Abstract
As a bacteriostatic agent, nitrite has been used in food preservation for centuries. When used in combination with antibiotics, nitrite is reported to work either cooperatively or antagonistically. However, the mechanism underlying these effects remains largely unknown. Here we show that nitrite mediates tolerance to aminoglycosides in both Gram-negative and Gram-positive bacteria, but has little interaction with other types of antibiotics. Nitrite directly and mainly inhibits cytochrome heme-copper oxidases (HCOs), and by doing so, the membrane potential is compromised, blocking uptake of aminoglycosides. In contrast, reduced respiration (oxygen consumption rate) resulting from nitrite inhibition is not critical for aminoglycoside tolerance. While our data indicate that nitrite is a promising antimicrobial agent targeting HCOs, cautions should be taken when used with other antibiotics, aminoglycosides in particular.
Similar content being viewed by others
Introduction
Nitrite, a ubiquitous nitrogen species, is the central player in the nitrogen biogeochemical cycle, linking nitrate to gas nitrogen or ammonium1. During its reduction to nitrogen, nitrite can be converted to nitric oxide (NO), a reactive free radical that interferes with protein cofactors, such as Fe–S clusters, heme, and lipoamide, or promotes the formation of reactive nitrogen species2,3,4. Although it has been widely accepted that antibacterial effects of nitrite, as used in food preservation for centuries, are attributable to NO formation5, many studies in recent years have revealed that nitrite has NO-independent effects6,7,8.
Nitrite (nebulized sodium nitrite) is currently in a phase 2b clinical trial as a drug for pulmonary hypertension9. The study has demonstrated that nitrite could be safely applied to reach millimolar concentrations in the airway surface liquid, paving the road for the agent to be a previously unidentified antimicrobial therapy. Nitrite has also been found to be either cooperative or antagonistic when used in combination with antibiotics in Pseudomonas aeruginosa8,10,11. These effects are a result of the bacteriostatic action, which does not require NO as an intermediate1. Antagonistic effects are commonly associated with aminoglycosides and also observed with ciprofloxacin (Cip), one of the quinolones that interfere with bacterial DNA gyrase10,12. In contrast, cooperative antimicrobial activities have been observed when used with polymyxins, polycationic lipopeptide antibiotics that interact with negatively charged lipopolysaccharide on the cell surface of Gram-negative bacteria8.
Tolerance induced by nitrite to aminoglycosides and Cip is proposed to be attributed to impaired respiration (oxygen consumption rate, hereafter referred to as respiration) given similar effects of other respiratory inhibitors, such as cyanide and carbonyl cyanide m-chlorophenylhydrazone (CCCP)10,11. As nitrite is bacteriostatic, this proposal is in perfect agreement with the recent report that antibiotic efficacy is linked to bacterial cellular respiration13. However, given that aminoglycosides have long been proposed to depend on proton motive force (PMF) for uptake14, it is possible that nitrite-mediated aminoglycoside tolerance may not be a direct result of reduced respiration. Unfortunately, in most, if not all, of these previous studies10,11,12,13, respiration is virtually an equivalent of PMF, because the observations were from the proton gradient inhibitors, with CCCP being the most commonly used one. Thus, although reduced respiration likely compromises PMF formation, the direct evidence is lacking. More importantly, despite these findings, the antimicrobial mechanisms of nitrite, especially those through which respiration and/or PMF are impaired, is only partially understood11.
Given that dependence on PMF for uptake of aminoglycosides is commonly found in both Gram-negative and Gram-positive bacteria, we hypothesize that nitrite must inhibit proteins that are well conserved. Moreover, we also hypothesize that the targets are likely to be out of the cytoplasm, because nitrite as a charged molecule could not be efficiently imported in at least some of these bacteria. By this logic, we focus on the respiratory chain, especially terminal oxidases. This coincides with the findings of systematic investigations into nitrite and NO physiology in Shewanella oneidensis, a facultative Gram-negative γ-proteobacterium renowned for respiratory versatility15,16,17,18,19,20. In this bacterium, the primary target of nitrite is heme-copper oxidase (HCO) cbb3 (hereafter referred to as cbb3), whereas NO indistinguishably interacts with all cytochromes c15,16,19,20.
The bacterial respiratory chain is generally branched after the quinone pool, linking to multiple terminal oxidases (oxygen reductases) that carry out reduction of O2 into H2O and simultaneously form PMF21. Prokaryotic terminal oxidases are characterized into two major groups: HCOs and the bd-type quinol oxidases22. Representative HCOs include cytochrome aa3 oxidases (mitochondrial-like oxidases, hereafter referred to as aa3) as in Bacillus subtilis and Staphylococcus aureus, cytochrome bo3 oxidases (hereafter referred to as bo3) as in Escherichia coli, and cbb3 as in S. oneidensis and P. aeruginosa.
In this study, we tested our hypotheses that targets of nitrite are conserved proteins in the respiration chain whose inhibition underlies increased tolerance to aminoglycosides with E. coli, S. oneidensis, P. aeruginosa. B. subtilis, and S. aureus. We found that although nitrite inhibits killing of aminoglycosides in all of these bacteria, the efficacy varies from one species to another. Effects of nitrite on individual HCOs were then assessed in the same genetic background and results revealed that cbb3 is much sensitive than bo3 and aa3. We further provided evidence to suggest that nitrite blocks the activity of aminoglycosides but not of other antibiotics and this action is dependent on PMF rather than impaired respiration. Overall, these data suggest that nitrite is a promising antimicrobial agent targeting HCOs, but cautions should be taken when used with other antibiotics, aminoglycosides in particular.
