Abstract
Proteins covalently attached to DNA, also known as DNA–protein crosslinks (DPCs), are common and bulky DNA lesions that interfere with DNA replication, repair, transcription and recombination. Research in the past several years indicates that cells possess dedicated enzymes, known as DPC proteases, which digest the protein component of a DPC. Interestingly, DPC proteases also play a role in proteolysis beside DPC repair, such as in degrading excess histones during DNA replication or controlling DNA replication checkpoints. Here, we discuss the importance of DPC proteases in DNA replication, genome stability and their direct link to human diseases and cancer therapy.
Similar content being viewed by others
Introduction
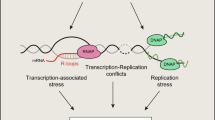
Our genome is constantly exposed to various forms of DNA damage. DNA–protein crosslinks (DPCs) are common lesions and form when a protein—of any size and nature—becomes covalently bound to DNA after exposure to a physical or chemical crosslinker. They are bulky and impose physical obstacles to various DNA metabolic processes, such as DNA replication, repair, transcription and recombination (Fig. 1). Thus, the fidelity of DNA replication across DPCs and the accuracy of DPC repair pathways are pivotal in avoiding genomic instability, which can lead to ageing-associated diseases such as cancer. Whereas the consequences of unrepaired DPCs are well appreciated, the components, regulation and dynamics of the DPC repair pathway(s) are far from being understood.
a DPCs on the leading or lagging strand can pose impediments for helicase and/or polymerase progression. b Proteolysis removes the bulk of this obstacle (DPC), and reduces the crosslinked protein to a peptide (remnant) that can be bypassed by TLS polymerase, thus resuming DNA replication. c DPCs can block transcription, but the mechanisms of transcription-dependent DPC proteolysis have not been explored.
DPCs are essentially removed by either canonical nucleases or dedicated proteases that degrade the protein component of the DPC (DPC proteolysis)1,2. DPC proteolysis has gained recognition with the discovery of the metalloproteases Wss1 (weak suppressor of Smt3-1) in yeast and SPRTN (SprT-like N-terminal domain, also known as Spartan or DVC-1) in higher eukaryotes3,4,5,6. Both proteases possess an intrinsic metalloprotease active center in specialized but phylogenetically distinguished domains (Fig. 2): the WLM (Wss1p-like metalloproteases) domain in Wss1 and SprT domain in SPRTN1. Wss1 and SPRTN depend on DNA binding for their proteolytic activity but have no defined sequence specificity. This feature is advantageous considering the heterogeneous nature of crosslinked proteins. However, it also exposes chromatin-associated and potentially functional proteins to the risk of undesired proteolysis. Thus, DPC proteolysis must be tightly regulated to minimize such risks. A comprehensive understanding of these regulatory modes is still lacking7. Remarkable progress in the field of DPC repair has also been made in recent studies on emerging DPC proteases with functional overlap to Wss1 and SPRTN. These proteolytic activities are primarily employed in DPC repair.
Domains/motifs for interactions are color-coded. Red: protease domains. Orange: Cdc48/p97 interaction. Green: PCNA interaction. Ochre: DNA binding. Blue: SUMO/ubiquitin/proteasome binding. The catalytic residues in the protease domains are underlined. The sites of disease-associated mutations in SPRTN (RJALS)60, FAM111A (KCS2)118,119, and FAM111B (POIKTMP)47 are marked in red. The asterisk in SPRTN indicates the site of nonsense mutations in RJALS patients that generate the protein SPRTN-ΔC60. RVP, retroviral protease. U1 and U2, potential UBLs in FAM111A61.
Besides DPCs, other types of substrates have been described for these proteases, for example tightly bound, albeit not crosslinked, chromatin proteins with the potential to disrupt the fidelity of DNA replication, like trapped poly (ADP-ribose) Polymerase-1 (PARP-1). It is therefore becoming increasingly clear that DPC proteases guard multiple aspects of genome maintenance.
In this review, we briefly discuss DPC formation, and expand on DPC proteases and the first and essential step of DPC repair: proteolysis of the protein component. We revise how DPC proteases contribute to genome maintenance, especially during DNA replication, and why defects in their activities lead to human disease.
DNA–protein crosslinks
DPCs originate when proteins become crosslinked to DNA after exposure to physical or chemical agents, such as UV light or aldehydes, respectively (non-enzymatic DPCs), or as a result of faulty enzymatic reactions (enzymatic DPCs)8. Enzymatic DPCs are well exemplified by Topoisomerase-1 and Topoisomerase-2 cleavage complexes (Topo-1ccs, Topo-2ccs). During the physiological reaction of Topoisomerase on DNA, a transient, covalent intermediate (i.e., cleavage complex) forms between the catalytic tyrosine residue and the DNA phosphate group (phosphotyrosyl linkage). Stabilization of the cleavage complex (and formation of a DPC) can happen spontaneously if DNA is damaged, but is enhanced in the presence of poisons, e.g., camptothecin (CPT) or etoposide, for Topo-1 or Topo-2, respectively9,10. Notably, Topo-1/2 poisons are widely exploited in cancer chemotherapy2,11. Enzymatic DPCs also include crosslinks of DNMT1 (DNA methyltransferase 1) to the DNA methylation inhibitor 5-aza-2’-deoxycytidine (5azadC) incorporated into DNA12,13, and of HMCES (5-Hydroxymethylcytosine binding, ES-cell-specific) to abasic sites in single-stranded DNA14.
In the case of non-enzymatic DPCs, virtually any protein—of variable size, structure and nature—in the vicinity of DNA can be crosslinked. One of the most potent crosslinkers, formaldehyde (FA), is heavily present in the environment and produced endogenously via processes like lipid peroxidation, and DNA, RNA or histone demethylation15,16,17,18,19. Hence, FA release can occur in the surroundings of DNA, implying that DPCs form continuously and cells must constantly overcome DPC-induced toxicity.
Defective DPC repair leads to sensitivity to crosslinking agents, faulty DNA replication and cell cycle abnormalities, which pave the way for chromosomal instability and carcinogenesis in humans and mice20. Hence, multiple pathways work to ensure a proper response to these insults. Nuclease-dependent mechanisms, like nucleotide excision repair (NER) and homologous recombination, operate in both bacteria and eukaryotic cells by excising the DNA flanking the DPC21,22,23,24,25. However, NER seems to have a fairly limited role in overall DPC repair, as it can only remove small DPCs (8–10 kDa in size in mammalian cells)5,23,26. Topo-1/2ccs can be excised by dedicated tyrosyl-DNA phosphodiesterases, TDP1 and TDP2, which cleave the phosphotyrosyl linkage27; this is normally shielded and becomes accessible to TDPs after partial proteolysis or structural changes of Topoisomerases28,29,30,31,32.
A much more pliable, less discriminate way to process DPCs is through proteolysis of the protein component operated by specialized DPC proteases, namely DPC proteolysis repair.
DPC proteases
Wss1 and SPRTN: the first members of a class of repair enzymes discovered
DPC proteolysis repair was discovered in yeast with the metalloprotease Wss13. A concomitant study in Xenopus egg extract hinted at a similar pathway in metazoans33. The dedicated enzyme in metazoans was later found to be SPRTN4,5,6. However, phylogenetic analysis has revealed that Wss1 and SPRTN are not orthologs but functional homologs1,11,34. The similarity between their sequences is limited within the protease domains—WLM and SprT—and around the active center typical of metalloproteases (HExxH), and they show 24% identity5 (Fig. 2).
Wss1 and SPRTN confer resistance to FA, showing a general role in DPC repair3,4,5,6. This is also supported by in vitro cleavage studies, which demonstrate that Wss1 and SPRTN can process DNA-binding proteins of variable size and structure, exemplified by the smaller histone H3 and the larger Topo-23,4,5,6. Although Wss1 is capable of processing purified Topo-1, WSS1 deletion alone does not sensitize yeast cells to CPT, in striking contrast with the effects of SPRTN depletion in mammalian cells3,5. This is due to the existence of a yeast protease with a similar function and will be discussed below. A good explanation for this promiscuity came from structural studies of the yeast WLM domain35. As the protease domain lacks a substrate-binding pocket, it can accommodate a vast range of structures. DPC proteolysis leaves a remnant peptide, still of unidentified size, attached to DNA. Although less problematic for DNA replication and transcription, this remnant must be repaired via other activities (e.g., nucleases, tyrosyl-DNA phosphodiesterases) in coordination with or post-proteolysis. How this is achieved is not well understood.
