Abstract
Guanosine 3′,5′-bis(pyrophosphate) (ppGpp) functions as a second messenger in bacteria to adjust their physiology in response to environmental changes. In recent years, the ppGpp-specific hydrolase, metazoan SpoT homolog-1 (Mesh1), was shown to have important roles for growth under nutrient deficiency in Drosophila melanogaster. Curiously, however, ppGpp has never been detected in animal cells, and therefore the physiological relevance of this molecule, if any, in metazoans has not been established. Here, we report the detection of ppGpp in Drosophila and human cells and demonstrate that ppGpp accumulation induces metabolic changes, cell death, and eventually lethality in Drosophila. Our results provide the evidence of the existence and function of the ppGpp-dependent stringent response in animals.
Similar content being viewed by others
Introduction
Organisms must adjust their physiology in response to environmental changes. The stringent response is one of the most important starvation/metabolic control responses in bacteria and is controlled by the hyper-phosphorylated nucleotide guanosine 3′,5′-bis(pyrophosphate) (ppGpp)1,2. ppGpp accumulates in bacterial cells upon exposure to various stresses and the accumulated ppGpp functions as an alarmone that can alter transcription3,4,5,6, translation7,8, and certain enzymatic activities6,9,10 to overcome a stress3. In Escherichia coli, ppGpp level is regulated by two distinct enzymes, namely RelA and SpoT11. Both RelA and SpoT catalyze pyrophosphorylation of GDP (or GTP) by using ATP to produce ppGpp12. However, only SpoT (and not RelA) hydrolyzes ppGpp13. The RelA/SpoT homologs (RSHs) are universally conserved in bacteria14 and have pivotal roles in various aspects of bacterial physiology, including the starvation response1,2, growth rate control15, antibiotic tolerance16, and darkness response17. RSHs are also found in eukaryotes, including the green algae Chlamydomonas reinhardtii18 and land plant Arabidopsis thaliana19, as well as metazoa, including humans and Drosophila melanogaster14,20.
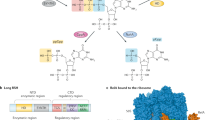
The RSH superfamily is classified into three types: long RSHs, small alarmone synthases (SASs), and small alarmone hydrolases (SAHs)14. Long RSHs have ppGpp synthase and hydrolase domains near their N terminus and a regulatory domain near their C terminus, which associates with variable interactors such as the ribosome21 and acyl carrier proteins22. SASs contain a single ppGpp synthase and SAHs contain a single hydrolase domain23. SASs are found not only in the bacterial phyla Firmicutes and Actinobacteria but also in archaea and some eukaryotes14. SASs in Firmicutes have been well characterized and include YwaC/RelP and YjbM/RelQ14,23,24. YwaC and YjbM have weak and strong ppGpp synthesis activity, respectively, for GTP homeostasis control6,23. SAHs are found in some bacterial species and in metazoa14. Metazoan SpoT homolog-1 (Mesh1) is one class of SAH found in metazoa (Fig. 1a). Mesh1 hydrolyzes ppGpp in vitro with comparable efficiency to bacterial long RSHs25,26. The Drosophila Mesh1 loss-of-function mutant (Mesh1 LOF) exhibits retarded growth, especially during starvation25, suggesting that the ppGpp-dependent stringent response has been conserved in metazoa. However, this hypothesis is still under investigation, because no known ppGpp synthase has been identified, and the existence of ppGpp in metazoa has never been confirmed.
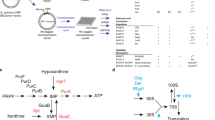
a Schematics of the primary structures of RSHs. A.A.: amino acids; ACT: aspartate kinase chorismite mutase TyrA domain; EF hand: Ca2+ binding helix E and F motif; Hyd: (p)ppGpp hydrolase domain; Syn: (p)ppGpp synthase domain; TGS: threonyl-tRNA synthetase, GTPase, SpoT domain. b Multiple-reaction-monitoring chromatograms for guanosine tetraphosphate ions in extracts from w− third instar larvae. The symbol “+” denotes the exogenous addition of the ppGpp standard to the sample. c Detection of ppGpp from various cultured cells and germ-free (GF) flies. Guanosine tetraphosphate-specific MS chromatograms of nucleotide pools extracted from larvae grown under the normal (GF control) and GF conditions, human cultured HeLa cells, human cultured PEAKrapid cells, and Drosophila cultured S2 cells.
Many studies have been conducted to attempt to detect ppGpp from metazoan cells25,27. The first attempt to detect ppGpp in eukaryotic cells was in 1970, in which nucleotides in HeLa cells were universally labeled with [32P]orthophosphate and the cell lysates were subjected to thin-layer chromatography; however, no ppGpp signal was detected, even after starvation28. Several years later, a report was published describing the presence of ppGpp along with other highly phosphorylated nucleotides in Chinese hamster ovary cells and baby hamster kidney cells29; however, this result could not be reproduced30. More recently, Sun et al.25 attempted to measure endogenous ppGpp in Drosophila by use of the high-performance liquid chromatography (LC) and matrix-assisted laser desorption/ionization-time of flight (TOF) mass spectrometry (MS); however, no ppGpp signals could be detected25. Therefore, it has been assumed that ppGpp is absent or, if present, accumulates very low levels in Metazoa25,27,29,30.
Recently, human Mesh1 (hMESH1) was shown to function as a cytosolic NADPH phosphatase31. The enzymatic efficiency (kcat/Km) of NADPH phosphatase activity of hMESH1 was calculated to be 14.4 mM−1 s−1, which is comparable to that of ppGpp hydrolase activity (9.46 mM−1 s−1)25,31. Expression of hMESH1 in human cells was upregulated by cystine deprivation or addition of ferroptosis-inducing erastin and hMESH1 overexpression (OE) resulted in lowered NADPH levels with increased sensitivity to ferroptosis31, suggesting that the NADPH phosphatase activity of hMESH1 is important in the regulation of ferroptosis. On the other hand, the physiological significance of the ppGpp-specific hydrolase activity of Mesh1 is currently unclear.
Here we were able to detect ppGpp in Drosophila and human cells, and demonstrate that Mesh1 modulates the endogenous ppGpp concentration in vivo, indicating that ppGpp metabolism is conserved in metazoa.
Results
Detection of ppGpp in Drosophila
To check for the existence of ppGpp in animals, we employed ultra-performance LC coupled with a tandem quadrupole MS (MS/MS)-based ppGpp quantification method, which we previously established for the characterization of ppGpp function in plants32. We extracted nucleotide pools from Drosophila late third instar larvae, in which Mesh1 expression was higher than at other stages as demonstrated by northern blotting25. Using the guanosine tetraphosphate-specific MS/MS mode (m/z transition from 602 to 159), three peaks were observed at 5.4, 6.2, and 6.4 min (Fig. 1b, blue), similar to what was observed for the detection of ppGpp in Arabidopsis tissues32. To determine which peak, if any, contained ppGpp in the Drosophila extract, we added ppGpp to the larvae extract (final concentration 16.7 nM) and subjected the sample to MS/MS analysis. The elution profile of the mixed sample revealed an increased area of the 6.4 min peak (Fig. 1b, orange), which matched the ppGpp standard peak (Fig. 1b, pink), strongly suggesting that it corresponds to ppGpp elution, as also seen in Arabidopsis32. As we fragmented the possible ppGpp elution for MS/MS detection, we could not collect separated nucleotides after LC to verify the identity of the ppGpp peak by other methods. Instead, we analyzed multiple fragmented ions derived from the elution and the ppGpp standard. The ppGpp standard and the 6.4 min elution peak of the Drosophila extract showed similar fragmented ion patterns at m/z = 150, 159, and 177, which correspond to guanine, deoxy-pyrophosphate, and pyrophosphate, respectively (Supplementary Fig. 1a, b), strongly suggesting that the 6.4 min peak represents the ppGpp elution. The 5.4 min peak was assigned as GTP (owing to cross-talk of the product ion) and the 6.2 min peak represented an unknown molecule (Supplementary Fig. 1a). It is noteworthy that a small amount of the unknown molecule could be detected in the ppGpp (Fig. 1b, pink) and GTP standards (Supplementary Fig. 1a), suggesting that it was also contaminated in the manufacturing process of the products. The unknown molecule is likely a guanosine tetraphosphate, as it could be passed through the first selected MS (MS1) at m/z = 602 and has a diphosphate moiety that could be detected in the second selected MS (MS2) at m/z = 159 after fragmentation in the tandem mass spectrometer. The pattern of fragmented ions of the unknown molecule is very similar to that of ppGpp (Supplementary Fig. 1b), suggesting that the structure of the unknown molecule resembles that of ppGpp. The intensity of the unknown peak varied with different developmental stages of Drosophila (Fig. 1b, c and Supplementary Fig. 2), implying the physiological function of the unknown molecule that may be involved in developmental processes. These results indicated that our LC-MS/MS-based ppGpp quantification method allows detection of ppGpp and an unknown molecule from Drosophila tissues with high sensitivity.
