Abstract
Iron is essential for many biological functions and iron deficiency and overload have major health implications. We performed a meta-analysis of three genome-wide association studies from Iceland, the UK and Denmark of blood levels of ferritin (N = 246,139), total iron binding capacity (N = 135,430), iron (N = 163,511) and transferrin saturation (N = 131,471). We found 62 independent sequence variants associating with iron homeostasis parameters at 56 loci, including 46 novel loci. Variants at DUOX2, F5, SLC11A2 and TMPRSS6 associate with iron deficiency anemia, while variants at TF, HFE, TFR2 and TMPRSS6 associate with iron overload. A HBS1L-MYB intergenic region variant associates both with increased risk of iron overload and reduced risk of iron deficiency anemia. The DUOX2 missense variant is present in 14% of the population, associates with all iron homeostasis biomarkers, and increases the risk of iron deficiency anemia by 29%. The associations implicate proteins contributing to the main physiological processes involved in iron homeostasis: iron sensing and storage, inflammation, absorption of iron from the gut, iron recycling, erythropoiesis and bleeding/menstruation.
Similar content being viewed by others
Introduction
Iron is an essential element for a wide variety of metabolic processes such as oxygen transport, cellular respiration, and redox reactions in numerous metabolic pathways. For this reason, iron homeostasis is tightly regulated on cellular and systemic levels to ensure a balance between uptake, transport, storage, and utilization. Iron deficiency is one of the five leading causes of disability worldwide, especially among children and women of childbearing age1,2. Similarly, iron overload is associated with an increased risk of several major chronic conditions, including diabetes and liver disease1,3.
Four iron biomarkers are used for clinical assessment of iron status: serum ferritin, serum iron, and total iron-binding capacity (TIBC) are measured directly, while transferrin saturation (TSAT) is derived as serum iron divided by TIBC. While serum ferritin correlates well with body iron stores in non-inflamed individuals4, TSAT measures the proportion of iron-binding sites of transferrin that are occupied by iron. TSAT indicates the availability of iron for erythropoiesis and is low in iron deficiency and high during iron overload. In some forms of anemia (e.g., anemia of inflammation) the iron is not transported efficiently to the bone marrow for erythropoiesis, despite adequate iron stores. Since in this situation there is adequate ferritin but low TSAT, it is useful to evaluate TSAT in addition to ferritin4,5.
Genome-wide association studies (GWAS) have previously investigated the association between sequence variants and iron homeostasis biomarkers6,7,8. The largest study to date yielded 11 loci: ABO, ARNTL, FADS2, HFE, NAT2, SLC40A1, TEX14, TF, TFR2, TFRC, and TMPRSS6 associating with one or more iron homeostasis biomarkers (ferritin, iron, TIBC or TSAT)6. To search for additional sequence variants associated with iron homeostasis, we performed a GWAS meta-analysis of ferritin, serum iron, TIBC, and TSAT in Iceland and blood donor studies from the UK (INTERVAL study) and Denmark (Danish Blood Donor Study). This was followed by cross-referencing of iron-associated loci with clinically relevant phenotypes (including iron deficiency anemia (IDA), iron overload, and red blood cell indices). We report associations with iron homeostasis biomarkers for 62 independent sequence variants at 56 loci, including 46 novel loci. Based on a literature review, we categorize 25 of these loci as involved in iron sensing or storage, inflammation, gut absorption, iron recycling, erythropoiesis, and bleeding/menstruation.
Results
Overview
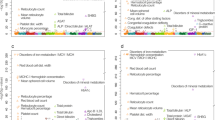
We performed a meta-analysis of four iron-related biomarkers: ferritin (N = 246,139), serum iron (N = 163,511), TIBC (N = 135,430), and TSAT (N = 131,471), combining GWAS results from Iceland, the UK, and Denmark (Fig. 1, Supplementary Data 1). We found associations with iron homeostasis biomarkers represented by 62 sequence variants at 56 loci, of which 46 have not been reported in the previous GWAS on iron homeostasis and are therefore considered novel (Table 1, Table 2, Fig. 2, and Supplementary Data 2). For each locus, we report the lead variant (lowest P value) and additional uncorrelated variants (r2 < 0.1) within the locus with genome-wide significance. Our criteria for statistical significance have been previously described9 (see “Methods”). A variant-to-gene mapping algorithm that takes into account gene location, variant effect (for coding variants), and effect on gene expression (eQTL) for each variant (lead variant and LD class) was used to choose a single candidate gene for each locus (see “Methods”). Twenty-five of the 62 iron homeostasis-associated sequence variants have a high-confidence predicted causal gene, 23 variants have multiple top-scoring genes, 36 variants have at least one coding variant or eQTL in the LD class, and 13 variants have more than one gene with coding variants and/or eQTL in the LD class (Supplementary Data 3). The LD class of a variant is defined as all variants having r2 > 0.8 with the variant. Linkage disequilibrium (LD) (r2) is estimated based on the Icelandic population. In cases where variants had more than one top-scoring gene, the gene closest to the lead variant was selected, except for two loci where likely candidate genes were present among the top-scoring genes (FTL (ferritin light chain) and HAMP (hepcidin)) (Supplementary Data 3). Fourteen of the variants associated with more than one biomarker, bringing the total number of observed associations to 87 (Supplementary Data 2). All our associations have P < 3.0 × 10−8. We replicated the association of all 11 previously reported variants6, 10 at genome-wide significance (Supplementary Data 2). In addition, we found six rare variants (minor allele frequency (MAF) < 1%), six low-frequency variants (1% ≤ MAF < 5%), and 37 common variants that have previously not been reported to associate with iron homeostasis biomarkers (Supplementary Data 2). Forty-six variants associate with a single iron biomarker (ferritin, 34; iron, 8; TIBC, 4), while 12 variants associate with more than one biomarker (Fig. 3). Only three variants associate with all four iron biomarkers: p.His678Arg in DUOX2 (rs57659670[C]; MAF = 7.53%), p.Cys282Tyr in HFE (rs1800562[A]; MAF = 6.77%), and p.Val749Ala in TMPRSS6 (rs855791[A]; MAF = 43.1%). The missense variant at DUOX2, a dual oxidase involved in the generation of H2O210, has not previously been associated with iron homeostasis.
We calculated the correlation between the iron biomarkers and selected other biomarkers related to iron metabolism (red blood cell indices, platelet count, erythrocyte sedimentation rate, and C-reactive protein) (Supplementary Fig. 1) and the genetic correlation between the four iron biomarkers (Supplementary Data 4). Among the iron homeostasis biomarkers, the strongest correlation was between iron and TSAT (0.86) and the strongest genetic correlation was also between these biomarkers (Iceland TSAT vs. UK iron: 0.53 (SE = 0.19), P = 0.0059; Iceland iron vs. UK TSAT: 0.54 (SE = 0.17), P = 0.0020) (Supplementary Data 4). Furthermore, we estimated the heritability of the iron homeostasis biomarkers to be between 0.16 and 0.32 using parent–offspring and sibling correlations, suggesting that heritability explains 16–32% of the variance of the four iron homeostasis markers studied (Supplementary Data 5).
We tested for heterogeneity between the results from the Icelandic, UK, and Danish cohorts (Supplementary Data 2). Of the 87 associations, 79 are with markers present in two or more populations and of these, 19 show nominally significant heterogeneity (P < 0.05). For all associations, the effects are in the same direction in all three populations, and for 79 of the 87 associations, effects are nominally significant (P < 0.05) in all three populations (Supplementary Fig. 2, Supplementary Fig. 3). Eight associations are reported with five rare variants at three loci found only in Iceland (MAF = 0.12–0.47%): three coding variants (two missense, one stop-gained) in STAB1, a stop-gained variant in TF, and a stop-gained variant in TMPRSS6. Common variants associating with iron biomarkers are reported in all three populations for each of these loci, providing additional evidence for these associations (Supplementary Data 2).
Because of the well-known difference in iron homeostasis between the sexes11, we tested for sexual dimorphism in iron biomarker associations (Supplementary Data 6). We found differences in the ferritin effect (using a test for heterogeneity with P value threshold P < 0.05/62 = 8.1 × 10−4) of five of the 62 variants. In addition, we identified one additional variant that only associates with ferritin in women: a missense variant in VWF (p.Tyr1584Cys/rs1800386[C]), a likely pathogenic type 2 von Willebrand disease (VWD) mutation12 (β = −0.17 standard deviation (SD) [−0.23, −0.12], P = 3.0 × 10−10). Of the six variants, four have greater effects in women (F5: six times greater effect, SLC25A37: three times greater effect, DUOX2: 36% greater effect, and VWF: 13 times greater effect) and two in men (HK1: four times greater effect, HFE p.Cys282Tyr: 51% greater effect) (Supplementary Data 6). The four variants with larger effects in women also have stronger effects in premenopausal than postmenopausal women (Supplementary Data 7). In addition, we find sex differences in variants in the well-known iron regulatory genes HFE (ferritin, iron, and TSAT) and TMPRSS6 (iron) (Supplementary Data 6), again with stronger effects in pre- vs. postmenopausal women (Supplementary Data 7). For the variants at F5, SLC25A37, DUOX2, and VWF that show a greater effect on women, the difference does not persist when comparing only men and postmenopausal women (Supplementary Data 8).
