005).
Michalska, K. et al. Functional plasticity of antibacterial EndoU toxins. Mol. Microbiol. 109, 509–527 (2018).
Bhardwaj, K. et al. Structural and functional analyses of the severe acute respiratory syndrome coronavirus endoribonuclease Nsp15. J. Biol. Chem. 283, 3655–3664 (2008).
Ivanov, K. A. et al. Major genetic marker of nidoviruses encodes a replicative endoribonuclease. Proc. Natl Acad. Sci. USA 101, 12694–12699 (2004).
Davies, D. R. & Hol, W. G. The power of vanadate in crystallographic investigations of phosphoryl transfer enzymes. FEBS Lett. 577, 315–321 (2004).
Deng, H., Burgner, J. W. & Callender, R. H. Structure of the ribonuclease·uridine−vanadate transition state analogue complex by Raman difference spectroscopy: mechanistic implications. J. Am. Chem. Soc. 120, 4717–4722 (1998).
Messmore, J. M. & Raines, R. T. Pentavalent organo-vanadates as transition state analogues for phosphoryl transfer reactions. J. Am. Chem. Soc. 122, 9911–9916 (2000).
Veenstra, T. D. & Lee, L. NMR study of the positions of His-12 and His-119 in the ribonuclease A-uridine vanadate complex. Biophys. J. 67, 331–335 (1994).
Wlodawer, A., Miller, M. & Sjolin, L. Active site of RNase: neutron diffraction study of a complex with uridine vanadate, a transition-state analog. Proc. Natl Acad. Sci. USA 80, 3628–3631 (1983).
Cuchillo, C. M., Nogues, M. V. & Raines, R. T. Bovine pancreatic ribonuclease: fifty years of the first enzymatic reaction mechanism. Biochemistry 50, 7835–7841 (2011).
Harris, M. E., Piccirilli, J. A. & York, D. M. Integration of kinetic isotope effect analyses to elucidate ribonuclease mechanism. Biochim Biophys Acta 1854, 1801–1808 (2015).
Nogues, M. V., Vilanova, M. & Cuchillo, C. M. Bovine pancreatic ribonuclease A as a model of an enzyme with multiple substrate binding sites. Biochim. Biophys. Acta 1253, 16–24 (1995).
Raines, R. T. Ribonuclease A. Chem. Rev. 98, 1045–1066 (1998).
Fontecilla-Camps, J. C., de Llorens, R., le Du, M. H. & Cuchillo, C. M. Crystal structure of ribonuclease A.d(ApTpApApG) complex. Direct evidence for extended substrate recognition. J. Biol. Chem. 269, 21526–21531 (1994).
Athmer, J. et al. In situ tagged nsp15 reveals interactions with coronavirus replication/transcription complex-associated proteins. mBio 8, https://doi.org/10.1128/mBio.02320-16 (2017).
Bianchi, E. M., Sajadi, S. A., Song, B. & Sigel, H. Stabilities and isomeric equilibria in aqueous solution of monomeric metal ion complexes of adenosine 5’-diphosphate (ADP3-) in comparison with those of adenosine 5′-monophosphate (AMP2-). Chemistry 9, 881–892 (2003).
Shi, Y., Zhao, G. & Kong, W. Genetic analysis of riboswitch-mediated transcriptional regulation responding to Mn2+ in Salmonella. J. Biol. Chem. 289, 11353–11366 (2014).
Ramesh, A., Wakeman, C. A. & Winkler, W. C. Insights into metalloregulation by M-box riboswitch RNAs via structural analysis of manganese-bound complexes. J. Mol. Biol. 407, 556–570 (2011).
England, T. E., Bruce, A. G. & Uhlenbeck, O. C. Specific labeling of 3′ termini of RNA with T4 RNA ligase. Methods Enzymol. 65, 65–74 (1980).
Sherlin, L. D. et al. Chemical and enzymatic synthesis of tRNAs for high-throughput crystallization. RNA 7, 1671–1678 (2001).
Zwieb, C., Guven, S. A., Wower, I. K. & Wower, J. Three-dimensional folding of the tRNA-like domain of Escherichia coli tmRNA. Biochemistry 40, 9587–9595 (2001).