We also attempted to detect another bacterial alarmone, guanosine 5′-triphosphate 3′-diphosphate (pppGpp)1,2 from Drosophila; however, no stable pppGpp elution peak could be observed using the LC-MS/MS-based ppGpp quantification method described above, even when a pppGpp standard was used (for details, see “Methods”). This may result from reduced affinity of pppGpp for the reversed-phase column in the chosen solvent, compared to that of ppGpp, due to extra phosphate moiety of pppGpp. In future, selection of an appropriate ion-pair reagent and precise determination of the reagent concentration should be investigated for establishing the pppGpp-specific quantification method from Drosophila tissues.
We next tested stage-specific accumulation of ppGpp to investigate whether ppGpp levels modulate during Drosophila development. Specifically, using the MS/MS method, we quantified ppGpp concentrations from wild-type (WT) Drosophila (w–) at various developmental stages: eggs (from the strain Canton-S: CS), late third instar larvae (male and female), pupae (day 2), virgins (male and female), and day 4 adults (male and female). ppGpp was detected in extracts from all stages (Fig. 1b and Supplementary Fig. 2a–d) at 50–250 pmol g−1 fresh weight (FW) (Supplementary Table 1), which is comparable to that found in Arabidopsis (~100 pmol g−1 FW)32, and to the previous study that reported that ppGpp concentration in Drosophila, if any, should be less than 1000 pmol g−1 FW25. Interestingly, the amount of ppGpp in pupae was greater than that at other stages (Fig. 2a, Supplementary Fig. 3a, and Supplementary Table 1). Given that Mesh1 is highly expressed in larvae and suppressed in pupae25, highly expressed Mesh1 in larvae seems to promote degradation of ppGpp and lowered Mesh1 expression in pupae could suppress ppGpp degradation. However, it is unknown whether ppGpp in Drosophila tends to be degraded or induced at each stage. The concentration of ppGpp (per FW) in female adults was slightly, but significantly, higher than in male adults (Fig. 2a); however, the relative proportion of ppGpp to GTP in females was ~50% less than that observed in males (Supplementary Fig. 3a). The GTP concentration in mating females was also higher (~3-fold) than in mating males (Supplementary Fig. 3b), although this concentration difference was not found in adult virgins (Fig. 2a and Supplementary Fig. 3a), suggesting that the increased ppGpp and GTP in mating females was derived from fertile eggs.
ppGpp levels per fresh weight (FW) in various stages in w− (a), w− instar larvae grown under normal and germ-free (GF) conditions (b), Mesh1 LOF (c), and Mesh1 GOF (d). Values represent the mean ± SD (n = 3 biologically independent samples). *p = 0.0008, **p = 0.0384 (compared to w−), ***p = 0.0392 (compared to w−); two-sided Student’s t-test.
Bacterial cells contain a large amount of ppGpp (~470 nmol g−1 FW)33,34 and animals harbor symbiotic bacteria35, suggesting that the ppGpp detected in Drosophila may have originated from such bacteria. To investigate this hypothesis, we quantified ppGpp in germ-free Drosophila, which lack bacterial symbionts36. Indeed, ppGpp was definitively detected in germ-free Drosophila, although the concentration was significantly lower than in non-germ-free control flies (Figs. 1c and 2b). These results indicated that although ~50% of the detected ppGpp might have originated from symbiotic bacteria (Fig. 2b), some ppGpp is synthesized in Drosophila by an unknown enzyme(s). To confirm this, we tested for ppGpp in extracts of germ-free human cells (HeLa and PEAKrapid) and Drosophila S2 cells in culture. The chromatograms showed peaks at the same retention times as those of late third instar larvae of Drosophila (Fig. 1c). The ppGpp concentration in HeLa cells was ~40 pmol g−1 FW and the relative concentration of ppGpp to GTP in both HeLa and S2 cells was ~2.5 × 10–3 (Supplementary Table 1), which is comparable to that determined in Drosophila larvae. These results indicated that ppGpp is present in metazoa and its concentration varies during development.
Effect of Mesh1 function on ppGpp level
Mesh1 expressed in E. coli exhibits ppGpp hydrolase activity25,26. However, Mesh1 activity in vivo has never been tested because ppGpp has not been detected in metazoa25. To verify that metazoan Mesh1 has ppGpp hydrolase activity, we quantified ppGpp in Drosophila extracts of Mesh1 LOF and the Mesh1-overexpressing transformant (Mesh1 GOF: gain of function)25. The chromatograms of ppGpp extracted from Mesh1 LOF had a sharp ppGpp peak (Supplementary Fig. 2e, f) and the concentration of ppGpp in larvae was significantly (~7-fold) higher than in WT (Fig. 2c and Supplementary Table 1). The Mesh1 LOF pupae contained ~3-fold more ppGpp than did WT (Supplementary Fig. 3c). To induce Mesh1 expression in Drosophila, we applied a GAL4/UAS system37. We crossed UAS-Mesh1 transformants with flies expressing GAL4 and used an actin promoter, Act5c, for systemic and constitutive OE of Mesh1 (Mesh1 GOF). We then measured the amount of ppGpp in pupae. The concentration of ppGpp in Mesh1 GOF was approximately 347 pmol/g FW and in the control was 652 pmol g−1 FW (Supplementary Table 1), suggesting that ppGpp degradation was enhanced in Mesh1 GOF compared to that in the control strain, although no clear significance was observed (P = 0.21, t-test) (Fig. 2d). The ppGpp level in the control strain of Mesh1 GOF was higher than in w− (Supplementary Table 1). The reason for the difference is currently unclear; however, it might be caused by different genotypes. These results implied that Mesh1 has ppGpp hydrolase activity in vivo. We also found that GTP levels in Mesh1 LOF larvae and pupae were ~10% and 20% lower, respectively, than those in WT (w–) (Supplementary Fig. 3d). Given that GTP synthesis is suppressed by ppGpp in Gram-positive bacteria6, ppGpp may play a role in GTP homeostasis in metazoa, although GTP levels in Mesh1 GOF pupae were the same as those in WT (Supplementary Fig. 3e).
