Abstract
Overcoming tumor-mediated immunosuppression and enhancing cytotoxic T-cell activity within the tumor microenvironment are two central goals of immuno-oncology (IO) drug discovery initiatives. However, exploratory assays involving immune components are often plagued by low-throughput and poor clinical relevance. Here we present an innovative ultra-high-content assay platform for interrogating T-cell-mediated killing of 3D multicellular tumor spheroids. Employing this assay platform in a chemical genomics screen of 1800 annotated compounds enabled identification of small molecule perturbagens capable of enhancing cytotoxic CD8+ T-cell activity in an antigen-dependent manner. Specifically, cyclin-dependent kinase (CDK) and bromodomain (BRD) protein inhibitors were shown to significantly augment anti-tumor T-cell function by increasing cytolytic granule and type II interferon secretion in T-cells in addition to upregulating major histocompatibility complex (MHC) expression and antigen presentation in tumor cells. The described biotechnology screening platform yields multi-parametric, clinically-relevant data and can be employed kinetically for the discovery of first-in-class IO therapeutic agents.
Similar content being viewed by others
Introduction
It is well established that the types and frequencies of immune infiltrates in solid tumors serve as prognostic indicators for progression-free survival in patients1,2. Comprehensively, it can be stated that increasing infiltration and/or activity of desirable anti-tumor T-cells (effector T-cells) within the tumor microenvironment results in more durable patient response. Identification of new proteins that confer an immunomodulatory response in malignancy is critically important, in part due to the enormous clinical successes of the checkpoint inhibitors Pembrolizumab (anti-PD-1) and Ipilimumab (anti-CTLA-4)3,4. However, efforts to identify new drug targets in this space have been hampered by a lack of reliable and robust in vitro assays coupled with poor predictive powers of in vivo rodent models5.
There are many ways to enhance anti-tumor T-cell function. Generation and expansion of tumor-killing effector CD8+ T-cells are crucial for anti-tumor immunity, and this can only result from appropriate T-cell priming (cross-presentation). In cancer, much of the T-cell priming events occur in the tumor’s draining lymph node where antigen-presenting cells (APCs) such as dendritic cells (DCs) and macrophages present tumor neoantigens to naive T-cells6. It is also desirable to increase penetration of T-cells into the tumor microenvironment, which is often poorly vascularized and characterized by dense extracellular matrix (ECM) and desmoplastic tissue7. A third way to increase T-cell tumor cytotoxicity is to facilitate the recognition of the tumor cells by the T-cells. The binding event between the T-cell receptor (TCR) and the peptide-containing major histocompatibility complex (pMHC) on tumor cells is biologically critical. Although a variety of peptides can be recognized by T-cells in the context of cancer8, tumor cells will often downregulate the expression of Human Leukocyte Antigen (HLA) genes which encode MHC proteins, thus rendering them invisible to effector T-cells9. Therefore, increasing tumor cell “visibility” would be desirable for new immuno-oncology therapeutics.
The necessity for scrutinizing cellular interactions in a 3-dimensional (3D) cell model lies in the physical and chemical cues present within the tumor microenvironment that cannot be recapitulated in conventional 2-dimensional cellular assays. Specifically, the multicellular tumor spheroid model is characterized by gradients of oxygen and nutrient penetration, large internal sections of hypoxia and acidic cores10, pathologies that are similarly observed in a clinical setting. Tumor cells often exhibit an altered metabolic phenotype called aerobic glycolysis (Warburg effect); the result of which is the production and secretion of lactic acid, which lowers the pH of the extracellular microenvironment and leads to areas of necrosis11. Tumor lactic acid production has been shown to be negatively correlated to T-cell proliferation and anti-tumor function12. The 3D multicellular tumor spheroid model is a faithful mimic of both hypoxia and lactic acid secretion and, thus, is a useful tool for the interrogation of these pathologies on the ability of effector T-cells to induce tumor cytotoxicity13. T-cell tumor infiltration is essential for effector function. Therefore, a 3D scaffolded tumor structure is a prerequisite for observing and modulating this biology14,15. Comprehensively, 3D tumor spheroid cell models may represent more faithful surrogates for the study of clinically relevant T-cell anti-tumor function than conventional 2D in vitro methods16.
The efforts described here identify new classes of targeted small molecules that enhance anti-tumor T-cell function. Further, we used viral peptide antigens in an HLA-matched co-culture system to mimic a tumor antigen-specific T-cell response. By miniaturizing the assay we were able to profile a large collection of compounds and identify several target proteins that can be modulated to augment cytotoxic T-cell activity. Specifically, CDK4/6 inhibitors and nonspecific BRD inhibitors were shown to act on both T-cell and tumor cell compartments by (1) increasing the release of T-cell cytolytic granules and interferon-gamma (IFNg) and (2) upregulating tumor cell MHC expression, antigen presentation, and IFNg secretion. This resulted in canonical T-cell-mediated tumor killing which was conserved over a variety of different HLA-matched patient donors. Several reports have identified methods that can be used to enhance anti-tumor T-cell function using both 2D17 and 3D cell models16, and this work aims to expand on those efforts in an industrialized drug discovery setting.
Results
Development of the T-cell tumor spheroid-killing platform and screening workflow
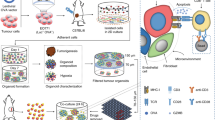
In developing an assay platform to enable the identification of agents that enhance T-cell anti-tumor function the aim was to detect compounds that elicit effects on T-cells either intrinsically or extrinsically via tumor cells. There are several conceivable ways experimental compounds might enhance T-cell-mediated anti-tumor function in our model system. Compounds might act directly on T-cells by enhancing anti-tumor cytokine pathways (e.g., IFNg, TNFa), agonizing cytolytic pathways, or enhancing perforin or cytolytic granzyme synthesis/secretion (Fig. 1a: 1 and 2). Alternatively, compounds might act via tumor cells to affect T-cell killing; by increasing MHC class I expression and/or antigen processing/presentation or upregulation of co-stimulatory molecules (Fig. 1a: 3).
a Three different mechanisms by which small molecules might enhance anti-tumor T-cell function: (1) upregulate biosynthesis/secretion of anti-tumor cytokines (e.g., IFNg, TNFa), (2) increase production/secretion of T-cell cytolytic granules or perforin proteins or (3) upregulate antigen processing/presentation on tumor cells. b Workflow of IO screening platform. Human HLA-A02 PBMCs are isolated and cultured in the presence of CEF viral peptides (day −9). Activated T-cells are expanded first in IL-2 (2 days), then in IL-2, IL-4, and IL-7 (4 days) to generate the desired memory T-cell population. Human HCT116-EGFP colorectal cancer cells are dispensed into 1536-well spheroid plates (day −2) and on day 0 T-cells are stained red and added to spheroid cultures. Experimental compounds are added and co-cultures are incubated for 3 days before the first imaging read and then a further 2 days before the final imaging read.