Borah, B. et al. Nuclear magnetic resonance and neutron diffraction studies of the complex of ribonuclease A with uridine vanadate, a transition-state analogue. Biochemistry 24, 2058–2067 (1985).
Ladner, J. E., Wladkowski, B. D., Svensson, L. A., Sjolin, L. & Gilliland, G. L. X-ray structure of a ribonuclease A-uridine vanadate complex at 1.3 A resolution. Acta Crystallogr. D Biol. Crystallogr. 53, 290–301 (1997).
Huynh, K. & Partch, C. L. Analysis of protein stability and ligand interactions by thermal shift assay. Curr. Protoc. Protein Sci. 79, 28.9.1–28.9.14 (2015).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. Sect. D-Biol. Crystallogr. 62, 859–866 (2006).
French, S. & Wilson, K. Treatment of negative intensity observations. Acta Crystallogr. Sect. A 34, 517–525 (1978).
Padilla, J. E. & Yeates, T. O. A statistic for local intensity differences: robustness to anisotropy and pseudo-centering and utility for detecting twinning. Acta Crystallogr. Sect. D-Biol. Crystallogr. 59, 1124–1130 (2003).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. Sect. D-Biol. Crystallogr. 67, 235–242 (2011).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. Sect. D-Biol. Crystallogr. 66, 22–25 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. Sect. D-Biol. Crystallogr. 53, 240–255 (1997). S0907444996012255.
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect D-Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D-Biol. Crystallogr. 66, 213–221 (2010).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallgr. 26, 283–291 (1993).
Copeland, R. A. Enzymes: A Practical Introduction to Structure, Mechanism, and Data Analysis, 2nd Edition. 305-317 (John Wiley & Sons, Inc., 2000).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Davis, I. W., Murray, L. W., Richardson, J. S. & Richardson, D. C. MOLPROBITY: structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res. 32, W615–619 (2004).
Acknowledgements
We truthfully thank the members of the SBC at Argonne National Laboratory, especially Darren Sherrell and Alex Lavens for their help with setting beamline and data collection at beamline 19-ID. Funding for this project was provided in part by federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services, under Contract HHSN272201700060C (A.J.) and by the DOE Office of Science through the National Virtual Biotechnology Laboratory, a consortium of DOE national laboratories focused on response to COVID-19, with funding provided by the Coronavirus CARES Act (A.J.). The use of SBC beamlines at the Advanced Photon Source is supported by the U.S. Department of Energy (DOE) Office of Science and operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357. J.W. was supported by the Hatch program of the National Institute of Food and Agriculture, U.S. Department of Agriculture.
Author information
Authors and Affiliations
Contributions
A.J. initiated the project, R.J. cloned, expressed, and purified protein. J.W. performed all enzyme activity and RNA binding assays. N.M. crystallized protein and complexes with ligands and with Y.K. collected diffraction data. Y.K. together with C.C. determined, refined, and analyzed structures. M.W. measured thermal stability of Nsp15. S.K. and V.N. performed virus assays and G.R. analyzed the data. Finally, A.J., Y.K., J.W., and G.R. conceived of and directed the research as well as wrote the manuscript, while K.M. analyzed structural data and also wrote portions of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, Y., Wower, J., Maltseva, N. et al. Tipiracil binds to uridine site and inhibits Nsp15 endoribonuclease NendoU from SARS-CoV-2. Commun Biol 4, 193 (2021). https://doi.org/10.1038/s42003-021-01735-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-01735-9
This article is cited by
-
COVID-19 drug discovery and treatment options
Nature Reviews Microbiology (2024)
-
Crystal structure, molecular docking with SARS-CoV-2 receptors, and potential drug property of tetrahedral Zn(II) complexes
Structural Chemistry (2024)
-
Therapeutic strategies for COVID-19: progress and lessons learned
Nature Reviews Drug Discovery (2023)
-
COVID-19 therapies: do we see substantial progress?
Cellular & Molecular Biology Letters (2022)
-
Structural biology of SARS-CoV-2: open the door for novel therapies
Signal Transduction and Targeted Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.