Abstract
We previously showed that mice lacking pituitary adenylate cyclase-activating polypeptide (PACAP) exhibit attenuated light-induced phase shift. To explore the underlying mechanisms, we performed gene expression analysis of laser capture microdissected suprachiasmatic nuclei (SCNs) and found that lipocalin-type prostaglandin (PG) D synthase (L-PGDS) is involved in the impaired response to light stimulation in the late subjective night in PACAP-deficient mice. L-PGDS-deficient mice also showed impaired light-induced phase advance, but normal phase delay and nonvisual light responses. Then, we examined the receptors involved in the response and observed that mice deficient for type 2 PGD2 receptor DP2/CRTH2 (chemoattractant receptor homologous molecule expressed on Th2 cells) show impaired light-induced phase advance. Concordant results were observed using the selective DP2/CRTH2 antagonist CAY10471. These results indicate that L-PGDS is involved in a mechanism of light-induced phase advance via DP2/CRTH2 signaling.
Similar content being viewed by others
Introduction
The mammalian circadian clock system comprises the endogenous master pacemaker located within the suprachiasmatic nucleus (SCN) in the hypothalamus and coordinates physiology and behavior in relation to the 24 h day/night cycle. At the molecular level, it has been established that interlocked feedback loops of transcriptional activation by the CLOCK/BMAL1 complex and repression by the PER/CRY complex integrate with diverse environmental and metabolic stimuli to generate internal 24 h timing1,2,3,4. The primary input stimulus of the mammalian circadian clock is light, but feeding, temperature, and social cues can also be entrainment factors3. Clock disruption is primarily associated with sleep disorders but is also implicated in various diseases, including mental, metabolic, cardiovascular, and inflammatory disorders. Studies on clock dysfunctions therefore aim to unravel mechanisms and reveal potential novel therapeutic targets for these disorders5,6.
Previously, we demonstrated that mice lacking pituitary adenylate cyclase-activating polypeptide (PACAP)7,8 (PACAP−/− mice) show an impairment in the light-dependent synchronization of diurnal rhythms (photic entrainment)9,10. A subset of retinal ganglion cells that express the circadian photopigment melanopsin also coexpress PACAP and glutamate, both of which have phase-shifting effects on the endogenous rhythm via regulation of clock gene expression in the SCN11,12. Thus, PACAP−/− mice are considered to be a unique animal model to further investigate mechanisms of the generation and entrainment of the circadian rhythm. However, fundamental insight into the molecular basis of photic entrainment by PACAP signaling remains elusive.
Here, we performed gene chip analysis of laser capture-microdissected SCNs from PACAP−/− and wild-type mice that had been kept in constant dark (DD) or exposed to light stimulation in the late subjective night to identify which genes were differentially expressed and categorize them by expression patterns. The analysis revealed that an increase in lipocalin-type prostaglandin (PG) D synthase (L-PGDS)13,14 expression in response to light stimulation in the late subjective night was not observed in PACAP−/− mice. In the brain, L-PGDS and its enzymatic product PGD2 have been well established to play a crucial role in the regulation of physiological sleep15,16. However, the function of L-PGDS, as well as two subtypes of the PGD2 receptor, namely, DP1 and DP2/CRTH2 (chemoattractant receptor-homologous molecule expressed on Th2 cells; also known as GPR44), in the circadian clock remains unknown. In the present study, we therefore examined circadian rhythms and light-induced phase shifts in mice lacking L-PGDS (L-PGDS−/−), DP1 (DP1−/−), or DP2/CRTH2 (CRTH2−/−). L-PGDS and DP2/CRTH2-deficient mice showed impaired phase advance under low intensity light. These results indicate that L-PGDS is involved in a mechanism of light-induced phase advance via DP2/CRTH2 signaling.