To further investigate Mesh1 and ppGpp function in Drosophila, we characterized the Mesh1 LOF mutant phenotype during hatching from larvae to pupae under starvation conditions. On rich medium (control), most larvae hatched at the second day after transfer and hatching rates from larvae to pupae were 82% for w– and 92% for Mesh1 LOF (Fig. 3a). On the starvation medium, the hatching date of Mesh1 LOF and w− larvae were around the fourth and the second day, respectively after transfer. This result is consistent with a previous study that reported that Mesh1 LOF larvae showed retarded growth25. Despite the delay of hatching of Mesh1 LOF, the hatching rate from larvae to pupae of Mesh1 LOF (75%) was the same as that of w− (74%) (Fig. 3a). To investigate the physiological importance of ppGpp for the starvation response, we measured the amount of ppGpp in Drosophila under rich and starved conditions. We used late third instar larvae (shown in Fig. 2c) as the control and larvae were grown in rich medium for 4 days after egg-laying (AEL), and then transferred to the starvation medium and incubated for one day for comparison with late third instar larvae. The starved Mesh1 LOF larvae weighed significantly more than the w– larvae (Supplementary Fig. 4a). Interestingly, the concentration of ppGpp in starved w− larvae did not differ from that of larvae in the control medium, whereas the concentration of ppGpp in Mesh1 LOF larvae was lower, i.e., the same as w− upon starvation (Fig. 3b and Supplementary Fig. 4b, c), indicating that ppGpp levels in w− larvae were normal under starvation conditions, but increased in rich medium in Mesh1 LOF. On the other hand, GTP levels (per FW) in Mesh1 LOF remained lower than that in w– upon starvation, although the GTP levels in w– were increased upon starvation (Supplementary Fig. 4d). These results suggested that the pupation delay of Mesh1 LOF (Fig. 3a) is not correlated with the ppGpp level itself, but is potentially due to the lowered GTP level in the mutant.
a Hatching rate of the larvae (4 days after egg-laying) of w– and Mesh1 LOF, which were initially grown on standard medium and then transferred to normal (Control) or starvation conditions. The colors indicate the number of flies from each stage (%); red: pupae, green: adults. b Concentrations of ppGpp in Mesh1 LOF under normal and starvation conditions. Values represent the mean ± SD (n = 3 biologically independent samples). Different letters (a and b) indicate significant differences between groups (p < 0.05; Tukey’s test).
ppGpp overaccumulation induces cell death in Drosophila
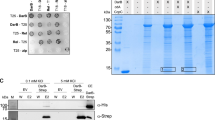
To further investigate the function of ppGpp in Drosophila, we heterologously expressed the Bacillus subtilis SAS YjbM (Fig. 1a), which has strong ppGpp synthase activity23. We produced a UAS-YjbM line, which drives yjbM expression constitutively when it is crossed with the actin-specific driver line Act5c-GAL4. We found that offspring having both UAS-yjbM and Act5c-GAL4 constructs were lethal, and progeny could not be produced. As an alternative, we transiently expressed yjbM under the control of the heat-shock-driven FLPase system (hs-yjbM) at day 3 AEL, when larvae show strong resistance against environmental changes38. hs-yjbM larvae could survive for several days after the induction of yjbM. The ppGpp chromatogram of the hs-yjbM exhibited a strong peak at 6.4 min (Fig. 4a), which corresponds to the retention time of the ppGpp standard (Fig. 1b), and indicated ~1200-fold higher ppGpp than in the control (Fig. 4b). After the heat shock, all the hs-yjbM died during the larvae and pupae stages (Fig. 4c), indicating that hyper-accumulation of ppGpp induces lethality in Drosophila, similar to that observed in the E. coli spoT mutant39.
a Guanosine tetraphosphate-specific MS chromatograms of nucleotides extracted from hs-yjbM OE (a heat-inducible yjbM overexpression line) and control larvae. b The concentration of ppGpp extracted from hs-yjbM OE larvae and the control (1 day after induction). Values represent the mean ± SD (n = 3 biologically independent samples). *p = 0.0227; two-sided Student’s t-test. c Drosophila development after induction of ppGpp production. Values represent the mean (n = 3 independent experiments). The colors indicate each stage of flies; blue: larvae, red: pupae, green: adults, light gray: dead larvae, gray: dead pupae.
As systemic expression of yjbM is lethal in Drosophila, we could not further investigate the influence of ppGpp accumulation. Thus, we conducted site-specific expression of yjbM by crossing with the eye-specific driver line Glass multimer reporter (GMR)-GAL4 (GMR-YjbM). We also used the temperature-dependent GAL4 repressor GAL80ts to reduce leaky expression of YjbM. Before induction of yjbM expression, the surface of the eyes was smooth, but abnormally glossy (Fig. 5a), suggesting partial induction of cell death perhaps owing to leaky expression of yjbM. After induction of yjbM, the centers of the eyes became more yellow, supporting the cell-death hypothesis. Microscopic analysis of the optic nerves inside the eyes revealed fragmentation of the axons (Fig. 5b). Similar axon fragmentation has been reported in axon degeneration and neural cell death40, suggesting that excess ppGpp is cytotoxic. On the other hand, the eye morphology of Mesh1 LOF did not differ from that of w–, as seen in both optical images and confocal images of dissected eyes (Supplementary Fig. 5), suggesting that the cell-death phenotype observed in the yjbM OE line may not be achieved with physiological concentrations of ppGpp.
a Optical images of eyes (bar = 100 μm) and b confocal fluorescence images of axons (visualized with red fluorescent protein; white bar = 20 μm, yellow bar = 5 μm) of flies overexpressing yjbM by the eye-specific GMR promoter. Flies incubated at 18 °C 1 day after emergence (before induction; upper) were transferred to 31 °C, to induce overexpression of yjbM in the eye. Images were acquired 1 day after induction.
Metabolic changes induced by altered Mesh1 expression
We next performed metabolome analysis of Mesh1 LOF and Mesh1 GOF to investigate Mesh1 function on metabolism. The levels of certain metabolites were significantly altered in the mutants compared with controls (Fig. 6, Supplementary Fig. 6, and Supplementary Tables 2 and 3). In Mesh1 LOF, tricarboxylic acid (TCA) cycle metabolites including 4-hydroxybenzoate, fumarate, and malate were higher than in WT (CS). In Mesh1 LOF, the level of sedoheptulose 7-phosphate (S7P), an intermediate in the pentose phosphate pathway, was lower than that in WT (Supplementary Table 2). Intermediates in the transsulfuration pathway, CySSG was higher, and taurine and NADP+ levels were lower in Mesh1 LOF than in the WT (Fig. 6a and Supplementary Table 2). In Mesh1 GOF, two arginine metabolites, citrulline and creatine, were lower than in the control (Fig. 6b and Supplementary Table 3). Some intermediates in the methionine cycle and the transsulfuration pathway were also altered in Mesh1 GOF. Specifically, methionine and cystathionine levels were lower, and adenosine and pyroglutamic acid levels were higher in Mesh1 GOF than in the control (Fig. 6b and Supplementary Table 3). The levels of six metabolites were altered in Mesh1 LOF and Mesh1 GOF, and three of them, including the two TCA cycle metabolites, malate and fumarate, were higher in Mesh1 LOF and lower in Mesh1 GOF (Fig. 6 and Supplementary Fig. 6). These results imply that Mesh1 influences both the TCA cycle and urea cycle in mitochondria, as well as the transsulfuration pathway and pentose phosphate pathway in the cytosol. In Mesh1 GOF, carboxymethyl lysine and pyroglutamic acid, one of the advanced glycoxidation end products and a glutathione metabolite, respectively, were increased (Fig. 6b and Supplementary Table 3). Advanced glycoxidation end products are involved in aging and in the development of many degenerative diseases in humans41. These results suggested that Mesh1 is important for maintaining some metabolic homeostasis in animal cells42.
Statistically significant changes in the concentrations of metabolites in Mesh1 LOF larvae compared with concentrations in w– larvae (p < 0.05, n = 6 biologically independent samples; two-sided Student’s t-test) (a), and in Mesh1 GOF larvae compared with those in the control (p < 0.05, n = 6 biologically independent samples; two-sided Student’s t-test) (b). Fold-change values in metabolite accumulation are expressed along a color gradient. 3PG: 3-phosphoglyceric acid; 5-OxoPro: pyroglutamic acid; CML: carboxymethyl lysine; CysSG: cysteine-glutathione disulfide; GAA: guanidinoacetic acid; GABA: gamma-aminobutyric acid, GSH: glutathione; GSSG: glutathione disulfide; PEP: phosphoenolpyruvate; SAH: S-adenosylhomocysteine; SAM: S-adenosylmethionine.