Results
Nitrite induces bacterial tolerance to aminoglycosides
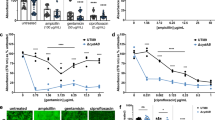
Nitrite has been found to block the activity of aminoglycosides in P. aeruginosa10. To test whether this phenomenon is common among bacteria, in this study we examined impacts of nitrite on aminoglycoside tolerance in S. oneidensis, E. coli, B. subtilis, and S. aureus, along with P. aeruginosa as the control. Susceptibility of all test strains to nitrite was first evaluated with the spot dilution test, to determine the proper concentration range for the study (Supplementary Fig. 1). S. oneidensis was much more sensitive to nitrite than E. coli and S. aureus as reported before19, whereas P. aeruginosa and B. subtilis showed nitrite tolerance in between. Time-kill kinetics of these strains were assessed by using the same approach for E. coli, P. aeruginosa, and S. aureus described previously10,13. Cultures for each strain grown aerobically in lysogeny broth (LB) to the mid-exponential phase (∼0.4 of OD600) were exposed to gentamicin (Gent) or streptomycin (Str) at 5× minimum inhibitory concentration (MIC) in the absence and presence of nitrite. For Gram-negative bacteria under test, the addition of nitrite 1.5–12 mM induced elevated tolerance to Gent (Fig. 1a–c). Although the induction appeared to be dose-dependent with nitrate at low concentrations (<3 mM), saturating effects were observed when higher concentrations were applied. The effects of nitrite on tolerance of Gram-positive bacteria, B. subtilis and S. aureus, to Gent, albeit visible only at much higher concentrations, were also dose-dependent (Fig. 1d–e). Apparently, nitrite was much less effective on these two bacteria, especially S. aureus, when compared with the three Gram-negative counterparts. Similar results were obtained from Str (Supplementary Fig. 2), suggesting that the phenomenon is likely common to aminoglycosides.
a–e Time-kill analysis of Gent (5× MIC) and nitrite combination for S. oneidensis, E. coli, P. aeruginosa, B. subtilis, and S. aureus. For all strains, cultures at the mid-exponential phase were used and their cell numbers before the treatment were as follows: S. oneidensis, ∼2 × 108 CFU/ml; E. coli, ∼2 × 108 CFU/ml; P. aeruginosa, ∼2 × 108 CFU/ml; B. subtilis, ∼5 × 107 CFU/ml, and S. aureus, ∼2 × 108 CFU/ml. Values shown were the number of viable cells 4 h after the treatment began. Impacts of nitrite on these bacteria were assessed and presented in Supplementary Fig. S1. Combination therapy was compared against monotherapy with Gent, between which statistically significant difference caused by nitrite were given (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001). f The spot dilution assay of Gent (5× MIC) and nitrite combination for indicated bacteria. Cultures prepared to contain ~109 CFU/ml were regarded as the undiluted (dilution factor, 0), which were subjected to tenfold series dilution. Five microliters of each dilution was dropped on LB agar plates containing Gent and nitrite, whose concentrations were the same as shown in a–f. Results were recorded after incubation of 18 h. Experiments were performed independently at least three times and representative data were presented.
For confirmation, we performed the spot dilution test. The droplets of exponential-phase cultures containing different numbers of cells were applied onto LB agar plates supplemented with Gent and nitrite. Clearly, nitrite was very effective in inducing tolerance of the three Gram-negative bacteria to Gent, up to 4-log increase in the number of viable cells (Fig. 1f). On the contrary, the ability of nitrite to modulate Gent tolerance of two Gram-positive bacteria was relatively weak. Although nitrite at the most effective concentration improved viability of B. subtilis cells by up to 2-log, it rescued cell survival <1-log in S. aureus. The contrast between Gram-negative and Gram-positive bacteria regarding the effect of nitrite on modulation of aminoglycoside tolerance suggests that physiological differences matter.
We then made attempts to determine whether the observed effects of nitrite are via NO. As this has been comprehensively addressed in the case of P. aeruginosa10, here we only repeated time-kill assays in the presence of PTIO (2-Phenyl-4,4,5,5-tetramethylimidazoline-1-oxyl-3-oxide), an NO scavenger. With all bacteria tested, results obtained in the presence of excessive PTIO were comparable to those in its absence (Supplementary Fig. 3). Moreover, the differences in data from the experiments without and with PTIO were statistically insignificant. As S. oneidensis could not produce NO19, it served as a perfect control for eliminating impacts of excessive PTIO on physiology. These data were in perfect agreement with the previous finding that nitrite induces tolerance to aminoglycoside antibiotics without involving NO.
Nitrite modulates aminoglycosides tolerance by inhibiting cbb 3
It is well known that the uptake of aminoglycoside antibiotics by bacterial cells is an energy-requiring process that depends on PMF14. The S. oneidensis genome encodes three terminal oxidases, HCOs caa3 and cbb3, as well as non-HCO bd, among which cbb3 and bd oxidases are produced and exhibit activity under normal conditions16 (Fig. 2a). For oxygen respiration, cbb3 is the predominant oxidase, whereas bd functions as an accessory oxidase and, more importantly, supports aerobic growth in the absence of cbb316. Given that cbb3 is proposed to be the primary target of nitrite in S. oneidensis15, we hypothesized that nitrite modulates aminoglycoside tolerance by inhibiting terminal oxidases.
a Model illustrating oxygen respiration in S. oneidensis. S. oneidensis possesses two HCOs, cbb3, and caa3, which obtain electrons from the quinol pool via the bc1 complex, and a quinol oxidase, bd. For oxygen respiration, cbb3 dictates and bd is its subordinate, whereas activity of caa3 is not detected under normal conditions. b Time-kill analysis of Gent (5× MIC) and nitrite (given in mM; −, no nitrite) combination for relevant S. oneidensis mutants. Con, no addition of Gent and nitrite. WT, wild type; Δcco, cbb3-less; Δcox, caa3-less; Δcyd, bd-less; Δpet, bc1-less. Asterisks indicate statistically significant difference between values being compared (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001). c The spot dilution assay of Gent (5× MIC) and nitrite (3 mM) combination for indicated bacteria. Experiments were performed independently at least three times and representative data were presented.
To test this, we examined Gent susceptibility of the cbb3-deficient strain (Δcco) constructed, verified, and characterized previously, which loses cytochrome c oxidase activity completely16. Time-kill assays revealed that the cbb3 loss induced tolerance to Gent substantially, more effective than the addition of nitrite at any concentrations (Fig. 2b). A comparable effect was observed from a mutant lacking the cytochrome bc1 complex (Δpet) (Fig. 2b), through which cbb3 obtains electrons from the quinone pool (Fig. 2a), supporting that the induction of Gent tolerance is attributable to the deficiency in cbb3-based respiration. In contrast, the deletion of cox (encoding caa3) genes (Δcox) did not elicit any detectable difference in Gent tolerance without or with nitrite when compared with the wild type, which is in perfect agreement with the fact that the enzyme is not produced to physiological relevant levels16,23. In the case of bd, its absence (Δcyd) did not affect Gent tolerance without nitrite (Fig. 2b), suggesting that bd activity is not critically implicated in Gent tolerance. However, it should be noted that the effect of nitrite on Gent tolerance could not be assessed directly, because the depletion of bd vastly sensitized S. oneidensis cells15,24.