Results
Identification of candidate genes involved in phase advance
We have previously observed that PACAP−/− mice show severe dysfunctions of photic entrainment10, particularly in the abolishment of phase advance with low intensity (20 lx) light stimulation9. Therefore, in this study, we addressed downstream pathways that underpin circadian entrainment and performed gene chip analysis of laser capture-microdissected SCNs (Supplementary Fig. 1a) from four mouse groups: PACAP−/− and wild-type mice, which were either illuminated with light in the late subjective night, circadian time (CT) 21, or kept without light (SCNs from 3 mice per group).
Among the ~22,000 genes represented on the oligonucleotide array, 539 genes with hybridization signal ratios with a more than 1.7-fold change compared to the values for wild-type mice kept without light were selected and regarded as differentially expressed genes (Fig. 1a). Of these 593 genes, we specifically analyzed genes that were upregulated (>1.7-fold change) or downregulated (>0.6-fold change) by light stimulation in PACAP−/− and wild-type mice. Of these genes, 108 (wild type only: 89 genes, PACAP−/− only: 28 genes, both: 9 genes) were upregulated, while 331 genes (wild type only: 296 genes, PACAP−/− only: 62 genes, both: 27 genes) were downregulated, under light stimulation (Fig. 1b).
a mRNA expression levels of the 539 differentially expressed genes as measured by hybridization signal intensity. Clustering dendrograms show the relative expression values according to the scale shown on the bottom left side (magenta, high expression level; light green, low expression level). b Venn diagram illustrating pairwise overlap of the genes. Data in each genotype comparison with light stimulation represent upregulated (>1.7-fold change) or downregulated (>0.6-fold change) genes. The 18 genes annotated to the term “extracellular region” are indicated by a purple dotted circle. These genes were upregulated by light stimulation in wild-type mice and are listed in Table 1. Real-time quantitative PCR analysis for L-Pgds and Cryab in the SCN upon light stimulation at CT 21 (c) and CT 15 (d). The Per1 and Prok2 genes were used as positive controls. ND indicates not detected. The values are shown as the mean ± SEM (n = 4–7 per group). Statistically significant differences were assessed using two-way ANOVA followed by Tukey–Kramer tests. *p < 0.05; **p < 0.01, vs kept without light stimulation.
Using the k-means clustering algorithm, the 539 genes were classified into 6 clusters on the basis of the similarity of the expression profiles (Supplementary Fig. 1b). Genes in cluster 1 were slightly downregulated in PACAP−/− mice kept without light and upregulated by light stimulation compared with the levels in wild-type mice kept without light. Genes in cluster 2 show increased expression in PACAP−/− mice kept without light and in both wild-type and PACAP−/− mice under light stimulation compared with the expression in wild-type mice kept without light. Genes in clusters 3 and 4 show decreased expression in both wild-type and PACAP−/− mice by light stimulation compared with the expression in wild-type mice kept without light. The difference between these clusters is the differential expression levels in PACAP−/− mice under light stimulation. Genes in cluster 3 show decreased expression levels in both wild-type and PACAP−/− mice under light stimulation. Genes in cluster 4 show decreased expression levels in wild-type mice compared with those in PACAP−/− mice under light stimulation. Genes in cluster 5 were slightly upregulated in PACAP−/− mice kept without light and downregulated by light stimulation in both wild-type and PACAP−/− mice compared with wild-type mice kept without light. Cluster 6 includes genes that do not belong to cluster 1–5. To gain further insight into the clusters, we analyzed the informative gene ontology terms for each cluster (Supplementary Table 1). We found that genes in these clusters were categorized into partially overlapping but different gene ontology classifications.
In addition, gene ontology enrichment analysis for functional classification of light-responsive genes differentially expressed in wild-type mice showed that genes annotated to the term “binding” were the vast majority (48%), including immediate early response genes such as Per1 and fos (Supplementary Fig. 1c). Gene ontology enrichment analysis for cellular components of light-responsive genes in wild-type and PACAP−/− mice showed that the ratio of the number of light-responsive genes annotated to the term “extracellular region” to the total number of light-responsive genes was the most different between PACAP−/− and wild-type mice (Supplementary Fig. 1d). Interestingly, most of the “extracellular region” genes were the 18 genes (indicated by a purple dotted circle in Fig. 1b and listed in Table 1) which were upregulated by light stimulation in wild-type mice but not in PACAP−/− mice. Of these 18 genes, L-Pgds (Ptgds) showed the second largest change in expression in response to light stimulation in wild-type mice (Table 1).