3-dimensional (3D) tumor spheroids were used in this phenotypic screening platform to mimic clinically relevant tumor physiology. Although small tumor spheroids do not typically display necrotic cores comparable to larger-sized spheroids10, 1536-well spheroids did display substantial areas of hypoxia and acidic pH (Supplementary Fig. 1a). T-cell anti-tumor function was quantitated by a reduction of spheroid size or complete dissociation of the 3D tumor spheroid structure. The assay began on day −9 when normal human donor peripheral blood mononuclear cells (PBMCs) were thawed and cultured in the presence of a 1 μM CEF peptide pool (Fig. 1b). The CEF peptide pool is a mix of 32 different peptides derived from either Cytomegalovirus (CMV), Epstein–Barr virus (EBV), or Influenza virus (Flu) and induces T-cell IFNg secretion and expansion in approximately 80% of tested donor PBMCs (Supplementary Fig. 1b). PBMC cultures were then expanded in the presence of low dose interleukin-2 (IL-2) for a further 3 days to increase the proliferation of CD8+CD127+ memory T-cells18. PBMC cultures, which consisted mainly of CD4+ and CD8+ T-cells, were then cultured in a “resting” medium for a further 3 days in the presence of IL-2, IL-4, and IL-7 (Fig. 1b)19. This resting period was necessary to mitigate nonspecific, antigen-independent cytolytic properties of the activated T-cells (Supplementary Fig. 1c). On day −2, 1000 human colorectal tumor cells (HCT116 expressing the enhanced green fluorescent protein-EGFP) were plated into 1536w spheroid plates in the presence of either 1 μM CEF peptide pool or a negative control CEF Scramble pool (to act as a counter screen for antigen-dependence). On day 0, T-cells were stained red and added to the tumor spheroids in a 3:1 ratio of T-cells to tumor cells (effector: target) (Fig. 1b). Experimental test compounds were then added at 30 μM single point dose. The assay proceeded for 3 days before the first imaging readout was collected and then another 2 days before the final readout was collected by high-content confocal microscopy (Fig. 1b).
Characterization of antigen presentation and reactivity
In order to ensure an antigen-dependent mechanism of T-cell anti-tumor function, it was necessary to quantitate tumor cell MHC expression and antigen presentation as well as T-cell antigen recognition. HCT116 colorectal cancer cells harbor the HLA-A02 allele (TRON Cell Line Portal: celllines.tron-mainz.de), and to confirm expression we used an antibody binding capacity (ABC) assay to quantitate HLA-A02 expression on HCT116 cells as ~81,677 molecules per cell (Supplementary Fig. 2a), indicating a high level of expression. HCT116 cells expressed high quantities of MHC on their cell surface membranes and treatment with IFNg did not meaningfully increase MHC presentation (Supplementary Fig. 2b). Next, we examined the exogenous loading of tumor cell HLA-A02 by first quantitating the amount of viral peptide expressed on MHC Class I molecules. HCT116 tumor spheroids were incubated with a fluorescently tagged CMV peptide [CEF 20: CMVpp65(495–503)-BODIPY-488], then treated with either a protein transport inhibitor (Brefeldin) or an unlabeled CMV cold competitor. Brefeldin was used as a positive control treatment in order to arrest protein transport within the cells, thus preventing intracellular MHC molecules from presenting peptides on cell surface membranes. The localization of the CMV-BODIPY peptide to the cell membranes of tumor cells was visualized by high-content confocal microscopy (Fig. 2a). Spheroids were holistically displayed as maximum intensity projections of the entire spheroid (Z-Stack Merge) and then as focused sections within the middle of the spheroid (single plane merge) to characterize CMV-BODIPY cellular localization. Tumor spheroids treated with the drug vehicle (DMSO) displayed punctate and cell membrane-localized CMV-BODIPY peptides, consistent with MHC-based cell membrane localization (Fig. 2a, DMSO, blue arrows). Treatment of spheroids with Brefeldin resulted in accumulation of CMV-BODIPY within cells and poor trafficking of the peptides to cell membranes (Fig. 2a, Brefeldin, white arrows). Treatment of spheroids with an unlabeled CMV cold competitor did not affect CMV-BODIPY transport to cell membranes to a noticeable degree, though we did observe some accumulation of CMV-BODIPY peptides within cells (Fig. 2a, CMV cold competitor). To functionally quantitate CD8+ T-cells specifically reactive to a common CMV peptide within the CEF pool (CMVpp65 495–503), we used antibody tetramers, soluble MHC molecules loaded with the CMVpp65 peptide antigen. T-cells stimulated with CEF pool were quantitated through tetramer flow cytometry and shown to be highly reactive against CMVpp65 peptide, but not against the negative control EBV peptide (Fig. 2b). PBMC donor #4666 was highly reactive to the CMVpp65 viral peptide and was chosen to be used in the subsequent high-content screening campaign.
a CEF viral antigens are presented on the cell surface membranes of HCT116 cells within a tumor spheroid. HCT116 tumor spheroids were loaded with CMVpp65 viral antigen conjugated to a BODIPY-488nm fluorochrome and treated with either Brefeldin (a protein transport inhibitor), DMSO (vehicle control), or an unlabeled CMV competitor peptide (CMV cold competitor). Spheroids were stained with a cell membrane dye (CellBrite) and a nuclear dye (Draq5) to identify the cellular localization of CMV-BODIPY peptides. Spheroids were imaged comprehensively in a large z-stack and then single planes were more closely scrutinized for CMV-BODIPY membrane localization. Blue arrows indicate membrane-localized CMV-BODIPY peptides and white arrows indicate cytoplasmic CMV-BODIPY peptides. Scale bar equals 60 µm. b Antigen-specific T-cell priming reveals pMHC-reactive memory T-cells. Human donor PBMCs were stimulated with either CMVpp65 or EBV-BMFL1 viral peptides and expanded for 7 days in culture. Flow cytometry analyses of CD3+CD8+ T-cell populations reveal binding of TCRs only to CMV tetramers and not EBV tetramers, indicating a CMV-reactive memory T-cell population had been selected and expanded. c Phenotyping of T-cells used for the spheroid-killing screen. PBMC cultures stimulated with CEF viral peptides and expanded in IL-2, then in IL-2, IL-4, and IL-7 were subjected to flow cytometry T-cell phenotyping at days 3, 6, and 9 post-stimulation. CD8+CD127+ T-cells indicate desirable memory effector T-cell population for screening.
To prepare cytotoxic T-cells, we expanded T-cells from PBMC cultures in three phases. First, T-cells were stimulated using the CEF viral peptide pool which initiated CD8+CD127+ T-cell expansion (Fig. 2c, day 3). T-cells were then further proliferated in the presence of IL-2 for three days which selectively increased CD8+ T-cell expansion over CD4+ T-cell expansion (Fig. 2c, day 6). The third phase of T-cell expansion required culturing T-cells in “resting” medium, which contained IL-2, IL-4, and IL-7 (Fig, 2c, day 9) and resulted in the further expansion of the desired CD8+CD127+ population. Failure to expand T-cells in a resting medium resulted in aggressive cytolytic behavior upon exposure to tumor spheroids in a non-antigen-dependent manner (Supplementary Fig. 1c). However, a 4-day expansion in T-cell resting media mitigated these antigen-independent cytolytic effects enabling the identification of T-cell-enhancing therapeutics.
To ensure that antigen alone was responsible for this observed T-cell anti-tumor function, we performed a cell-killing assay using T-cells that had been primed with the CMVpp65 viral peptide and HCT116 colorectal cancer cells engineered to express the Firefly Luciferase (FL) reporter gene. HCT116-FL cells were grown in the presence of a limiting dilution of either CMVpp65 peptide or a scramble negative control peptide. T-cells were added in a 3:1 ratio (T-cells: tumor cells) and co-culture proceeded for 48 h. Analysis of HCT116-mediated Firefly Luciferase enzyme indicated T-cells killed tumor cells in a CMV-dose-dependent manner; this cytotoxicity was not observed in tumor cells that had been loaded with scramble control peptides (Supplementary Fig. 3).