Active DPC proteases are potentially dangerous and cells adopt strategies to restrain their activities7. First, Wss1 and SPRTN bind to and are activated by single-stranded (ss) and double-stranded (ds) DNA3,4,5,6,36,37,38; this feature protects nuclear soluble proteins from unwanted proteolysis. The structure of the SprT domain from human SPRTN suggests that the DNA dependence relies on the active site being shielded in the absence of DNA; DNA binding exposes the narrow groove with catalytic residues for peptide cleavage37. Second, Wss1 and SPRTN are capable of in trans self-cleavage in the presence of DNA; this property is a regulatory mechanism to shut down the protease activity when at the chromatin4. Third, the activity of DPC proteases is further sharpened by post-translational modifications (phosphorylation, ubiquitylation, SUMOylation, acetylation) of the DPC proteases and/or their substrates, and interaction with partner proteins, for instance the sliding clamp PCNA (proliferating cell nuclear antigen), the ATPase p97 or the Topo-1 binding protein TEX264. The latter point will be expanded in a separate paragraph.
Emerging DPC proteases in genome stability
Recently, proteases other than SPRTN have emerged as potential DPC repair enzymes. These are ACRC/GCNA, FAM111A and B, DDI1 and DDI2 in human, and their respective orthologs. Among the human enzymes, SPRTN is the only essential gene in variety of cancer cell lines (Table 1; www.depmap.org).
These emerging DPC proteases have been linked to processing of DPCs and tightly bound proteins (Figs. 2 and 3). Besides the protease domain, where conserved catalytic residues map (Fig. 2), they share remarkable similarities in overall domain organization, which underlies potentially similar patterns of regulation. Nonetheless, some proteases might be more specific for a certain cell cycle phase, a developmental stage, or a class of substrates.
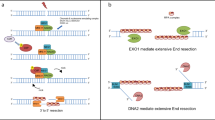
a Wss1 and Ddi1 are both required for the repair of non-enzymatic DPCs and Topo-1ccs during replication. b FAM111A and SPRTN have both been implicated in Topo-1cc resolution, based on CPT resistance. c Sensitivity to PARP-1 inhibitors and DNA combing assays suggest that FAM111A, but not SPRTN, is involved in the resolution of trapped PARP-1. d Human GCNA resolves DNMT1 crosslinks at 5-azadC incorporation sites (red arrowhead), behind the replication fork. e GCNA resolves Topo-2ccs in flies, worms and zebrafish.
Germ cell nuclear antigen; also known as acid repeat-containing protein (ACRC)]
Germ cell nuclear antigen (GCNA) contains a SprT metalloprotease domain conserved across eukarya11,39. In metazoans, it is expressed in germ cells, where it was recently shown to target Topo-2ccs40,41. GCNA ectopically expressed in mammalian culture cells has also been involved in the resolution of 5-azadC-induced DNMT1 crosslinks42 (Fig. 3). GCNA is not essential in mammalian cell lines (Table 1), but its knock-out (KO) increases the chances of embryonic lethality in flies, worms and zebrafish40.
DNA damage inducible 1 (DDI1) and DDI2 proteins
DDI1 and DDI2 proteins are aspartic proteases that interact with ubiquitin and the proteasome via a ubiquitin-like domain (UBL)43,44 (Fig. 2). They promote proteasome-dependent replication fork restart after replicative stress through degradation of replication termination factor 2 (RTF2)44. A direct involvement in DPC repair has not yet been explored, however the S.cerevisiae homolog Ddi1 was recently shown to aid Wss1 in resolution of CPT-induced and FA-induced DPCs45,46.
Family with sequence similarity 111 member A (FAM111A)
FAM111A and its homolog FAM111B are serine proteases. Not much is known about FAM111B, except for its association with the etiology of a form of poikiloderma with pulmonary fibrosis (POIKTMP)47. FAM111A was previously described as a PCNA interactor and for its role in restricting viral replication in host cells48,49,50. Only recently has it been implicated in the repair of Topo-1ccs and trapped PARP-151 (Figs. 3 and 4).
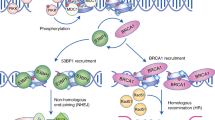
a Wss1 cleaves excess histones binding to ssDNA after fork stalling. ssDNA is formed after HU. b PARP-1 is recruited to DNA to repair different types of damage, including base adducts generated by alkylating agents. In the presence of PARP inhibitors, self-PARylation and chromatin dissociation are blocked. Trapped PARP-1 represents a barrier for replication. FAM111A confers resistance to PARP inhibitors, but whether it acts proteolytically on PARP-1 awaits formal verification. c During DNA replication SPRTN cleaves the C-terminus of Chk1 and releases the active kinase domain (K) from chromatin.
Evidence that GCNA, DDIs and FAM111A cleave DPC substrates is currently lacking. However, they do possess catalytic activity. FAM111A self-cleavage has been shown in vitro and in cells51,52; ScDdi1 and HsDDI2 process substrates modified with long ubiquitin chains in vitro53,54. Moreover, complementation studies show that mutations in the putative catalytic residues of GCNA40, ScDdi145, and FAM111A51,52 generate non-functional proteins. For ScDdi1, the in vitro activity has been tested in the presence of dsDNA, an activator of Wss1 and SPRTN, without success45. Although DNA-dependent activity has been established as a regulatory mechanism for Wss1 and SPRTN, different proteases might respond to different modes of regulation.
A common feature among DPC proteases is DNA binding, a confirmation of their chromatin-related function. Minimal DNA binding domains (DBDs, Fig. 2) have been mapped by testing truncated versions in either DNA binding assays (e.g., EMSA)—as for Wss1, SPRTN, and FAM111A3,4,5,37,51,55—or complementation studies in cells—as for Ddi1’s HDD (helical domain of Ddi1)45,46. Like SPRTN and Wss1, FAM111A can bind ssDNA, and the integrity of this domain is necessary for in vitro self-cleavage, as well as its function in cells51. A possible interpretation for the ssDNA dependent-proteolysis will be elaborated later.
Overlapping functions of DPC proteases
The existence of different DPC proteases indicates that cells have invested in multiple enzymes to counteract DPC-dependent toxicity (Fig. 3). This comes as little surprise considering the heterogeneous nature of crosslinked proteins. Although SPRTN does not have strict sequence specificity, it preferentially cleaves DNA-binding proteins in poorly structured regions rich in lysine, arginine and serine residues5. The existence of alternative proteases would ensure that crosslinked proteins lacking these features are efficiently processed to prevent genomic instability. Here, we highlight the functional overlap between Wss1, SPRTN and the other proteases.
In metazoans, GCNA is predominantly expressed in germ cells and early embryos39. Therefore, most of the studies have been conducted in animal models rather than human cells. In Drosophila and C.elegans, GCNA mutation exposes germ cells and embryos to replication stress—formation of RPA/γH2Ax foci and hydroxyurea (HU) sensitivity—and genomic instability, e.g., chromosome segregation defects and micronuclei, which overall limit reproductive success40,41. C.elegans fertility defects are exacerbated by concomitant mutations in the SPRTN homolog dvc-140,41. Drosophila embryos mutated in both Gcna and mh (maternal haploid, mh, is the Drosophila SPRTN homolog) do not complete embryogenesis40. Setting a role in DPC repair along with DVC-1/mh, GCNA deletion increases total DPCs in germ cells and early embryos of flies, worms and zebrafish40, with Topo-2cc being among the most abundant40,41. Dvc-1 and gcna-1 mutant worm embryos are equally sensitive to formaldehyde42. Overall, this indicates an overlap between GCNA and DVC-1 at the organismal level. Germ cells and embryos are particularly vulnerable to the genomic instability resulting from DPC accumulation because mistakes would be inherited. Also, changes in gene expression and histone demethylation during embyogenesis56 might especially expose germ cells to FA release17 and DPCs formation, which explains the need for having multiple DPC proteases.
S.cerevisiae Ddi1 and human FAM111A are required for tolerance to Topo-1ccs, a common target of yeast Wss1 and human SPRTN. Unlike wss1 mutation alone, the deletion of both WSS1 and DDI1 sensitizes yeast cells to CPT and identifies Ddi1 as the elusive, redundant protease for Topo-1cc repair3,45. Genetic data additionally indicate that Ddi1 gives resistance to the DPC-inducing agent FA, along with Wss145,46.