Iron homeostasis variants and protein quantitative loci (pQTL)
To gain further insight into the biological pathways involved in iron homeostasis, we tested for association of the 62 iron homeostasis variants (including all variants with r2 ≥ 0.8 with any iron homeostasis variants) with an expression of 4792 proteins in serum using the SomaLogic Somascan platform based on samples from 35,559 Icelanders (Methods, Supplementary Data 9). Among the 62 variants, 30 have at least one associated pQTL, where we use r2 > 0.8 as the limit for considering variants as associated. The variants at ABO, SERPINA1, FUT2, ABCA5, GCKR, and ASGR1 each have over 50 pQTL, with 24 other variants have between one and 14 pQTL. Interestingly, variants at or close to HFE (rs55925606[G], β = −0.153 SD [−0.186, −0.120], P = 2.8 × 10−20), MTMR4 (rs34523089[T], β = 0.062 SD [0.040, 0.084], P = 2.3 × 10−8) and TMPRSS6 (rs885791[A], β = 0.060 SD [0.044, 0.076], P = 9.5 × 10−13) all associate with protein levels of hepcidin. The variant at TMPRSS6 also associates with increased protein levels of erythropoietin (β = 0.066 SD [0.050, 0.082], P = 1.6 × 10−15) and transferrin receptor protein 1 (β = 0.127 [0.111, 0.143], P = 1.7 × 10−52). Furthermore, variants at LEPR and IL6R associate with decreased levels of the inflammatory mediator’s serum amyloid A-1 and A-2 proteins, with the variant at LEPR also associating with reduced levels of C-reactive protein. The rs762752083[T] stop-gained variant at STAB1 associates with increased levels of von Willebrand factor (β = 0.510 SD [0.347, 0.673], P = 7.9 × 10−10). Finally, the rs199138[A] intron variant at DUOX2 associates with decreased levels of ferritin light chain (β = −0.121 SD [−0.150, −0.092], P = 3.2 × 10−16). This variant is in strong LD (r2 = 0.97) with the DUOX2 His678Arg missense variant found to associate with a decrease in serum ferritin.
The loci in the context of systemic iron homeostasis
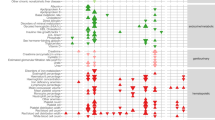
Based on a literature review, we placed 24 of the 56 candidate genes, as well as the female-specific candidate gene VWF, into 6 categories representing the main physiological processes involved in iron homeostasis: hepcidin regulation and iron storage (FTL, HAMP, HFE, TMPRSS6, TFR2, TFRC, TF, MTMR4, and SERPINA1), inflammation (IL6R, NOD1, and IKZF1), gut absorption (SLC11A2, SLC40A1, EGLN3, and DUOX2), iron recycling (SLC11A2, SLC40A1, STAB1, TRIB1, and MAFB), erythropoiesis (ERFE, SLC25A37, MYB, and HK1) and bleeding/menstruation (F5 and VWF) (Fig. 4).
Novel loci are in bold font. *Gene with a predicted probability of being causal (based on a variant-to-gene algorithm, see “Methods”) larger than 50%. The liver, blood spot, and erythropoiesis/hematopoiesis cartoons were bought from Shutterstock (standard license), the macrophage is from Wikimedia Commons (https://commons.wikimedia.org/wiki/File:Macrophage.svg).
Hepcidin regulation and iron storage: Synthesis of the iron homeostasis hormone hepcidin (HAMP) is under tight regulation by the liver iron sensing and signaling cascade involving several proteins, including those encoded by HFE, TMPRSS6, TF, TFR2, and TFRC13. Hepcidin as the major iron homeostasis hormone regulates iron transport from cells through inhibition (and degradation) of ferroportin in cells, such as intestinal epithelial and liver cells and macrophages13. HAMP, HFE, TMPRSS6, TF, TFR2, and TFRC along with the iron storage protein ferritin light chain (encoded by FTL) all have variants associated with iron biomarkers. Furthermore, the MTMR4 variant rs34523089 (MAF = 14.1%), associates with ferritin (β = 0.069 SD [0.059, 0.078], P = 3.2 × 10−48). MTMR4 has been shown to localize to early endosomes where it interacts with and dephosphorylates activated R-Smads, thus negatively regulating transforming growth factor β (TGFβ) signaling14 and TGFβ1 has been shown to activate hepcidin mRNA expression15. The MTMR4 variant also associates with hepcidin protein levels in our pQTL study, similar to what was seen with variants in the known hepcidin regulators, TMPRSS6, and HFE (Supplementary Data 9). The SERPINA1 p.Glu288Val variant (rs17580[A], MAF = 3.79%) associates with increased TIBC (β = 0.076 SD [0.053, 0.099], P = 1.2 × 10−10). SERPINA1 encodes the protease inhibitor (PI) alpha-1-antitrypsin (A1AT) and the p.Glu288Val variant—also known as the PI S allele—is associated with A1AT-deficiency (A1ATD)16. Liver disease in A1ATD has been linked to liver iron overload17, and recently A1AT was shown to increase hepcidin expression through proteolytic cleavage and inhibition of TMPRSS618.
Inflammation: IL6 and its receptor IL6R are important inflammatory mediators positively regulating liver hepcidin during inflammation19,20,21. The IL6R p.Asp358Ala variant (rs2228145[C], MAF = 41%) that associates with decreased risk of rheumatoid arthritis22,23 associates with an increase in serum iron (β = 0.026 SD [0.018, 0.034], P = 8.4 × 10−11). Leptin and its receptor LEPR, in addition to its central role as an adipokine, have been shown to control cellular immune responses in several pathological situations including rheumatic diseases24. The intergenic variant rs35945185[A] (MAF = 36.5%) linked to LEPR associates with iron (β = 0.031 SD [0.023, 0.039], P = 1.54 × 10−13. The IL6R and LEPR associated variants (rs2228145[C], rs35945185[A]) both are negatively associated with inflammatory markers serum amyloid A-1/A-2 proteins, and furthermore, the LEPR associated variant is also negatively associated with C-reactive protein (Supplementary Data 9). Also, the rs2529440[T] intron variant (MAF = 45%) at NOD1, encoding an intracellular innate immune pattern recognition sensor for bacterial cell components25, associates with a reduction in ferritin levels (β = −0.035 SD [−0.014, −0.028], P = 4.6 × 10−23). Furthermore, the rs12718598[C] intron variant in IKZF1, encoding the lymphocyte specification and differentiation transcription factor Ikaros, shown to play a role in auto-immune diseases26, associates with increased serum iron levels (β = 0.027 SD [0.019, 0.034], P = 3.7 × 10−11).
Gut absorption: Iron absorption is mediated by the two iron transporters DMT1 (encoded by SLC11A2) at the luminal side and ferroportin (encoded by SLC40A1) basolaterally, both regulated by hepcidin signals and both harboring variants associated with iron homeostasis biomarkers13. Recently, hepcidin blocking of intestinal ferroportin was shown to inhibit HIF-2α expression, through increased intracellular iron and subsequent activation of iron-dependent prolyl hydroxylases, leading to reduced expression of iron absorptive proteins27. Mammalial HIF-α prolyl hydroxylases are encoded by the three genes EGLN1-328. The rs996347[C] intron variant (MAF = 35%) at EGLN3 associates with increased ferritin (β = 0.049 SD [0.042, 0.056], P = 3.0 × 10−41). EGLN3 is a likely candidate to mediate the inhibition of intestinal HIF2α expression, as it specifically inhibits HIF-2α rather than HIF-1α 29,30. The DUOX2 p.His678Arg variant (rs57659670[C], MAF = 7.5%) associates with reduced ferritin (β = −0.14 SD [−0.16, −0.13], P = 1.1 × 10−113), serum iron (β = −0.042 SD [−0.056, −0.028], P = 1.1 × 10−8), and TSAT (β = −0.058 SD [−0.074, −0.041], P = 5.7 × 10−12) and increased TIBC (β = 0.077 SD [0.060, 0.094], P = 3.7 × 10−19). DUOX2 is expressed in the upper intestinal mucosa and may play a role in innate mucosal immunity10,31. Furthermore, in mouse models, DUOX1 and DUOX2 knockouts have a greater susceptibility to Helicobacter felis infection and inflammation32 and epidemiological studies have indicated that H. pylori infections in humans are associated with reduced iron stores33.
Iron recycling: recycling of heme–iron takes place in the reticuloendothelial system in the spleen and liver, where old red cells are taken up and iron recycled back to the bone marrow, providing over 90% of the iron needed for the generation of heme in red cell precursors1. DMT1 and ferroportin also transport iron from endocytic vesicles and export iron out of the macrophage, respectively34. Furthermore, three uncorrelated rare variants (MAF < 1%) in STAB1 (p.Glu117Ter/rs762752083[T], p.Gly189Ser/rs750717575[A] and p.Glu527Lys/rs745795585[A]) and a variant in LD with a STAB1 variant (GNL3 p.Ser451Thr/rs34216132[C], r2 > 0.99 with the STAB1 variant p.Ser1089Gly/rs41292856[G]) (Supplementary Fig. 4) all associate with increased ferritin, with effects ranging from 0.17 to 0.35 SD (P = 2.2 × 10−8 to 2.6 × 10−19). STAB1 is primarily expressed in M2-macrophages and sinusoidal endothelial cells35 and has been shown to affect phosphatidylserine-mediated uptake of aged red blood cells36,37. We also report associations of the intergenic variants rs2954029[T] (MAF = 48%) and rs6029148[A] (MAF = 7.1%) with reduced and increased ferritin (rs2954029[T]: β = −0.024 SD [−0.031, −0.018], P = 1.4 × 10−12; rs6029148[A]: β = 0.046 SD [0.033, 0.058], P = 5.6 × 10−12). Their closest protein-coding genes, TRIB1 (for rs2954029) and MAFB (for rs6029148), have both been shown to control the differentiation of macrophages38,39.