We then made attempts to verify these observations with the spot dilution test. The loss of cbb3, albeit only modestly, reduces aerobic growth rates, whereas neither caa3 nor bd affects growth when cbb3 is present16. Consistent results were obtained from LB agar plates (Fig. 2c), although the growth difference between the wild-type and Δcco (the same for Δpet) strains appeared even smaller. Clearly, the cco and pet mutants exhibited substantially elevated Gent tolerance, whereas the strains lacking either caa3 or bd behaved as the wild type. Importantly, the Δcyd strain was hypersensitive to nitrite, while all other strains under test had comparable nitrite susceptibility. This hypersensitivity prevented Δcyd cells from growing on LB agar plates containing 3 mM nitrite, a condition that blocked Gent activity in the wild-type and Δcox strains. These data, all together, manifest that nitrite modulates aminoglycoside susceptibility by compromising cbb3-based respiration, implying that cbb3, the bc1 complex, or both could be targets of nitrite. This study focused on cbb3, because it has been shown to be susceptible to nitrite15,16,17,18,19,20.
HCOs are susceptible to nitrite
Given that other HCOs, including caa3, aa3, and bo3, are capable of proton-pumping during oxygen respiration as cbb3, we reasoned that they modulate aminoglycoside susceptibility through a similar mechanism. To test this, genes encoding caa3 (S. oneidensis cox operon), bo3 (E. coli cyo operon), and aa3 (S. aureus qox operon), and the required accessary proteins were expressed in the S. oneidensis wild-type and Δcco strains (Supplementary Fig. 4a), and their influences on aminoglycoside susceptibility without or with nitrite in S. oneidensis were examined. By this way, all HCOs are compared in the same genetic background to avoid interference of physiological differences of bacteria from which each HCO comes (Fig. 1).
When isopropyl β-d-1-thiogalactoside (IPTG) was supplemented up to 1 mM, the expression of S. oneidensis caa3 did not significantly affect growth of either the wild-type or Δcco strain (Fig. 3a and Supplementary Fig. 4b). In contrast, E. coli bo3 and S. aureus aa3 produced with 0.1 mM IPTG substantially inhibited growth. We then determined whether these proteins have oxidase activity in S. oneidensis. The Nadi assay was employed for assessing caa3 activity, because the approach specifically detects cytochrome c oxidase-dependent activity25. Using N, N-dimethyl-p-phenylenediamine monohydrochloride as an exogenous electron donor, cytochrome c oxidase catalyzes the rapid formation of indophenol blue from colorless a-naphthol. As reported before16, the Δcco strain expressing caa3 with IPTG up to 0.5 mM barely showed cytochrome c oxidase activity, contrasting the wild-type and the Δcco strains expressing cco with 0.02 mM IPTG (Fig. 3b). However, when overproduced with 1 mM IPTG, caa3 did exhibit residual activity.
HCOs, including S. oneidensis caa3, E. coli bo3, and S. aureus aa3, were expressed in the S. oneidensis Δcco strain as described in Supplementary Fig. S4A. Expression was driven by IPTG-inducible promoter Ptac. a HCOs in excess inhibit growth of S. oneidensis. Shown were cell densities (by CFU) of the Δcco strain expressing one of HCOs with IPTG at indicated concentrations, which were derived from 6 h cultures presented in Supplementary Fig. S4B. Asterisks indicate statistically significant difference between values being compared (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001). b Activity of cbb3 and caa3 revealed by the Nadi assay. The method is based on the rapid formation of indophenol blue from colorless α-naphtol catalyzed by cytochrome c oxidase, using DMPD as an exogenous electron donor. Results shown were photographed 2 min after the reaction started. ∆cco/cbb3 represented the genetically complemented strain, in which a copy of cco was driven by Ptac. c Nitrite susceptibility of HCOs. Respiration rates of membranes containing one of the indicated oxidases prepared from cells grown with 0.1 mM IPTG were measured as a function of nitrite concentrations. Lines represent single exponential fits to the data (Table 1) for IC50 values. d Nitrite susceptibility of the S. oneidensis Δcyd strain expressing one of HCOs with 0.1 mM IPTG. Experiments were performed independently at least three times and representative data were presented.
To determine the activity of E. coli bo3 and S. aureus aa3 produced in S. oneidensis, membrane preparations of the ΔccoΔcyd strain expressing each of HCOs were used to measure oxygen respiration in a cell-free amperometric assay3,26. It should be noted that no aerobic growth was observed from the ΔccoΔcyd strain expressing caa3, even with 1 mM IPTG, whereas both E. coli bo3 and S. aureus aa3 supported growth, albeit slowly (Supplementary Fig. 4c). Oxygen consumption was examined with the membranes (∼200 μg/ml) using 5 mM NADH as the electron donor. Respiratory activities of the membranes of S. oneidensis cells expressing E. coli bo3 in the presence of 0.02 and 0.1 mM IPTG were recorded as ~256 and 564 μM O2/mg protein/s, respectively (Table 1). From S. oneidensis cells expressing S. aureus aa3 in the presence of 0.02 and 0.1 mM IPTG, activities were recorded as ∼481 and 814 μM O2/mg protein/s, respectively. To compare the relative sensitivities to nitrite inhibition, we measured the half-maximal inhibitory concentration for nitrite (IC50(NO2−)) for both E. coli bo3 and S. aureus aa3 in the S. oneidensis membranes (Table 1). At ∼70 μM O2, the IC50(NO2−) values for E. coli bo3 and S. aureus aa3 were ∼120 ± 13 and 137 ± 15 μM, respectively (Fig. 3c). These values are substantially higher than that for S. oneidensis cbb3 (57 ± 6 μM) but still lower than that for bd (207 ± 18 μM). We then examined nitrite susceptibility of S. oneidensis Δcyd strains expressing E. coli bo3 and S. aureus aa3. Indeed, these foreign oxidases, when expressed with 0.1 mM IPTG, elevated nitrite tolerance, albeit still more sensitive to nitrite than the wild type (Fig. 3d). All of these data indicate that bacterial HCOs are generally substantially more sensitive to nitrite than bd.