SPRTN and FAM111A also have overlapping functions. Both proteins have been found at nascent DNA5,50,57 and both prevent replication fork stalling in the presence of crosslinking agents, specifically, FA for SPRTN5,58 and CPT for both FAM111A and SPRTN51. FAM111A depletion does not appear to reduce fork speed in unchallenged conditions, in contrast to SPRTN mutation5,51,59,60,61. Thus, FAM111A might come into play when the DPC overload exceeds the capacity of SPRTN, which is expressed at low levels. Such interplay is purely speculative at present. Interestingly, FAM111A KO but not SPRTN hypomorphic cells are sensitive to PARP-1 inhibitors, which were shown to trap PARP to DNA, suggesting that some DPC proteases might have preference for certain substrates51.
Regulation of DPC proteases by post-translational modifications
Post-translational modifications (PTMs) have great regulatory potential, and most of the proteases discussed here can bind ubiquitin, via ubiquitin-associated domain (UBA) or ubiquitin-binding zinc finger (UBZ), or SUMO (Small ubiquitin-like modifier) via SUMO-interacting motif (SIM) (Fig. 2). Treatment of cells with DPC inducing agents, like FA and 5-azadC, induces signaling cascades, which culminate with the modification of the crosslinked proteins by ubiquitin and SUMO42,62,63. We can predict that recruitment to or persistence on the damage site is modulated by PTMs. GCNA, for example, localizes to DNMT1 foci that form after 5-azadC treatment; this relocation depends on SUMOylation and GCNA’s SIMs42 (Fig. 2). SPRTN forms nuclear foci after exposure to FA and fails to do so upon pharmacological inhibition of the ubiquitylation pathway42,62. Whether this relocation depends on its UBZ remains to be demonstrated. This initial evidence, substantiated by the predominance of SUMO-interacting and ubiquitin-interacting domains, proves that DPC proteases have the potential to be regulated via interaction with post-translationally modified proteins.
DPC proteases can themselves be modified for regulatory purposes. SPRTN exists in a mono-ubiquitylated form, but exposure to FA leads to its de-ubiquitylation—two different studies have identified the de-ubiquitylating enzymes VCPIP and USP11—and acetylation to allow recruitment to chromatin4,64,65.
Wss1 and SPRTN interact with the ATPase Cdc48/p97 also known as valosin-containing protein (VCP) in mammals, via SHP (suppressor of high copy PP1)-box and VCP-interacting motif (VIM)3,36,66,67,68 (Fig. 2). Cdc48/p97 is a ubiquitin-dependent and SUMO-dependent chaperone acting with specific cofactors to unfold substrates for proteasomal degradation or disassembly from various macromolecules69,70. The unfolding activity of p97 directly aids repair of Topo-1ccs, to which p97 is recruited via its novel cofactor TEX264 (Testis Expressed 264) before SPRTN-mediated proteolysis68. Whether and how unfoldases are a constitutive requirement of DPC proteolysis repair remains to be established.
DPC proteases and DNA replication
DPCs can stall the CMG (Cdc45-Mcm2-7-Gins) replicative DNA helicase complex and/or DNA polymerase during DNA replication, depending on their size and location (leading or lagging strand)33,71 (Fig. 1). Actively replicating cells are more susceptible to FA and CPT than non-replicating cells5,72,73. If left unrepaired, these replication obstacles can cause prolonged fork stalling that leads to fork collapse, formation of double-strand breaks (DSBs) and genomic instability74.
In yeast, Wss1 has been proposed to allow completion of DNA replication after FA exposure3, and Ddi1 is recruited to the damage site for removal of a model DPC in synchronized, S phase yeast cells45. However, repair of DPCs in a DNA replication-dependent manner has been more directly shown for SPRTN, both in human cells5 and in Xenopus egg extract75,76.
In line with S phase repair, SPRTN levels are higher in S/G2 phase—due to APC (Anaphase promoting complex)-Cdh1-dependent degradation in G1 phase66—when it localizes directly at the replication fork5,57. SPRTN interacts with PCNA via a PIP (PCNA-interacting protein)-box (Fig. 2), which is dispensable for SPRTN recruitment to the replication fork. A truncation variant lacking the C-terminus (and therefore the PIP-box) still localizes at nascent DNA in iPOND (isolation of proteins on nascent DNA) experiments57 and rescues DNA fiber length60. Thus, it is unclear how SPRTN interacts with the replication machinery for DPC repair.
In vitro, substrate processing by SPRTN is most effectively stimulated by ssDNA—with the potential to form hairpins—or dsDNA structures with nicks or gaps close to the crosslinked protein4,37,38. This is consistent with results obtained in Xenopus egg extracts using a DPC plasmid, where SPRTN cleaves a model DPC without ongoing replication when there is a gap in the complementary DNA filament75. During DNA replication in cells, uncoupling between helicase and stalled DNA polymerase exposes ssDNA and generates a dsDNA/ssDNA ‘hybrid’ that is compatible with the in vitro model. This specific DNA context presumably protects non-crosslinked chromatin-associated proteins from SPRTN activity38.
GCNA is also active in S phase. Its role during DNA synthesis is supported by the replication stress (formation of RPA/γH2Ax foci) in mitotically active germ cells of GCNA mutant flies and worms40, and by the fact that the Mcm2-7 components of the CMG helicase complex figure among the DPCs in GCNA mutant fly embryos40. Another study links GCNA to the resolution of DNMT1 crosslinks, although this was shown with ectopically expressed GCNA in human somatic cells42. DNMT1 crosslinks form behind the replication fork, when DNMT1 is recruited to newly synthesized DNA to restore the DNA methylation pattern, where it can get crosslinked in the presence of 5-azadC. It remains to be determined whether effective DNMT1 crosslink repair needs GCNA catalytic activity.
FAM111A levels increase in late S phase and remain high during G2/M48. FAM111A interacts with PCNA via a PIP-box (Fig. 2)50,52. It is unclear to what extent FAM111A depletion impairs DNA replication. EdU incorporation studies have produced conflicting results50,52,61. DNA combing assays in the absence of exogenous damage have either shown no reduction in the length of nascent DNA in FAM111A-depleted cells or a slight increase in tract length to compensate for the decrease in origin firing51,61. Instead, FAM111A KO reduces replication speed upon treatment with CPT and PARP inhibitors and increases cellular sensitivity to these poisons51. Consistently, Topo-1cc foci accumulate in KO cells; an intact PIP-box and an intact protease domain are required to rescue these phenotypes51. Although in vitro proteolysis of Topo-1 has not been shown so far, FAM111A may be required for the removal of replication barriers. In this respect, the extent of overlap with SPRTN remains to be determined.
Strikingly, FAM111A overexpression causes EdU incorporation and cell cycle defects, and consequently increases the rate of DNA damage and apoptosis; this is probably due to unspecific cleavage of replication fork proteins, e.g., PCNA and RFC (Replication factor C)50,52,61,77. However, overexpression of the PIP-box mutated variant recapitulates these phenotypes to some extent, arguing that binding to PCNA might not be the cause of the replication problems50,52,61. These replication defects are exacerbated by overexpression of FAM111A variants carrying disease-causing mutations (Kenny–Caffey syndrome type 2, see below and Fig. 2)52,61,77. Overexpression of FAM111B mutant variants, causative of POIKTMP (Fig. 2), largely mimics the phenotype of FAM111A overexpression52. In both cases, concomitant mutations in the protease domains abolish these defects, showing that the observed phenotypes are the direct cause of FAM111A/B-dependent proteolysis52. These data suggest that deregulated FAM111A/B enzymatic activity is detrimental to the cells. Thus, tight regulation of the level and activity of these proteases is critical for the successful completion of DNA replication.
Downstream pathway and translesion DNA synthesis
Translesion DNA synthesis (TLS) is a damage tolerance mechanism where the conventional DNA polymerase is exchanged for an error-prone TLS polymerase that bypasses the lesion. This switch depends on PCNA and its ubiquitylation by the E3 ubiquitin ligase Rad1878. SPRTN was linked to TLS regulation after UV damage before its role in DPC repair was established66,67,79,80,81,82. SPRTN interacts with both PCNA and ubiquitin, but how SPRTN regulates TLS is not entirely clear. Some studies suggest it is a positive regulator of TLS79,81,82, while others propose that SPRTN prevents UV-induced, TLS-dependent mutagenesis66,67,80.
