Abstract
Ancient remains found in permafrost represent a rare opportunity to study past ecosystems. Here, we present an exceptionally well-preserved ancient bird carcass found in the Siberian permafrost, along with a radiocarbon date and a reconstruction of its complete mitochondrial genome. The carcass was radiocarbon dated to approximately 44–49 ka BP, and was genetically identified as a female horned lark. This is a species that usually inhabits open habitat, such as the steppe environment that existed in Siberia at the time. This near-intact carcass highlights the potential of permafrost remains for evolutionary studies that combine both morphology and ancient nucleic acids.
Similar content being viewed by others
Introduction
Permafrost deposits containing both animal and plant material represent a unique opportunity to reconstruct paleoenvironments1,2. In recent years, permafrost sites in the Arctic have revealed a wealth of frozen animal carcasses from the last glaciation, including mammoths, woolly rhinoceroses, horses, bisons, and wolverines3,4. These remains are of great interest to paleontology since they enable a better understanding of the impact of climate change on species, populations, and communities1,2. Moreover, because such frozen carcasses are often extremely well preserved, they allow for studies of morphological traits, as well as the ecology and evolution of a range of extinct and extant animal species (e.g.5,6,7). The high degree of preservation of both tissue and DNA in permafrost remains has been particularly beneficial to the field of ancient DNA (aDNA). For instance, sequencing of DNA from Pleistocene and Holocene mammal remains has provided a better understanding of the behaviour of extinct species5,8, while the sequencing of complete Pleistocene genomes has allowed studies of evolutionary rates, extinction dynamics and inter-specific hybridisation9,10,11. However, most examples of this type of frozen permafrost tissues have been from large mammals. To our knowledge, no frozen bird carcasses have yet been described from late Pleistocene permafrost deposits.
Here, we present a complete bird specimen located within an undisturbed permafrost sequence in the Belaya Gora area of north-eastern Siberia. We radiocarbon dated this specimen and used high-throughput shotgun sequencing to identify the species through reconstruction of its complete mitogenome. This unique find highlights the importance of preserving newly discovered permafrost remains for future genomic and morphometric studies of Pleistocene fauna.
Results
Radiocarbon dating and DNA sequencing of bird carcass found in permafrost
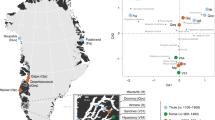
The frozen carcass of a near-complete passerine (Fig. 1) was recovered in permafrost from a site 30 km east from the village of Belaya Gora, Yakutia (N68.57887, E147.16055; Fig. 2). The site comprises a series of tunnels that have been hydraulically mined into the permafrost by fossil ivory hunters, and is located on the small Tirekhtyakh River, which is a tributary to the Indigirka River. The bird carcass was found ~150 lateral meters into one of the tunnels, at a depth of roughly 7 meters from the surface.
The bird was radiocarbon dated to 42,600 +/− 1100 years BP (OxA-38572), which corresponds to a calibrated age between 44,163–48,752 Cal years BP12. We subsequently used ~50 mg of the bird tissue for DNA extraction and whole genome shotgun-sequencing. To identify the species, we then mapped the shotgun-sequencing data to the chicken (Gallus gallus; GenBank: NC_001323) mitogenome. Out of a total of 85,408,658 reads, 3,663 (<0.01%) mapped against the chicken mitogenome. While the reconstructed mitogenome was incomplete, we extracted a partial COI gene (232 bp), which is routinely used for species identification. We then searched for matches with this gene in GenBank avian genetic databases using BLASTn13,14 and found a 100% identity match with the horned lark (Eremophila alpestris).
To estimate the overall endogenous DNA content of the ancient horned lark extract, we then mapped the shotgun-sequencing data to a nuclear genome assembly from a modern horned lark (Sigeman et al.15; N50 = 20 kb; subspecies E. a. flava). Out of 8,5408,658 reads, 757,883 (0.89%) reads mapped to this nuclear genome. The average genome depth was of 0.032X (min = 0; max = 116), with 2.5% of the assembly having a coverage ≥1X. We also found that there were on average 37 and 78 reads per 100 kb mapping to Z-linked and autosomal contigs, respectively. This corresponds to a ratio of Z-linked/autosomal reads of 0.47. Since female birds are the heterogametic sex (ZW), and since the female-specific W chromosome is highly degraded and differentiated to the Z chromosome over most of its length15, this ratio indicates that the specimen was a female.