HCOs regulate nitrite-mediated aminoglycoside susceptibility
To assess the roles of caa3, bo3, and aa3 in nitrite-mediated aminoglycoside susceptibility, we examined Gent susceptibility of the Δcco strain expressing one of these oxidases. In the presence of 1 mM IPTG, caa3 could elicit a slight decrease in Gent tolerance but failed to do so when IPTG was supplemented at 0.5 mM or lower (Fig. 4a). In the presence of 0.02 mM IPTG, the Δcco strain expressing either E. coli bo3 or S. aureus aa3 became more sensitive to Gent (Fig. 4b, c). Similarly, these two oxidases produced with 0.1 mM or above IPTG sensitized cells, although in the meantime they impaired growth. We then examined the effect of nitrite on Gent tolerance of the Δcco strain expressing one of these oxidases (IPTG concentrations for caa3, 1 mM, and for bo3 and aa3, 0.1 mM). Expressions of E. coli bo3 and S. aureus aa3 substantially decreased Gent tolerance of the Δcco strain, even with 1.5 mM nitrite (Fig. 4d). Expectedly, S. oneidensis caa3 exhibited extremely low effectiveness: statistically significant differences were observed only from nitrite at 6 and 12 mM. All of these data suggest that these oxidases play a role in terms with aminoglycoside susceptibility similar to that of cbb3 in S. oneidensis.
Cell numbers before the treatment were ∼2 × 108 CFU/ml. Time-kill analysis of Gent (5× MIC) for the Δcco strain expressing one of HCOs with IPTG at indicated concentrations. a caa3, b bo3, and c aa3. Asterisks indicate statistically significant difference between values being compared (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001). d Time-kill analysis of Gent (5× MIC) and nitrite combination for the Δcco strain expressing one of HCOs. IPTG for caa3, 1 mM; for bo3 and aa3, 0.1 mM. Asterisks indicate statistically significant difference between values being compared (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001).
PMF but not respiration dictates uptake of aminoglycosides
It is well known that the uptake of aminoglycoside antibiotics by bacterial cells is an energy-requiring process that depends on PMF14. However, recent lines of evidence have suggested a link between antibiotic efficacy and bacterial cellular respiration13. Thus, it is possible that nitrite induces tolerance to aminoglycosides by impairing bacterial cellular respiration. To test this, we measured oxygen consumption rates of strains lacking each of three oxidases. As shown in Fig. 5a, the wild-type, Δcox, and Δcyd strains exhibited indistinguishable oxygen consumption and CCCP expectedly stimulated oxygen consumption substantially. In contrast, the Δcco strain was modestly defective, in comparison with the effect of electron transport inhibitor antimycin A, which abolished oxygen consumption (Fig. 5a). Thus, the cbb3 loss only mildly affects oxygen consumption rate, in line with the finding that it slightly impairs aerobic growth16. We then directly measured the generation of the membrane potential (ΔΨ) of these strains. After de-energization of the membrane potential by starvation, cells incubated in LB were stained with DiOC2 and fluorescence intensities were assayed. The wild-type cells were clearly energized but became depolarized by the addition of protonophore CCCP (Fig. 5b). Compared with the wild type, both the Δcco and Δpet strains demonstrated a dramatic loss of ΔΨ, whereas the Δcox and Δcyd strains displayed residual depolarization, signifying maintenance of a ΔΨ (Fig. 5b). Moreover, impacts of caa3, bo3, and aa3 on ΔΨ were assessed in the Δcco strain and results demonstrated that bo3 and aa3 elevated ΔΨ of Δcco cells, whereas caa3 was ineffective (Fig. 5c).
a Oxygen consumption in whole cells. Oxygen consumption was measured with a cell density of 108 cells/ml immediately after the addition of indicated chemicals. Antimycin A (Ant A) and CCCP were added at final concentrations of 10 μg/ml and 50 μM, respectively. AU, arbitrary units. b Impact of oxidases and exposure to ionophores on membrane potentials. Measurement of fluorescence intensity using dye DiOC2. The averaged fluorescence intensity of the mutants was normalized to that of the wild type, which was set to 1, giving relative Δψ. Statistics values were deduced on the basis of comparisons with the wild type. CCCP, 50 μM. Asterisks indicate statistically significant difference between values with asterisks and that of the wild type (n ≥ 3, ***P < 0.001). c Impact of oxidases and exposure to ionophores on membrane potentials. Asterisks indicate statistically significant difference between values being compared (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001). d Time-kill analysis of Gent (5× MIC) and KCN (red), BDQ (green) or CCCP (purple) combination for WT (∼2 × 108 CFU/ml before the treatment). Chemi, one of the chemicals with superscript L and H representing low and high concentrations (100 and 500 μM for KCN, 1 and 5 μM for BDQ, 10 and 50 μM for CCCP, respectively). Asterisks indicate statistically significant difference between values being compared (n ≥ 3, *P < 0.05; **P < 0.01; ***P < 0.001). Experiments were performed independently at least three times and data were presented as means ± SEM.
To further support that PMF is the primary factor for aminoglycoside tolerance. Respiration inhibitor KCN and respiration stimulator bedaquiline (BDQ), which target HCOs and the ATPase, respectively27, as well as PMF destroyer CCCP, were compared in terms of their impacts on Gent tolerance of the S. oneidensis wild-type and Δcco strains. Treatments of the wild-type strain with KCN and CCCP resulted in substantially increased tolerance to Gent, whereas no significant effect was observed from BDQ (Fig. 5d). In contrast, the Δcco strain was not responsive to any of these drugs (Supplementary Fig. 5). As BDQ differs from the other two inhibitors in that it stimulates respiration without significantly altering ΔΨ28, these data support that PMF rather than respiration efficacy dictates the uptake of aminoglycoside antibiotics.