Erythropoiesis: The bone marrow relays signals inhibiting liver hepcidin synthesis under a state of stress erythropoiesis to make iron available to erythroid precursors40. Variants located close to two known iron regulators within the erythropoiesis compartment, the intergenic variant rs13253974[A] (MAF = 32%) near SLC25A37 (mitoferrin-1)41 and the intron variant rs13007705[T] at ERFE (erythroferrone)40,42 associate with increased ferritin (rs13253974[A]: β = 0.024 SD [0.017, 0.032], P = 2.5 × 10−11), serum iron (rs13007705[T]: β = 0.029 SD [0.021, 0.037], P = 2.0 × 10−12) and TSAT (rs13007705[T]: β = 0.033 SD [0.024, 0.042], P = 1.1 × 10−12). In addition, the rs9399136[C] variant in the HBS1L-MYB intergenic region associates with increased serum iron (β = 0.057 SD [0.049, 0.066], P = 1.1 × 10−36) and TSAT (β = 0.067 SD [0.057, 0.077], P = 5.3 × 10−39) and reduced TIBC (β = −0.033 SD [−0.044, −0.023], P = 2.7 × 10−10), while the rs17476364[C] intron variant in HK1 associates with increased ferritin (β = 0.043 SD [0.032, 0.054], P = 3.6 × 10−14). Variants in the HBS1L-MYB intergenic region are known to associate strongly with fetal hemoglobin levels43,44. Fetal hemoglobin levels are induced during stress erythropoiesis45,46, a condition also involving ERFE signaling40. Furthermore, HK1 mutations are associated with reduced red cell survival47.
Bleeding/menstruation: Loss of iron occurs primarily through epithelial desquamation and blood loss48. Two iron homeostasis variants show sexual dimorphism: The F5 p.Arg534Gln variant (rs6025[T], MAF = 2.8%; also known as factor V Leiden) associates with increased ferritin (β = 0.15 SD [0.12, 0.17], P = 6.9 × 10−37) and reduced TIBC (β = −0.093 SD [−0.13, −0.061], P = 1.8 × 10−08) and has a six times stronger effect on ferritin in females than in males, while the VWF p.Tyr1584Cys variant (rs1800386[C], MAF = 0.94%) associates with reduced ferritin in females only (β = −0.17 SD [−0.23, −0.12], P = 3.0 × 10−10). The factor V Leiden variant is associated with a hypercoagulable state49 and the VWF p.Arg534Gln variant is associated with type 2 VWD, a common inherited bleeding disorder12. Since both variants show stronger effects in premenopausal than postmenopausal women, and both variants affect clotting, they are likely working through blood loss and primarily menstrual bleeding. In addition, in a meta-analysis using data from Iceland, Denmark, and the UK, the factor V Leiden variant is protective against menorrhagia (OR = 0.82 SD [0.76–0.88], P = 2.1 × 10−7). To further address the effects of variants affecting bleeding and thrombosis we carried out a candidate gene analysis for association with iron homeostasis markers. We screened for coding variants in 375 genes associated with abnormal bleeding (HP:0001892) and 76 genes associated with venous thrombosis (HP:0004936) listed in Human Phenotype Ontology (https://hpo.jax.org/). Nineteen new variants in 14 genes associated with at least one iron homeostasis marker (Supplementary Data 10, Bonferroni corrected P value threshold of 1.7 × 10−5, ~3000 variants). Only one of these variants in ARHGAP31 showed sexual dimorphism (effect on ferritin; women (β = 0.230, P = 8.4 × 10−8), men (β = −0.044 SD, P = 0.389), Phet = 4.0 × 10−5) (Supplementary Data 10). One additional missense variant in IRF2BP2 associated with iron in premenopausal women only (premenopausal women (β = −0.149 SD, P = 4.63 × 10−5), postmenopausal (β = 0.091 SD, P = 0.036), Phet = 2.40 × 10−5) (Supplementary Data 10).
Iron homeostasis variants and red blood cell traits
To better understand the effect of the sequence variants on iron homeostasis and iron usage, we tested for association with the red blood cell indices hemoglobin (N = 286,622), mean corpuscular hemoglobin (N = 286,245), mean corpuscular volume (N = 286,248), and reticulocyte count (N = 19,031) and compared the effects of variants on them and the four iron biomarkers (Supplementary Fig. 5, Supplementary Data 11). Normally, as body iron stores fall, the hemoglobin concentration, mean corpuscular volume, and mean corpuscular hemoglobin concentration also fall. The p.Cys282Tyr variant at HFE (rs1800562) strongly affects all iron and red blood cell biomarkers except reticulocyte count. Variants at DUOX2, F5, and TRIB1 have a similar pattern of effects on iron and red blood cell biomarkers, with a negative effect on TIBC and mainly positive effects on the red cell indices (Supplementary Fig. 5). The variant showing the strongest effect on ferritin is a stop-gained variant in STAB1 (Stabilin-1) (β = 0.35 SD [0.26, 0.44], P = 3.2 × 10−14) (Supplementary Data 2). This variant also shows an unusual pattern with decreased hemoglobin along with increased ferritin, indicating that body stores of iron are sufficient but the recycling of iron from stores is abnormally reduced (Supplementary Fig. 5).
IDA and iron overload
The two extremes of iron homeostasis, iron deficiency, and iron overload, are clinically important and associated with high disease burden4,50. In iron deficiency, depletion of iron stores is followed by reduced iron availability for erythropoiesis, leading to IDA, presenting as hypochromic, microcytic anemia with low ferritin and/or low TSAT48. Increased TSAT, most commonly defined as a saturation above 50%, is used as a screening marker for hemochromatosis and iron overload51. To understand how the 62 iron homeostasis variants affect either IDA or iron overload, we tested for association with IDA (defined as ever simultaneously having hemoglobin < 120 g/L for women, <130 g/L for men, MCV < 80 fl, MCH < 27 pg and either ferritin < 10 mcg/L or TSAT < 16%; Ncases = 6476, Ncontrols = 362,706)5 and iron overload (defined as TSAT ever >50%4, Ncases = 4156, Ncontrols = 342,647) (Fig. 5, Supplementary Data 12), correcting for 2 × 62 = 124 performed tests. The missense variants in DUOX2 (p.His678Arg; rs57659670[C]) and F5 (p.Arg534Gln, rs6025[T]) associate with IDA (DUOX2 p.His678Arg: OR = 1.29 [1.20–1.39], P = 2.0 × 10−11; F5 p.Arg534Gln: OR = 0.60 [0.49–0.73]; P = 3.4 × 10−7). The variants showing sexual dimorphism for the effect on ferritin also showed similar trends with regard to IDA (Supplementary Fig. 6, Supplementary Data 13). In addition, a 3.55 kb deletion in the SLC11A2 3′ untranslated region (3′ UTR) and its downstream intron associates with IDA through a recessive mode of inheritance (OR = 32.5 [10.0–105]; P = 6.4 × 10−9) (Fig. 5, Supplementary Data 12, Supplementary Fig. 7). A rare frameshift mutation in TMPRSS6 (p.Asn473ThrfsTer63, rs773570300) only detected in the Icelandic cohort (MAF = 0.16%) also associated with IDA (OR = 3.0 [2.1–4.4]; P = 1.2 × 10−8). The rs9399136[C] variant in the intergenic HBS1L/MYB region is the only variant to associate with both IDA (OR = 0.84 [0.80–0.89], P = 4.7 × 10−11) and iron overload (OR = 1.13 [1.07–1.20], P = 1.4 × 10−5). This variant has not been associated with iron homeostasis but has been associated with hematological traits52 and variants in the same region have been associated with fetal hemoglobin expression43,53. Additionally, variants in the iron homeostasis regulatory genes HFE, TMPRSS6, TF, and TFR2 associate with iron overload (Fig. 5, Supplementary Data 12).
A forest plot showing the odds ratio (error bars showing 95% confidence intervals) for each of the genetic variants associated with either iron deficiency anemia (IDA) or iron overload (IO). *Results for the SLC11A2 deletion variant are shown for the recessive model while results for other variants are for the additive model. Min/Maj minor/major allele, MAF minor allele frequency, OR odds ratio, CI confidence interval.