Although TLS regulation by SPRTN was initially linked to UV lesion repair, it is plausible that bypass of the remnant peptide after bulk DPC proteolysis also requires TLS (Fig. 1). Some lines of evidence support this: (i) UV light can crosslink proteins to DNA83,84,85; (ii) epistasis analysis shows that Rad18 and SPRTN work together58; (iii) in Xenopus egg extracts, TLS polymerases are required to complete replication of a plasmid carrying a model DPC after digestion by SPRTN75; (iv) in yeast, WSS1 deletion lowers mutation rates after FA3. Therefore, TLS might act immediately after DPC proteolysis to avoid fork stalling, since mutations are still more desirable than replication fork collapse.
There is one case where the accumulation of mutations can be beneficial. This is the hypermutation of immunoglobulin loci. SPRTN has been shown to favor mutations (DNA polymerase eta and zeta-dependent) at abasic sites and template switch-mediated gene conversion during immunoglobulin gene diversification in chicken B cells86. Notably, abasic sites are a common source of DPC formation, and the hypermutation phenotype is consistent with TLS acting downstream of DPC proteolysis. In agreement, lower mutation rates at the immunoglobulin locus are observed in SPRTN-depleted cells86. This suggests that SPRTN plays a role in generating the diversity of immunoglobulin repertoire and in the DNA damage tolerance pathway when DNA replication forks get stalled.
DPC repair outside S phase
Despite the numerous bodies of evidence linking SPRTN to replication-coupled DPC proteolysis, a replication-independent function of SPRTN has not been ruled out4,5. SPRTN mutations or insufficiency manifest into hepatocellular carcinoma57,60. The liver might be more exposed than other organs to DPCs because inhaled and ingested compounds can be catabolized into formaldehyde. However, the great majority of hepatocytes are quiescent87, therefore it is plausible that SPRTN is operating outside of S phase, and that malfunctioning would lead to cancer. It is also possible that liver damage observed in SPRTN-mutated patients or SPRTN hypomorphic mice initiates during embryogenesis’ S phase but this defect emerges later.
Starvation-arrested worm larvae, where replication is not happening (cells are arrested in G1/S), are sensitive to formaldehyde4, indicating that processes other than DNA synthesis are affected. Another example of SPRTN activity outside S phase comes from Drosophila. The expression of a mutated mh (the SPRTN homolog) from the maternal genome in the zygote leads to loss of the paternal DNA during the first mitotic division and results in the lethality of haploid embryos—hence the name mh, “maternal haploid”88,89. This loss is caused by the lack of condensation of the paternal DNA during mitosis, and it is inherently unlinked to DNA replication and S phase88. The paternal DNA remodeling still relies on mh catalytic activity, but the mechanism is unclear88,89.
GCNA also works outside S phase. Its expression peaks in mitosis, where it localizes on chromosomes41. Topo-2ccs are prominent GCNA targets40,41. Topo-2ccs are abundant in mitosis because Topo-2 activity is required for separation of sister chromatids. Supporting a role for GCNA in Topo-2cc repair, GCNA and Topo-2 physically interact and colocalize, and mutant worms are sensitive to etoposide but not to camptothecin41.
Overall, these data confirm that DPC protease activity is essential to maintain genome integrity during DNA replication, as well as outside of S phase.
DPC proteases and transcription
DPCs are also potential obstacles for transcription20. Although transcriptional stalling and mutagenesis can be threatening for genome stability90, the effects of DPCs on the progression of RNA polymerases are a largely neglected field.
Some evidence linking DPC proteases and transcription has emerged. Rpb1, the largest and catalytic subunit of the RNA polymerase II, is degraded via Ddi1 and Wss1 after exposure to HU or UV light45. Both proteases interact with Rpb1. However, this evidence provides insufficient proof that Ddi1 and Wss1 target Rpb1 as a crosslinked protein. Its degradation might be a consequence of transcription stalling due to roadblocks ahead of RNA polymerase II. In fact, under these circumstances Rpb1 degradation was described to rely on the ubiquitin-proteasome system, the ATPase Cdc48/p97 and SUMO91,92. At any rate, a stalled, non-crosslinked Rpb1 would be equally “eligible” for degradation by DPC proteases, since other “non-DPC” substrates exist and will be discussed later in this review.
A link between transcription and FAM111A in human cells also exists. FAM111A overexpression reduces Rpb1 levels at the chromatin and, consequently, EU incorporation52. FAM111A also interacts with Rpb1 upon overexpression52. However, it is unclear if these phenotypes are related to direct Rpb1 proteolysis.
DPCs and proteasome
The most studied proteolytic machinery in the cell is the 26S proteasome (henceforth proteasome), which degrades proteins that have been previously modified by ubiquitin and unstructured by unfoldases93,94. It is not surprising that proteasome activity has been tested towards DPCs26. The involvement of the proteasome implies that the DPC must be modified with ubiquitin. Early sources of evidence showed that proteasome inhibition delayed Topo-1/2 degradation following exposure to Topo-1 and −2 poisons, suggesting that the proteasome is required for Topo-cc repair95,96,97. However, these initial experiments failed to substantiate the formation of ubiquitylated Topo-ccs. Topoisomerase ubiquitylation was shown in whole-cell extracts – rather than on chromatin or Topo-ccs – and there was no increase in the topoisomerase ubiquitylation state after proteasomal inhibition95,97. Moreover, these studies used very high doses of CPT (μM range)96,97, which crosslink 90% of Topo-198 and might cause an overload of Topo-1ccs beyond the repair capacity of the other, replication-dependent, proteases.
More recent studies have directly shown that both enzymatic and non-enzymatic DPCs are modified by ubiquitin14,42,62,63,75. However, whether this modification leads to proteasomal degradation seems controversial, as ubiquitylation might rather have a signaling role62.
Crosslinked proteins can be concomitantly modified by ubiquitin and SUMO (SUMO-1 or SUMO-2/3)42,62,63,75. Modification via SUMO-2/3 prior to ubiquitylation hints at the involvement of SUMO-targeted ubiquitin ligases (STUbLs), which ubiquitylate SUMO-modified substrates for proteasomal degradation42,63. However, scenarios have been described where these two modifications are independent, although the function of DPC SUMOylation in these cases remains unclear62,75.
S.cerevisiae Ddi1 is a proteasome adapter99 (Fig. 2). Ddi1’s UBL binds the proteasome subunit Rpn1100,101, and, unlike conventional UBL domains, it also binds ubiquitin, along with UBA43,102. However, there is no evidence that Ddi1 functions in DPC repair with the aid of the proteasome. In fact, overexpression of mutated versions lacking UBL and UBA domains complements DDI1 deletion in FA sensitivity experiments, while mutation of the putative catalytic domain does not45. A different scenario is very likely for the human homologs DDI1 and DDI2. DDI1/2 bind the proteasome via their UBLs43,44 (Fig. 2). DDI1/2 recruit the proteasome at stalled replication forks to remove RTF2, whose persistence would prevent fork restart after HU and cause chromosome instability44.
As for proteasome-dependent DPC repair during replication, experiments in Xenopus egg extracts have provided the most compelling evidence75,76. Instead, in mammalian cells, ubiquitylation of Topo-ccs via the STUbL RNF4 prior to proteasomal degradation is independent of replication63. A recent report claims that the enzyme HMCES crosslinks to abasic sites ahead of the replication fork. In this way, HMCES shields the abasic sites from replication by TLS polymerases and prevents mutagenesis14. HMCES crosslinks are ubiquitylated and processed by the proteasome, but the role of other proteases has not been tested14. Although physiological crosslinking must happen during DNA synthesis to prevent switching to TLS polymerases, it has not been established that HMCES-DPC repair by the proteasome happens during replication. Post-replicative repair is plausible, as (i) HCMES-DPC per se represents an impediment to DNA polymerase progression and (ii) proteolysis by the proteasome would likely leave a peptide that can halt the DNA polymerase. In either case, a tolerance mechanism other than TLS must be in place to bypass the DPC or the remnant peptide (e.g., template switching). Therefore, post-replicative proteasome-dependent proteolysis remains a formal possibility in this case.