Discussion
It is well established that ppGpp is a bacterial and plastidial second messenger1,2,3. However, ppGpp function in metazoa has not been determined because ppGpp had not been detected in metazoan cells in spite of many trials27. Here we developed a high-sensitivity quantification method for ppGpp in metazoan cells by modification of our previous ppGpp quantification method used for plants and bacteria32. In the method, total nucleotide pools extracted with 2 M formic acid are subjected to a reversed-phase/weak anion exchange column in buffer containing ethylenediaminetetraacetic acid (EDTA), by which low amounts of ppGpp extracted from metazoa were efficiently concentrated. It is possible that chelating excess cations by EDTA in the solvent helps ppGpp adsorb to the anion exchange column. A chloroform extraction step was also added before LC-MS/MS, which resulted in enhancement of the ppGpp peak. The established method allowed us to stably detect small amounts of ppGpp and the unknown guanosine tetraphosphate molecule from Drosophila and human cells by LC-MS/MS (Fig. 1b and Supplementary Figs. 1 and 2). Given fragmentation patterns of ppGpp and the unknown molecule are nearly identical (Supplementary Fig. 1b), the structures of the two must be very similar. One possible candidate for the unknown molecule is guanosine 2′,5′-bis(pyrophosphate), although this molecule has never been detected in cells. Further structural characterization of the unknown molecule should be performed in the future.
By use of the established method, we succeeded in detecting ppGpp in metazoan cells (Fig. 2 and Supplementary Figs. 2 and 3). ppGpp levels in Drosophila pupae were significantly higher than those in other stages (Fig. 2a and Supplementary Fig. 3a), implying that ppGpp is involved in metamorphosis. Mesh1 LOF showed higher levels of ppGpp (Fig. 2c and Supplementary Fig. 3c), suggesting that Mesh1 works as a ppGpp hydrolase in vivo in Drosophila, and that ppGpp metabolic systems have been conserved in metazoa. One could expect that ppGpp would be accumulated upon starvation in Metazoa as it is in bacteria1; however, ppGpp levels in w− before and after starvation treatment were the same (Fig. 3b and Supplementary Fig. 4b), suggesting that ppGpp synthesis is not upregulated upon starvation in Drosophila. Alternatively, ppGpp synthesis may be upregulated only in small areas of specific organs under starvation conditions. Notably, the ppGpp levels in Mesh LOF were significantly reduced upon starvation (Fig. 3b), suggesting the existence of a ppGpp-specific hydrolase other than Mesh1. In Arabidopsis, ppGpp is rapidly accumulated upon light-to-dark transition and the accumulated ppGpp is degraded in several hours in the dark32,43. The efficient degradation of ppGpp in plants is achieved by not only RSHs, but also several nucleotide diphosphate linked some moiety X (Nudix) hydrolase enzymes, similar to the process that occurs in bacteria44,45. Given that Nudix enzymes are conserved in Metazoa46, ppGpp levels in Metazoa are also likely controlled by multiple enzymes including Mesh1 and Nudix hydrolases, as well as unknown ppGpp synthases.
Although the physiological role of Mesh1-dependent ppGpp control in Metazoa is still largely unknown, our metabolome analyses suggested that enzymes involved in purine metabolism47 are regulated by Mesh1 activity. In Mesh1 GOF, adenosine and adenine levels were higher and hypoxanthine level was lower than those in the control (Supplementary Table 3). In B. subtilis, (p)ppGpp directly inhibits certain enzymes involved in GTP synthesis, including guanylate kinase and hypoxanthine phosphoribosyltransferase3, and this inhibition reduces the cellular GTP pool and retards transcription by reducing the concentrations of factors (e.g., ribonucleotides) necessary for transcription48. The reduced levels of hypoxanthine observed in Mesh1 GOF (Supplementary Table 3) were likely due to the reduction of ppGpp (Fig. 2d) and metazoan hypoxanthine phosphoribosyltransferase conserves the amino acid residues necessary for ppGpp binding in bacterial enzymes49. Together with the increased concentration of GTP in the Mesh1 LOF (Supplementary Fig. 3d), these results imply that Mesh1-dependent ppGpp control in metazoans regulates the GTP/ATP ratio by controlling purine nucleotide metabolism, as in bacteria. Moreover, other GTP-binding proteins could be ppGpp targets50.
hMESH1 was recently found to have NADPH phosphatase activity31. The NADP(H) concentration in the hMESH1-knockdown line was ~2 times higher than that in the control31, suggesting that Mesh1 has a dual function to sustain levels of intracellular ppGpp and NADPH. However, NADP+ concentrations in the Drosophila Mesh1 LOF were about half of that in WT (P = 0.050) (Fig. 6a and Supplementary Table 2); this result is the opposite of what is observed in the hMESH1-knockdown line. Although the reason for this variation is not known, one possibility is that null mutation of Mesh1 induces a decrease of the NADP(H) pool size to compensate for the constitutive increase of NADP(H) levels upon loss of NADPH phosphatase activity of Mesh1.
Recently, the impact of artificial accumulation of ppGpp in the yeast Saccharomyces cerevisiae was examined by heterologous expression of E. coli relA, which revealed that accumulated ppGpp induces expression of some mitochondrial proteins such as cytochrome oxidase and the succinate dehydrogenase homolog SHH4, the latter of which catalyzes fumarate synthesis from succinate in the TCA cycle51. In this study, we found that Mesh1 LOF accumulates fumarate and malate (Fig. 6a and Supplementary Table 2), suggesting that highly accumulated ppGpp in Mesh1 LOF influences TCA cycle metabolisms in mitochondria. Previous microarray analysis of the Drosophila Mesh1 LOF indicated that expression of genes for mitochondrial function, lipid and carbohydrate metabolisms were altered in the mutant25, supporting the hypothesis. The ppGpp-accumulating S. cerevisiae also has upregulated expression of 6-phosphogluconolactonase SOL4, transketolase TKL2, and transaldolase NQM1, all of which function in the pentose phosphate pathway51. TKL2 catalyzes the synthesis of S7P and glyceraldehyde 3-phosphate (GAP), and NQM1 catalyzes synthesis of fructose 6-phosphate and erythrose 4-phosphate from S7P and GAP52. The concentration of S7P in Mesh1 LOF was significantly decreased compared to that in the control (Supplementary Table 2), suggesting that accumulated ppGpp changes metabolism in the pentose phosphate pathway in Drosophila. Given the pentose phosphate pathway requires NADP+ 53, one could assume that inhibition of ppGpp hydrolase and NADP(H) phosphatase activities of Mesh1 result in increasing ppGpp and NADP(H) pool sizes that could consequently upregulate pentose phosphate pathway activity. This further supports the physiological bases of the dual activities of Mesh1 as ppGpp hydrolase and NADPH phosphatase to control specific metabolisms in metazoa.
We observed retarded growth of Mesh1 LOF under starvation conditions (Fig. 3a), as reported previously25. The pupation rate of Mesh1 LOF was the same as that of WT, suggesting that the viabilities of Mesh1 LOF and WT were comparable (Fig. 3a). On the other hand, the viability of Mesh1 LOF was reported to be 70% of that in WT under starvation conditions in the previous study25. This inconsistency may be due to different timing of transfer of the larvae to the starved medium. In the previous study, larvae were transferred to starvation medium at 40 h AEL, whereas we transferred larvae to the starvation medium at 4 days AEL. These results suggest that Mesh1 function in Drosophila is more important to endure the starvation at early larval stages than in later periods. It should be noted that it is still not possible to distinguish phenotypes caused by loss of ppGpp hydrolase and/or NADPH phosphatase activities in Mesh1 LOF and GOF. Clearly, further studies are needed to determine how ppGpp works in metazoan cells.
Overall, our method for quantifying ppGpp in animal cells should provide opportunities for investigating the stringent response in metazoa. Since our research revealed that ppGpp and its metabolic system are conserved in metazoan cells, which was a decade-old enigma, this result should provide useful insights for future studies such as understanding how ppGpp is synthesized and how it functions in metazoan cells.