HCOs are not involved in tolerance to other antibiotics
We further tested whether nitrite also mediates susceptibility of the S. oneidensis wild-type and Δcco strains to chloramphenicol (Cam, bacteriostatic) and rifampin (Rif, bactericidal), whose targets are in the cytoplasm as aminoglycosides. In addition, polymyxin and Cip, with which nitrite shows cooperative and antagonistic activity, respectively, against P. aeruginosa biofilms, were also included8,11. The time-kill assay revealed that nitrite had little effect on the killing of Rif, polymyxin, or Cip, regardless of the presence of cbb3 or not (Supplementary Fig. 6a). Similar results were observed with Cam (Supplementary Fig. 6a), which is expected, because this assay is not effective for bacteriostatic antibiotics. We then compared the influence of nitrite (3 and 6 mM) on growth of these strains in the presence of these antibiotics at the sublethal concentrations. Clearly, nitrite inhibited growth of the wild-type and Δcco strains comparably, and this inhibition was independent of Rif (Fig. 6a, b). In contrast, nitrite aggravated the inhibitory effect of Cam on growth of both strains, implicating that their impacts are additive (Fig. 6c). For further verification, we assessed growth of these two strains with nitrite and spectinomycin (Spect), a bacteriostatic aminoglycoside. Expectedly, nitrite improved growth of the wild type in the presence of Spect (1 μg/ml) (Fig. 6d). Importantly, this effect was not evident for the Δcco strain (Supplementary Fig. 6b), further supporting that HCOs are essential for nitrite-mediated uptake of aminoglycosides.
Discussion
Nitrite has emerged as an attractive pharmaceutical molecule in recent years, although it has been used as a bacteriostatic agent in food preservation for centuries. Nebulized sodium nitrite, a hypoxia-sensitive NO-dependent selective pulmonary vasodilator29, has been advanced into randomized trials in pulmonary arterial hypertension (PAH) patients9. Although further efforts are required to demonstrate the efficacy of nebulized sodium nitrite in PAH patients, the data at the current stage provide sufficient evidence to support the safe use of inhaled nitrite in the clinical setting. This serves great motivation for attempts to use nitrite as an antimicrobial agent alone and in combination with other antibiotics, and not surprisingly, many have been made6,8,10,11.
There have been a myriad of investigations into the mechanisms by which nitrite inhibits growth of bacteria. The prevailing theory is that the process is NO-dependent: nitrite is converted to NO in cells, which in turn directly inhibits many cellular targets5. An overwhelming majority of the NO targets identified to date are cytoplasmic metabolic enzymes with essential redox-active centers that comprise of Fe–S clusters, hemes, or protein thiols2,3,4. In this study, we show that HCOs are the most vulnerable cellular target of nitrite in bacteria, as proposed before15. Despite this, the possibility that the bc1 complex is an equally critical target of nitrite could not be ruled out and effects to test this are underway. Growth inhibition of nitrite in vivo, however, appears very modest, if not undetectable, in many bacteria, because they are additionally equipped with non-HCO oxidases, such as bd and alternative oxidases (AOXs), which are highly resistant to nitrite and NO3,15,24,30.
The data presented here manifest that nitrite induces tolerance to aminoglycosides in both Gram-negative and Gram-positive bacteria. However, it is clear that nitrite is more effective on Gram-negative than on Gram-positive bacteria. Given that all bacterial HCOs are susceptible to nitrite, other physiological aspects may be accountable for the difference. Recent studies have linked to aminoglycoside susceptibility with various metabolites31,32. Although some metabolites potentiate aminoglycoside sensitivity in both Gram-negative and Gram-positive bacteria, others can enable aminoglycoside activity in Gram-negative (E. coli) but not Gram-positive bacteria (S. aureus)31. We thus speculate that that nitrite may impact distinct metabolic pathways and/or same metabolic pathways to varying extent, modulating PMF and thereby aminoglycoside susceptibility.
In addition to cbb3, three other HCOs that are widely found in bacteria, caa3, bo3, and aa3, were examined for their roles in nitrite-mediated tolerance to aminoglycosides. In E. coli and S. aureus, bo3 and aa3 are the primary oxidases for oxygen respiration, respectively. In line with this, expression of either of these two HCOs impacts S. oneidensis physiology. Both HCOs produced at proper levels support growth of strains lacking all native oxidases, largely restore ΔΨ of the S. oneidensis cbb3-less mutant, and sensitize the cells to aminoglycosides, in comparison with the wild type. When in excess, either bo3 or aa3 severely inhibits growth of the S. oneidensis cbb3-less mutant, a widely reported phenomenon whose underpinning mechanism remains unanswered33. Moreover, our data demonstrated that both E. coli bo3 and S. aureus aa3 are more tolerant to nitrite and their expression confers the S. oneidensis bd-less mutant increased tolerance to nitrite. Nevertheless, it is conceivable that these oxidases play a dispensable role in tolerance of their native hosts to nitrite given the presence of bds and AOXs.
In the case of caa3, its physiological role in S. oneidensis and also in P. aeruginosa is negligible under normal conditions because of extremely low expression16,33,34. In this study, we detected cytochrome c oxidase activity from the S. oneidensis cbb3-less mutant expressing caa3 in the presence of 1 mM IPTG, supporting that the enzyme could function when expressed to physiologically relevant levels23,35. However, given that S. oneidensis and P. aeruginosa mutants that lack all other oxidases are unable to grow aerobically, even with caa3 forcibly produced at substantially elevated levels, it is possible that the oxidase may be only residually active, as suggested before16,35. Therefore, we were not surprised to find that the contribution of S. oneidensis caa3 to respiration, PMF generation, nitrite tolerance, or nitrite-mediated aminoglycoside tolerance is hardly detected even when overproduced.
Nowadays, it is quite common to use antibiotic combinations to treat bacterial infections. However, many in vitro studies have demonstrated attenuation of bactericidal activity across a range of drugs and organisms when used with a bacteriostatic agent36,37,38. It has been proposed that this is due to reduction of respiratory/metabolic rates by bacteriostatic agents, which lowers bactericidal antibiotic efficacy13,39. However, despite a bacteriostatic agent, nitrite does not modulate aminoglycoside tolerance mainly by impairing respiration/metabolism; instead, nitrite compromises PMF generation by directly inhibiting HCOs. We found that tolerance to aminoglycosides is not affected by BDQ, which stimulates respiration without significantly affecting ΔΨ28. Moreover, by expressing cbb3, bo3, and aa3 in the same genetic background, we were able to show that the efficacy of these enzymes in PMF generation is comparable. The correlation between the restoration of ΔΨ by bo3 and aa3 in the cbb3-less mutant and lowered tolerance to aminoglycosides supports that PMF is critical for uptake of aminoglycosides. Interestingly, with respect to respiration, non-HCO oxidases almost suffice, based on the observation that the HCO-deficient mutants grow only marginally slower than the wild type15,16,20. In contrast, non-HCO oxidases are inferior to HCOs in generating PMF, because they could not pump protons21,22. These differences therefore further support that PMF rather than respiration is critical for nitrite-modulated tolerance to aminoglycosides.