In conclusion, replication-coupled proteolysis by a large complex like the proteasome remains a matter of future investigation in mammalian cells. For example, it will be informative to determine whether proteasome subunits appear at nascent chromatin after treatment with DPC-inducing agents (e.g., by iPOND), since assessing the proteins enriched at stalled forks has been useful for other kinds of replication stress103.
DPC proteases in processing of non-crosslinked, tightly bound substrates
Mass spectrometry has been performed on DPCs isolated from cells depleted of SPRTN5 and GCNA40 to identify the most abundant crosslinked proteins. These screenings have proved that Topoisomerases, histones and Mcm subunits of the CMG helicase complex are the major DPCs in SPRTN-depleted cells and GCNA KO flies5,40. Nucleic acid-binding proteins are expected to come up in such screenings5 since proteins acting in the vicinity of the DNA are most likely crosslinked. However, in vitro cleavage experiments are mainly performed on substrates that have the propensity for DNA binding but are not crosslinked to DNA3,4,5,6,37. Hence DNA association rather than crosslinking per se is a requisite for cleavage. Recent studies have proven that this is true in cells as well. The three studies below illustrate that proteases process non-covalently DNA-associated proteins to protect cells from DNA replication errors.
Histones: A recent study postulates that Wss1 degrades unassembled, yet non-covalently bound, histones during replication stress104. This Wss1 activity would protect cells from the unspecific binding of excess histones to the ssDNA accumulating after HU exposure, which can interfere with DNA metabolism104 (Fig. 4A). The sensitivity of wss1 mutant yeast cells to HU is greatly enhanced by concomitant deletion of DDI1, and a catalytically inactive Ddi1 does not rescue cell survival45,46. Thus, the protease Ddi1, like Wss1, might be required to cope with replication stress beyond DPCs. More interestingly, complementation studies in yeast suggest that hDDI1/2 retain the same function46.
PARP-1: Tightly bound proteins can be as dangerous as DPCs for replication fork progression105. This scenario is well illustrated by PARP-1 trapping. PARP-1 is an enzyme participating in DNA replication and many DNA repair pathways106,107. PARP-1 catalyses the formation of poly-ADP ribose (PAR) chains—a reaction known as PARylation—that recruit other repair proteins; self-PARylation triggers an electrostatic dissociation from chromatin, allowing the repair reaction to proceed106. Pharmacological inhibition causes PARP to become tightly bound—or trapped—on chromatin because of the defective self-PARylation. Chromatin persistence due to PARP inhibitors is more problematic than the catalytic inhibition per se108. In fact, trapped PARP-1 can cause replication fork collapse and DSBs and has the potential of killing BRCA-deficient cells, a strategy that is used for treatment of cancer patients with BRCA mutations109. FAM111A depletion sensitizes cells to niraparib and talazoparib51, two PARP inhibitors with a strong trapping capacity108,110. Niraparib treatment reduces replication fork speed in FAM111A-depleted cells and causes replication stress. Rescue of the replication defect depends on FAM111A catalytic residues, PIP-box and DNA binding, implying that FAM111A uses its proteolytic activity to remove obstacles imposed by trapped PARP-1 to replication forks51 (Fig. 4B). Whether other proteases participate in the removal of tightly bound proteins remains to be established.
Chk1: The degradation of a non-crosslinked substrate echoes DPC removal: in either case, proteolysis eliminates potential obstacles for DNA replication and metabolism, which threaten cell viability51,104. In striking contrast is the cleavage of the checkpoint kinase Chk1 by SPRTN during DNA synthesis59. During S phase, SPRTN cleaves Chk1 at the chromatin, in a loosely structured region at the C-terminus, and releases truncated Chk1 N-terminal fragments with stronger kinase activity than the full-length protein59 (Fig. 4C). This way SPRTN ensures a basal and physiological Chk1 activation to support DNA replication progression in the absence of exogenous damage, i.e., in the absence of insults that cause ssDNA accumulation and robust ATR-dependent Chk1 activation cascade59,111. Overexpression of Chk1 N-terminal fragments restores normal DNA replication in SPRTN-depleted cells (i.e., replication fork speed and origin firing) and partially rescues developmental defects of SPRTN-deficient zebrafish embryos59.
Besides uncovering an important role for Chk1 during unperturbed replication, this study also shows that the function of DPC proteases goes well beyond the proteolysis of replication fork barriers.
DPC proteases and diseases
Defective DNA replication, often referred to as DNA replication stress, is one of the major causes of cancer112,113,114. DNA replication stress can be triggered by obstacles ahead of the replication fork, such as those imposed by crosslinked or tightly bound proteins. Since the activity of DPC proteases is linked to DNA replication, strong association exists between defective DPC proteases and human diseases40,57,60.
Ruijs-Aalfs syndrome (RJALS)
Biallelic and monogenic mutations in SPRTN cause a rare autosomal recessive progeroid disease known as RJALS60. This was the first disease to be linked to defective DPC proteolysis repair. The loss-of-function mutations identified in patients so far are a missense mutation that generates the catalytically inactive SPRTN-Y117C variant, and non-sense mutations that generate a truncated SPRTN, SPRTN-ΔC (Fig. 2), which retains partial functionality due to intact protease activity but defective cellular localization4,5,57,60. RJALS patients show signs of premature aging—including cataracts, graying of the hair, lipodistrophy—and develop early-onset hepatocellular carcinoma (HCC)60. These phenotypes have been recapitulated in SPRTN hypomorphic mice57,115, and definitively prove that defective SPRTN alone is responsible for the RJALS phenotype.
Aging and cancer are seemingly conflicting manifestations, but truly two outcomes of the same underlying cellular defects—the accumulation of DNA damage and genomic instability116,117. RJALS patient cells (patient-derived lymphoblastoid cell lines and fibroblasts) display DNA replication stress, G2/M leakage, increased number of DSBs, chromosomal aberrations and increased level of total DPCs. It is unclear why human patients and mice develop HCC while the rest of the body ages.
Kenny–Caffey syndrome type 2 (KCS2) and gracile bone dysplasia (GCLEB)
FAM111A mutations cause KCS2, a rare autosomal dominant disease characterized by short stature, hypoparathyroidism, hypocalcemia and abnormal bone development. KCS2 patients do not show any predisposition to develop cancer118,119. Heterozygous mutations in FAM111A have also been found in patients with GCLEB, or osteocraniostenosis119. GCLEB is lethal in newborns because of severe skeletal abnormalities.
KCS2-causing and GCLEB-causing mutations alter FAM111A proteolytic activity. Substitution of R569 to Histidine (H) is a recurrent mutation in KCS2118,119 and falls in the proximity of the catalytic residue S541 (Fig. 2). FAM111A-R569H and other disease-causing mutations enhance FAM111A self-cleavage in vitro, showing that they are gain-of-function mutations51,52. Expression of FAM111A-R569H or other mutated variants causes replication and transcription defects and apoptosis52,61,77. Thus, it seems plausible that the toxicity of the hyperactive protein arises from uncontrolled degradation of replication and transcription proteins (e.g., RFC, Rpb1)52; however, it is unknown how these mutations result in the observed phenotypes.
Pediatric germ cell tumors (GCTs)
Mutations in GCNA associate with GCTs. Downregulation of GCNA expression—due to copy number loss and promoter hypermethylation—is found in 66% of GCTs40 and correlates with a poor prognosis. These alterations in GCTs show that GCNA protease is critical for the genomic stability of germ cells.
Conclusions and perspectives
The topic of DPC proteolysis repair has undergone an extraordinary advance in the past year. The list of specific repair enzymes has come to include novel DPC proteases working with Wss1 and SPRTN. In most cases, direct proteolysis of DPC substrates remains to be formally tested with in vitro assays; however, experiments conducted in cells, such as sensitivity to crosslinking agents, have established that an intact catalytic domain is always essential. We anticipate that more rigorous and systematic in vitro studies will elucidate the optimal requirements for DPC proteolysis.
The fidelity of DNA replication and genome stability are compromised by loss or misregulation of these proteases. This is not only due to defective DPC repair but also to defective removal of non-covalent replication barriers. The latter exemplifies other important functions of DPC proteases in the maintenance of genome integrity.