Next, to improve the quality of the ancient horned lark mitogenome, we remapped the shotgun-sequencing data against a modern horned lark de novo mitogenome15, which we generated using Mitobim v1.9.016. This resulted in an ancient consensus mitogenome with an average depth of 134X (min = 18; max = 264) and 40,685 (0.05%) reads mapping to the modern horned lark mitogenome. To confirm species identity, we then extracted the complete COI (683 bp) and cytb (987 bp) genes, which had a 99% and 95% identity match with horned lark, respectively.
Phylogenetic placement of the ancient bird specimen
A comprehensive study on lark phylogeny resolving the complex relationships among larks (Alaudidae) indicated dramatic morphological divergence in some lineages as well as multiple examples of parallel morphological evolution17. Within larks, the systematics and evolution of the genus Eremophila were until recently poorly understood, with many subspecies described (owing in part to the variation and divergence in plumage18,19). However, a recent study from Ghorbani et al.20 based on ND2 and cytb genes identified four distinct Eremophila lineages that diverged in the late Pliocene to early Pleistocene. Furthermore, E. alpestris may have originated in north-eastern Siberia during the middle Pleistocene18,20, before diverging into Eurasian and North American clades.
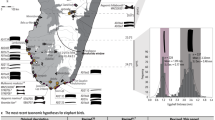
Specimens such as this ancient lark present an opportunity to understand the evolution and biogeography of this poorly resolved Pleistocene species. We thus examined the phylogenetic placement of the ancient horned lark within the diversity of extant larks using the modern ND2 and cytb (2034 bp) datafrom Ghorbani et al.20 and a Bayesian relaxed clock model implemented in BEAST221. The ancient horned lark was placed close to the node between E. a. flava and E. a. brandti (Fig. 3), which are nowadays distributed across the north Palearctic (Scandinavia, Northern Russia) and central Palearctic aridlands (Kazakhstan, Mongolia, China), respectively19,22. Pollen records from the Yakutia region indicate that the habitat was dominated by a mix of tundra and steppe environments between 50 and 30 ka BP23, making E. a. flava the most likely ecological analogue to this ancient specimen18. Notably there was low posterior branch support (<0.5) for assigning the ancient specimen to either of these subspecies. Based on this, we hypothesize that the ancient specimen may have belonged to an ancestral population for E. a. flava and E. a. brandti, and that these two lineages subsequently evolved into separate subspecies as a consequence of environmental changes during the Pleistocene/Holocene transition. Such a Holocene divergence into tundra and steppe forms is consistent with earlier work by Edwards et al.24 proposing a general stratification from a mixed steppe-tundra into separate tundra and steppe biomes at the end of the last glaciation.
The phylogeny is inferred from the Bayesian analysis of concatenated ND2 and cytb (2034 bp) for 31 horned lark from Ghorbani et al.20. The ancient horned lark highlighted in gray. Nodes with <0.5 support should be considered collapsed.
Discussion
The methods used by fossil ivory hunters may cause damage to animal remains of special scientific value, particularly those with smaller body size that are not easily identified during hydraulic mining. Nevertheless, the exceptionally high degree of morphological preservation of this specimen and of similarly well-preserved animal remains from the area around Belaya Gora holds great promise for further research into the region’s evolutionary history. For instance, molecular identification of the sex of animal remains, which is not always possible based on skeletal remains, allows for investigations into the behavioural ecology of extinct species5,8, while sequencing of mitochondrial or nuclear genomes can enable studies of temporal range shifts associated with past climatic fluctuations25, as well as reconstructing the past demography in extant and extinct species11,26,27. Moreover, because estimates of substitution rates from modern pedigree data can be biased28, whole genome sequencing of permafrost specimens could provide more accurate estimation of molecular clocks and thus allow improved studies of the micro-evolution of a range of species (e.g., 29,30). Finally, one especially important advantage of frozen carcasses lies in the preservation of several different tissues and organs. Sequencing RNA from these tissues would give information beyond what is achievable from genomic data alone, enabling studies of gene expression, as shown recently in a study on ancient wolf remains dating back some 14 ka BP31. Pleistocene tissue remains recovered from permafrost thus have the potential to become instrumental in better understanding processes such as biological regulation and gene expression in relation to past climate change.