We tested the 62 iron homeostasis variants for association with the following eleven clinical manifestations of iron overload and/or iron deficiency54 based on various meta-analyses performed in Iceland using data from Iceland, UK, Denmark, and the USA: hemochromatosis, liver fibrosis/cirrhosis, liver cancer, type 2 diabetes, impotence, cardiomyopathy, osteoporosis, osteoarthritis, hyperpigmentation, amenorrhea, and restless leg syndrome (Supplementary Data 14). Taking all 62 × 11 = 682 tests into account using Bonferroni correction, the TMPRSS6 p.Val749Ala variant (rs855791[A]) associates with less risk of hemochromatosis (Ncases = 719, Ncontrols = 497,001; OR = 0.80 [0.72–0.89], P = 6.1 × 10−5). The HFE p.Cys282Tyr variant (rs1800562[A]), the main variant associating with recessive hereditary hemochromatosis (type 1) associates with a higher risk of hemochromatosis (additive model: OR = 25.7 [21.6–30.5], P < 10−300; recessive model: OR = 218.9 [164.6–291.0], P < 10−300), liver fibrosis/cirrhosis (Ncases = 1043, Ncontrols = 705,646; additive model: OR = 1.50 [1.30–1.74], P = 4.5 × 10−8; recessive model: OR = 5.54 [3.58–8.58], P = 1.54 × 10−14) and liver cancer (Ncases = 844, Ncontrols = 792,550; additive model: OR = 1.53 [1.28–1.82], P = 2.3 × 10−6; recessive model: OR = 8.19 [5.04–13.29], P = 1.8 × 10−17), consistent with previous reports3. Furthermore, the GCKR p.Leu446Pro variant (rs1260326[T]) associates with a lower risk of type 2 diabetes (Ncases = 36,710, Ncontrols = 663,962; OR = 0.94 [0.93–0.96], P = 1.4 × 10−10) (Supplementary Data 14). We also generated polygenic risk scores (PRS) for ferritin and TSAT and regressed the scores against the same eleven clinical manifestations of iron overload and/or iron deficiency (Supplementary Data 15). The PRS for ferritin and TSAT only associated with hemochromatosis (ferritin OR = 2.71 [2.49–2.95], P = 9.4 × 10−119; TSAT OR = 3.75 [3.51–4.00], P < 1 × 10−300). Restless leg syndrome has repeatedly been associated with iron deficiency55,56 and iron supplementation recommended in select cases57. We confirm that in the Icelandic restless leg syndrome cohort iron biomarkers suggest increased iron deficiency (lower ferritin (β = −0.07 SD [−0.12, −0.02] SD; P = 0.0037) and TSAT (β = −0.06 SD [−0.12, −0.001], P = 0.045) and higher TIBC (β = 0.14 SD [0.08, 0.20] SD; P = 8.5 × 10−6) and there is increased incidence of IDA compared to population controls (OR = 1.39 [1.03–1.84], P = 0.0244) (Supplementary Data 16). The lack of genetic association seen with either individual iron homeostasis variants or PRS argues against a simple causal relationship between iron deficiency and restless leg syndrome.
Novel SLC11A2 deletion variant
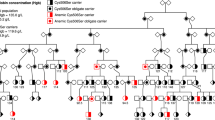
Rare loss-of-function mutations in SLC11A2 (solute carrier family 11 member 2 encoding DMT1, divalent metal transporter 1) have been associated with a microcytic anemia with iron overload under the recessive mode of inheritance58,59,60 demonstrating a role of DMT1 in both iron absorption and recycling. We identified 14 homozygous carriers of the abovementioned deletion in SLC11A2 in the Icelandic cohort, seven of whom had been diagnosed with IDA (microcytic anemia with low ferritin and/or low TSAT) and one with transfusion-dependent anemia; two had required transfusions and one intravenous iron (Supplementary Data 17).
Transcription of SLC11A2 leads to four major mRNAs with differing tissue-specific expression patterns61. These messages differ both in their usage of 5′ exons 1a or 1b and usage of alternative 3′ translated and untranslated regions (UTRs) (Fig. 6A). These alternative UTRs differ in that one contains an iron-response element (IRE), denoted IRE+, while the other UTR lacks such a motif, denoted IRE−. The IRE+ UTR is primarily expressed in duodenal and kidney epithelium, mediates iron absorption, and is regulated directly by cellular iron status through interaction with IRE-binding proteins62,63. Of the four highest expressed transcripts in blood, two contain the IRE− UTRs, and two contain the IRE+ UTRs. The SLC11A2 deletion extends from within the IRE+ containing 3′ UTR and into the downstream intron (Fig. 6A). Heterozygotes (N = 251) and homozygotes (N = 2) express 40% (95% CI: 58–62%, P = 2.2 × 10−16) and 81% (95% CI: 73–87%, P = 0.015) less IRE+ transcripts than wildtype (N = 12,828), respectively (Fig. 6B). When comparing allele-specific transcription in heterozygotes there was a 3.7-fold (P = 2.2 × 10−17) preference for wildtype allele in IRE+ containing alleles. The deletion removes the native 3′ UTR polyadenylation signal, likely resulting in an unstable mRNA. The IRE− transcripts are expressed at 29% greater levels by heterozygotes than noncarriers (95% CI: 26–33%, P = 2.2 × 10−16) and at 133% greater levels by homozygotes (95% CI: 48–205%; P = 0.016). These data suggest that the SLC11A2 deletion causes isoform-specific effects, suppressing the expression of IRE+ containing transcripts, that are primarily expressed in absorptive duodenal and kidney epithelium62 leading to reduced absorption. This leads to a recessive hereditary IDA. Hepcidin levels based on proteomics samples from 35,559 Icelanders are reduced in SLC11A2 deletion carriers (β = −0.172 SD [−0.257, −0.088], P = 5.9 × 10−5), consistent with systemic iron deficiency64. A single homozygous carrier of the deletion has a hepcidin value 2.17 SDs below average. The only previously described genetic IDA, iron refractory IDA, due to homozygous loss-of-function variants in TMPRSS6, is associated with hepcidin dysregulation and inappropriately high hepcidin values64.
A Coverage plot of RNA-sequenced reads from whole blood showing the median normalized expression at the 3′ UTR end of SLC11A2 in wild-type (AA, green; n = 12,828) heterozygous (ADel, blue; n = 251) and homozygous (DelDel, red; n = 2) individuals for the different SLC11A2 deletion genotypes. B Comparison of expression levels of the four major SLC11A2 transcripts (two transcripts without iron response elements (IRE) in their 3′ UTR (IRE-) and two with IRE (IRE+) in their 3′ UTR in whole blood RNA using a mixed violin- and boxplot. The violin plot shows the density where the width represents the frequency of the log2 normalized expression levels. The white boxes show the distribution statistics (interquartile range and median) and the whiskers represent ±1.5× the interquartile range. The filled circles correspond to individual expression values or outliers that lie beyond the extremes of the whiskers.
Discussion
Through a GWAS meta-analysis of the iron homeostasis biomarkers ferritin, serum iron, iron-binding capacity, and TSAT in Iceland, Denmark, and the UK, we have identified 56 loci harboring variants associating with one or more of these biomarkers, 46 of which are novel (including six rare variants, six low-frequency variants, and 37 common variants). Among the novel loci, variants in DUOX2 and SLC11A2 associate with increased risk of IDA, while the F5 rs6025[T] variant protects against IDA. Furthermore, the rs9399136[C] variant at the HBS1L/MYB locus is protective against IDA while increasing the risk of iron overload.
While most of these iron homeostasis variants show similar effects in Iceland, UK, and Denmark, the observed heterogeneity for a subset of the variants may reflect demographic, clinical, and environmental differences. In clinical populations, iron homeostasis markers are more frequently measured in individuals with suspected iron deficiency or overload. Blood donors, on the other hand, are screened for anemia and several other diseases at every donation. Therefore, people with a previous history of iron deficiencies are underrepresented in blood donor studies, although a substantial proportion of blood donors will develop low or reduced iron stores65. These differences in cohort characteristics could partially explain heterogeneity in effect sizes between populations for a subset of the variants. The sex-specific heterogeneity reported highlights the differences between the sexes in iron homeostasis. The two sequence variants showing the strongest heterogeneity are both variants in coagulation factors, the well-known factor V-Leiden mutation49 and a mutation in VWF known to cause type 2 VWD12. These variants are likely to mediate their effects through increased (VWF) or decreased (F5) blood loss, in women being mainly mediated through menstruation, supported by the finding that factor V Leiden variant protects against menorrhagia. Thirty-four of the 62 reported novel variants only associate with ferritin. Possible explanations of this high number could be that ferritin as a biomarker is affected by a more broad array of processes, such as inflammation and tissue damage (e.g., liver injury)66, and also that we have more ferritin measurements (~246 K vs. ~131–163 K).
Iron deficiency is a major global health problem, especially for children and women2. A worldwide survey in 2010 showed that one-third of the world population is anemic with iron deficiency being responsible for approximately half of that cases50. In addition to the nonspecific symptoms of IDA, it also may contribute globally to reduced cognitive performance in children67, adverse outcomes of pregnancies68, and decline in cognition in the elderly48,69. Despite the importance of iron deficiency and IDA, no systematic genetic studies looking at iron deficiency or IDA have been performed. Sequence variants that are common (at DUOX2 and the HBS1L-MYB intergenic region), low-frequency (at F5), and rare (at TMPRSS6 and SLC11A2) associate with IDA (Fig. 5, Table 1). The association of the missense DUOX2 variant with all iron homeostasis markers, as well as with IDA is striking. That this association was seen in all three populations studied but not observed in previous GWAS of iron homeostasis is intriguing, however, it should be noted that Benyamin et al.6 reported a subgenome-wide significant association with ferritin near this locus (rs16976620). Our study is significantly larger and also benefits from more comprehensive imputation panels made available since then, which likely enabled us to not only detect an association at genome-wide significance but also map this to the likely causal gene with high confidence.