Screen for enhancers of T-cell-mediated anti-tumor function
To find target proteins whose functions might be modulated to enhance T-cell-mediated tumor cytotoxicity, we employed the spheroid-killing IO platform in a chemical genomics screen of 1770 annotated compounds with known or suspected targets or mechanisms of action (MoAs)20. The compound collection screened during this initiative was a subset of the library described in20 and consisted of low molecular weight targeted perturbagens covering more than 16 different protein and enzyme classes (Supplementary Fig. 4a). The majority of abnormal spheroid phenotypes observed could be broadly categorized as either Hit Class I or Hit Class II (Fig. 3a). Hit Class I compounds reduced the size of the spheroid but maintained its overall shape factor compared to the vehicle control (DMSO; control picture); whereas treatment with Hit Class II compounds resulted in the complete loss of both spheroid shape factor and spheroid area (Fig. 3a). Hit Class II compounds, such as CDK4/6 inhibitors, demonstrated severe deterioration of spheroid structural integrity compared to the DMSO (vehicle) negative control in which the spheroid grew to a large and characteristically rounded shape (Fig. 3b). Interestingly, T-cell proliferation was often correlated to anti-tumor activity indicated by increased numbers of T-cells observed in hit compound-treated samples versus non-hit compound-treated samples (Supplementary Fig. 4b). Increased T-cell proliferation was often observed in DMSO-treated samples, which is why T-cell count calculations were performed against the median average of the screened compounds and not the DMSO vehicle controls (Supplementary Fig. 4b).
a Scatter plot showing the results of the 1770 compound high-throughput phenotypic screen for enhancers of anti-tumor T-cell function. The majority of compounds did not elicit any change in tumor spheroid phenotype and clustered with the DMSO vehicle control samples. Hit compounds that yielded desirable anti-tumor phenotypes were broadly classified by their ability to reduce tumor size (Hit Class I) or size and structure (Hit Class II). Solid lines indicate the mean for either the spheroid area (y axis) or the spheroid shape factor (x axis); dotted lines indicate three standard deviations from the mean. b Example of a hit compound from the screen. Tumor spheroids treated with the CDK4/6 inhibitor are shown in three different channels alongside the DMSO negative control. Scale bars indicate 100 µm.
The high-throughput screen was run concurrently with a counter screen for the purpose of distinguishing compounds whose activities were dependent on tumor cell antigen presentation and T-cell recognition. The primary screen was run using both T-cells and tumor cells loaded with identical tumor antigens (CEF Peptide Pool), the counter screen was run using negative control antigens (CEF Scramble Pool), and all of the hit compounds were profiled on tumor spheroids alone to identify those compounds that were not dependent at all on the presence of T-cells (No T-cells).
Predictably, most of the compounds that demonstrated spheroid cytotoxicity were not at all dependent upon the presence of T-cells. These nonspecific cytotoxic compounds, such as microtubule inhibitors and histone deacetylase (HDAC) inhibitors, were able to dissociate tumor spheroids and kill tumor cells (Fig. 4a, Cytotoxic). Other classes of compounds that killed tumor spheroids were those that interfered with MAPK signaling. HCT116 cells are mutant KRAS-driven colon cancer, so compounds that antagonize RAF, MEK, or ERK signaling would be expected to act selectively against this tumor type21. Intriguingly, many compounds whose targets and mechanisms act through ECM remodeling could be distinguished by their unique phenotypes. Compounds that antagonize matrix metalloproteinases (MMPs) or integrin receptors led to the complete structural dissociation of the tumor spheroids without any noticeable associated cellular toxicities (Fig. 4a, ECM). To distinguish between spheroid destruction and tumor cell killing we profiled hit compounds on T-cell: HCT116-FL spheroid co-cultures. Luciferase assays performed on treated HCT116-FL spheroids revealed consistencies between the phenotypic imaging data and the spheroid viability data. Compounds that inhibited HDAC or ERK proteins severely affected tumor cell viability at low doses (Fig. 4b). Tubulin inhibitor compounds were less toxic to HCT116-FL spheroids and MMP inhibitors demonstrated minimal viability effects on spheroids, consistent with the imaging analyses (Fig. 4b).
a Hit compounds from the primary screen were tested on T-cell/spheroid co-cultures where both cell types were primed with the same antigen (CEF Peptide Pool), cell types were primed with different antigens (CEF Scramble) or spheroid monocultures with no T-cells (No T-cells). Cytotoxic compounds that targeted key cellular machinery exhibited anti-tumor properties independent of T-cell presence (MTi β-tubulin inhibitor, HDACi histone deacetylase inhibitor, MAPKi mitogen-activated protein kinase inhibitor). ECM-targeting compounds could be distinguished by their unique, though non-toxic, phenotypes (MMPi matrix metalloproteinase inhibitor). Spheroid size is quantitated in bar graphs on right, relative to DMSO controls (ns = not significant and error bars indicate standard deviation of three biologically independent samples). b Viability readout of spheroids from a using HCT116-Luc reporter spheroids in the presence of T-cells and hit compounds with matching viral antigens (CEF Peptide Pool). HDAC inhibitors and β-tubulin inhibitors demonstrated sigmoidal dose–response curves, ERK inhibitors demonstrated high potencies and MMP inhibitors did not significantly affect tumor spheroid cell viability.
There were, however, a number of compounds from the screen that demonstrated T-cell-dependent activities. For example, compounds that negatively affected NF-κB or IL-4 signaling showed very strong antigen-dependent anti-tumor activities, as scramble antigen controls did not produce these phenotypes (Fig. 5a, Anti-inflammatory). As mentioned previously, another class of compounds that was shown to enhance T-cell tumor cytotoxicity was the cyclin-dependent kinase (CDK) group of kinase inhibitors. Compounds that inhibited CDK4 and/or CDK6 were able to slow tumor spheroid growth in the absence of T-cells, consistent with the behavior of cytostatic cell-cycle-targeting compounds (Fig. 5a, Cytostatic, No T-cells). However, in the presence of T-cells, CDK4/6 inhibitors were able to significantly reduce tumor spheroid size and these phenotypes were exacerbated when T-cells were added to tumors loaded with their cognate viral antigens (Fig. 5a, Cytostatic). In addition to CDK inhibitors, bromodomain (BRD) inhibitors also demonstrated antigen-dependent anti-tumor phenotypes. Compounds targeting other proteins with previously-unreported T-cell-modifying activities also scored as hits in the screen. Many of these compounds acted through nonspecific (non-antigen-related) T-cell mechanisms and demonstrated tumor-killing behaviors in the CEF Peptide Pool, as well as the CEF Scramble Pool negative control and to the best of our knowledge, have not been previously reported to have immunomodulatory properties (Fig. 5a, Novel Targets). To confirm these compounds were inducing T-cell-mediated cytotoxicity on tumor spheroids we performed identical assays using HCT116-FL spheroids loaded with matched CEF Pool peptides. All of the T-cell-dependent compound classes identified from the phenotypic screen demonstrated potent viability effects on HCT116-FL spheroids, indicating tumor cell death in addition to tumor spheroid destruction (Fig. 5b).
a Hit compounds from the primary screen were tested on T-cell/spheroid co-cultures where both cell types were primed with the same antigen (CEF Peptide Pool), cell types were primed with different antigens (CEF Scramble) or spheroid monocultures with no T-cells (No T-cells). Anti-inflammatory compounds (NF-κB and IL-4 inhibitors) were shown to enhance T-cell spheroid-killing properties, where antigen-dependent interactions significantly increased compound efficacies. Other compounds, such as CDK4/6 or BRD inhibitors, demonstrated enhanced T-cell anti-tumor function in a mainly antigen-independent manner (Cytostatic). Several compounds targeting novel proteins not reportedly associated with enhanced T-cell cytotoxicity demonstrated potent antigen-dependent phenotypes. Spheroid sizes are quantitated in bar graphs on right, relative to DMSO controls. Scale bars indicate 100 µm. An independent sample t-test was used to calculate probability where *p < 0.05, **p < 0.01; n = 3 biologically independent samples and error bars represent standard deviation. b Viability readout of spheroids from a using HCT116-Luc reporter spheroids in the presence of T-cells and hit compounds with matching viral antigens (CEF Peptide Pool). NF-κB inhibitor demonstrated toxicity but limited potency, while the CDK4/6 inhibitor showed a predictably cytostatic response. Sonic Hedgehog inhibitor (SHHi, Cyclopamine), as well as a compound targeting an unidentified protein (GNF001), demonstrated high potencies and sigmoidal dose–response curves.