DPC toxicity opens an interesting perspective for cancer treatment as well. The most commonly used chemotherapy agents are Topoisomerase poisons that induce enzymatic DPCs known as Topo-1/2ccs2,11. Specific DPC protease inhibitors could synergize with DPC-inducing agents and be instrumental to chemotherapy. The recent finding that FAM111A confers resistance to PARP inhibitors51 has potential implications in cancer treatment, not only in light of the development of a FAM111A inhibitor. In fact, FAM111A expression varies among cancer cell lines51, thus the efficacy of PARP inhibitors might change. These types of information should help design better treatment strategies.
We expect that research will focus more imminently on the identification of substrates and the regulatory mechanisms, including interaction and coordination with other repair proteins for timely activation of the DPC protease activity. This basic knowledge of DPC proteases will pave the way for a better understanding of human diseases and cancer therapy.
References
Vaz, B., Popovic, M. & Ramadan, K. DNA–protein crosslink proteolysis repair. Trends Biochem. Sci. 42, 483–495 (2017).
Stingele, J., Bellelli, R. & Boulton, S. J. Mechanisms of DNA-protein crosslink repair. Nat. Rev. Mol. Cell Biol. 18, 563–573 (2017).
Stingele, J., Schwarz, M. S., Bloemeke, N., Wolf, P. G. & Jentsch, S. A DNA-dependent protease involved in DNA-protein crosslink repair. Cell 158, 327–338 (2014). This seminal study has uncovered the existence of a specialised enzyme for DPC repair.
Stingele, J. et al. Mechanism and regulation of DNA-protein crosslink repair by the DNA-dependent metalloprotease SPRTN. Mol. Cell 64, 688–703 (2016). This article has described the first layer of regulation for the protease SPRTN, which is its de-ubiquitylation after DPC formation.
Vaz, B. et al. Metalloprotease SPRTN/DVC1 orchestrates replication-coupled DNA-protein crosslink repair. Mol. Cell 64, 704–719 (2016). This article has discovered SPRTN as the first DNA-dependent metalloprotease in metazoans for removal of DNA-protein crosslinks during DNA replication.
Lopez-Mosqueda, J. et al. SPRTN is a mammalian DNA-binding metalloprotease that resolves DNA-protein crosslinks. Elife 5, 1–19 (2016).
Ruggiano, A. Biochemical sciences spotlight the trinity of SPRTN protease regulation. Trends Biochem. Sci. https://doi.org/10.1016/j.tibs.2020.10.007. (2020).
Stingele, J. & Jentsch, S. DNA-protein crosslink repair. Nat. Rev. Mol. Cell Biol. 16, 455–460 (2015).
Pommier, Y. DNA topoisomerase I inhibitors: chemistry, biology, and interfacial inhibition. Chem. Rev. 109, 2894–2902 (2009).
Nitiss, J. L. Targeting DNA topoisomerase II in cancer chemotherapy. Nat. Rev. Cancer 9, 338–350 (2009).
Fielden, J., Ruggiano, A., Popović, M. & Ramadan, K. DNA protein crosslink proteolysis repair: from yeast to premature ageing and cancer in humans. DNA Repair. 71, 198–204 (2018).
Santi, D. V., Norment, A. & Garrett, C. E. Covalent bond formation between a DNA-cytosine methyltransferase and DNA containing 5-azacytosine. Proc. Natl Acad. Sci. USA 81, 6993–6997 (1984).
Maslov, A. Y. et al. 5-Aza-2′-deoxycytidine-induced genome rearrangements are mediated by DNMT1. Oncogene 31, 5172–5179 (2012).
Mohni, K. N. et al. HMCES maintains genome integrity by shielding abasic sites in single-strand DNA. Cell 176, 144–153.e13 (2019).
De Groot, A. C., Flyvholm, M. A., Lensen, G., Menné, T. & Coenraads, P. J. Formaldehyde-releasers: Relationship to formaldehyde contact allergy. Contact allergy to formaldehyde and inventory of formaldehyde-releasers. Contact Dermat. 61, 63–85 (2009).
Pontel, L. B. et al. Endogenous formaldehyde is a hematopoietic stem cell genotoxin and metabolic carcinogen. Mol. Cell 60, 177–188 (2015).
Hou, H. & Yu, H. Structural insights into histone lysine demethylation. Curr. Opin. Struct. Biol. 20, 739–748 (2010).
Ayala, A., Muñoz, M. F. & Argüelles, S. Lipid peroxidation: production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxid. Med. Cell. Longev. 2014, 1–31 (2014).
Traube, F. R. & Carell, T. The chemistries and consequences of DNA and RNA methylation and demethylation. RNA Biol. 14, 1099–1107 (2017).
Ide, H., Nakano, T., Salem, A. M. H. & Shoulkamy, M. I. DNA–protein cross-links: formidable challenges to maintaining genome integrity. DNA Repair 71, 190–197 (2018).
Nakano, T. et al. Nucleotide excision repair and homologous recombination systems commit differentially to the repair of DNA-protein crosslinks. Mol. Cell 28, 147–158 (2007).
de Graaf, B., Clore, A. & McCullough, A. K. Cellular pathways for DNA repair and damage tolerance of formaldehyde-induced DNA-protein crosslinks. DNA Repair 8, 1207–1214 (2009).
Nakano, T. et al. Homologous recombination but not nucleotide excision repair plays a pivotal role in tolerance of DNA-protein cross-links in mammalian cells. J. Biol. Chem. 284, 27065–27076 (2009).
Hoa, N. N. et al. Mre11 is essential for the removal of lethal topoisomerase 2 covalent cleavage complexes. Mol. Cell 64, 580–592 (2016).
Deshpande, R. A., Lee, J. H., Arora, S. & Paull, T. T. Nbs1 converts the human Mre11/Rad50 nuclease complex into an endo/exonuclease machine specific for protein-DNA adducts. Mol. Cell 64, 593–606 (2016).
Quievryn, G. & Zhitkovich, A. Loss of DNA–protein crosslinks from formaldehyde-exposed cells occurs through spontaneous hydrolysis and an active repair process linked to proteosome function. Carcinogenesis 21, 1573–1580 (2000).
Pommier, Y. et al. Tyrosyl-DNA-phosphodiesterases (TDP1 and TDP2). DNA Repair 19, 114–129 (2014).
Yang, S. W. et al. A eukaryotic enzyme that can disjoin dead-end covalent complexes between DNA and type I topoisomerases. Proc. Natl Acad. Sci. USA 93, 11534–11539 (1996).
Debethune, L. Processing of nucleopeptides mimicking the topoisomerase I-DNA covalent complex by tyrosyl-DNA phosphodiesterase. Nucleic Acids Res. 30, 1198–1204 (2002).
Interthal, H., Chen, H. J. & Champoux, J. J. Human Tdp1 cleaves a broad spectrum of substrates, including phosphoamide linkages. J. Biol. Chem. 280, 36518–36528 (2005).
Schellenberg, M. J. et al. ZATT (ZNF451)–mediated resolution of topoisomerase 2 DNA-protein cross-links. Science 357, 1412–1416 (2017).
Gao, R. et al. Proteolytic degradation of topoisomerase II (Top2) enables the processing of Top2·DNA and Top2·RNA covalent complexes by tyrosyl-DNA-phosphodiesterase 2 (TDP2). J. Biol. Chem. 289, 17960–17969 (2014).
Duxin, J. P., Dewar, J. M., Yardimci, H. & Walter, J. C. Repair of a DNA-protein crosslink by replication-coupled proteolysis. Cell 159, 346–357 (2014). This study has described for the first time that DPC proteolysis repair happens during DNA replication.
Reinking, H. K., Hofmann, K. & Stingele, J. Function and evolution of the DNA-protein crosslink proteases Wss1 and SPRTN. DNA Repair 88, 102822 (2020).
Yang, X. et al. Structural analysis of Wss1 protein from saccharomyces cerevisiae. Sci. Rep. 7, 1–9 (2017).
Balakirev, M. Y. et al. Wss1 metalloprotease partners with Cdc48/Doa1 in processing genotoxic SUMO conjugates. Elife 4, e06763 (2015).
Li, F., Raczynska, J. E., Chen, Z. & Yu, H. Structural insight into DNA-dependent activation of human metalloprotease spartan. Cell Rep. 26, 3336–3346.e4 (2019).
Reinking, H. K. et al. DNA structure-specific cleavage of DNA-protein crosslinks by the SPRTN protease. Mol. Cell 80, 102–113.e6 (2020). This article has described a second layer of regulation for the protease SPRTN, which is activated by a specific combination of ssDNA and dsDNA.