Methods
Sample collection and radiocarbon dating
Following retrieval of the near-complete bird carcass (Figs. 1 and 4) from the permafrost tunnel located along the Tirekhtyakh River (Fig. 2), we excised ~120 mg of feather from the specimen and used this for radiocarbon dating at the Oxford Radiocarbon Accelerator Unit (ORAU) using the Oxcal computer program (v4.3)12. In addition, a small piece of tissue (~300 mg) was collected and stored at −20 °C for subsequent DNA analysis. The carcass itself has been accessioned into the collections at the Sakha Academy of Sciences in Yakutsk (accession ID: 2018-Tir-Shorelark-01).
DNA extraction and sequencing
After an overnight digestion of ~50 mg of bird tissue in a buffer optimised to digest keratin-rich tissues32, we extracted DNA using the DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany). We then built a double-stranded Illumina library according to Meyer & Kircher33. We used 20 μl of DNA extract in a 40 μl blunt-end repair reaction with the following final concentration: 1×buffer Tango, 100 μM of each dNTP, 1 mM ATP, 20 U T4 polynucleotide kinase (Thermo Scientific), and 3U USER enzyme (New England Biolabs). Treatment with USER enzyme was performed to excise uracil residues resulting from post-mortem damage34,35. The sample was incubated for 3 h at 37 °C, followed by the addition of 0.8 μl T4 DNA polymerase (Thermo Scientific), and incubation at 25 °C for 15 min and 12 °C for 5 min and was then cleaned using MinElute (Qiagen, Hilden, Germany) spin columns following the manufacturer’s protocol and eluted in 20 μl EB Buffer. Next, we performed an adapter ligation step where DNA fragments of the library were ligated to a combination of incomplete, partially double-stranded P5- and P7-adapters. This reaction was performed in a 40 μl reaction volume using 20 μl of blunted DNA from the clean-up step and 2 μl P5–P7 adapter mix (0.5 μM final concentration for each adapter) per sample with a final concentration of 1×T4 DNA ligase buffer, 5% PEG-4000, 5U T4 DNA ligase (Thermo Scientific). The sample was incubated for 30 min at room temperature and cleaned using MinElute spin columns as described above. Next, we performed an adapter fill-in reaction in 40 μl final volume using 20 μl adapter-ligated DNA with a final concentration of 1× Thermopol Reaction Buffer, 250 μM of each dNTP, 12U Bst Polymerase (Thermo Scientific), Long Fragments. The library was incubated at 37 °C for 20 min and heat-inactivated at 80 °C for 20 min.
This library was then used as stock for indexing PCR amplification using one indexing PCR amplification with double-indexed P5–P7 primers. The amplification was performed in 25 μl volumes with 3 μl of adapter-ligated library as template, with the following final concentrations: 1x AccuPrime reaction mix, 0.3 μM P7-P5 indexing primer mix, 7 U AccuPrime Pfx (Thermo Scientific), and the following cycling protocol: 95 °C for 2 min, 12 cycles at 95 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min, and a final extension at 72 °C for 5 min.
Purification and size selection of the library was then performed using Agencourt AMPure XP beads (Beckman Coulter, Brea, CA, USA), first using 0.5X bead:DNA ratio and secondly 1.8X to remove long and short (i.e., adapter dimers) fragments, respectively. Library concentration was measured with a high-sensitivity DNAchip on a Bioanalyzer 2100 (Agilent, Santa Clara, CA, USA). Finally, the library was sequenced on an Illumina NovaSeq6000 SP lane with a 2 × 50 bp setup, incl. Xp kit (validated method) at the SciLifeLab sequencing facility in Stockholm.
Extractions and library preparation were conducted in a separate aDNA lab and appropriate precautions were taken to minimize the risk of contamination of the ancient sample36.