Characteristics of lead compounds and T-cell phenotypes
Few compounds that scored in the screen demonstrated antigen-dependent T-cell mechanisms. Therefore, to expand on these compounds and target classes, we selected small molecules from our compound libraries that have similar structures or similar targets as the hits from the high-throughput screen. 72 of these SAR compounds (structure–activity relationship) were tested for T-cell-mediated anti-tumor activity. One particularly interesting compound that elicited antigen-dependent anti-tumor T-cell function was cyclopamine, a well-known Sonic Hedgehog (SHH) pathway inhibitor, and Smoothened (SMO) antagonist. Cyclopamine treatment of T-cell/tumor spheroid cultures led to robust killing and dissociation of tumor spheroids in a dose-dependent manner (Fig. 6a). Further, antigen-matched cultures demonstrated a 7-fold increase in cyclopamine potency over non-antigen-matched (Fig. 6a, IC50 curves). The activity of cyclopamine in this context was T-cell-dependent, as monoculture spheroids demonstrated no sensitivity to cyclopamine treatment (Fig. 6a, No T-cells). Finally, several experimental therapeutics with unknown targets exhibited potent anti-tumor T-cell efficacies when tested kinetically and in dose–response. GNF001, a small molecule with an unknown target, yielded potent T-cell-mediated tumor killing in a time- and antigen-dependent manner (Fig. 6b). This compound showed slight anti-tumor activity after 3 days but marked anti-tumor efficacy after 5 days (Fig. 6b, IC50 curves).
a Dose response profile of Cyclopamine, an inhibitor of SHH and Smoothened antagonist, at two-time points. Day 5 images and IC50 curves reveal antigen-dependent anti-tumor T-cell behavior, where CEF Scramble or no T-cell control samples did not elicit significant phenotypes. b Dose–response profile of GNF001, an inhibitor of an unknown protein target, at two-time points. Day 5 images and IC50 curves reveal a 22-fold increase in compound potency when T-cells and tumor cells were primed with identical antigens (CEF Peptide Pool) compared to mismatched viral antigens (CEF Scramble).
When performing cellular studies involving multiple cell types, it is critical to establish which cellular compartment is responsible for eliciting any observed phenotypes. Therefore, we interrogated the effects of our lead compounds on both monoculture T-cells and tumor spheroids. Compounds were applied to T-cells or tumor spheroids for 48 h, then washed out, before combining cells into the co-culture model described earlier. Compounds that antagonized CDK4/6 activity were observed to elicit their activities via both cell types, as T-cell anti-tumor activity was observed under both tumor spheroid-only treatment and T-cell-only treatment (Supplementary Fig. 5). However, a dose–response for tumor cytolytic phenotypes was observed for CDK4/6 inhibitors only under tumor spheroid treatment; only the highest dose of this compound on T-cells was able to enhance T-cell-mediated anti-tumor function (Supplementary Fig. 5). BRD inhibitors and the proprietary compound GNF001 likely worked through tumor spheroid-mediated mechanisms as T-cell anti-tumor function was seen only in the spheroid-treated samples compared to vehicle-treated cultures (DMSO) (Supplementary Fig. 5). Compounds that antagonized NF-κB or IL-4 inflammatory pathways did not elicit potent phenotypes under monoculture treatment conditions accept at the highest potencies, indicating the necessity for both cell types to be present for efficacy (Supplementary Fig. 5).
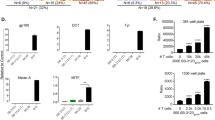
To understand the mechanism of action of these compounds, we treated either T-cells or tumor spheroids in monoculture with lead compounds for 48 h and quantitated cytokine secretion pathways and cell surface markers by FACS. Although none of the compounds except for cyclopamine were able to significantly increase the proliferation of CD3+ T-cells, we did observe the significant expansion of the CD8+/CD25+ T-cell fraction under CDK4/6 inhibitor, NF-κB inhibitor, and cyclopamine treatment (Fig. 7a). More importantly, however, were the increases in T-cell reactivity to the CMV tetramer antibody (Fig. 7a, CD8+/CMVtet+) observed for nearly all compounds tested. These T-cell phenotypes indicated increased recognition of the CMV viral peptide, which may explain a potential mechanism by which tumor spheroids were more susceptible to T-cell cytolytic function under compound treatment. Intriguingly, treating T-cells with CDK4/6 inhibitors led to a drastic increase in interferon-gamma secretion, even in the absence of tumor spheroid target cells, indicating significant agonism of this anti-tumor cytokine pathway (Fig. 7a, CD8+/IFNg+). In conjunction with the FACS-based T-cell phenotyping experiments, we also interrogated HLA-A02 cell surface expression and proinflammatory cytokine secretion by tumor spheroids under monoculture compound treatment conditions. Nearly all compounds tested in this manner were observed to increase the expression of HLA-A02 molecules on tumor cell surface membranes (Fig. 7b, HLA-A02+). Additionally, CDK4/6 inhibitors and GNF001 also were shown to significantly increase interferon-gamma secretion in tumor spheroids (Fig. 7b, IFNg+). Surprisingly, TNF-alpha secretion was reduced in tumor spheroids in nearly all samples indicating antagonism of this cytokine pathway under compound treatment (Fig. 7b, TNFa+). How reduced TNFa secretion in tumor spheroids affects T-cell anti-tumor function has yet to be determined.
a T-cells were treated with hit compounds or DMSO vehicle control for 48 h and phenotypically profiled by FACS. b Tumor spheroid cultures were treated with hit compounds or DMSO vehicle control for 48 h and phenotypically profiled by FACS. An independent sample t-test was used to calculate probability where *p < 0.05, **p < 0.01, ***p < 0.001; n = 3 independent experiments and error bars indicate standard deviation.