The phenotype of recessive IDA with low iron stores that we report with the rare 3.5 kb deletion within SLC11A2 is different from the previously reported recessive hypochromic anemia with iron overload associated with this gene55,56,57. Further studies to define the pathways mediating the effects of the variants associating with IDA could help shed light on the pathophysiology of iron deficiency. Notably, neither any individual iron homeostasis variants nor the PRS for ferritin or TSAT associate with the risk of restless leg syndrome, a neurological disorder suggested being exacerbated by iron deficiency70. Although this argues against a simple causal relationship between the two, a more complex relationship, e.g., through brain iron concentations71 cannot be ruled out. Even though hereditary hemochromatosis is most often associated with HFE p.Cys282Tyr homozygosity, the penetrance is only 28% in males and much lower in females72. The common missense variant in TMPRSS6 (rs855791[A], MAF = 43.1%) protects against hereditary hemochromatosis (OR = 0.80 [0.72–0.89], P = 6.1 × 10−5) and could thus be a modifying gene in this disease.
In summary, we have identified 46 novel loci affecting iron homeostasis. Many of the novel candidate genes have roles in homeostasis through mechanisms, such as absorption, iron recycling, erythropoiesis, and hepcidin regulation. Furthermore, we show an association of five of these loci with IDA, a major clinical entity that hitherto has not been studied thoroughly from a genetic point of view. This study reveals a substantial catalog of possible iron regulatory genes, awaiting further inquiry to fully elucidate their functional role.
Methods
Study subjects from Iceland
The Icelandic data (where around one-half of all individuals had repeated measurements) include the vast majority of all clinical laboratory results in Iceland from 1990 to 2017. Serum iron and TIBC were measured with colorimetric methods and serum ferritin was measured with an electrochemiluminescence immunoassay using reagents and calibrators and Cobas 6000 and 8000 modular instruments from Roche Diagnostics, Mannheim, Germany. Hemoglobin concentration measurements, as well as other basic hematology parameters used, were measured on EDTA anticoagulated blood using the Sysmex XN-1000 hematology analyzer.
All participants who donated samples gave informed consent and the National Bioethics Committee of Iceland approved the study (VSN-15-198) which was conducted in agreement with conditions issued by the Data Protection Authority of Iceland. Personal identities of the participant’s data and biological samples were encrypted by a third-party system (Identity Protection System), approved, and monitored by the Data Protection Authority.
Study subjects from the UK
The INTERVAL study is a prospective cohort study of approximately 45,000 blood donors, representative of the wider donor population, nested in a randomized control trial. Participants, aged 18 years or older, were recruited between 2012 and 2014 from 25 National Health Service Blood and Transplant static donor centers in England. All participants provided written, informed consent, and the study was approved by the Cambridge (East) Research Ethics Committee (ref: 11/EE/0538).
Ferritin measurement was based on the immunological agglutination principle with the enhancement of the reaction by latex. Latex particles coated with anti-ferritin antibodies agglutinated with ferritin and the precipitate was determined turbidimetrically at 570/800 nm. Serum iron was measured using a colorimetric method (FerroZine) without deproteinization. Under acidic conditions, iron was liberated from transferrin. Acrobat reduced Fe3+ to Fe2+ which then reacted with FerroZine to form a colored complex. The color intensity is directly proportional to the iron concentration and was measured photometrically. TIBC was calculated by summing up serum iron and unsaturated iron-binding capacity, which was also measured photometrically. TSAT was calculated by dividing serum iron by TIBC concentration. All data points lying more than 4.5 interquartile range from the median were considered outliers and removed (591 for ferritin, 7 for transferrin, 65 for TSAT, and 37 for serum iron).
The genotyping protocol and quality control procedures for INTERVAL study samples have been described in detail previously73. Briefly, DNA extracted from buffy coat was used to assay approximately 820,000 variants and short insertions/deletions on the Affymetrix Axiom «Biobank» genotyping array (Affymetrix, Santa Clara, California, US). Genotyping was performed in multiple batches of approximately 4800 samples each. Sample QC was performed including exclusions for sex mismatches, low call rates, duplicate samples, extreme heterozygosity, and non-European descent. We carried out high-resolution multiple imputations using a joint UK10K and 1,000 Genomes Phase 3 (May 2013 release) reference panel and retained variants with a MAF ≥ 0.1% and/or INFO score ≥0.4 for analysis.
The meta-analyses of hemochromatosis, liver fibrosis/cirrhosis, liver cancer, type 2 diabetes, osteoarthritis, impotence, cardiomyopathy, osteoporosis, hyperpigmentation, and amenorrhea (Supplementary Data 14) include data from the UK Biobank, accessed under Application Number 56270.
Study subjects from Denmark
The Danish Blood Donor Study (DBDS), initiated in 2010 as collaborative blood donor-oriented and generic research platform and is an on-going nation-wide prospective cohort with inclusion sites at all Danish blood collection facilities. Currently, more than 110,000 blood donors are participating, and more than 95% of invited blood donors are willing to participate74. Due to the step-wise roll-out of DBDS, an enrichment of individuals from the greater Copenhagen region (the capital) and the central region of Jutland (the second largest city) are present in this study. DBDS has secured necessary permissions and approval from the Danish Data Protection Agency (2007-58-0015) and the Scientific Ethical Committee system (M-20090237). Briefly, regarding the DBDS genomic cohort DNA is purified from whole blood and subsequently stored at −20 degrees Celsius. DBDs participants in this study has been genotyped in 1 batch at Decode genetics using the Global Screening Array by Illumina optimized for comparison with the Illumina Omni Express chip75. Ferritin was measured on fresh EDTA-anticoagulated plasma samples using two commercially available assays: for 30,903 individuals using Ortho Vitros 5600 (Ortho Clinical Diagnostics, Rochester, NY, USA), and for 2851 individuals using Abbott Architect i2000SR (Abbott Laboratories, Abbott Park, IL, USA), including 27 individuals that had measurements taken using both methods.
Whole-genome sequencing
The process used to whole-genome sequence the 28,075 Icelanders, as well as the subsequent imputation, has been described in recent publications76,77. In summary, we sequenced the whole genomes of 28,075 Icelanders using Illumina technology to a mean depth of at least 10× (median 32×). Single-nucleotide polymorphism (SNPs) and indels were identified and their genotypes called using joint calling with Graphtyper78. In total, 155,250 Icelanders were genotyped using Illumina SNP chips and their genotypes were phased using long-range phasing79. All sequenced individuals were also chip-typed and long-range phased, providing information about haplotype sharing that was subsequently used to improve genotype calls. Genotypes of the 32 million high-quality sequence variants were imputed into all chip-typed Icelanders. Variants in the Icelandic and Danish cohorts were imputed based on the IMPUTE HMM model80 as previously described81. Variants in INTERVAL were imputed using the Sanger Imputation Server (https://imputation.sanger.ac.uk) which implements the Burrows–Wheeler transform imputation algorithm PBWT on whole chromosomes. A combined UK10K and the 1000 Genomes Phase 3 reference panel was used82. Using genealogic information, the sequence variants were also imputed into relatives of the chip-typed further increasing the sample size for association analysis and the power to detect associations. All of the variants tested had imputation information over 0.8. The GWAS from Denmark was performed using 19 million markers identified through whole-genome sequencing of 2816 Danes that were subsequently imputed into 84,386 chip-typed individuals. The GWAS from the UK was performed with 19 million markers from the UK10K and 1000 Genomes Phase 3 reference panel, imputed into 43,059 chip-typed individuals participating in the INTERVAL study. In total, 40 million markers were tested in the meta-analysis.
Association testing and meta-analysis
The four iron homeostasis biomarkers (ferritin, serum iron, TIBC, and TSAT) were each rank-based inverse normal transformed to a standard normal distribution (separately for each sex) and adjusted for age using a generalized additive model. In addition, for the UK cohort, the biomarkers were adjusted for menopausal status, ABO blood group, body mass index, smoking levels, alcohol levels, and iron supplementation status. For each sequence variant, the mixed model implemented in the software BOLT-LMM v2.383, using the genotype as an additive covariate and the transformed quantitative trait as a response, was used to test for association with quantitative traits. Logistic regression was used to test for association between variants and case-control phenotypes, using software developed at deCODE genetics76.
We used LD score regression to account for distribution inflation in the dataset due to cryptic relatedness and population stratification84. LD score regression intercepts were as follows: ferritin: 1.032 (SE = 0.011), iron: 1.016 (SE = 0.025), TIBC: 1.030 (SE = 0.039), TSAT: 1.025 (SE = 0.020). We used logistic regression to test for association between sequence variants and binary traits, regressing trait status against expected genotype count. In the Icelandic data, we adjusted for sex, age, and county of birth by including these variables in the logistic regression model. In the UK and Danish data we adjusted for sex and age, as well as principal components in order to adjust for population stratification.