Data processing, genome reconstruction and analyses
First, raw aDNA data was demultiplexed using bcl2Fastq v2.17.1 with default settings (Illumina Inc.). We then trimmed adapters and merged paired-end reads using SeqPrep v1.137 with default settings but with a minor modification in the source code, allowing us to choose the best quality scores of bases in the merged region instead of aggregating the scores following Palkopoulou et al.11. We then merged sequencing reads and mapped them against the reference mitogenomes for chicken (Gallus gallus; GenBank: NC_001323) mitogenome using the BWA v0.7.13 aln algorithm38 and slightly modified default settings with deactivated seeding (−l 16,500), allowing more substitutions (−n 0.01) and allowing up to two gaps (−o 2). The BWA samse command was then used to generate alignments in SAM format. The resulting reads were then processed in SAMtools v1.939, converted to BAM format coordinate sorted and indexed. We removed duplicates from the alignments using a custom python script to avoid inflation of length distribution for loci with deep coverage11. Next, we used Picard v1.141 (http://broadinstitute.github.io/picard) to assign read group information including library, lane and sample identity to each bam file. Reads were then re-aligned around indels using GATK v3.4.040. Only reads/alignments with mapping quality ≥30 were kept for subsequent analysis. We then estimated the endogenous mitochondrial DNA content (i.e., the proportion of reads mapping to the mitogenome) and coverage with SAMtools.
Secondly, we reconstructed a consensus mitogenome using a majority consensus rule and a minimum of 5X coverage in Geneious® v7.0.33641 and the raw data mapped to chicken. We extracted a partial COI gene (232 bp) from this mitogenome and used it as input for species identification in BLASTn13,14.
Third, after identifying a 100% identity match with the horned lark (Eremophila alpestris), we generated a de novo mitogenome for a modern horned lark (E. a. flava), using whole genome sequencing data from Sigeman et al.15, with Mitobim v1.9.016 using the ‘quick’ option and the Mongolian lark (Melanocorypha mongolica; GENBANK: NC_036760), which is the closest-related species with a compete mitogenome17 as bait reference. Reconstruction of this mitogenome was done after four iterations. Next, we remapped the raw data for the ancient horned lark to the modern lark mitogenome, built its consensus genome in Geneious® using the same parameters as above and estimated the endogenous mitochondrial DNA content, as described above. We also extracted the cytb (987 bp) and COI (683 bp) genes to confirm species identity in BLASTn13,14.
Fourth, we investigated the phylogenetic relationships between the ancient horned lark and modern horned larks by including it in a mitochondrial data set consisting of 2034 bp (996 bp cytochrome b and 1038 bp ND2) from Ghorbani et al.20. We only included specimens without missing data, making for a total of 31 modern larks. The evolutionary relationship was inferred by implementing a Bayesian relaxed clock model in BEAST2 v2.4.021. We ran Markov chain Monte Carlo chains for 80 million generations (sampling every 100 generations) using a relaxed lognormal distribution for the molecular clock model and assuming a birth-death speciation process for the tree prior. We used a GTR + G and HKY + G model for cytb and for ND2, respectively based on Ghorbani et al.20. The tree was calibrated by using the mean radio carbon date value for ancient horned lark as tip-date. We checked for convergence between runs and analysis performance using Tracer v1.642 and accepted the results if the values of the estimated sample size (ESS) were larger than 200, suggesting little autocorrelation between samples. The resulting trees were combined in TreeAnnotator v1.7.543 and the consensus tree with the divergence dates was visualized in FigTree v1.4.344.
Fifth, to estimate the overall endogenous DNA content, we mapped all raw data to the complete nuclear genome of the modern horned lark (Sigeman et al.15; N50 = 20 kb, 243,805 contigs) using the same mapping parameters as described above for mapping data to mitogenomes.
Finally, to determine the sex of the bird, we counted the number of reads mapping to autosomal (0.95 Gb) and Z-linked contigs (60.7Mb15) and calculated the average number of reads mapping to each contig groups per block of 100 kb. In birds, males are the homogametic sex (ZZ) while females are the heterogametic sex (ZW). Thus, a male should have the same proportion of reads mapping to autosomal and Z-linked contigs, making for a Z/autosomal ratio of ~1. Conversely, a female should have half the number of reads mapping to the Z-link contigs compared to autosomal contigs, making for a Z/autosomal ratio of ~0.5. A total of 22,356 and 735,448 reads mapped to the to Z-linked (n = 2006) and autosomal (n = 241,800) contigs, respectively. This corresponded to an average of 37 and 78 reads per 100 kb mapped to Z-linked and autosomal contigs, respectively.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw fastq files and bam files mapped to the de novo horned lark mitogenome are deposited at the European Nucleotide Archive (ENA; Proj ID PRJEB35255). Concatenated cytb+ND2 sequences are provided in Supplementary Data 1.