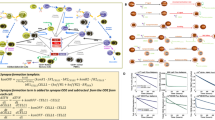
Establishing conserved efficacies over patient cohorts is of critical importance with experimental therapies. Therefore, we sought to characterize the effects of one lead compound over a variety of normal donor PBMCs. Kinetic studies using the CDK4/6 inhibitor (CDK4/6i) compound over 9 HLA-A02 normal donor PBMCs yielded heterogeneous results. Donor T-cell/spheroid co-cultures were treated with either 50 or 5 μM CDK4/6i for 96 h and tracked for spheroid size under antigen-matched (CEF) or antigen-mismatched (Scr) conditions. Three donors demonstrated anti-tumor efficacies only under high dose of compound treatment and only under antigen-matched conditions (Fig. 8a, Antigen-Dependent Responders). Three donors exhibited anti-tumor cytotoxicity under high dose treatment, independent of antigen recognition (Supplementary Fig. 6a). Two donors showed no response to CDK4/6i treatment under any conditions (Fig. 8a, Non-Responders). These data reflect a heterogeneous response rate of immunomodulatory agents comparable to clinical settings.
a Kinetic response of 6 normal human donor T-cells over 96 h under either high dose (50 μM) or low dose (5 μM) CDK4/6i treatment with matched peptides (CEF) or mismatched peptides (Scr). Data normalized to equivalent DMSO negative control samples; n = 3 independent experiments. b T-cell phenotyping data from five patients from a showing populations of CD8+-gated cells under all four CDK4/6i growth conditions. An independent sample t-test was used to calculate probability where *p < 0.05, **p < 0.01, ***p < 0.001 compared to Scr negative control samples; n = 3 independent experiments and error bars indicate standard deviation.
For a more detailed analysis of T-cell phenotypes during co-culture with tumor spheroids under CDK4/6i treatment, we extracted T-cells from these cultures and profiled them for functional markers of active tumor killing. Nearly all of the CD8+ T-cells were positive for the degranulation marker CD107 indicating a high degree of tumor killing (Fig. 8b). We observed modest yet significant increases in Granzyme-B (GranB) and Interferon-γ (IFNg) in the CEF-treated samples over the Scr-treated samples indicating potent cytokine expression under antigen-matched T-cell conditions. Further T-cell FACS-based phenotyping (Supplementary Fig. 6b) revealed an increase in CD8+CD107+, CD8+CMVtet+, CD8+GranB+ and/or CD8+IFNg+ cells in the CEF-treated spheroids over Scr-treated spheroids in three donors that also showed an antigen-mediated T-cell anti-tumor response (Fig. 8b; D17044427, D19054173, D17044460, CEF). T-cells from two donors that did not demonstrate CEF-specific T-cell killing or any anti-tumor efficacy did not display similar T-cell phenotypes (Fig. 8b; D19054445, D19054242). These data indicate that donor T-cells which have contributed to anti-tumor cytolytic function display canonical T-cell degranulation and cytokine markers which are proportional to the observed level of tumor cell killing.
Discussion
Augmenting anti-tumor immunity through the enhancement of CD8+ cytolytic T-cell function is a central initiative in the field of immuno-oncology (IO). The study of functional immunity in vitro can be challenging, and IO cellular assay platforms may not take into account important characteristics of the tumor microenvironment, such as hypoxia and low interstitial pH, which may drastically affect anti-tumor T-cell function. Further, the only way to truly mimic clinical T-cell-tumor cell interaction ex vivo is through the use of primary patient tumor material and autologous PBMCs; a limited prospect when conducting industrial drug discovery screening campaigns that require large amounts of cells. The assay platform presented here attempts to address both of these shortcomings through the use of 3D tumor spheroids and HLA-A02-matched T-cells and tumor cells. Additionally, the assay platform relies on an antigen-dependent mechanism of T-cell-tumor cell interaction, mimicking T-cell education in the draining lymph node, thereby making it highly mimetic of clinical tumor immunology.
When profiling an advanced library of annotated small molecules, it can be expected that compounds targeting key cellular housekeeping proteins will elicit cytotoxic phenotypes; for example, the large number of tubulin inhibitors that scored as hits in the screen. Similarly, given the KRAS-dependency of the HCT116 human colorectal cancer cell type used, we expected the high proportion of MAPK pathway-targeting kinase inhibitors to show anti-tumor activity.
We focused our attention on those compounds that elicited a tumor cytotoxic phenotype only in the presence of T-cells and only within an antigen-matched cellular context. Several compounds targeting the NF-κB pathway as well as antagonists of proinflammatory cytokines (e.g., IL-4) demonstrated potent anti-tumor phenotypes in an antigen-dependent manner. Although much is known about NF-κB signaling and its antagonism in CD4+ regulatory T-cells, NF-κB signaling is less characterized in tumor-associated cytotoxic CD8+ T-cells. NF-κB inhibition has been shown to decrease Foxp3 expression on CD4+ T-cells and increase IFNg and IL-2 secretion22, so NF-κB inhibition in CD8+ T-cells might act through similar cytokine release mechanisms23. Surprisingly, NF-κB inhibitors demonstrated anti-tumor efficacy only in the co-culture cellular model, where no anti-tumor activities were observed when either T-cells or tumor spheroids were treated separately. Consistent with this observation, treating T-cells with NF-κB inhibitors did not significantly affect their proliferation, differentiation, or proinflammatory cytokine secretion. Treatment of tumor spheroids with NF-κB inhibitors slightly enhanced HLA-A02 expression on cell surface membranes and decreased IFNg and TNFa secretion, though these phenotypes would not explain the strong anti-tumor efficacies observed during T-cell/spheroid co-culture treatment. It is tantalizing to speculate that co-culture of T-cells with tumor spheroids may activate NF-κB signaling in one of these cell types that is not present in monoculture and that activation of this paracrine signaling mechanism is responsible for the observed efficacies of NF-κB-targeting compounds. IL-4, a key component of chronic inflammation, has been shown to be pro-tumorigenic and inhibition has been shown to promote the anti-tumor activity of cytotoxic T lymphocytes24. Therefore, our observation that small molecule-mediated IL-4 inhibition enhances T-cell cytolytic function is consistent with reported results.
Strikingly, a number of characteristically cytostatic compounds were able to significantly enhance anti-tumor T-cell function. Particularly, treatment with CDK4/6 inhibitors had a mild cytostatic effect on monoculture tumor spheroids but marked cytotoxic effects were observed when those compounds were combined with T-cells. Given the recent finding that CDK4/6 inhibition augments tumor immunity by increasing the functional capacity of tumor cells to present antigen and reducing the activity of immunosuppressive regulatory T-cells25, we explored whether those phenotypes were present in our cellular model during CDK4/6 inhibitor treatment. Consistent with previous reports, we detected significant increases in HLA-A02 expression on cell surface membranes of tumor spheroids when exposed to CDK4/6 inhibitory compounds and this activity may have been a consequence of enhanced IFNg secretion which was concomitantly increased in these cells during drug treatment. Similarly, CDK4/6 inhibitors were able to significantly enrich T-cell reactivity to the CMV tetramer antibody as well as agonizing IFNg expression, indicating this class of compound may simultaneously increase antigen presentation by target cells and antigen recognition by cytolytic T-cells.
In a mechanism similar to CDK inhibition, bromodomain (BRD) protein inhibitors displayed strong anti-tumor function when combined with T-cells. Reported immunomodulatory activities of BRD inhibitors include (1) expansion and enhancement of cytotoxic CD8+ T-cell populations26 and (2) increased CD8+ T-cell penetration, IFNg secretion, and degranulation in vivo27. Although we did not observe the expansion of cytolytic T-cell populations or IFNg expression under BRD inhibitor treatment, we did detect significant increases in both tumor cell HLA-A02 cell surface expression and T-cell CMVpp65 antigen reactivity. These data indicate that similar to CDK4/6 inhibitors, BRD antagonists may simultaneously enhance T-cell-mediated surveillance of antigen-presenting cells while decreasing immune escape by tumor cells.