Due to the considerable biological interest in L-PGDS, we performed real-time quantitative PCR using amplified RNA from laser-captured microdissected SCNs. We observed that L-Pgds was the only gene whose expression was increased by light at CT 21 in wild-type mice but not in PACAP−/− mice [two-way analysis of variance (ANOVA), gene effect: F(1,19) = 0.089, p = 0.77; light effect: F(1,19) = 3.86, p = 0.06; interaction: F(1,19) = 1.083, p = 0.31]. In addition, light-induced upregulation of L-Pgds expression was not observed in either wild-type or PACAP−/− mice under light stimulation at CT 15 [two-way ANOVA, gene effect: F(1,14) = 3.83, p = 0.07; light effect: F(1,14) = 0.19, p = 0.67; interaction: F(1,14) = 0.094, p = 0.76] (Fig. 1c, d). On the other hand, the changes in the expression of Cryab were not significant in wild-type and PACAP−/− mice under light stimulation at CT 15 and CT 21 or without stimulation [two-way ANOVA, CT 15, gene effect: F(1,14) = 0.52, p = 0.48; light effect: F(1,14) = 1.83, p = 0.198; interaction: F(1,14) = 1.83, p = 0.198; CT 21, gene effect: F(1,19) = 2.38, p = 0.14; light effect: F(1,19) = 0.98, p = 0.33; interaction: F(1,19) = 0.10, p = 0.75]. The discrepancy between the microarray and RT-PCR results may be explained by the extremely low expression level of Cryab in the SCN. As expected, the expression of Per1 and Prok2, which were positive targets, was significantly increased in wild-type and PACAP−/− mice under light stimulation either at CT 15 [two-way ANOVA, Per1, gene effect: F(1,14) = 7.18, p = 0.018; light effect: F(1,14) = 24.96, p = 0.0002; interaction: F(1,14) = 1.24, p = 0.28; Prok2, not detected] or at CT 21 [two-way ANOVA, Per1, gene effect: F(1,18) = 0.44, p = 0.52; light effect: F(1,18) = 12.89, p = 0.002; interaction: F(1,18) = 0.014, p = 0.91; Prok2, gene effect: F(1,18) = 2.43, p = 0.14; light effect: F(1,18) = 18.35, p = 0.0004; interaction: F(1,18) = 2.63, p = 0.12].
L-PGDS localization in the SCN
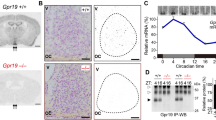
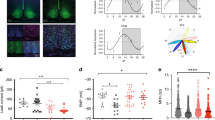
Of the genes altered at CT 21, only L-Pgds showed differences in expression between wild-type and PACAP−/− mice (Fig. 1c). Under DD conditions, the expression of L-Pgds showed a tendency to be high at CT 15 and low at CT 21 in wild-type mice, while the opposite trend was observed in PACAP−/− mice (Supplementary Fig. 2a). The lack of induction of L-Pgds by light at CT 21 in PACAP−/− mice was considered to result from the elevated basal expression of L-Pgds in the SCN in these mutant mice. L-PGDS is known to produce PGD2, which is the most abundant prostanoid in the brains16. In situ hybridization of L-Pgds signals was detected as intense clusters of black grains under bright-field illumination (Fig. 2a, b) as well as white grains under dark-field illumination (Fig. 2d). We counted the number of L-Pgds signals on individual cell bodies that were merged with Nissl-stained neurons in the SCN using ImageJ software (NIH) (Fig. 2b). Although the number of L-Pgds-expressing cells tended to increase upon illumination at CT 21 in wild-type mice, the results did not show statistically significant differences [two-way ANOVA, gene effect: F(1,23) = 0.45, p = 0.51; light effect: F(1,23) = 1.60, p = 0.22; interaction: F(1,23) = 2.32, p = 0.14] (Fig. 2c). Then, we counted the cellular intensity of L-Pgds expression in the SCN. The cellular intensity of L-Pgds expression was significantly increased by light at CT 21 in wild-type mice [two-way ANOVA, gene effect: F(1,36) = 10.11, p = 0.003; light effect: F(1,36) = 12.97, p = 0.001; interaction: F(1,36) = 