Carmell, M. A. et al. A widely employed germ cell marker is an ancient disordered protein with reproductive functions in diverse eukaryotes. Elife 5, e19993 (2016).
Bhargava, V. et al. GCNA preserves genome integrity and fertility across species. Dev. Cell 52, 38–52.e10 (2020).
Dokshin, G. A. et al. GCNA interacts with Spartan and Topoisomerase II to regulate genome stability. Dev. Cell 52, 53–68.e6 (2020).
Borgermann, N. et al. SUMOylation promotes protective responses to DNA‐protein crosslinks. EMBO J. 38, e101496 (2019).
Nowicka, U. et al. DNA-damage-inducible 1 protein (Ddi1) contains an uncharacteristic ubiquitin-like domain that binds ubiquitin. Structure 23, 542–557 (2015).
Kottemann, M. C., Conti, B. A., Lach, F. P. & Smogorzewska, A. Removal of RTF2 from stalled replisomes promotes maintenance of genome integrity. Mol. Cell 69, 24–35.e5 (2018).
Serbyn, N. et al. The aspartic protease Ddi1 contributes to DNA-protein crosslink repair in yeast. Mol. Cell 77, 1066–1079.e9 (2020). This article has reported the existence of a second protease in S.cerevisiae with overlapping functions to Wss1 during DNA replication.
Svoboda, M., Konvalinka, J., Trempe, J. F. & Grantz Saskova, K. The yeast proteases Ddi1 and Wss1 are both involved in the DNA replication stress response. DNA Repair 80, 45–51 (2019).
Mercier, S. et al. Mutations in FAM111b cause hereditary fibrosing poikiloderma with tendon contracture, myopathy, and pulmonary fibrosis. Am. J. Hum. Genet. 93, 1100–1107 (2013).
Fine, D. A. et al. Identification of FAM111A as an SV40 host range restriction and adenovirus helper factor. PLoS Pathog. 8, e1002949 (2012).
Panda, D., Fernandez, D. J., Lal, M., Buehler, E. & Moss, B. Triad of human cellular proteins, IRF2, FAM111A, and RFC3, restrict replication of orthopoxvirus SPI-1 host-range mutants. Proc. Natl Acad. Sci. USA 114, 3720–3725 (2017).
Alabert, C. et al. Nascent chromatin capture proteomics determines chromatin dynamics during DNA replication and identifies unknown fork components. Nat. Cell Biol. 16, 281–291 (2014).
Kojima, Y. et al. FAM111A protects replication forks from protein obstacles via its trypsin-like domain. Nat. Commun. 11, 1–14 (2020). This study has discussed the existence of a second DPC protease in humans.
Hoffmann, S. et al. FAM111 protease activity undermines cellular fitness and is amplified by gain‐of‐function mutations in human disease. EMBO Rep. 21, e50662 (2020).
Yip, M. C. J., Bodnar, N. O. & Rapoport, T. A. Ddi1 is a ubiquitin-dependent protease. Proc. Natl Acad. Sci. USA 117, 7776–7781 (2020).
Dirac-Svejstrup, A. B. et al. DDI2 is a ubiquitin-directed endoprotease responsible for cleavage of transcription factor NRF1. Mol. Cell 79, 332–341.e7 (2020).
Toth, A., Hegedus, L., Juhasz, S., Haracska, L. & Burkovics, P. The DNA-binding box of human SPARTAN contributes to the targeting of Polη to DNA damage sites. DNA Repair. 49, 33–42 (2017).
Jambhekar, A., Dhall, A. & Shi, Y. Roles and regulation of histone methylation in animal development. Nat. Rev. Mol. Cell Biol. 20, 625–641 (2019).
Maskey, R. S. et al. Spartan deficiency causes accumulation of Topoisomerase 1 cleavage complexes and tumorigenesis. Nucleic Acids Res. 45, 4564–4576 (2017).
Mórocz, M. et al. DNA-dependent protease activity of human Spartan facilitates replication of DNA-protein crosslink-containing DNA. Nucleic Acids Res. 45, 3172–3188 (2017).
Halder, S. et al. SPRTN protease and checkpoint kinase 1 cross-activation loop safeguards DNA replication. Nat. Commun. 10, 1–18 (2019). This study has discovered one of the non-DPC-related functions of proteases.
Lessel, D. et al. Mutations in SPRTN cause early onset hepatocellular carcinoma, genomic instability and progeroid features. Nat. Genet. 46, 1239–1244 (2014). This article has described for the first time the link between SPRTN mutations and aging diseases in human.
Rios-Szwed, D. O. et al. FAM111A regulates replication origin activation and cell fitness. bioRxiv 2020.04.22.055574. https://doi.org/10.1101/2020.04.22.055574 (2020).
Vaz, B. et al. SPRTN protease and SUMOylation coordinate DNA-protein crosslink repair to prevent genome instability. bioRxiv 2020.02.14.949289. https://doi.org/10.1101/2020.02.14.949289 (2020).
Sun, Y. et al. A conserved SUMO-Ubiquitin pathway directed by RNF4/SLX5-SLX8 and PIAS4/SIZ1 drives proteasomal degradation of topoisomerase DNA-protein crosslinks. bioRxiv 707661. https://doi.org/10.1101/707661 (2019).
Perry, M. et al. USP11 deubiquitinates monoubiquitinated SPRTN to repair DNA-protein crosslinks. bioRxiv 2020.06.30.180471. https://doi.org/10.1101/2020.06.30.180471 (2020).
Huang, J. et al. Tandem deubiquitination and acetylation of SPRTN promotes DNA-protein crosslink repair and protects against aging. Mol. Cell 79, 824–835.e5 (2020).
Mosbech, A. et al. DVC1 (C1orf124) is a DNA damage-targeting p97 adaptor that promotes ubiquitin-dependent responses to replication blocks. Nat. Struct. Mol. Biol. 19, 1084–1092 (2012).
Davis, E. J. et al. DVC1 (C1orf124) recruits the p97 protein segregase to sites of DNA damage. Nat. Struct. Mol. Biol. 19, 1093–1100 (2012).
Fielden, J. et al. TEX264 coordinates p97- and SPRTN-mediated resolution of topoisomerase 1-DNA adducts. Nat. Commun. 11, 1–16 (2020). This article has identified the first DPC-specific cofactor for the ATPase p97.
van den Boom, J. & Meyer, H. VCP/p97-mediated unfolding as a principle in protein homeostasis and signaling. Mol. Cell 69, 182–194 (2018).
Franz, A., Ackermann, L. & Hoppe, T. Ring of change: CDC48/p97 drives protein dynamics at chromatin. Front. Genet. 7, 1–14 (2016).
Nakano, T. et al. Translocation and stability of replicative DNA helicases upon encountering DNA-protein cross-links. J. Biol. Chem. 288, 4649–4658 (2013).
Hsiang, Y. H., Lihou, M. G. & Liu, L. F. Arrest of replication forks by drug-stabilized topoisomerase I-DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 49, 5077–5082 (1989).
Kumari, A., Lim, Y. X., Newell, A. H., Olson, S. B. & McCullough, A. K. Formaldehyde-induced genome instability is suppressed by an XPF-dependent pathway. DNA Repair 11, 236–246 (2012).
Cortez, D. Replication-coupled DNA repair. Mol. Cell 74, 866–876 (2019).
Larsen, N. B. et al. Replication-coupled DNA-protein crosslink repair by SPRTN and the proteasome in Xenopus egg extracts. Mol. Cell 73, 574–588.e7 (2019).
Sparks, J. L. et al. The CMG helicase bypasses DNA-protein cross-links to facilitate their repair. Cell 176, 167–181.e21 (2019).
Nie, M. et al. Activation of FAM111A protease induces defects in nuclear function that likely underlie its roles in disease and viral restriction. bioRxiv 2020.05.04.077594. https://doi.org/10.1101/2020.05.04.077594 (2020).
Branzei, D. & Psakhye, I. DNA damage tolerance. Curr. Opin. Cell Biol. 40, 137–144 (2016).
Juhasz, S. et al. Characterization of human Spartan/C1orf124, an ubiquitin-PCNA interacting regulator of DNA damage tolerance. Nucleic Acids Res. 40, 10795–10808 (2012).