One compound that exhibited potent anti-tumor function in an antigen- and T-cell-dependent manner was cyclopamine. Cyclopamine is a well-known teratogen with anti-cancer properties28. Cyclopamine is a Sonic Hedgehog (SHH) pathway inhibitor, binding Smoothened (SMO) protein with high affinity, and shutting down SHH signaling29. Cyclopamine immunomodulatory activity has been shown to be dependent upon oxygenation state, where enhanced T-cell proliferation, tumor cytotoxicity, and IFNg secretion only occurred under hypoxic conditions30,31. During our study, we revealed a potent ability of cyclopamine to induce antigen-dependent T-cell-mediated tumor cytotoxicity. Through FACS-based T-cell phenotyping, we observed modest yet significant increases in both total CD3+ and CD8+/CD25+ cytolytic T-cell proliferation under cyclopamine treatment as well as upregulation of the checkpoint molecule PD-1. It is conceivable the hypoxic environment of spheroid cores was responsible for the observed efficacy of cyclopamine in our assay, a phenomenon that would not have been detected in a conventional 2D cell-killing assay, yet has far-reaching clinical implications.
Several small molecules with targets not previously reported to be involved in T-cell biology or immunomodulation also scored as hits on the screen. One of these compounds, GNF001, yielded a highly desirable therapeutic profile; showing no detectable anti-tumor function in the absence of T-cells. This compound is speculated to affect its anti-tumor function via the tumor cell compartment due to the significant increases in IFNg expression induced in tumor cells under compound exposure. In support of this hypothesis, we observed the anti-tumor efficacy of this compound only during tumor spheroid monoculture treatment, but not in T-cell monoculture treatment. More thorough MoA interrogations are needed to understand the mechanisms by which GNF001 enhances antitumor T-cell function via tumor cells.
Response to immunomodulatory chemotherapeutics in the clinic is heterogeneous. Appropriate patient stratification for clinical testing is crucial for maximizing therapy response. By profiling nine different PBMC donors in a mini “ex vivo clinical trial”, we were able to identify people that would potentially respond to CDK4/6 inhibition in a model of colorectal cancer. These donors could be categorized as either antigen-dependent responders, antigen-independent responders, or non-responders, similar to heterogeneous IO-response rates observed in clinical settings. Moreover, T-cell phenotyping from these donor cultures revealed a tight correlation between activation of T-cell killing markers and an anti-tumor response, indicating the utility of this assay platform for profiling patient T-cell response to therapy.
Taken together, our unique IO assay platform provides evidence that T-cell anti-tumor function can be profiled in a high-throughput industrial drug discovery setting and that small molecule targeted inhibitors can augment intrinsic properties of T-cells either by upregulation of antigen recognition machinery and/or secretion of IFNg. Alternatively, through this study, we revealed that many of these targeted inhibitors are also capable of upregulating tumor cell MHC expression and IFNg secretion, which may, consequently, enhance tumor immune surveillance and recognition by cytolytic T-cells. Comprehensively, these data reveal T-cell- and tumor cell-mediated immunomodulatory functions of targeted small molecules that may be exploited clinically to synergize with checkpoint inhibitors and improve patient care.
Methods
Ultra-HTS 3D T-cell tumor spheroid-killing assay platform
To mimic T-cell-tumor cell interactions that occur in a clinical setting we used human donor peripheral blood mononuclear cells (PBMCs) that were matched to human colorectal cancer cells by their HLA-A02 alleles. Further, in order to create an expanded population of effector CD8+ memory T-cells, PBMCs were stimulated with a pool of virally derived peptides that induce viral antigen-dependent T-cell expansion. The same viral peptides were used to load tumor cell HLA-A02 molecules, ensuring interactions between T-cells and tumor cells were dependent upon antigen presentation by the tumor cells and recognition by memory T-cells through their TCRs. This method served as a surrogate for tumor neoantigen presentation and recognition by T-cells that might occur in patients. Primary human peripheral blood mononuclear cells (PBMCs) were obtained from a healthy 38-year-old donor and were characterized by PCR as HLA-A02 positive (Donor ID 4666, Hemacare). Frozen PBMCs were thawed and cultured in RPMI + 10% human serum + 1× pen/strep/anti-fungal at ~1 Mil cells/ml. PBMCs were stimulated with 1 µM CEF Viral Peptide Pool (Anaspec) for 48 h. On day 2, recombinant human (rh) IL-2 (Peprotech) was added at 0.5 ng/ml for 72 h, and cultures were replenished with rhIL-2 every 24 h. On day 5, rhIL-2 (0.5 ng/ml), rhIL-4 (0.5 ng/ml), and rhIL-7 (10 ng/ml) (all from Peprotech) were added, and cultures were incubated for a further 4 days, with cytokine replenishment every 48 h. Two days prior to harvest of T-cells, HCT116-EGFP cells (human colorectal cancer cells expressing the enhanced green fluorescent protein; engineered from parental HCT116 cells acquired from ATCC) were dispensed into 1536-well Ultra-Low Attachment Spheroid Microplates (Corning) using an automated liquid dispenser (GNF systems) at 1000 cells per well in 4 µl of T-cell media with either 1 µM CEF Peptide Pool or a negative control CEF Scramble Pool (JPT). Spheroid formation proceeded for 48 h. After 9 days of culture, PBMCs (heretofore referred to as T-cells) were harvested, counted, and stained with 0.5 µM CellTracker Deep Red Dye (ThermoFisher Scientific) according to the manufacturer’s instructions. T-cells (3000 cells/3 µl in T-cell media) were then dispensed into 1536-well Ultra-Low Attachment Spheroid Microplates containing the HCT116-EGFP spheroids. At this time, 70 nl of experimental test therapeutic compounds were dispensed using Echo Acoustic Transfer (Labcyte). Plates were incubated in a temperature-controlled tissue culture incubator at 37 °C with 5% CO2. After 3 days, plates were imaged using the ImageXpress Micro Confocal High-Content Imaging System (Molecular Devices) with a 488 nm laser for the HCT116-EGFP spheroid and a 561 nm laser for the T-cell population. Wells were read using a Z-stack series of 27 individual planes over a 200 µm vertical distance and images were analyzed using a maximum intensity projection of all 27 planes. Plates were again imaged on day 5 using the same plate parameters.
During experiments when either T-cells or tumor spheroids were treated with compounds separately in monoculture, the following method was used. T-cells were treated with experimental compounds for 48 h, then cultures were washed 3× with PBS before being resuspended in T-cell media and applied to tumor spheroid cultures as described earlier in the methods. For tumor spheroid monoculture treatment, tumor spheroids were treated with experimental compounds for 48 h, then washed 3× with PBS before the addition of T-cell media and T-cell cultures as described earlier in the methods.
Confocal tomography was conducted with an ImageXpress Micro Confocal (Molecular Devices, San Jose, CA) using a 60 µm Nipkow spinning disk. GFP and Cy5 filter sets optimized for LED excitation were used to acquire eGFP and CellTracker DeepRed signals. A sampling depth of 220 µm was obtained with a 10×0.45NA Plan Apochromat objective. Maximum intensity projections were created for high-content analysis (HCA), which was performed using MetaXpress Custom Module Editor. The HCA pipeline measured the total area of tumor spheroids and T cells using the segmentation of HCT116-GFP and CellTracker DeepRed, respectively. Additionally, the pipeline measured the quantity of HCT116 spheroids detected. This facilitated the identification of improper cell seeding and spheroid fractures. Quantitative HCA results were passed through a custom KNIME (KNIME, Zurich, Switzerland) data pipeline for treatment annotation and calculation of CMV peptide treated to scramble peptide treated HCT116 tumor spheroid area ratio.