10.47, p = 0.003] but not in PACAP−/− mice (Fig. 2d, e). In contrast, there was no difference in the number of L-Pgds-positive neurons in the outside vicinity of the SCN with or without light stimulation between genotypes [L-PGDS−/−; light stimulant, 272 ± 35 cells; kept without light, 251 ± 24 cells; wild type; light stimulant, 308 ± 26 cells; kept without light, 251 ± 18 cells, not significant, two-way ANOVA] (Supplementary Fig. 2b). We also performed double immunostaining of L-PGDS with NeuN or Olig2 and observed that L-PGDS immunoreactivity was mainly localized in NeuN-positive neurons and partly in Olig2-positive oligodendrocytes (Supplementary Fig. 2c, d). In addition, double immunostaining of L-PGDS with vasoactive intestinal peptide (VIP) or vasopressin (AVP) (Fig. 2f, g) showed that VIP-immunoreactive neurons were localized exclusively in the ventrolateral core region in the SCN and their fibers were distributed throughout the SCN (Fig. 2f), while AVP-immunoreactive neurons were localized mostly in the dorsomedial shell part in the SCN (Fig. 2g). The roles of SCN core and shell are known to be functionally distinct17. L-PGDS-immunoreactivity was localized with both VIP and AVP throughout the SCN (Fig. 2d, e). The immunoreactivity of L-PGDS was localized in the perinucleus, such as the nuclear envelope, Golgi apparatus, and secretory vesicles. In contrast, the immunoreactivity of VIP or AVP was localized in the cytoplasm and fibers. Although the subcellular localization differed between L-PGDS and VIP or AVP, these signals partly merged and seemed to be in the same neurons. Therefore, we assume that L-PGDS and VIP or AVP were colocalized in the neurons (Fig. 2f, g).
a In situ hybridization with a [35S]CTP-labeled antisense probe for L-Pgds in the SCN at CT 21. Representative bright-field photomicrographs are shown for PACAP−/− and wild-type mice, which were either light stimulated (light+) at CT 21 or kept without light (light−). 3V third ventricle; OC optic chiasma. The dotted line indicates the borders of the SCN area that contain densely aggregated cresyl violet-stained nuclei. Red arrowheads in the right SCN region indicate L-Pgds signals that were merged with counterstained neurons. Bar, 100 μm. b Wild-type mice illuminated with light. Right panel, magnification of the area marked with a black box. Red arrowheads indicate L-Pgds signals that were merged with counterstained neurons. Bars, 10 μm. c The number of L-Pgds-positive cells quantified in the SCN. To quantitatively determine the L-Pgds-expressing neurons in the whole SCN, five coronal SCN sections every four sections per mouse were used for statistical analysis. The values are shown as the mean ± SEM (n = 5–9 per group). *p < 0.05. Statistically significant differences were assessed using two-way ANOVA followed by Tukey–Kramer tests. d Representative dark-field photomicrographs. Bar, 100 μm. Right panel, magnification of the area marked with a white box. Red arrowheads indicate L-Pgds signals on individual cell bodies that were merged with Nissl-stained neurons. Bar, 10 μm. e Intensity of in situ hybridization signals for L-Pgds in each cell in the SCN. The values are shown as the mean ± SEM (n = 10 per group). **p < 0.01. Statistically significant differences were assessed using two-way ANOVA followed by Tukey–Kramer tests. Double immunohistochemical staining for L-PGDS (magenta) and VIP (green, f) or AVP (green, g) in the SCN. Right panels, magnification of the areas marked with white boxes. White arrowheads, cells that were immunoreactive for L-PGDS and VIP or AVP. Bars, 100 μm.