Methods
Fly strains, genetics, and growth conditions
Flies were kept in standard Drosophila medium at 25 °C, except for activating GAL80ts (18 °C). To activate the heat-shock (hs) promoter, flies were transferred from 18 °C to 31 °C 1 day after emergence. To calculate the concentration of ppGpp in each stage of Drosophila, eggs were collected for 19 h on an apple juice medium, larvae at third instar larvae, pupae 48 h after pupation, virgin during 0–8 h after hatching, and adults 4 days after culture on the standard medium. To induce yjbM OE on day 3 AEL, hs-yjbM was heat-shocked for 30 min at 37 °C and then maintained for 10 min at room temperature. This heat-shock procedure was then repeated. To compare the pupation rate and ppGpp level under starvation conditions, eggs from flies of each genotype were collected over 2 h on an apple juice medium. Eggs were aged on standard Drosophila medium at room temperature (25 °C) and the standard medium was used as the normal growth condition (see below for recipe). To assess the effects of starvation, larvae at 4 days AEL under medium were transferred to starvation medium (see below for recipe). The following fly stocks and mutant alleles were used: CS, w1118 (w–), Mesh1 LOF mutant, UAS-Mesh1, UAS-yjbM, Act5C-Gal4, Act5c-Gal4-tubGAL80ts, hsFLP-AyGAL4-UAS-GFP, and GMR-myr-RFP-GMR-GAL4-tubGal80ts. The UAS-yjbM (residues 1–636 with a C-terminal FLAG tag) fly was produced as follows. At first, B. subtilis yjbM was amplified with a primer pair, 5′-AGGGAATTGGGAATTCAAAATGGATGACAAACAGTGG-3′ and 5′-ATCTGTTAACGAATTCTTACTTATCATCATCATCCTTATAATC-3′, and synthesized yjbM gene54 as a template. The amplified fragment was cloned into EcoRI-cut pUASTattB55 by In-fusion cloning kit (Clontech), which was integrated into the specific site (VK31) of chromosome III with the ΦC31 system55. Transformants were produced by BestGene, Inc. (USA). Mesh1 LOF and UAS-Mesh1 were kindly provided by J. Chung (Seoul National University)25. CS was used as the WT strain for the detection of ppGpp from eggs. w– was used as the control strain for comparison with mutants. Eye-specific expression was achieved using the GAL4-UAS expression system37. GMR-GAL4 was used to induce expression in eyes56. To analyze the yjbM OE line, we made +/+; +/+; Act5c-GAL4, Tub-Gal80ts/UAS-yjbM (or +) (yjbM OE), hs-FLP/w; AyGAL4, UAS-GFP/+; +/UAS-yjbM (or +) (hs-yjbM OE), and GMR-myr-RFP/w; GMR-GAL4, tub-Gal80ts/+; +/UAS-yjbM (or +) (GMR-yjbM OE). To analyze the Mesh1 GOF, we crossed UAS-Mesh1 flies with yw; Act5c-Gal4/TM6B and w− flies with yw; Act5c-Gal4/TM6B as a control. We sampled +/yw; UAS-Mesh1/+; +/ Act5c-Gal4 as the Mesh1 GOF line and w/yw; +/+; +/ Act5c-Gal4 as the control.
Gene expression
To evaluate the effects of ppGpp accumulation, we used the GAL4/UAS system37. We used transformants that had UAS-yjbM or UAS-Mesh1, which were silent in the absence of GAL4. They were crossed with flies expressing GAL4. In the progeny of this cross, GAL4 protein activated transcription of yjbM or Mesh1 by binding to a UAS37. GMR-GAL4 was used for induction of photoreceptor gene expression. The GMR sequence is an engineered transcription regulatory region containing a binding site for the protein gl, a photoreceptor-specific transcription factor56. myr is the sequence from D. melanogaster Src64B that includes the myristoylation signal and is used to target a protein to the plasma membrane. To induce yjbM expression at a specific developmental time, we used GAL80ts or the flippase (flp)/FRT system in conjunction with the GAL4/UAS system, which was controlled by the hs promoter38,57. GAL80ts is a temperature-sensitive GAL80ts gene. At 18 °C, GAL80ts binds GAL4 and this heterodimer works as a GAL4 repressor. Upon transfer to a temperature above 29 °C, GAL80ts is unable to bind GAL4 and its repressive function is lost57. Thus, from routine culture at 18 °C, flies were transferred to 31 °C when induction of yjbM was desired. With the flp/FRT system, the transformants had three genes to control gene expression: heat shock-flp fusion gene, the Act5C promoter-GAL4 fusion gene with a FRT cassette containing a transcription termination signal, and UAS-yjbM. Before heat shock, Act5C-GAL4 was interrupted by the transcription termination signal in the FRT cassette. Heat shock induced the expression of flp encoding the flippase, which excised the FRT cassette from the fusion gene. The GAL4 protein expressed from the rearranged fusion gene activated transcription of UAS-yjbM38.
Culture medium for Drosophila
For the standard medium, 15 g soybean flour, 10 g agar, 100 g cornmeal, 30 g yeast, 30 g malt, and 50 g glucose were mixed in 1 l water and then heated at 90 °C for 1.5 h. After the mixture had cooled, 2 ml propionic acid and 2.5 g nipagin in 5 ml 100% ethanol were added. The mixture was then dispensed into columned vials or bottles until it covered the bottom to a depth of ~2 cm. The starvation medium consisted of 20% (w/v) sucrose-agar medium containing 0.5% ethanol and 0.2% propionic acid in distilled/deionized water58.
Germ-free larvae
The w1118 flies were obtained from the Bloomington Stock Center (Bloomington, IN) and used as the standard strain. Adult flies were maintained in autoclaved standard yeast medium containing tetracycline (50 µg/ml) for three generations to eliminate the common intracellular bacteria Wolbachia pipientis. Embryos of the flies were then bleached using 2.7% sodium hypochlorite and washed twice in 70% ethanol, followed by three washes in sterile water. Washed embryos were transferred to autoclaved yeast vials without tetracycline. The third instar larvae were collected and bacterial contamination in larvae was assessed by PCR using 16S rDNA universal primers 8FE (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492 R (5′-GGMTAGCTTGTTACGACTT-3′).
Cell culture
Human cervical cancer cells (HeLa) and human embryonic kidney cells (PEAKrapid) were obtained from the American Type Culture Collection (Manassas, VA). Cells were cultured in Dulbecco’s modified Eagle medium (Sigma-Aldrich, St. Louis, MO) supplemented with 10% fetal bovine serum (Thermo Fisher Scientific, Waltham, MA), 100 U ml−1 penicillin (Nacalai Tesque, Kyoto, Japan), and 100 µg ml−1 streptomycin (Nacalai Tesque) at 37 °C in 5% CO2. Drosophila S2 cells (GIBCO, R690-07) were cultured at 25 °C in Schneider’s medium supplemented with 10% heat-inactivated fetal bovine serum (Thermo Fisher Scientific) and 1× antibiotic–antimycotic (Thermo Fisher Scientific). Plastic Petri dishes were used for cell culture, and harvested cells were frozen at −70 °C until use.
Phenotypic analysis of Drosophila
To analyze the effect of ppGpp on cell proliferation under starvation conditions, 50 flies of w– and those of Mesh1 LOF that had developed for 3 days (i.e., 3 days AEL) were transferred and kept on standard or starvation medium. The number of pupae and adults in the medium were counted every 24 h. hs-yjbM OE was used for the analysis of the effect of excess ppGpp. To induce yjbM OE at day 3 AEL, hs-yjbM OE was heat-shocked for 30 min at 37 °C and then held at 10 min at room temperature, and this heat-shock procedure was then repeated. After heat shock, the larvae were cultured for 24 h on standard medium containing Formula 4–24® Instant Drosophila Medium, Blue (Carolina Biological Supply Company) at 25 °C. Larvae for the two genotypes were then gathered after incubation: hs-FLP-GFP/w; AyGAL4, UAS-GFP/+, +/yjbM (or +). Twenty larvae were transferred to standard medium; every 24 h, all flies were assessed as living or dead and as larvae, pupae, or adults. To observe the effects of excess ppGpp on eye development, GMR-yjbM adults and other control genotypes were transferred from 18 °C to 31 °C 1 day after emergence. After 0 (as a control) or 1 day incubation, the eyes and dissected eyes were examined microscopically.