Analysis of tumor spheroid 3D pathologies
HCT116 parental cells were used to create ultra-HTS 1536-well spheroids as described earlier in the methods. Spheroids were then stained with Image-iT™ Green Hypoxia Reagent to highlight areas of low oxygen and pHrodo™ Red, SE to highlight areas of low pH (ThermoFisher Scientific) according to the manufacturer’s instructions. Spheroids were then imaged using high-content microscopy as described earlier in the methods.
CMV viral peptide tracking assay
HCT116 human colorectal cancer cells (ATCC) were cultured in McCoy’s 5 A media + 10% FBS + 1× pen/strep/anti-fungal according to standard growth conditions. Cells were harvested, counted, and dispensed into a 384-well Black/Clear Bottom Polystyrene Spheroid Microplates (Corning) at 10,000 cells/well in 30 µl media in the presence or absence of 10 µg/ml recombinant human interferon-gamma (IFNg). Cells were incubated in a temperature-controlled tissue culture incubator at 37 °C with 5% CO2 for 48 h. After 48 h of spheroid growth, 2 µg/ml Brefeldin-A (Sigma-Aldrich) or a volume equivalent of the dimethyl sulfoxide (DMSO) vehicle was added and spheroids were incubated for 30 min. After 30 min, 1 nM CMV-BODIPY-SE (Anaspec) viral peptide was added with or without a cold CMV peptide competitor (10 nM). Spheroids were then incubated for a further 2 h. Spheroids were then washed 3× with Hank’s Balanced Salt Solution (HBSS) and stained with CellBrite 555 Membrane Dye (Biotium) according to the manufacturer’s instructions. Spheroids were then washed 3× with HBSS + 0.1% BSA and fixed in 1.5% paraformaldehyde for 20 min at room temperature (RT). Spheroids were then washed 3× with HBSS + 0.1% BSA and stained with 0.5× Draq5 nuclear dye (ThermoFisher). Spheroids were then washed 3 x with HBSS + 0.1% BSA and stored in 30 µl HBSS for imaging. Spheroids were imaged using the ImageXpress Micro Confocal High-Content Imaging System (Molecular Devices) with a 601 nm laser for Draq5 nuclear dye, a 488 nm laser for CMV-BODIPY-SE, and a 561 nm laser for CellBrite 555 membrane dye. CMV tracking experiments were performed in triplicate.
Flow cytometry analyses of T-cells and tumor cells
For T-cell phenotyping analyses, PBMCs were harvested at three different time points during stimulation and expansion (days 3, 6, and 9) and T-cells were characterized using flow cytometry. Suspension PBMCs were harvested, washed with FACS buffer (PBS + 2% FBS), and stained with the following antibodies: anti-CD4-PE (L200, BD Biosciences), anti-CD8-AF488 (RPA-T8, BD Biosciences), anti-CD127-BV421 (Biolegend), and Propidium Iodide (Invitrogen). Cells were then run through a FACSCalibur flow cytometer (BD Biosciences) and cell populations were analyzed using FloJo v10 software (FloJo LLC).
T-cells were also analyzed for their ability to bind MHC-presented CMV and EBV viral peptides. PBMCs were stimulated with either CMVpp65 (495–503) (Anaspec) or EBV BMLF1 (280–288) (Anaspec) viral peptides for 2 days, expanded in rhIL-2 (0.5 ng/ml), and harvested after 7 days of proliferation. Cells were washed 2x with FACS buffer and stained with the following antibodies: anti-CD8-APC-Cy7 (SK1, BioLegend), anti-CD3-PerCP-Cy5.5 (OKT3, BioLegend) as well as iTAg MHC Tetramer/PE-HLA-A*02:01 CMVpp65 (495–503) and MHC Tetramer/BV421-HLA-A*02:01 EBV BMLF1 (280–288) (both from MBL International Corp.) and Propidium Iodide (Invitrogen). Cells were then analyzed by flow cytometry as described earlier in the methods. For experiments where T-cells were treated with experimental compounds and phenotyped by flow cytometry, the following method was used. T-cells were treated with experimental compounds for 48 h, then cells were washed 2× with FACS buffer and stained with the following antibodies: anti-CD3-PerCP-Cy5.5 (OKT3, BioLegend), anti-CD4-PE (L200, BD Biosciences), anti-CD8a-APC/Cy7 (RPA-T8, Biolegend), anti-CD25-APC (M-A251, Biolegend), anti-CD279-PE/Cy7 (PD-1, EH12.287, Biolegend), anti-CD107a-BV605 (H4A3, Biolegend), anti-human Granzyme-B-FITC (GB11, BD Biosciences), anti-human Interferon-gamma-PerCP (4 S.B3, Invitrogen), anti-human TNFa-PE (MAB11, Biolegend), iTAg MHC Tetramer/PE-HLA-A*02:01 CMVpp65 (495–503) (MBL International Corp.) and Zombie-UV live/dead dye (Biolegend).
HCT116 human colorectal cancer cells were profiled for MHC expression by flow cytometry. HCT116-EGFP cells were grown in the presence or absence of 1 µg/ml rhIFNg for 24 h, washed with PBS, and harvested. Cells were washed 1× with FACS buffer and stained with the following antibodies: anti-HLA-A02-APC (BB7.2, BD Biosciences), anti-HLA-A/B/C (W6/32, eBioscience), mIgG2b-APC isotype control (BD Biosciences), and mIgG2a-PE isotype control (eBioscience). Cells were then washed 2x with FACS buffer and analyzed by flow cytometry as described earlier in the methods.
ELISA Spot (ELISpot) analysis of donor T-cell memory
Human donor HLA-A02 PBMCs were acquired from Hemacare. Viral peptides chosen for T-cell memory and reactivity were the CEF Peptide Pool (Anaspec), CMVpp65 (495–503), EBV BMLF1 (280–288) and the CEF Scramble negative control peptide pool (JPT), as well as PMA/Ionomycin as a positive control. ELISpot was performed with the VECTASTAIN ABC HRP Kit (Vector Laboratories) according to the manufacturer’s instructions, using a primary anti-human-IFNg (Mabtech) monoclonal antibody and a secondary IFNg-Biotin antibody on 96-well ELISpot plates (Millipore). ELISPOT AEC Substrate (BD Biosciences) was used to resolve spots on membranes and IFNg+ cells were evaluated in a blinded fashion (ZellNet Consulting, Inc.) using an ELISpot reader (KS ELISpot reader, Zeiss) with KS ELISpot software version 4.9.16. The plate evaluation process, including the setup of optimal reading parameters, followed the International Guidelines on ELISpot plate evaluation32.
Analysis of antigen-dependent T-cell-mediated tumor cell killing
Human PBMCs were thawed and stimulated with 1 µM CMVpp65 (495–503) viral peptide, then expanded as described earlier in the methods. HCT116-FL cells (stable line expressing firefly luciferase transgene) were seeded into 384-well white bottom plates (10,000 cells in 30 µl T-cell media per well) and grown for 48 h in the presence of identical CMVpp65 (495–503) viral peptide or a scramble negative control peptide in dose–response. 60,000 T-cells were then added to HCT116 wells (~3:1 T-cells:tumor cells) and the co-cultures were incubated for a further 48 h. After 48 h of co-culture, Brite-Glo reagent (Promega) was added according to the manufacturer’s instructions, and plates were read on an EnVision Multimode Plate Reader (PerkinElmer). HCT116-derived firefly luciferase units were used as a surrogate for cell viability.