To date, there have been many reports regarding modulation of nitrite on bacterial tolerance to antibiotics, aminoglycosides in particular8,11. However, the effects observed are not consistent and even contradicting in some cases. Here we have carried out a comprehensive investigation into the mechanisms underlying nitrite-mediated aminoglycoside tolerance in multiple model bacteria. By comparing aminoglycoside susceptibility of strains lacking each of terminal oxidases and producing one of possible HCOs without and with nitrite, we discover that HCO-based respiration is essential to nitrite-mediated aminoglycoside tolerance. In contrast, HCOs are dispensable in modulating susceptibility to antibiotics other than aminoglycoside. Using biochemical inhibitors of the electron transport chain, we present evidence to show that nitrite-mediated aminoglycoside tolerance occurs through the generation of PMF rather than consumption of oxygen. Collectively, these data support nitrite as an antimicrobial agent targeting HCOs of bacteria but demand careful consideration of its application when used in combination with antibiotics.
Methods
Strains, plasmids, and culture conditions
E. coli and S. oneidensis strains and plasmids used in this study were listed in Table 2. In addition, P. aeruginosa strain PA14, S. aureus strain ATCC 25923, and B. subtilis strain 168 were used in this study. Sequences of the primers used in this study were available upon request. All chemicals are from Sigma-Aldrich, Co., unless otherwise noted. E. coli and S. oneidensis were grown aerobically in Lysogeny broth (LB, Difco, Detroit, MI) at 37 °C and 30 °C, respectively, for genetic manipulation. When appropriate, the growth medium was supplemented with the following: 2, 6-diaminopimelic acid (DAP), 0.3 mM; ampicillin (Sangon, Shanghai), 50 μg/ml; kanamycin, 50 μg/ml; and Gent, 15 μg/ml.
Antibiotic tolerance assessment
To prepare samples for antibiotic tolerance assessment, overnight cultures were inoculated into fresh LB by 200× dilution, shaken at 200 r.p.m. at 30 °C (S. oneidensis) and 37 °C (E. coli, P. aeruginosa, B. subtilis, and S. aureus) on a rotating shaker at 250 r.p.m. in flasks, and growth was recorded by measuring optical density at 600 nm (OD600). Cells grown to the mid-exponential phase (∼0.4 of OD600, the same throughout the study) were collected and adjusted to proper concentrations for subsequent analyses.
Time-kill assays were performed as described elsewhere13. Cells were plated in a six-well dish and antibiotics were added at 5× MIC for all bacteria under test, unless otherwise noted: Gent 2.5 μg/ml, Str 5 μg/ml, Cip 1 μg/ml, Rif 1 μg/ml, and colistin (polymyxin) 2.5 μg/ml. To generate biological equivalents of cell killing, 5 μg/ml Gent and 10 μg/ml Str were used for P. aeruginosa and 5 μg/ml Gent for S. aureus. Aliquots of 200 μl were taken at specified times, serially diluted, and spot-plated onto LB agar plates, to determine colony-forming units per ml (CFU/mL). Dilutions that grew 50–200 colonies were counted. For the spot dilution test, on LB agar plates.
The spot dilution test was employed to evaluate viability and growth inhibition on plates essentially the same as described previously40. In brief, cells at the mid-exponential phase were collected by centrifugation and adjusted to 109 cells/ml, which was set as undiluted (dilution factor 0). Tenfold serial dilutions were prepared with fresh LB medium and 5 μl of each dilution was dropped onto LB plates containing antibiotics MIC and/or nitrite, in the form of sodium nitrite (Sangon, Shanghai). The plates were incubated at 30 °C for 18 h or as indicated when different before being read. In addition, growth inhibition assays were used to evaluate impacts of bacteriostatic antibiotics at 0.2× MIC: Cam 2 μg/ml and Spect 8 μg/ml.
In-frame mutant construction
In-frame deletion strains were constructed using the att-based fusion PCR method as described previously41. In brief, the fusion fragments were generated from the first round of PCR products flanking the genes of interest and introduced into pHGM01, and the resulting vectors were transformed into and maintained in E. coli DAP auxotroph WM3064. After verification, the correct vectors were transferred into relevant S. oneidensis strains via conjugation, allowing integration of the fusion constructs into the chromosome. Verified transconjugants were screened for gent-sensitive and sucrose-resistant colonies. Mutants were verified by sequencing the mutated region.
Controlled gene expression
Plasmid pHGEN-Ptac was used for genetic complementation of the mutants generated in this study42. Genes of interest were generated by PCR, cloned into the vector, and the resultant vectors were transformed into E. coli WM3064. After verification by sequencing, the vectors were transferred into the relevant S. oneidensis strains via conjugation. Expression of the cloned genes was controlled by IPTG (Abcam Shanghai)-inducible promoter Ptac. For expressing S. aureus aa3, the qoxABCD operon was first cloned behind Ptac and then the ctaM gene was placed after the operon but before the terminator sequence.
Cytochrome oxidase activity assay
The Nadi assay was used for visual analysis of cytochrome c oxidase activity26. Five microliters of each culture at the mid-log phase under test was dropped onto LB plates and the plates were incubated for 24 h. A solution of 0.5% α-naphthol in 95% ethanol and 0.5% DMPD (N,N-dimethyl-ρ-phenylenediamine monohydrochloride) was applied to cover the droplets developed. Formation of indophenol blue was timed as an indicator of cytochrome c oxidase activity.