References
Schirrmeister, L. et al. Paleoenvironmental and paleoclimatic records from permafrost deposits in the Arctic region of Northern Siberia. Quat. Int. 89, 97–118 (2002).
Wetterich, S. et al. Palaeoenvironmental dynamics inferred from late Quaternary permafrost deposits on Kurungnakh Island, Lena Delta, Northeast Siberia, Russia. Quat. Sci. Rev. 27, 1523–1540 (2008).
Boeskorov, G. G. et al. New findings of unique preserved fossil mammals in the permafrost of Yakutia. Dokl. Biol. Sci. 452, 291–295 (2013).
Boeskorov, G. G. et al. A study of a frozen mummy of a wild horse from the Holocene of Yakutia, East Siberia, Russia. Mammal. Res. 63, 307–314 (2018).
Pečnerová, P. et al. Genome-based sexing provides clues about behavior and social structure in the woolly mammoth. Curr. Biol. 27, 3505–3510 (2017).
Tridico, S. R., Rigby, P., Kirkbride, K. P., Haile, J. & Bunce, M. Megafaunal split ends: Microscopical characterisation of hair structure and function in extinct woolly mammoth and woolly rhino. Quat. Sci. Rev. 83, 68–75 (2014).
Fisher, D. C. et al. Anatomy, death, and preservation of a woolly mammoth (Mammuthus primigenius) calf, Yamal Peninsula, northwest Siberia. Quat. Int. 255, 94–105 (2012).
Gower, G. et al. Widespread male sex bias in mammal fossil and museum collections. Proc. Natl. Acad. Sci. USA 201903275, https://doi.org/10.1073/pnas.1903275116 (2019).
Barlow, A. et al. Partial genomic survival of cave bears in living brown bears. Nat. Ecol. Evol. https://doi.org/10.1038/s41559-018-0654-8 (2018).
Orlando, L. et al. Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature 499, 74–78 (2013).
Palkopoulou, E. et al. Complete genomes reveal signatures of demographic and genetic declines in the woolly mammoth. Curr. Biol. 25, 1395–1400 (2015).
Ramsey, C. B. & Lee, S. Recent and planned developments of the program OxCal. Radiocarbon 55, 720–730 (2013).
Altschul, S. F., Gish, W. R., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36 (Suppl 2), W5–W9 (2008).
Sigeman, H. et al. Repeated sex chromosome evolution in vertebrates supported by expanded avian sex chromosomes. Proc. R. Soc. B 20192051, https://doi.org/10.1098/rspb.2019.2051 (2019).
Hahn, C., Bachmann, L. & Chevreux, B. Reconstructing mitochondrial genomes directly from genomic next-generation sequencing reads - A baiting and iterative mapping approach. Nucleic Acids Res. 41 e19 (2013).
Alström, P. et al. Multilocus phylogeny of the avian family Alaudidae (larks) reveals complex morphological evolution, non-monophyletic genera and hidden species diversity. Mol. Phylogenet. Evol. 69, 1043–1056 (2013).
Drovetski, S. V., Raković, M., Semenov, G., Fadeev, I. V. & Red’kin, Y. A. Limited phylogeographic signal in sex-linked and autosomal loci despite geographically, ecologically, and phenotypically concordant structure of mtDNA variation in the holarctic avian genus Eremophila. PLoS ONE 9, e87570 (2014).
de Juana, E., Suarez, F., Ryan, P., Alström, P. & Donald, P. in Handbook of the birds of the World, Volume 9: Cotingas to Pipits and Wagtails 496–601 (eds Hoyo, J., Elliott, A. & Christie, D.) https://doi.org/10.1676/0043-5643(2006)118[430:ol]2.0.co;2 (2004).
Ghorbani, F. et al. Mitochondrial phylogeography of the genus Eremophila confirms underestimated species diversity in the Palearctic. J. Ornithol. https://doi.org/10.1007/s10336-019-01714-2 (2019).
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Mullarney, K., Svensson, L., Zetterström, D. & Grant, P. J. Bird Guide. the Most Complete Field Guide to the Birds of Britain and Europe. (HarperCollins Publishers Ltd., London 2006).