Results from the Icelandic, UK, and Danish datasets were combined using a fixed-effect inverse-variance weighted meta-analysis, where different datasets were allowed to have different population frequencies for alleles and genotypes but assumed to have a common effect. Heterogeneity in effect estimates was assessed using a likelihood ratio test. Effects are always given in units of SD. The pooled SD using data from Iceland, UK, and Denmark are 1.08 μg/L for log(ferritin), 7.76 μmol/L for iron, 14.14 μmol/L for TIBC, and 13.25% for TSAT.
We accounted for multiple testing by means of a weighted Bonferroni correction, taking into account the higher prior probability of association of certain variant annotations while controlling the family wise error rate (FWER) at 0.059. The method has been described previously9 and results in stricter multiple testing correction than the commonly used threshold of 5 × 10−8 (which would not control FWER at 0.05 given that 40 million markers were tested) while being more powerful than simply correcting for 40 million tests using a fixed threshold of 0.05/40,000,000 = 1.25 × 10−9. The resulting significance thresholds were 2.0 × 10−7 for high-impact variants (including stop-gained, frameshift, splice-acceptor, or splice-donor variants, N = 11,723), 4.0 × 10−8 for “moderate-impact” variants (including missense, splice-region variants and in-frame indels, N = 202,336), 3.7 × 10−9 for “low-impact” variants (including upstream and downstream variants, N = 2,896,354) and 6.1 × 10−10 or for the “lowest-impact” variants (including intron and intergenic variants, N = 37,239,641).
Loci were defined based on physical proximity, where variants in a 500 kb window (lead variant ±250 kb) were considered to be at a single locus. We defined novel loci as loci not reported in previous GWAS of biomarkers of iron homeostasis.
Variant-to-gene mapping
To predict the most likely causal gene for each variant we used an algorithm taking into account the gene location with regard to LD class (defined as all variants with r2 > 0.8 with the lead variant), the variant effect for coding variants, and the effect on gene expression (eQTL, restricting to the top cis-eQTL). The algorithm, called variant-to-gene mapping, considers all genes within the LD class ±250 kb and outputs a score for each gene.
Often, the GWAS variant is not causal itself but in LD with the causal variant. To identify the likely causal gene, we defined all variants in linkage disequilibrium (r2 > 0.8) with the GWAS variant as the LD class. We assumed local effects, where genes overlapping the LD class interval receive a distance score of 5, while genes within 250 kb on each side of the LD class interval receive a distance score of 1. The variants in the LD class were then scored based on their capability to affect gene coding (i.e., transcription/translation): a variant with high impact (stop-gain and stop-loss, frameshift indel, donor and acceptor splice-site, and initiator codon variants) was given a coding score of 150, while a variant with moderate impact (missense, in-frame indel, splice region) was given a coding score of 30. For each gene, we summed up the coding scores of all coding variants affecting it, i.e., coding variants within the gene itself. Variants shown to be correlated with gene expression (eQTL) in any tissue received an eQTL score of 50. We restricted ourselves to the top cis-eQTL (lowest P value < 10−7, distance from gene < 1 Mb) for each gene and tissue. We assumed that eQTL in different tissues/different variants were due to the same signal. Therefore, we did not sum up the eQTL scores per gene but used the maximum eQTL score per gene. The total score per gene was computed as the maximum of its distance, coding, and eQTL scores. The normalized gene score was computed by scaling such that the sum of normalized scores for all candidate genes was 1, so to enable direct comparisons across genes. Note that this automatically takes the gene density into account. In cases where more than one gene share the maximum score (for example, if the LD class has four genes and they all have probability = 0.25), we chose the gene with the most significant eQTL if such information existed, otherwise the gene closest to the lead variant was selected. Relative values for the scoring for high- and moderate-impact values were based on enrichment analysis, as previously described9, while the score of 50 for eQTL was determined in order to make coding and eQTL equally informative overall. Values for proximity were set to have some degree of preference for closeby genes, given otherwise equal evidence, while at the same time giving stronger weight to coding and eQTL than to proximity alone. Data sources for eQTL data are listed in Supplementary Data 18.
Genetic overlap with other traits
We calculated genetic correlations between pairs of traits using the cross-trait LD score regression methods84 in our meta-analysis using summary statistics from traits in the Icelandic and UK datasets. We used results for about 1.2 million variants, well imputed in both datasets and for LD information we used precomputed LD scores for European populations (available from the Broad Institute).
Heritability estimation
Heritability was estimated in the following two ways: (1) 2 × parent–offspring correlation, (2) 2 × full sibling correlation, using the Icelandic data (where all family relationships are known).
Polygenic risk scores
We generated PRS for ferritin and TSAT and regressed the scores, along with sex, year of birth, and 20 principal components as covariates in logistic regression models against 11 clinical manifestations of iron deficiency or iron overload (restless legs, hemochromatosis, liver fibrosis/cirrhosis, liver cancer, type 2 diabetes, osteoarthritis, impotence, cardiomyopathy, osteoporosis, hyperpigmentation, and amenorrhea). Scores are based on a framework set of 620,000 high-quality SNPs covering the whole genome, adjusted for LD using LDpred85. The methods used to generate the PRS have been previously described in detail86. For restless legs, the phenotype data is from Iceland while for the other ten phenotypes, data is from UK Biobank (restless legs were not available in UK Biobank). To minimize bias and/or overfitting, the geographical population with the phenotype data is not included when generating the scores. Thus, for ferritin, the PRS for restless legs is based on a Denmark+UK GWAS meta-analysis, while the PRS for the other phenotypes is based on Iceland + Denmark GWAS meta-analyses. For TSAT, the PRS for restless legs is based on the UK GWAS, while the PRS for the other phenotypes is based on the Icelandic GWAS.
Protein measurements (pQTL)
During 2000–2019, we collected plasma samples from 40,004 Icelanders. Fifty-two percent of the samples were collected as part of the Icelandic Cancer Project, while the remaining samples (48%) were collected as part of various genetic programs at deCODE genetics, Reykjavík, Iceland. All samples were measured using the SOMAscan platform, containing 5284 aptamers providing a measurement of relative binding of the plasma sample to each of the aptamers in relative fluorescence units, corresponding to 4,792 proteins. After quality control, unique measurements for N = 35,559 individuals were used for genome-wide association analysis.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The Icelandic population WGS data have been deposited at the European Variant Archive under accession code PRJEB15197. The authors declare that the data supporting the findings of this study are available within the article, its Supplementary Data files, and upon request. Overall meta-analysis summary statistics have been shared at https://www.decode.com/summarydata/. The UK Biobank data can be obtained upon application (ukbiobank.ac.uk). For this study, UK-Biobank data was under project number 56270.
Code availability
We used publicly available software (URLs listed below) in conjunction with algorithms described in the Methods section: BWA 0.7.10 mem (https://github.com/lh3/bwa); GenomeAnalysisTKLite 2.3.9 (https://github.com/broadgsa/gatk/); Picard tools 1.117 (https://broadinstitute.github.io/picard/); SAMtools 1.3 (http://samtools.github.io/); Bedtools v2.25.0-76-g5e7c696z (https://github.com/arq5x/bedtools2/); Variant Effect Predictor (https://github.com/Ensembl/ensembl-vep).
References
Muckenthaler, M. U., Rivella, S., Hentze, M. W. & Galy, B. A red carpet for iron metabolism. Cell 168, 344–361 (2017).
Vos, T. et al. Global, regional, and national incidence, prevalence, and years lived with disability for 328 diseases and injuries for 195 countries, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet 390, 1211–1259 (2017).
Pilling, L. C. et al. Common conditions associated with hereditary haemochromatosis genetic variants: cohort study in UK Biobank. Br. Med. J. 364, k5222 (2019).
Adams, P. C. Epidemiology and diagnostic testing for hemochromatosis and iron overload. Int. J. Lab. Hematol. 37, 25–30 (2015).
Camaschella, C. Iron-deficiency anemia. N. Engl. J. Med. 372, 1832–1843 (2015).
Benyamin, B. et al. Novel loci affecting iron homeostasis and their effects in individuals at risk for hemochromatosis. Nat. Commun. 5, 4926 (2014).
Liao, M. et al. Genome-wide association study identifies variants in PMS1 associated with serum ferritin in a Chinese population. PloS ONE 9, e105844 (2014).
Raffield, L. M. et al. Genome-wide association study of iron traits and relation to diabetes in the Hispanic Community Health Study/Study of Latinos (HCHS/SOL): potential genomic intersection of iron and glucose regulation? Hum. Mol. Genet. 26, 1966–1978 (2017).
Sveinbjornsson, G. et al. Weighting sequence variants based on their annotation increases power of whole-genome association studies. Nat. Genet. 48, 314–317 (2016).
Sirokmany, G., Donko, A. & Geiszt, M. Nox/Duox family of NADPH oxidases: lessons from knockout mouse models. Trends Pharmacol. Sci. 37, 318–327 (2016).
Rushton, D. H. & Barth, J. H. What is the evidence for gender differences in ferritin and haemoglobin? Crit. Rev. Oncol. Hematol. 73, 1–9 (2010).
Freitas, S. D. S. et al. Genetic variants of VWF gene in type 2 von Willebrand disease. Haemophilia 25, e78–e85 (2019).
Wang, C. Y. & Babitt, J. L. Liver iron sensing and body iron homeostasis. Blood 133, 18–29 (2019).