Statistics and reproducibility
Independent sample t-tests were used to calculate probability between two different groups. Experiments typically consisted of three or more biological replicates and studies were often repeated three or more times to determine the robustness of data. For HTS assays, four or more biological replicates were often profiled per experiment.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data sets generated during and/or analyzed during HTS campaigns of the current study are not publicly available due to proprietary activities of compounds with known targets or molecular mechanisms of action but are available from the corresponding author on reasonable request. High-throughput compound screening data are currently stored on the GNF (Novartis) cloud-based data storage system. Non-HTS data has been supplied with the manuscript under Supplementary Data 1.
Change history
27 August 2021
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1038/s42003-021-02569-1
References
Jochems, C. & Schlom, J. Tumor-infiltrating immune cells and prognosis: the potential link between conventional cancer therapy and immunity. Exp. Biol. Med. 236, 567–579 (2011).
Mao, Y. et al. The prognostic value of tumor-infiltrating lymphocytes in breast cancer: a systematic review and meta-analysis. PLoS ONE 11, e0152500 (2016).
Brahmer, J. et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N. Engl. J. Med. 373, 123–135 (2015).
Hodi, F. S. et al. Improved survival with ipilimumab in patients with metastatic melanoma. N. Engl. J. Med. 363, 711–723 (2010).
Johnson, J. I. et al. Relationships between drug activity in NCI preclinical in vitro and in vivo models and early clinical trials. Br. J. Cancer 84, 1424–1431 (2001).
Sanchez-Paulete, A. R. et al. Antigen cross-presentation and T-cell cross-priming in cancer immunology and immunotherapy. Ann. Oncol. 28, xii74 (2017).
Peranzoni, E., Rivas-Caicedo, A., Bougherara, H., Salmon, H. & Donnadieu, E. Positive and negative influence of the matrix architecture on antitumor immune surveillance. Cell. Mol. Life Sci. 70, 4431–4448 (2013).
Coulie, P. G., Van den Eynde, B. J., van der Bruggen, P. & Boon, T. Tumour antigens recognized by T lymphocytes: at the core of cancer immunotherapy. Nat. Rev. Cancer 14, 135–146 (2014).
Marincola, F. M., Jaffee, E. M., Hicklin, D. J. & Ferrone, S. Escape of human solid tumors from T-cell recognition: molecular mechanisms and functional significance. Adv. Immunol. 74, 181–273 (2000).
Horman, S. R. et al. Functional profiling of microtumors to identify cancer associated fibroblast-derived drug targets. Oncotarget 8, 99913–99930 (2017).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Fischer, K. et al. Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood 109, 3812–3819 (2007).
Hirschhaeuser, F. et al. Multicellular tumor spheroids: an underestimated tool is catching up again. J. Biotechnol. 148, 3–15 (2010).
Ramgolam, K. et al. Melanoma spheroids grown under neural crest cell conditions are highly plastic migratory/invasive tumor cells endowed with immunomodulator function. PLoS ONE 6, e18784 (2011).
Budhu, S. et al. CD8+ T cell concentration determines their efficiency in killing cognate antigen-expressing syngeneic mammalian cells in vitro and in mouse tissues. J. Exp. Med. 207, 223–235 (2010).
Herter, S. et al. A novel three-dimensional heterotypic spheroid model for the assessment of the activity of cancer immunotherapy agents. Cancer Immunol. Immunother. 66, 129–140 (2017).
Schmittnaegel, M. et al. Committing cytomegalovirus-specific CD8 T cells to eliminate tumor cells by bifunctional major histocompatibility class I antibody fusion molecules. Cancer Immunol. Res. 3, 764–776 (2015).
Kennell, A. S., Gould, K. G. & Salaman, M. R. Proliferation assay amplification by IL-2 in model primary and recall antigen systems. BMC Res. Notes 7, 662 (2014).
Vella, A., Teague, T. K., Ihle, J., Kappler, J. & Marrack, P. Interleukin 4 (IL-4) or IL-7 prevents the death of resting T cells: stat6 is probably not required for the effect of IL-4. The. J. Exp. Med. 186, 325–330 (1997).
Canham, S. M. et al. Systematic chemogenetic library assembly. Cell Chem. Bio. 27, 1124–1129 (2020).
Thakuri, P. S., Gupta, M., Plaster, M. & Tavana, H. Quantitative size-based analysis of tumor spheroids and responses to therapeutics. Assay. Drug Dev. Technol. 17, 140–149 (2019).
Bettelli, E., Dastrange, M. & Oukka, M. Foxp3 interacts with nuclear factor of activated T cells and NF-kappa B to repress cytokine gene expression and effector functions of T helper cells. Proc. Natl Acad. Sci. USA 102, 5138–5143 (2005).
Mineharu, Y. et al. Gene therapy-mediated reprogramming tumor infiltrating T cells using IL-2 and inhibiting NF-kappaB signaling improves the efficacy of immunotherapy in a brain cancer model. NeuroTherapeutics 9, 827–843 (2012).
Ito, S.-e, Shirota, H., Kasahara, Y., Saijo, K. & Ishioka, C. IL-4 blockade alters the tumor microenvironment and augments the response to cancer immunotherapy in a mouse model. Cancer Immunol., Immunother. 66, 1485–1496 (2017).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Riganti, C. et al. Bromodomain inhibition exerts its therapeutic potential in malignant pleural mesothelioma by promoting immunogenic cell death and changing the tumor immune-environment. Oncoimmunology 7, e1398874 (2018).
Adeegbe, D. O. et al. Synergistic immunostimulatory effects and therapeutic benefit of combined histone deacetylase and bromodomain inhibition in non-small cell lung cancer. Cancer Discov. 7, 852–867 (2017).
Taipale, J. et al. Effects of oncogenic mutations in smoothened and patched can be reversed by cyclopamine. Nature 406, 1005–1009 (2000).
Chen, J. K., Taipale, J., Cooper, M. K. & Beachy, P. A. Inhibition of Hedgehog signaling by direct binding of cyclopamine to smoothened. Genes Dev. 16, 2743–2748 (2002).
Onishi, H. et al. The Hedgehog inhibitor cyclopamine impairs the benefits of immunotherapy with activated T and NK lymphocytes derived from patients with advanced cancer. Cancer Immunol. Immunother 62, 1029–1039 (2013).
Onishi, H. et al. Hedgehog signaling regulates PDL-1 expression in cancer cells to induce anti-tumor activity by activated lymphocytes. Cell. Immunol. 310, 199–204 (2016).
Janetzki, S. et al. Guidelines for the automated evaluation of Elispot assays. Nat. Protoc. 10, 1098–1115 (2015).
Acknowledgements
We would like to thank Kyle Scharkss and Chris Trussell for technical assistance with flow cytometry and Tiffany Chu for technical assistance with image analyses. We would like to thank Angelica Mendiola for preparing the virus. We would also like to thank John Hoerter and Tony Orth for their intellectual contributions. Lastly, we would like to thank Christopher McAllister for assistance with figure esthetics.
Author information
Authors and Affiliations
Contributions
J.T. performed all experimental procedures and wrote the manuscript. D.Q. designed high-content imaging screening and metrics. E.R. contributed to the design of the platform. L.L. and C.R. performed much of the flow cytometry studies. F.L. analyzed the imaging data and SRH designed the platform, analyzed results, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail:https://doi.org/10.1038/s42003-021-02569-1
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
To, J., Quackenbush, D., Rowell, E. et al. RETRACTED ARTICLE: A biomimetic assay platform for the interrogation of antigen-dependent anti-tumor T-cell function. Commun Biol 4, 56 (2021). https://doi.org/10.1038/s42003-020-01565-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01565-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.