Immunohistochemistry and imaging
Brains of the 1-day-old adult flies (or those of 1 day after induction of YjbM) was dissected in phosphate-buffered saline and fixed by 4% paraformaldehyde solution. After several washes, the brains were incubated with primary (o/n) and secondary (2 h) antibodies sequentially. After immersing in the Vectashield mounting medium (Vector Laboratories Inc.), the brains were mounted vertically on the slide glass59. The primary antibody for the photoreceptor cell-specific membrane protein, chaoptin, was mAb24B10 (1:50, DSHB). The secondary antibodies were conjugated to Alexa488 (1:400, Life Technologies). Images were obtained with Nikon C2+ confocal microscopes and processed with NIS-elements AR and Adobe Photoshop.
Quantification of ppGpp and GTP by LC-ESI-MS/MS
ppGpp was extracted from cells and quantified as described by Ihara et al.32 with slight modifications as detailed below. Standards for ppGpp (≥85%), GTP (92%), and GTP (>99%) were purchased from Trilink, TaKaRa, and Bio Basic, Inc., respectively, and used without further purification. Fifty flies grown under normal conditions or ~65 mg of flies grown under starvation conditions were collected for all quantifications, except that only one larva was collected for quantification of ppGpp in YjbM OE. The collected samples were frozen in liquid nitrogen, broken into pieces using a pestle and mortar, and then extracted in 3 ml of 2 M formic acid/1 mM EDTA pH 4.5. Extracts were incubated for 30 min on ice. Subsequently, 50 mM ammonium acetate/1 mM EDTA pH 4.5 was added to the crushed tissue to a volume of 6 ml and the extracts were divided into two 3 ml portions. To estimate the ppGpp recovery rate32, 25 µl of 2 µM ppGpp was added to one of the two solutions (the standard mixed sample). For GTP quantification, this standard mixing step was omitted, because the GTP recovery rate was constant, perhaps because of its high concentration. To remove hydrophobic debris, 1.5 ml chloroform was added, the mixtures shaken three times, centrifuged at 1600 × g for 1 min at room temperature, and each aqueous (upper) layer transferred to a new tube.
The samples were then subjected to solid-phase extraction. A 1-ml OASIS WAX column 30 mg (Waters) was rinsed with 1 ml of methanol followed by 1 ml ammonium acetate pH 4.5. Each sample solution was loaded onto the column, which was then washed with 1 ml of ammonium acetate pH 4.5 followed by 1 ml of methanol. Nucleic acids were eluted from the column with 1 ml methanol/deionized H2O/25% aqueous ammonia (20:70:10) solution. The eluate was lyophilized in a Proteosave SS 15 ml Conicaltube (Sumitomo Bakelite), dissolved in 200 µl in Milli-Q water, and then filtered through a FavorPrepTM Plasmid DNA Extraction Column (FAVORGEN). A 94 µl portion of the eluate was then mixed with 6 µl acetonitrile in a PSVial (AMR).
A 10 µl aliquot of this solution was injected into an Acquity UPLC system (Waters) and separated on an ACQUITY UPLC® BEH C18 column (2.1 × 50 mm, 1.7 µm particle size; Waters). Nucleotides were resolved with a linear gradient of two mobile phases: solvent A (500 ml of 8 mM N,N-Dimethylhexylamine with 80 µl of acetic acid) and B (acetonitrile) at a flow rate of 0.3 ml min−1, as follows: initially 0% B; 0–40% B over 10 min; 40–0% B over 10.5 min; reequilibration at 0% B over 20 min. Effluents from UPLC were introduced online into an ACQUITY TQD tandem quadrupole mass spectrometer with an electrospray ionization (ESI) interface (Waters). Argon was used as collision gas at 0.3 ml min−1. Negative ESI-MS/MS detection was carried out in the multiple-reaction monitoring mode or product ion scan mode using the following parameters: capillary voltage, 2.5 kV; source temperature, 150 °C; desolvation temperature, 400 °C; cone gas flow, 50 l h−1; desolvation flow, 800 l h−1; LM1 resolution, 13; LM 2 resolution, 15. The cone voltage and collision energy were 44 V and 44 eV, respectively, for ppGpp, and 40 V and 32 eV, respectively, for GTP. The multiple-reaction monitoring transitions were 602→159 for ppGpp and 522→159 for GTP.
The ESI source was set to negative-ion mode with a capillary voltage of 2.5 kV. The source and desolvation temperatures were set to 150 °C and 400 °C, respectively. ppGpp was detected in multiple-reaction monitoring mode. The cone voltage and collision energy were 44 V and 44 eV, respectively. The mass of the parent ion (602 m/z) was selected by the first quadrupole and fragmented in the collision cell to the target ion (159 m/z).
ppGpp and GTP levels were quantified against standard curves. Standard solutions consisted of 20 µl of 2 µM ppGpp or 0.4 mM GTP mixed with 12 µl of acetonitrile and 168 µl of Milli-Q water in a PSVial (AMR). The standard curves were linear at concentrations between 1.5625 nM and 200 nM for ppGpp, and 2500 nM and 40,000 nM for GTP. The calibration curves were used when the r2 values of the plotted calibration data were >0.99. The final ppGpp concentration in each sample was corrected based on its recovery rate calculated by the ppGpp standard mixed sample, as reported32. Obtained data are shown in Supplementary Data File S1.
The LC-MS/MS-based ppGpp quantification method described above was used to detect pppGpp with the multiple-reaction monitoring transitions at 682→159 and 234. For the experiment, pppGpp standard12 was prepared as follows. At first, B. subtilis yjbM was amplified with a primer pair, 5′-AAGGAGATATACATATGGATGACAAACAGTGGG-3′ and 5′-GGTGGTGGTGCTCGAGTTGCTGCTCAGATCCTTTC-3′, and synthesized yjbM gene54 as a template. The amplified fragment was cloned into NdeI-XhoI-cut pET29a (Merck) by In-fusion cloning kit (Clontech). The C-terminal His-tagged YjbM was expressed in the E. coli strain BL21(DE3) and purified by His-bind resin (Novagen) according to the manufacturer’s recommendations. The purified YjbM was mixed with 5 mM GTP and 5 mM ATP in a buffer containing 50 mM Tris/HCl pH 7.5, 200 mM NaCl, 20 mM MgCl2, and 20 mM KCl, and incubated for 30 min at room temperature, followed by chloroform extraction. The aqueous phase that contained the nucleotides was subjected to anion exchange chromatography (HitrapQ HP, 1 mL; GE Healthcare). A gradient ranging from 0–2 M NaCl was used to separately elute the nucleotides.
Metabolome analysis by CE-TOF MS
Ten flies of late third instar larvae were collected for metabolome analysis. Each sample was extracted in 500 µl methanol containing 8 µM of each of two reference compounds, namely methionine sulfone for the cation analysis and camphor 10-sulfonic acid for the anion analysis, using a Retsch mixer mill MM310 at a frequency of 27 Hz for 1 min. The extracts were then centrifuged at 15,000 × g for 3 min at 4 °C. The supernatant was transferred into a tube, and 500 µl chloroform and 200 µl water were added to the tube to perform liquid–liquid extraction. The upper layer (water) was transferred to a new tube, and evaporated for 30 min at 45 °C by a centrifugal concentrator to obtain two layers. The resulting partially evaporated liquid was then centrifugally filtered through a PALL Nanosep 3-kDa cutoff filter at 9100 × g for 90 min at 4 °C to remove high-molecular-weight compounds such as oligo-sugars. The filtrate was evaporated to dryness for 120 min using a centrifugal concentrator. The residue (ca. 25 mg for each sample) was dissolved in 20 µl water containing 200 µM of each of the internal standards, namely 3-aminopyrrolidine for the cation analysis and trimesic acid for the anion analysis, that were used to compensate for differences in migration time in the peak annotation step.