Yu, J. et al. MTMR4 attenuates transforming growth factor beta (TGFbeta) signaling by dephosphorylating R-Smads in endosomes. J. Biol. Chem. 285, 8454–8462 (2010).
Chen, S. et al. Transforming growth factor beta1 (TGF-beta1) activates hepcidin mRNA expression in hepatocytes. J. Biol. Chem. 291, 13160–13174 (2016).
Stoller, J. K. & Aboussouan, L. S. A review of alpha(1)-antitrypsin deficiency. Am. J. Respir. Crit. Care Med. 185, 246–259 (2012).
Elzouki, A. N., Hultcrantz, R., Stal, P., Befrits, R. & Eriksson, S. Increased PiZ gene frequency for alpha 1 antitrypsin in patients with genetic haemochromatosis. Gut 36, 922–926 (1995).
Schaefer, B. et al. Impaired hepcidin expression in alpha-1-antitrypsin deficiency associated with iron overload and progressive liver disease. Hum. Mol. Genet. 24, 6254–6263 (2015).
Nemeth, E. et al. IL-6 mediates hypoferremia of inflammation by inducing the synthesis of the iron regulatory hormone hepcidin. J. Clin. Investig. 113, 1271–1276 (2004).
Verga Falzacappa, M. V. et al. STAT3 mediates hepatic hepcidin expression and its inflammatory stimulation. Blood 109, 353–358 (2007).
Weiss, G., Ganz, T. & Goodnough, L. T. Anemia of inflammation. Blood 133, 40–50 (2019).
Kim, K. et al. High-density genotyping of immune loci in Koreans and Europeans identifies eight new rheumatoid arthritis risk loci. Ann. Rheum. Dis. 74, e13 (2015).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376 (2014).
Abella, V. et al. Leptin in the interplay of inflammation, metabolism and immune system disorders. Nat. Rev. Rheumatol. 13, 100–109 (2017).
Keestra-Gounder, A. M. & Tsolis, R. M. NOD1 and NOD2: beyond peptidoglycan sensing. Trends Immunol. 38, 758–767 (2017).
Chen, Q. et al. Multiple functions of Ikaros in hematological malignancies, solid tumor and autoimmune diseases. Gene 684, 47–52 (2019).
Schwartz, A. J. et al. Hepatic hepcidin/intestinal HIF-2alpha axis maintains iron absorption during iron deficiency and overload. J. Clin. Investig. 129, 336–348 (2019).
Place, T. L. & Domann, F. E. Prolyl-hydroxylase 3: evolving roles for an ancient signaling protein. Hypoxia 2013, 13–17 (2013).
Appelhoff, R. J. et al. Differential function of the prolyl hydroxylases PHD1, PHD2, and PHD3 in the regulation of hypoxia-inducible factor. J. Biol. Chem. 279, 38458–38465 (2004).
Taniguchi, C. M. et al. Cross-talk between hypoxia and insulin signaling through Phd3 regulates hepatic glucose and lipid metabolism and ameliorates diabetes. Nat. Med. 19, 1325 (2013).
Ha, E. M., Oh, C. T., Bae, Y. S. & Lee, W. J. A direct role for dual oxidase in Drosophila gut immunity. Science 310, 847–850 (2005).
Grasberger, H., El-Zaatari, M., Dang, D. T. & Merchant, J. L. Dual oxidases control release of hydrogen peroxide by the gastric epithelium to prevent Helicobacter felis infection and inflammation in mice. Gastroenterology 145, 1045–1054 (2013).
Hudak, L., Jaraisy, A., Haj, S. & Muhsen, K. An updated systematic review and meta-analysis on the association between Helicobacter pylori infection and iron deficiency anemia. Helicobacter 22, https://doi.org/10.1111/hel.12330 (2017).
Drakesmith, H., Nemeth, E. & Ganz, T. Ironing out ferroportin. Cell Metab. 22, 777–787 (2015).
Kzhyshkowska, J., Gratchev, A. & Goerdt, S. Stabilin-1, a homeostatic scavenger receptor with multiple functions. J. Cell. Mol. Med. 10, 635–649 (2006).
Lee, S. J., Park, S. Y., Jung, M. Y., Bae, S. M. & Kim, I. S. Mechanism for phosphatidylserine-dependent erythrophagocytosis in mouse liver. Blood 117, 5215–5223 (2011).
Park, S. Y. et al. Stabilin-1 mediates phosphatidylserine-dependent clearance of cell corpses in alternatively activated macrophages. J. Cell Sci. 122, 3365–3373 (2009).
Satoh, T. et al. Critical role of Trib1 in differentiation of tissue-resident M2-like macrophages. Nature 495, 524 (2013).
Aziz, A., Soucie, E., Sarrazin, S. & Sieweke, M. H. MafB/c-Maf deficiency enables self-renewal of differentiated functional macrophages. Science 326, 867–871 (2009).
Kautz, L. et al. Identification of erythroferrone as an erythroid regulator of iron metabolism. Nat. Genet. 46, 678–684 (2014).
Troadec, M. B. et al. Targeted deletion of the mouse Mitoferrin1 gene: from anemia to protoporphyria. Blood 117, 5494–5502 (2011).
Arezes, J. et al. Erythroferrone inhibits the induction of hepcidin by BMP6. Blood 132, 1473–1477 (2018).
Danjou, F. et al. Genome-wide association analyses based on whole-genome sequencing in Sardinia provide insights into regulation of hemoglobin levels. Nat. Genet. 47, 1264 (2015).
Thein, S. L. et al. Intergenic variants of HBS1L-MYB are responsible for a major quantitative trait focus on chromosome 6q23 influencing fetal hemoglobin levels in adults. Proc. Natl Acad. Sci. USA 104, 11346–11351 (2007).
Desimone, J., Biel, M. & Heller, P. Maintenance of fetal hemoglobin (Hbf) elevations in the baboon by prolonged erythropoietic stress. Blood 60, 519–523 (1982).
Lavelle, D., Desimone, J., Heller, P., Zwiers, D. & Hall, L. On the mechanism of Hb F-elevations in the baboon by erythropoietic stress and pharmacological manipulation. Blood 67, 1083–1089 (1986).
Bianchi, M. & Magnani, M. Hexokinase mutations that produce nonspherocytic hemolytic-anemia. Blood Cells Mol. Dis. 21, 2–8 (1995).
Camaschella, C. Iron deficiency. Blood 133, 30–39 (2019).
Van Cott, E. M., Khor, B. & Zehnder, J. L. Factor V Leiden. Am. J. Hematol. 91, 46–49 (2016).
Kassebaum, N. J. et al. A systematic analysis of global anemia burden from 1990 to 2010. Blood 123, 615–624 (2014).
McLaren, C. E. et al. Distribution of transferrin saturation in an Australian population: relevance to the early diagnosis of hemochromatosis. Gastroenterology 114, 543–549 (1998).
Kanai, M. et al. Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat. Genet. 50, 390–400 (2018).
Menzel, S. et al. A QTL influencing F cell production maps to a gene encoding a zinc-finger protein on chromosome 2p15. Nat. Genet. 39, 1197–1199 (2007).
Barton, J. C. & Adams, P. C. Clinical guidelines: HFE hemochromatosis-screening, diagnosis and management. Nat. Rev. Gastroenterol. Hepatol. 7, 482–484 (2010).
O’Keeffe, S. T., Gavin, K. & Lavan, J. N. Iron status and restless legs syndrome in the elderly. Age Ageing 23, 200–203 (1994).
Sun, E. R., Chen, C. A., Ho, G., Earley, C. J. & Allen, R. P. Iron and the restless legs syndrome. Sleep 21, 371–377 (1998).
Allen, R. P. et al. Evidence-based and consensus clinical practice guidelines for the iron treatment of restless legs syndrome/Willis-Ekbom disease in adults and children: an IRLSSG task force report. Sleep Med. 41, 27–44 (2018).
Beaumont, C. et al. Two new human DMT1 gene mutations in a patient with microcytic anemia, low ferritinemia, and liver iron overload. Blood 107, 4168–4170 (2006).
Iolascon, A. et al. Microcytic anemia and hepatic iron overload in a child with compound heterozygous mutations in DMT1 (SCL11A2). Blood 107, 349–354 (2006).
Mims, M. P. et al. Identification of a human mutation of DMT1 in a patient with microcytic anemia and iron overload. Blood 105, 1337–1342 (2005).
Yanatori, I. & Kishi, F. DMT1 and iron transport. Free Radic. Biol. Med. 133, 55–63 (2019).
Hubert, N. & Hentze, M. W. Previously uncharacterized isoforms of divalent metal transporter (DMT)-1: implications for regulation and cellular function. Proc. Natl Acad. Sci. USA 99, 12345–12350 (2002).
Hentze, M. W., Muckenthaler, M. U. & Andrews, N. C. Balancing acts: molecular control of mammalian iron metabolism. Cell 117, 285–297 (2004).
Heeney, M. M. & Finberg, K. E. Iron-refractory iron deficiency anemia (IRIDA). Hematol. Oncol. Clin. North Am. 28, 637–652 (2014). v.