Andreev, A. A. et al. Vegetation and climate history in the Laptev Sea region (Arctic Siberia) during Late Quaternary inferred from pollen records. Quat. Sci. Rev. 30, 2182–2199 (2011).
Edwards, M. E., Brubaker, L. B., Lozhkin, A. V. & Anderson, P. M. Structurally novel biomes: a response to past warming in Beringia. Ecology 86, 1696–1703 (2005).
Palkopoulou, E. et al. Synchronous genetic turnovers across Western Eurasia in Late Pleistocene collared lemmings. Glob. Chang. Biol. 22, 1710–1721 (2016).
Ersmark, E. et al. Population Demography and Genetic Diversity in the Pleistocene Cave Lion. Open Quat. 1, 1–14 (2015).
Shapiro, B. et al. Rise and fall of the Beringian steppe bison. Science 306, 1561–1565 (2004).
Ho, S. Y. W., Phillips, M. J., Cooper, A. & Drummond, A. J. Time dependency of molecular rate estimates and systematic overestimation of recent divergence times. Mol. Biol. Evol. 22, 1561–1568 (2005).
Lambert, D. M. et al. Rates of evolution in ancient DNA from Adélie penguins. Science 295, 2270–2273 (2002).
Millar, C. D. et al. Mutation and evolutionary rates in adélie penguins from the Antarctic. PLoS Genet. 4, e1000209 (2008).
Smith, O. et al. Ancient RNA from Late Pleistocene permafrost and historical canids shows tissue-specific transcriptome survival. PLoS Biol. 17, e3000166 (2019).
Gilbert, M. T. P. et al. Whole-genome shotgun sequencing of mitochondria from ancient hair shafts. Science 317, 1927–1930 (2007).
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 5, pdb.prot5448 (2010).
Briggs, A. W. et al. Removal of deaminated cytosines and detection of in vivo methylation in ancient DNA. Nucleic Acids Res. 38, 1–12 (2010).
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res. 40, e3 (2012).
Knapp, M., Clarke, A. C., Horsburgh, K. A. & Matisoo-Smith, E. A. Setting the stage - Building and working in an ancient DNA laboratory. Ann. Anat. 194, 3–6 (2012).
John JS. SeqPrep 1.1, https://github.com/jstjohn/SeqPrep. (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Kearse, M. et al. Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Rambaut, A., Suchard, M., Xie, D. & Drummond, A. ‘Tracer’ 1.6. http://beast.bio.ed.ac.uk/Tracer (2014).
Rambaut, A. & Drummond, A. TreeAnnotator v1.7.0. Univ. Edinburgh, Inst. Evol. Biol. (2013).
Rambaut, A. FigTree, a graphical viewer of phylogenetic trees. https://tree.bio.ed.ac.uk/software/figtree (2007).
Acknowledgements
Genome sequencing and radiocarbon dating was funded through a grant from FORMAS (grant no. 2018-01640). The authors acknowledge Boris Berezhnov and Spartak Khabrov for access to the bird remains. The authors also acknowledge support from the Uppsala Multidisciplinary Centre for Advanced Computational Science for assistance with massively parallel sequencing and access to the UPPMAX computational infrastructure. Sequencing was performed by the Swedish National Genomics Infrastructure (NGI) at the Science for Life Laboratory, which is supported by the Swedish Research Council and the Knut and Alice Wallenberg Foundation. The field work was supported by Renegade Pictures UK Ltd. Open access funding provided by Stockholm University.
Author information
Authors and Affiliations
Contributions
N.D. and L.D. conceived the study. D.W.G.S. performed DNA extractions and library preparation. N.D. and P.G.P.E. analysed the data. N.D. wrote the manuscript. N.D., D.W.G.S., H.S., P.G.P.E., J.G., D.C.F., A.V.P., V.L.H., V.P., B.H., and L.D. contributed to the final version of the manuscript and approved it for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dussex, N., Stanton, D.W.G., Sigeman, H. et al. Biomolecular analyses reveal the age, sex and species identity of a near-intact Pleistocene bird carcass. Commun Biol 3, 84 (2020). https://doi.org/10.1038/s42003-020-0806-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-0806-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.