Solubilized membranes were prepared for quantitative analysis of the cytochrome oxidase activity as described previously24. In brief, cell pellets were resuspended in 20 mM Tris-HCl (pH 7.6) supplemented with DNase I and protease inhibitors, and disrupted by French Press. After debris and unbroken cells removing, the membranes were pelleted by ultracentrifugation for 1 h at 230,000 × g at 4 °C and subsequently resuspended in 20 mM Tris-HCl pH 7.6 with 5% glycerol to a protein concentration of 10 mg/ml. Solubilization was performed with n-dodecyl β-d-maltoside (DDM) to a final concentration of 1% (w/v) on a rotary tube mixer for 2 h at 4 °C. The DDM-solubilized membranes were obtained by collecting the supernatant after ultracentrifuging for 1 h at 230,000 × g at 4 °C. Pellets, containing 30–40 mg/ml of protein, were resuspended in sodium phosphate buffer pH 7.5, 50 mM NaCl, 10% glycerol, and the suspension was stored at −80 °C if not immediately used. The cytochrome oxidase activity was assayed as a measure of oxygen consumption rates using an OxyGraph oxygen electrode (Hansatech), using NADH as an electron donor according to the methods described previously3. The IC50 values of the cytochrome bd and cbb3-HCO for nitrite were obtained from plots of rates against nitrite concentrations.
The OxyGraph oxygen electrode (Hansatech) was also used to measure oxygen consumption rates of whole bacterial cells, according to the manufacturer’s instructions. S. oneidensis cells collected from cultures grown to the mid-exponential phase in LB by centrifugation were suspended to 1 × 108 cells/ml in sterile distilled water and oxygen consumption was measured at 25 °C.
Membrane potential measurements
Membrane potential (ΔΨ) measurements were carried out with the BacLight™ Bacterial Membrane Potential Kit (ThermoFisher). Cells of bacteria under test were grown to the mid-exponential phase in LB were collected, washed and resuspended in PBS buffer for 2 h at 30 °C. After starvation, cells were adjusted to the same cell density (∼107 cells/ml), incubated with 30 μM DiOC2 in LB at room temperature for 30 min in the dark. ΔΨ was analyzed using a FACSCalibur flow cytometer (Becton Dickinson) according to the manufacturer’s guidelines. As a depolarized control, the protonophore carbonyl cyanide m-chlorophenylhydrazone (CCCP) was added at a final concentration of 5 μM where indicated.
Statistics and reproducibility
Each experiment was performed at least three times independently with identical experimental settings, to ensure reproducibility. The experiments were not randomized and the investigators were not blinded to bacteria strain genotypes during experiments. Sample sizes for each experiment are indicated in the figures and figure legends. Statistical tests were performed in Microsoft Excel and GraphPad Prism 7 software. Statistical significance was presented as follows: *p < 0.05, **p < 0.01, and ***p < 0.001.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this paper.
Data availability.
The data that support the findings of this study are included in the paper or available from the corresponding author upon request. The source data presented in the main figures are provided in Supplementary Data 1.
References
Maia, L. B. & Moura, J. J. G. How biology handles nitrite. Chem. Rev. 114, 5273–5357 (2014).
Hyduke, D. R., Jarboe, L. R., Tran, L. M., Chou, K. J. Y. & Liao, J. C. Integrated network analysis identifies nitric oxide response networks and dihydroxyacid dehydratase as a crucial target in Escherichia coli. Proc. Natl Acad. Sci. USA 104, 8484–8489 (2007).
Mason, M. G. et al. Cytochrome bd confers nitric oxide resistance to Escherichia coli. Nat. Chem. Biol. 5, 94–96 (2009).
Richardson, A. R. et al. Multiple targets of nitric oxide in the tricarboxylic acid cycle of Salmonella enterica Serovar Typhimurium. Cell Host Microbe 10, 33–43 (2011).
Reddy, D., Lancaster, J. & Cornforth, D. Nitrite inhibition of Clostridium botulinum: electron spin resonance detection of iron-nitric oxide complexes. Science 221, 769–770 (1983).
Major, T. A. et al. Sodium nitrite-mediated killing of the major cystic fibrosis pathogens Pseudomonas aeruginosa, Staphylococcus aureus, and Burkholderia cepacia under anaerobic planktonic and biofilm conditions. Antimicrob. Agents Chemother. 54, 4671–4677 (2010).
Bueno, M., Wang, J., Mora, A. L. & Gladwin, M. T. Nitrite signaling in pulmonary hypertension: mechanisms of bioactivation, signaling, and therapeutics. Antioxid. Redox Signal. 18, 1797–1809 (2013).
Zemke, A. C. et al. Nitrite modulates bacterial antibiotic susceptibility and biofilm formation in association with airway epithelial cells. Free Radic. Biol. Med. 77, 307–316 (2014).
Rix, P. J. et al. Pharmacokinetics, pharmacodynamics, safety, and tolerability of nebulized sodium nitrite (AIR001) following repeat-dose inhalation in healthy subjects. Clin. Pharmacokinet. 54, 261–272 (2015).
Zemke, A. C., Gladwin, M. T. & Bomberger, J. M. Sodium nitrite blocks the activity of aminoglycosides against Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 59, 3329–3334 (2015).
Zemke, A. C., Kocak, B. R. & Bomberger, J. M. Sodium nitrite inhibits killing of Pseudomonas aeruginosa biofilms by ciprofloxacin. Antimicrob. Agents Chemother. 61, e00448–16 (2016).
Correia, S., Poeta, P., Hébraud, M., Capelo, J. L. & Igrejas, G. Mechanisms of quinolone action and resistance: where do we stand? J. Med. Microbiol. 66, 551–559 (2017).
Lobritz, M. A. et al. Antibiotic efficacy is linked to bacterial cellular respiration. Proc. Natl Acad. Sci. USA 112, 8173–8180 (2015).
Taber, H. W., Mueller, J. P., Miller, P. F. & Arrow, A. S. Bacterial uptake of aminoglycoside antibiotics. Microbiol. Rev. 51, 439–457 (1987).
Fu, H. et al. Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis. Environ. Microbiol. 15, 2198–2212 (2013).
Zhou, G. et al. Combined effect of loss of the caa3 oxidase and Crp regulation drives Shewanella to thrive in redox-stratified environments. ISME J. 7, 1752–1763 (2013).
Jin, M., Fu, H., Yin, J., Yuan, J. & Gao, H. Molecular underpinnings of nitrite effect on cyma-dependent respiration in Shewanella oneidensis. Front. Microbiol. 7, 1154 (2016).