All CE-TOF MS experiments were performed using a Model G7100A CE Instrument (Agilent Technologies, Sacramento, CA), an Agilent G6224A TOF LC/MS system, an Agilent 1200 Infinity series G1311C Quad Pump VL, an G1603A Agilent CE-MS adapter, and a G1607A Agilent CE-ESI-MS sprayer kit. The G1601BA 3D-CE ChemStation software for CE and G3335-64002 MH Workstation were used. Separations were carried out using a fused silica capillary (50 µm i.d. × 100 cm total length) filled with 1 M formic acid for cation analyses or with 20 mM ammonium formate pH 10.0 for anion analyses as the electrolyte. The capillary temperature was maintained at 20 °C. The 15 nl sample solutions were injected at 50 mbar for 15 s. The sample tray was cooled below 10 °C. Prior to each run, the capillary was flushed with electrolyte for 5 min. The applied voltage for separation was set at 30 kV. Methanol/water (50% v/v) containing 0.5 µM reserpine was delivered as the sheath liquid at 10 µl min−1. ESI-TOF MS was conducted in the positive-ion mode for cation analyses or in the negative-ion mode for anion analyses, and the capillary voltage was set at 30 kV. A flow rate of heated dry nitrogen gas (heater temperature 300 °C) was maintained at 10 l min−1. The fragmentor, skimmer, and Oct RFV voltages were automatically set at optimum values. Automatic recalibration of each acquired spectrum was performed using the masses of reference standards. The methanol dimer ion ([2 M + H]+, m/z 65.0597) and reserpine ([M + H]+, m/z 609.2806) for cation analyses or the formic acid dimer ion ([2M-H]–, m/z 91.0037) and reserpine ([M-H]–, m/z 607.2661) for anion analyses provided the lock mass for exact mass measurements. Exact mass data were acquired at a rate of 1.5 cycles s−1 over a 50–1000 m/z range. In every single sequence analysis (maximum 36 samples) with our CE-TOF MS system, we analyzed the standard compound mixture both before and after each analysis of a sample. The detected peak area of the standard compound mixture was compared at regular intervals to assess sensitivity and reproducibility. The standard compound mixture was composed of major detectable metabolites, including amino acids and organic acids, and this mixture was freshly prepared at least once every 6 months. For all analyses, there were no differences in the sensitivity of detection of the standard compounds mixture.
An original data file (d) was converted to a unique binary file (.kiff) using in-house software. Peak picking and alignment were performed using another in-house software package in which peaks were picked and aligned among samples automatically. Using the detected m/z and migration time values of standard compounds, including internal standards, peaks were annotated automatically using the same software. For normalization, the individual areas of the detected peaks were divided by the peak area of the relevant internal reference standards. Using the calibration curves for standard compounds, peak area values were converted into values corresponding to amounts. Obtained data are shown in Supplementary Data File 1.
Statistics and reproducibility
Statistical significance of data was tested by two-sided Student’s t-test and/or Tukey’s test by used of Excel and MEPHAS (http://www.gen-info.osaka-u.ac.jp/MEPHAS/tukey-e.html), respectively. The sample size (n) and the nature of replicates have been given wherever relevant.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The dataset generated and analyzed in the current study is available as Supplementary Data File 1. All other data (if any) are available from the corresponding author upon request.
References
Potrykus, K. & Cashel, M. (p)ppGpp: still magical? Annu. Rev. Microbiol. 62, 35–51 (2008).
Ronneau, S. & Hallez, R. Make and break the alarmone: regulation of (p)ppGpp synthetase/hydrolase enzymes in bacteria. FEMS Microbiol. Rev. 43, 389–400 (2019).
Liu, K., Bittner, A. N. & Wang, J. D. Diversity in (p)ppGpp metabolism and effectors. Curr. Opin. Microbiol. 24, 72–79 (2015).
Ross, W., Vrentas, C. E., Sanchez-Vazquez, P., Gaal, T. & Gourse, R. L. The magic spot: a ppGpp binding site on E. coli RNA polymerase responsible for regulation of transcription initiation. Mol. Cell 50, 420–429 (2013).
Ross, W. et al. ppGpp binding to a site at the RNAP-DksA interface accounts for its dramatic effects on transcription initiation during the stringent response. Mol. Cell 62, 811–823 (2016).
Kriel, A. et al. Direct regulation of GTP homeostasis by (p)ppGpp: a critical component of viability and stress resistance. Mol. Cell 48, 231–241 (2012).
Feng, B. et al. Structural and functional insights into the mode of action of a universally conserved Obg GTPase. PLoS Biol. 12, e1001866 (2014).
Mitkevich, V. A. et al. Thermodynamic characterization of ppGpp binding to EF-G or IF2 and of initiator tRNA binding to free IF2 in the presence of GDP, GTP, or ppGpp. J. Mol. Biol. 402, 838–846 (2010).
Liu, K. et al. Molecular mechanism and evolution of guanylate kinase regulation by (p)ppGpp. Mol. Cell 57, 735–749 (2015).
Gallant, J., Irr, J. & Cashel, M. The mechanism of amino acid control of guanylate and adenylate biosynthesis. J. Biol. Chem. 246, 5812–5816 (1971).
Cashel, M. & Gallant, J. Two compounds implicated in the function of the RC gene of Escherichia coli. Nature 224, 488–490 (1969).
Steinchen, W. et al. Catalytic mechanism and allosteric regulation of an oligomeric (p)ppGpp synthetase by an alarmone. Proc. Natl Acad. Sci. USA 112, 13348–13353 (2015).
Ernst-August, H., Monika, G. & Dietmar, R. Degradation of guanosine 3′ diphosphate 5′ diphosphate in vitro by the spoT gene product of Escherichia coli. Eur. J. Biochem. 89, 125–131 (1978).
Atkinson, G. C., Tenson, T. & Hauryliuk, V. The RelA/SpoT homolog (RSH) superfamily: distribution and functional evolution of ppGpp synthetases and hydrolases across the tree of life. PLoS ONE 6, e23479 (2011).
Potrykus, K., Murphy, H., Philippe, N. & Cashel, M. ppGpp is the major source of growth rate control in E. coli. Environ. Microbiol. 13, 563–575 (2011).
Hobbs, J. K. & Boraston, A. B. p)ppGpp and the stringent response: an emerging threat to antibiotic therapy. ACS Infect. Dis. 5, 1505–1517 (2019).
Hood, R. D., Higgins, S. A., Flamholz, A., Nichols, R. J. & Savage, D. F. The stringent response regulates adaptation to darkness in the cyanobacterium Synechococcus elongatus. Proc. Natl Acad. Sci. USA 113, E4867–E4876 (2016).
Kasai, K. et al. A RelA-SpoT homolog (Cr-RSH) identified in Chlamydomonas reinhardtii generates stringent factor in vivo and localizes to chloroplasts in vitro. Nucleic Acids Res. 30, 4985–4992 (2002).
Maekawa, M. et al. Impact of the plastidial stringent response in plant growth and stress responses. Nat. Plants 1, 15167 (2015).
Ito, D., Ihara, Y., Nishihara, H. & Masuda, S. Phylogenetic analysis of proteins involved in the stringent response in plant cells. J. Plant Res. 130, 625–634 (2017).
Brown, A., Fernández, I. S., Gordiyenko, Y. & Ramakrishnan, V. Ribosome-dependent activation of stringent control. Nature 534, 277–280 (2016).
Battesti, A. & Bouveret, E. Acyl carrier protein/SpoT interaction, the switch linking SpoT-dependent stress response to fatty acid metabolism. Mol. Microbiol. 62, 1048–1063 (2006).
Nanamiya, H. et al. Identification and functional analysis of novel (p)ppGpp synthetase genes in Bacillus subtilis. Mol. Microbiol. 67, 291–304 (2008).
Steinchen, W. et al. Structural and mechanistic divergence of the small (p)ppGpp synthetases RelP and RelQ. Sci. Rep. 8, 1–10 (2018).
Sun, D. et al. A metazoan ortholog of SpoT hydrolyzes ppGpp and functions in starvation responses. Nat. Struct. Mol. Biol. 17, 1188–1194 (2010).
Zhu, M. & Dai, X. Growth suppression by altered (p)ppGpp levels results from non-optimal resource allocation in Escherichia coli. Nucleic Acids Res. 47, 4684–4693 (2019).