Rigas, A. S., Pedersen, O. B., Magnussen, K., Erikstrup, C. & Ullum, H. Iron deficiency among blood donors: experience from the Danish Blood Donor Study and from the Copenhagen ferritin monitoring scheme. Transfus. Med. 29, 23–27 (2019).
Kell, D. B. & Pretorius, E. Serum ferritin is an important inflammatory disease marker, as it is mainly a leakage product from damaged cells. Metallomics 6, 748–773 (2014).
McCann, J. C. & Ames, B. N. An overview of evidence for a causal relation between iron deficiency during development and deficits in cognitive or behavioral function. Am. J. Clin. Nutr. 85, 931–945 (2007).
Achebe, M. M. & Gafter-Gvili, A. How I treat anemia in pregnancy: iron, cobalamin, and folate. Blood 129, 940–949 (2017).
Andro, M., Le Squere, P., Estivin, S. & Gentric, A. Anaemia and cognitive performances in the elderly: a systematic review. Eur. J. Neurol. 20, 1234–1240 (2013).
Guo, S. Y. et al. Restless legs syndrome: from pathophysiology to clinical diagnosis and management. Front. Aging Neurosci. 9, 171 (2017).
Ward, R. J., Zucca, F. A., Duyn, J. H., Crichton, R. R. & Zecca, L. The role of iron in brain ageing and neurodegenerative disorders. Lancet Neurol. 13, 1045–1060 (2014).
Allen, K. J. et al. Iron-overload-related disease in HFE hereditary hemochromatosis. N. Engl. J. Med. 358, 221–230 (2008).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429 e19 (2016).
Pedersen, O. B. et al. The Danish Blood Donor Study: a large, prospective cohort and biobank for medical research. Vox Sang. 102, 271 (2012).
Hansen, T. F. et al. DBDS Genomic Cohort, a prospective and comprehensive resource for integrative and temporal analysis of genetic, environmental and lifestyle factors affecting health of blood donors. BMJ Open 9, e028401 (2019).
Gudbjartsson, D. F. et al. Large-scale whole-genome sequencing of the Icelandic population. Nat. Genet. 47, 435–U20 (2015).
Jonsson, H. et al. Data Descriptor: Whole genome characterization of sequence diversity of 15,220 Icelanders. Sci. Data 4, 170115 (2017).
Eggertsson, H. P. et al. Graphtyper enables population-scale genotyping using pangenome graphs. Nat. Genet. 49, 1654 (2017).
Kong, A. et al. Detection of sharing by descent, long-range phasing and haplotype imputation. Nat. Genet. 40, 1068–1075 (2008).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Gudbjartsson, D. F. et al. Sequence variants from whole genome sequencing a large group of Icelanders. Sci. Data 2, 150011 (2015).
Huang, J. et al. Improved imputation of low-frequency and rare variants using the UK10K haplotype reference panel. Nat. Commun. 6, 8111 (2015).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Vilhjalmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Kong, A. et al. Selection against variants in the genome associated with educational attainment. Proc. Natl Acad. Sci. USA 114, E727–E732 (2017).
Di Angelantonio, E. et al. Efficiency and safety of varying the frequency of whole blood donation (INTERVAL): a randomised trial of 45 000 donors. Lancet 390, 2360–2371 (2017).
Acknowledgements
We thank all the study subjects for their valuable participation, the staff from all studies, and the participating physicians. Participants in the INTERVAL randomized controlled trial were recruited with the active collaboration of NHS Blood and Transplant England (www.nhsbt.nhs.uk), which has supported fieldwork and other elements of the trial. DNA extraction and genotyping were co-funded by the National Institute for Health Research (NIHR), the NIHR BioResource (http://bioresource.nihr.ac.uk/) and the NIHR Cambridge Biomedical Research Centre (BRC) [*]. The academic coordinating centre for INTERVAL was supported by core funding from NIHR Blood and Transplant Research Unit in Donor Health and Genomics (NIHR BTRU-2014-10024), UK Medical Research Council (MR/L003120/1), British Heart Foundation (SP/09/002; RG/13/13/30194; RG/18/13/33946) and the NIHR Cambridge BRC [*]. The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care. A complete list of the investigators and contributors to the INTERVAL trial is provided in reference87. The academic coordinating centre would like to thank blood donor centre staff and blood donors for participating in the INTERVAL trial. Professor John Danesh holds a British Heart Foundation Professorship and a National Institute for Health Research Senior Investigator Award. Steven Bell and Dragana Vuckovic were funded by the NIHR Blood and Transplant Research Unit in Donor Health and Genomics (NIHR BTRU-2014-10024). Stephen Kaptoge is funded by a BHF Program Grant (RG/18/13/33946). Will Astle, Joanna Howson, and Tao Jiang are funded by the NIHR Cambridge BRC. Angela M. Wood and Elias Allara are supported by the EC-Innovative Medicines Initiative (BigData@Heart). Praveen Surendran is supported by a Rutherford Fund Fellowship from the Medical Research Council grant MR/S003746/1. This work was supported by Health Data Research UK, which is funded by the UK Medical Research Council, Engineering and Physical Sciences Research Council, Economic and Social Research Council, Department of Health and Social Care (England), Chief Scientist Office of the Scottish Government Health and Social Care Directorates, Health and Social Care Research and Development Division (Welsh Government), Public Health Agency (Northern Ireland), British Heart Foundation and Wellcome. The Novo Nordisk Foundation (NNF14CC0001 and NNF17OC0027594). The Innovative Medicines Initiative 2 Joint Undertaking under grant agreement No. 115881 (RHAPSODY) (Karina Banasik and Søren Brunak). The Danish Administrative Regions; The Danish Administrative Regions’ Bio- and Genome Bank; The authors thank all the blood banks in Denmark for both collecting and contributing data to this study. Danish Blood Donor Research Fund. Aarhus University, Copenhagen University Hospital Research Fund.
Author information
Authors and Affiliations
Consortia
Contributions
Conception or design of the work: S.B., A.S.R., M.K.M., E.F., U.T., P.S., D.G., K.S., N.S., W.H.O., D.J.R., A.S.B., J.D., E.D.A., and H.U.; the acquisition of data: V.T., B.V., P.T.O., I.O., G.R., O.S., H.S., I.J., H.S., H.H., E.A., A.R., N.S., W.H.O., D.J.R., A.S.B., J.D., E.D.A., A.S.R., E.S., K.S.B., L.W.T., C.E., K.M.D., O.B.V.P., H.P., K.R.N., He.H., T.F.H., D.G.C., K.B., S.B., H.U., and D.B.R.; analysis of data: M.K.M., E.F., G.B., J.K.S., G.H., H.P.E., S.K., S.H.L., K.J., M.I.M., M.L.F., A.O., J.K.S., G.T., D.G., S.B., E.A., and D.W.l.; interpretation of data: M.K.M., E.F., G.T., U.T., P.S., D.G., K.S., S.B., E.A., A.R., D.S.P., J.M.M.H., W.J.A., D.V., Pr.S., T.J., P.A., S.K., A.M.W., J.E.P., N.S., W.H.O., D.J.R., A.S.B., J.D., E.D.A., A.S.R., and H.U.; drafting the paper: M.K.M., E.F., P.S., D.G., K.S., S.B., E.A., E.D.A., and A.S.R.; and all authors were involved in editing the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: Henrik Ullum received an unrestricted research grant from Novartis. Cristian Erikstrup received an unrestricted research grant from Abbott. Søren Brunak reports grants from Innovation Fund Denmark, grants from Novo Nordisk Foundation during the conduct of the study; and personal fees from Intomics A/S and Proscion A/S, outside the submitted work. John Danesh sits on the International Cardiovascular and Metabolic Advisory Board for Novartis (since 2010); the Steering Committee of UK Biobank (since 2011); the MRC International Advisory Group (ING) member, London (since 2013); the MRC High Throughput Science ‘Omics Panel Member, London (since 2013); the Scientific Advisory Committee for Sanofi (since 2013); the International Cardiovascular and Metabolism Research and Development Portfolio Committee for Novartis; and the Astra Zeneca Genomics Advisory Board (2018). Adam S. Butterworth has received grants outside of this work from AstraZeneca, Biogen, BioMarin, Bioverativ, Merck, and Novartis and personal fees from Novartis. For the authors who are affiliated with deCODE genetics/Amgen, we declare competing financial interests as employees.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bell, S., Rigas, A.S., Magnusson, M.K. et al. A genome-wide meta-analysis yields 46 new loci associating with biomarkers of iron homeostasis. Commun Biol 4, 156 (2021). https://doi.org/10.1038/s42003-020-01575-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01575-z
This article is cited by
-
Do iron homeostasis biomarkers mediate the associations of liability to type 2 diabetes and glycemic traits in liver steatosis and cirrhosis: a two-step Mendelian randomization study
BMC Medicine (2024)
-
Iron status and sarcopenia-related traits: a bi-directional Mendelian randomization study
Scientific Reports (2024)
-
Hepatic immune regulation and sex disparities
Nature Reviews Gastroenterology & Hepatology (2024)
-
Genetically determined circulating micronutrients and the risk of nonalcoholic fatty liver disease
Scientific Reports (2024)
-
A Novel Homozygous Germline Mutation in Transferrin Receptor 1 (TfR1) Leads to Combined Immunodeficiency and Provides New Insights into Iron-Immunity Axis
Journal of Clinical Immunology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.