Jin, M., Zhang, Q., Sun, Y. & Gao, H. NapB in excess inhibits growth of Shewanella oneidensis by dissipating electrons of the quinol pool. Sci. Rep. 6, 37456 (2016).
Meng, Q., Yin, J., Jin, M. & Gao, H. Distinct nitrite and nitric oxide physiologies in Escherichia coli and Shewanella oneidensis. Appl. Environ. Microbiol. 84, e00559–18 (2018).
Meng, Q., Sun, Y. & Gao, H. Cytochromes c constitute a layer of protection against nitric oxide but not nitrite. Appl. Environ. Microbiol. 84, e01255–18 (2018).
Yoshikawa, S. & Shimada, A. Reaction mechanism of cytochrome c oxidase. Chem. Rev. 115, 1936–1989 (2015).
Borisov, V. B., Gennis, R. B., Hemp, J. & Verkhovsky, M. I. The cytochrome bd respiratory oxygen reductases. Biochim. Biophys. Acta 1807, 1398–1413 (2011).
Le Laz, S. et al. Expression of terminal oxidases under nutrient-starved conditions in Shewanella oneidensis: detection of the A-type cytochrome c oxidase. Sci. Rep. 6, 19726 (2016).
Yin, J. et al. Regulation of nitrite resistance of the cytochrome cbb3 oxidase by cytochrome c ScyA in Shewanella oneidensis. Microbiol. Open 4, 84–99 (2015).
Marrs, B. & Gest, H. Genetic mutations affecting the respiratory electron-transport system of the photosynthetic bacterium Rhodopseudomonas capsulata. J. Bacteriol. 114, 1045–1051 (1973).
Hammer, N. D., Schurig-Briccio, L. A., Gerdes, S. Y., Gennis, R. B. & Skaar, E. P. CtaM is required for menaquinol oxidase aa3 function in Staphylococcus aureus. mBio 7, e00823–16 (2016).
Andries, K. et al. A diarylquinoline drug active on the ATP synthase of Mycobacterium tuberculosis. Science 307, 223–227 (2005).
Lamprecht, D. A. et al. Turning the respiratory flexibility of Mycobacterium tuberculosis against itself. Nat. Commun. 7, 12393 (2016).
Hunter, C. J. et al. Inhaled nebulized nitrite is a hypoxia-sensitive NO-dependent selective pulmonary vasodilator. Nat. Med. 10, 1122–1127 (2004).
May, B., Young, L. & Moore, A. L. Structural insights into the alternative oxidases: are all oxidases made equal? Biochem. Soc. Trans. 45, 731–740 (2017).
Allison, K. R., Brynildsen, M. P. & Collins, J. J. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature 473, 216–220 (2011).
Meylan, S. et al. Carbon sources tune antibiotic susceptibility in Pseudomonas aeruginosa via tricarboxylic acid cycle control. Cell Chem. Biol. 24, 195–206 (2017).
Arai, H. et al. Enzymatic characterization and in vivo function of five terminal oxidases in Pseudomonas aeruginosa. J. Bacteriol. 196, 4206–4215 (2014).
Kawakami, T., Kuroki, M., Ishii, M., Igarashi, Y. & Arai, H. Differential expression of multiple terminal oxidases for aerobic respiration in Pseudomonas aeruginosa. Environ. Microbiol. 12, 1399–1412 (2010).
Osamura, T., Kawakami, T., Kido, R., Ishii, M. & Arai, H. Specific expression and function of the A-type cytochrome c oxidase under starvation conditions in Pseudomonas aeruginosa. PLoS ONE 12, e0177957 (2017).
Brown, T. H. & Alford, R. H. Antagonism by chloramphenicol of broad-spectrum beta-lactam antibiotics against Klebsiella pneumoniae. Antimicrob. Agents Chemother. 25, 405–407 (1984).
Ankomah, P., Johnson, P. J. T. & Levin, B. R. The pharmaco –, population and evolutionary dynamics of multi-drug therapy: experiments with S. aureus and E. coli and computer simulations. PLoS Pathog. 9, e1003300 (2013).
Johansen, H. K., Jensen, T. G., Dessau, R. B., Lundgren, B. & Frimodt-Møller, N. Antagonism between penicillin and erythromycin against Streptococcus pneumoniae in vitro and in vivo. J. Antimicrob. Chemother. 46, 973–980 (2000).
Rittershaus, E. S. C., Baek, S.-H. & Sassetti, C. M. The normalcy of dormancy: common themes in microbial quiescence. Cell Host Microbe 13, 643–651 (2013).
Jiang, Y. et al. Protection from oxidative stress relies mainly on derepression of OxyR-dependent KatB and Dps in Shewanella oneidensis. J. Bacteriol. 196, 445–458 (2014).
Jin, M. et al. Unique organizational and functional features of the cytochrome c maturation system in Shewanella oneidensis. PLoS ONE 8, e75610 (2013).
Meng, Q., Liang, H. & Gao, H. Roles of multiple KASIII homologues of Shewanella oneidensis in initiation of fatty acid synthesis and in cerulenin resistance. Biochim. Biophys. Acta 1863, 1153–1163 (2018).
Chen, H., Luo, Q., Yin, J., Gao, T. & Gao, H. Evidence for the requirement of CydX in function but not assembly of the cytochrome bd oxidase in Shewanella oneidensis. Biochim. Biophys. Acta 1850, 318–328 (2015).
Fu, H., Jin, M., Ju, L., Mao, Y. & Gao, H. Evidence for function overlapping of CymA and the cytochrome bc1 complex in the Shewanella oneidensis nitrate and nitrite respiration. Environ. Microbiol. 16, 3181–3195 (2014).
Acknowledgements
This research was supported by National Natural Science Foundation of China (31930003,41976087), by Ten Thousand Talent Program of China, and by the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Contributions
Y.Z. and H.G. designed the research and wrote the paper. Y.Z., K.G., and Q.M. performed research and analyzed data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Guo, K., Meng, Q. et al. Nitrite modulates aminoglycoside tolerance by inhibiting cytochrome heme-copper oxidase in bacteria. Commun Biol 3, 269 (2020). https://doi.org/10.1038/s42003-020-0991-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-0991-4
This article is cited by
-
A mini-review: environmental and metabolic factors affecting aminoglycoside efficacy
World Journal of Microbiology and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.