Silverman, R. H. & Atherly, A. G. The search for guanosine tetraphosphate (ppGpp) and other unusual nucleotides in eucaryotes. Microbiol. Rev. 43, 27–41 (1979).
Smulson, M. Amino add deprivation of human cells: effects on RNA synthesis, RNA polymerase, and ribonudeoside phosphorylation. Biochim. Biophys. Acta 199, 537–540 (1970).
Rhaese, H. J. Studies on the control of development synthesis of regulatory nucleotides, HPN and MS, in mammalian cells in tissue cultures. FEBS Lett. 53, 2–7 (1975).
Thammana, P., Buerk, R. R. & Gordon, J. Absence of ppGpp production in synchronised balb/c mouse 3T3 cells on isoleucine starvation. FEBS Lett. 68, 187–190 (1976).
Ding, C.-K. C. et al. MESH1 is a cytosolic NADPH phosphatase that regulates ferroptosis. Nat. Metab. 2, 270–277 (2020).
Ihara, Y., Ohta, H. & Masuda, S. A highly sensitive quantification method for the accumulation of alarmone ppGpp in Arabidopsis thaliana using UPLC-ESI-qMS/MS. J. Plant Res. 128, 511–518 (2015).
Traxler, M. F. et al. The global, ppGpp-mediated stringent response to amino acid starvation in Escherichia coli. Mol. Microbiol. 68, 1128–1148 (2008).
Glazyrina, J. et al. High cell density cultivation and recombinant protein production with Escherichia coli in a rocking-motion-type bioreactor. Microb. Cell Fact. 9, 1–11 (2010).
Shibata, T. et al. Crosslinking of a peritrophic matrix protein protects gut epithelia from bacterial exotoxins. PLoS Pathog. 11, 1–15 (2015).
Shibata, T. et al. Transglutaminase-catalyzed protein-protein cross-linking suppresses the activity of the NF-κB-like transcription factor relish. Sci. Signal. 6, ra61 (2013).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Ito, K., Awano, W., Suzuki, K., Hiromi, Y. & Yamamoto, D. The Drosophila mushroom body is a quadruple structure of clonal units each of which contains a virtually identical set of neurones and glial cells. Development 124, 761–771 (1997).
Xiao, H. et al. Residual guanosine 3′,5′-bispyrophosphate synthetic activity of relA null mutants can be eliminated by spoT null mutations. J. Biol. Chem. 266, 5980–5990 (1991).
Gerdts, J., Brace, E. J., Sasaki, Y., DiAntonio, A. & Milbrandt, J. SARM1 activation triggers axon degeneration locally via NAD+ destruction. Science 348, 453–457 (2015).
Vistoli, G. et al. Advanced glycoxidation and lipoxidation end products (AGEs and ALEs): an overview of their mechanisms of formation. Free Radic. Res. 47, 3–27 (2013).
Sarubbi, E., Rudd, K. E. & Cashel, M. Basal ppGpp level adjustment shown by new spoT mutants affect steady state growth rates and rrnA ribosomal promoter regulation in Escherichia coli. Mol. Gen. Genet. 213, 214–222 (1988).
Ono, S. et al. Plastidial (p)ppGpp synthesis by the Ca2+-dependent RelA-SpoT homolog regulates adaptation of chloroplast gene expression to darkness in Arabidopsis. Plant Cell Physiol. https://doi.org/10.1093/pcp/pcaa124 (2020).
Ooga, T. et al. Degradation of ppGpp by nudix pyrophosphatase modulates the transition of growth phase in the bacterium Thermus thermophilus. J. Biol. Chem. 284, 15549–15556 (2009).
Ito, D. et al. Enzymatic and molecular characterization of Arabidopsis ppGpp pyrophosphohydrolase, AtNUDX26. Biosci. Biotechnol. Biochem. 76, 2236–2241 (2012).
Srouji, J. R., Xu, A., Park, A., Kirsch, J. F. & Brenner, S. E. The evolution of function within the Nudix homology clan. Proteins. 85, 775–811 (2017).
Witowski, N. et al. A four-compartment metabolomics analysis of the liver, muscle, serum, and urine response to polytrauma with hemorrhagic shock following carbohydrate prefeed. PLoS ONE 10, e0124467 (2015).
Krásny, L. & Gourse, R. L. An alternative strategy for bacterial ribosome synthesis: Bacillus subtilis rRNA transcription regulation. EMBO J. 23, 4473–4483 (2004).
Anderson, B. W. et al. Evolution of (p)ppGpp-HPRT regulation through diversification of an allosteric oligomeric interaction. Elife 8, 1–31 (2019).
Verstraeten, N., Fauvart, M., Versees, W. & Michiels, J. The universally conserved prokaryotic GTPases. Microbiol. Mol. Biol. Rev. 75, 507–542 (2011).
Hesketh, A., Vergnano, M., Wan, C. & Oliver, S. G. Bacterial signaling nucleotides inhibit yeast cell growth by impacting mitochondrial and other specifically eukaryotic functions. MBio 8, 1–20 (2017).
Cole, L. & Kramer, P. R. Sugars, fatty acids, and energy biochemistry. Hum. Physiol. Biochem. Basic Med. 17–30, https://doi.org/10.1016/b978-0-12-803699-0.00019-0 (2016).
Werner, C., Doenst, T. & Schwarzer, M. Metabolic Pathways and Cycles. The Scientist’s Guide to Cardiac Metabolism, https://doi.org/10.1016/B978-0-12-802394-5/00004-2 (Elsevier, 2016).
Ihara, Y. & Masuda, S. Cytosolic ppGpp accumulation induces retarded plant growth and development. Plant Signal. Behav. 11, e1132966 (2016).
Bischof, J., Maeda, R. K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific φC31 integrases. Proc. Natl Acad. Sci. USA 104, 3312–3317 (2007).
Hay, B. A., Wolff, T. & Rubin, G. M. Expression of baculovirus P35 prevents cell death in Drosophila. Development 120, 2121–2129 (1994).
McGuire, S. E., Le, P. T., Osborn, A. J., Matsumoto, K. & Davis, R. L. Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302, 1765–1768 (2003).
Britton, J. S. & Edgar, B. A. Environmental control of the cell cycle in Drosophila: nutrition activates mitotic and endoreplicative cells by distinct mechanisms. Development 125, 2149–2158 (1998).
Hakeda-Suzuki, S. et al. Golden goal collaborates with flamingo in conferring synaptic-layer specificity in the visual system. Nat. Neurosci. 14, 314–323 (2011).
Acknowledgements
We thank J. Chung (Seoul National University) for proving fly strains. We acknowledge support from JSPS KAKENHI Grant Number 19K22418 (to S.M.). D.I. is supported by a Grant-in-Aid for JSPS Fellows Grant Number 18J12607.
Author information
Authors and Affiliations
Contributions
D.I. and S.M. designed the research. D.I. and Y.I. quantified ppGpp and GTP. D.I., H.K., and T. Suzuki constructed and characterized Drosophila mutants. A.O. performed metabolome analysis. T. Shibata and S.-I.K. grew and cultivated germ-free flies. N.N. and T.A. maintained and sampled cultured cells. D.I. and S.M. wrote the manuscript. All authors contributed to the discussion of the results.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ito, D., Kawamura, H., Oikawa, A. et al. ppGpp functions as an alarmone in metazoa. Commun Biol 3, 671 (2020). https://doi.org/10.1038/s42003-020-01368-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01368-4
This article is cited by
-
The immune response to RNA suppresses nucleic acid synthesis by limiting ribose 5-phosphate
The EMBO Journal (2024)
-
Significance of the plastidial stringent response for plant growth on soil
Plant Growth Regulation (2024)
-
Control of transcription elongation and DNA repair by alarmone ppGpp
Nature Structural & Molecular Biology (2023)
-
Feedforward growth rate control mitigates gene activation burden
Nature Communications (2022)
-
Metabolic changes contributing to large biomass production in the Arabidopsis ppGpp-accumulating mutant under nitrogen deficiency
Planta (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.