Machida, Y., Kim, M. S. & Machida, Y. J. Spartan/C1orf124 is important to prevent UV-induced mutagenesis. Cell Cycle 11, 3395–3402 (2012).
Ghosal, G., Leung, J. W. C., Nair, B. C., Fong, K. W. & Chen, J. Proliferating cell nuclear antigen (PCNA)-binding protein C1orf124 is a regulator of translesion synthesis. J. Biol. Chem. 287, 34225–34233 (2012).
Centore, R. C., Yazinski, S. A., Tse, A. & Zou, L. Spartan/C1orf124, a reader of PCNA ubiquitylation and a regulator of UV-induced DNA damage response. Mol. Cell 46, 625–635 (2012).
Pashev, I. G., Dimitrov, S. I. & Angelov, D. Crosslinking proteins to nucleic acids by ultraviolet laser irradiation. Trends Biochem. Sci. 16, 323–326 (1991).
Smith, K. C. Dose dependent decrease in extractability of DNA from bacteria following irradiation with ultraviolet light or with visible light plus dye. Biochem. Biophys. Res. Commun. 8, 157–163 (1962).
SHETLAR, M. D., CHRISTENSEN, J. & HOM, K. Photochemical addition of amino acids and peptides to DNA. Photochem. Photobiol. 39, 125–133 (1984).
Nakazato, A. et al. SPARTAN promotes genetic diversification of the immunoglobulin-variable gene locus in avian DT40 cells. DNA Repair 68, 50–57 (2018).
Grishaw, W. J. A morphologic study of deoxyribonucleic acid synthesis and cell proliferation in regenerating rat liver; autoradiography with thymidine-H-3. Cancer Res. 22, 842–849 (1962).
Delabaere, L. et al. The spartan ortholog maternal haploid is required for paternal chromosome integrity in the drosophila zygote. Curr. Biol. 24, 2281–2287 (2014).
Tang, X. et al. Maternal haploid, a metalloprotease enriched at the largest satellite repeat and essential for genome integrity in Drosophila embryos. Genetics 206, 1829–1839 (2017).
Brŕgeon, D. & Doetsch, P. W. Transcriptional mutagenesis: causes and involvement in tumour development. Nat. Rev. Cancer 11, 218–227 (2011).
Heckmann, I., Kern, M. J., Pfander, B. & Jentsch, S. A SUMO-dependent pathway controls elongating RNA Polymerase II upon UV-induced damage. Sci. Rep. 9, 17914 (2019).
Verma, R., Oania, R., Fang, R., Smith, G. T. & Deshaies, R. J. Cdc48/p97 mediates UV-dependent turnover of RNA Pol II. Mol. Cell 41, 82–92 (2011).
Bedford, L., Paine, S., Sheppard, P. W., Mayer, R. J. & Roelofs, J. Assembly, structure, and function of the 26S proteasome. Trends Cell Biol. 20, 391–401 (2010).
Collins, G. A. & Goldberg, A. L. The Logic of the 26S Proteasome. Cell 169, 792–806 (2017).
Mao, Y., Desai, S. D., Ting, C. Y., Hwang, J. & Liu, L. F. 26S proteasome-mediated degradation of topoisomerase II cleavable complexes. J. Biol. Chem. 276, 40652–40658 (2001).
Lin, C. P., Ban, Y., Lyu, Y. L., Desai, S. D. & Liu, L. F. A ubiquitin-proteasome pathway for the repair of topoisomerase I-DNA covalent complexes. J. Biol. Chem. 283, 21074–21083 (2008).
Desai, S. D., Liu, L. F., Vazquez-Abad, D. & D’Arpa, P. Ubiquitin-dependent destruction of topoisomerase I is stimulated by the antitumor drug camptothecin. J. Biol. Chem. 272, 24159–24164 (1997).
Hsiang, Y. H. & Liu, L. F. Identification of mammalian dna topoisomerase i as an intracellular target of the anticancer drug camptothecin. Cancer Res. 48, 1722–1726 (1988).
Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 78, 477–513 (2009).
Elsasser, S. et al. Proteasome subunit Rpn1 binds ubiquitin-like protein domains. Nat. Cell Biol. 4, 725–730 (2002).
Gomez, T. A., Kolawa, N., Gee, M., Sweredoski, M. J. & Deshaies, R. J. Identification of a functional docking site in the Rpn1 LRR domain for the UBA-UBL domain protein Ddi1. BMC Biol. 9, 33 (2011).
Trempe, J. F. et al. Structural studies of the yeast DNA damage-inducible protein Ddi1 reveal domain architecture of this eukaryotic protein family. Sci. Rep. 6, 1–13 (2016).
Sirbu, B. M. et al. Identification of proteins at active, stalled, and collapsed replication forks using isolation of proteins on nascent DNA (iPOND) coupled with mass spectrometry. J. Biol. Chem. 288, 31458–31467 (2013).
Maddi, K. et al. Wss1 promotes replication stress tolerance by degrading histones. Cell Rep. 30, 3117–3126.e4 (2020). This study has discovered one of the non-DPC-related functions of proteases.
Fu, Y. V. et al. Selective bypass of a lagging strand roadblock by the eukaryotic replicative DNA helicase. Cell 146, 931–941 (2011).
Pascal, J. M. The comings and goings of PARP-1 in response to DNA damage. DNA Repair 71, 177–182 (2018).
Hanzlikova, H. et al. The importance of poly(ADP-Ribose) polymerase as a sensor of unligated Okazaki fragments during DNA replication. Mol. Cell 71, 319–331.e3 (2018).
Murai, J. et al. Trapping of PARP1 and PARP2 by clinical PARP inhibitors. Cancer Res. 72, 5588–5599 (2012).
Lord, C. J. & Ashworth, A. PARP inhibitors: synthetic lethality in the clinic. Science 355, 1152–1158 (2017).
Murai, J. et al. Stereospecific PARP trapping by BMN 673 and comparison with olaparib and rucaparib. Mol. Cancer Ther. 13, 433–443 (2014).
Bruhn, C. & Foiani, M. A model of DNA damage response activation at stalled replication forks by SPRTN. Nat. Commun. 10, 16–18 (2019).
Gaillard, H., García-Muse, T. & Aguilera, A. Replication stress and cancer. Nat. Rev. Cancer 15, 276–280 (2015).
Macheret, M. & Halazonetis, T. D. DNA replication stress as a hallmark of cancer. Annu. Rev. Pathol. Mech. Dis. 10, 425–448 (2015).
Lukas, C. et al. 53BP1 nuclear bodies form around DNA lesions generated by mitotic transmission of chromosomes under replication stress. Nat. Cell Biol. 13, 243–253 (2011).
Maskey, R. S. et al. Spartan deficiency causes genomic instability and progeroid phenotypes. Nat. Commun. 5, 1–12 (2014).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194 (2013).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Isojima, T. et al. A recurrent de novo FAM111A mutation causes Kenny-Caffey syndrome type 2. J. Bone Miner. Res. 29, 992–998 (2014).
Unger, S. et al. FAM111A mutations result in hypoparathyroidism and impaired skeletal development. Am. J. Hum. Genet. 92, 990–995 (2013).
Acknowledgements
We deeply apologize to the researchers whose work was not cited. We thank a Medical Research Council (MRC) Programme grant to K.R. (MC_EX_MR/K022830/1). A.R. was supported by an EMBO LTF (ALTF 1109-2017). We would like to thank our DPhil student Gwendoline Hoslett for helping us with the text editing and her comments.
Author information
Authors and Affiliations
Contributions
A.R. wrote the draft and prepared the figures. K.R. initiated and supervised this work. A.R. and K.R prepared the final version of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ruggiano, A., Ramadan, K. DNA–protein crosslink proteases in genome stability. Commun Biol 4, 11 (2021). https://doi.org/10.1038/s42003-020-01539-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01539-3
This article is cited by
-
Transcription-coupled DNA–protein crosslink repair by CSB and CRL4CSA-mediated degradation
Nature Cell Biology (2024)
-
Transcription-coupled repair of DNA–protein cross-links depends on CSA and CSB
Nature Cell Biology (2024)
-
Targeting DNA damage response pathways in cancer
Nature Reviews Cancer (2023)
-
FAM111A is dispensable for electrolyte homeostasis in mice
Scientific Reports (2022)
-
Human topoisomerases and their roles in genome stability and organization
Nature Reviews Molecular Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.