Impaired light-induced phase advance in L-PGDS−/− mice
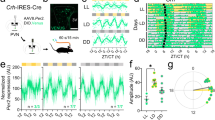
We subsequently examined circadian rhythms of locomotor activity in L-PGDS−/− mice. L-PGDS−/− mice were normally synchronized with a light cycle of 12 h of light and 12 h of dark (LD) and retained behavioral periodicity under DD conditions compared with wild-type mice (Fig. 3a, b, Table 2). However, under constant light (LL) condition, L-PGDS−/− mice showed slightly but significantly increased duration of free-running period (Fig. 3c, Table 2). The pattern and magnitude of locomotor activity during an LD cycle was not different between L-PGDS−/− and wild-type mice [two-way repeated measures ANOVA, gene effect, F(1,8) = 0.26, p = 0.62; time effect: F(23,184) = 20.59, p < 0.0001; interaction: F(23,184) = 0.80, p = 0.73] (Fig. 3d).
Representative double-plotted actograms of wild-type and L-PGDS−/− mice kept in the LD cycle (light, 12 h; dark, 12 h) (a) or transferred from LD to DD (b) or LL (c). d Daily variations in locomotor activities of wild-type and L-PGDS−/− mice under LD conditions. Intensity of illumination during the light phase, 100 lx. The values are expressed as the mean ± SEM (n = 5 per group). Statistically significant differences were assessed using two-way repeated measures ANOVA followed by Tukey–Kramer tests.
In contrast, L-PGDS−/− mice showed impaired photic entrainment function. A light pulse (20, 100, or 600 lx, 30 min) in the late subjective night (CT 21) induced phase advance of locomotor activity rhythms at significantly lower levels in L-PGDS−/− mice than in wild-type mice [two-way ANOVA, gene effect: F(1,23) = 32.65, p < 0.0001; light effect: F(2,23) = 4.54, p = 0.022; interaction: F(2,23) = 2.39, p = 0.11] (Fig. 4a, b and Supplementary Fig. 3), while a light pulse in the early subjective night (CT 15) induced phase delay in L-PGDS−/− and wild-type mice at similar levels [two-way ANOVA, gene effect: F(1,28) = 0.36, p = 0.55; light effect: F(2,28) = 0.27, p = 0.77; interaction: F(2,28) = 0.49, p = 0.62] (Fig. 4c, d).
Phase shift induced by light stimulation at CT 21 (a, b) and CT 15 (c, d) in L-PGDS−/− and wild-type mice. a, c Representative double-plotted actograms. Yellow arrowheads indicate light stimulation (20 lx, 30 min). Paired red lines represent the onset of activity. Quantification of the phase shift induced by light stimulation. The values are expressed as the mean ± SEM (n = 4–5 (b), n = 3–9 (d) per group). *p < 0.05; **p < 0.01. Statistically significant differences were assessed using two-way ANOVA followed by Tukey–Kramer tests. Resynchronization of circadian rhythms to time shifts, 8 h advance (e, f) and 8 h delay (g, h). e, g Representative patterns of wheel running activity. f, h Quantification of the phase shift. The values are expressed as the mean ± SEM (n = 6–8 per group). **p < 0.01. Statistically significant differences were assessed using two-way repeated measures ANOVA.
Using a jet lag model in which mice were exposed to an 8 h time shift (advance or delay), we examined the synchronization of circadian rhythms in L-PGDS−/− mice. When the light cycle was advanced by 8 h, L-PGDS−/− mice showed significantly delayed synchronization compared to wild-type mice [two-way repeated measures ANOVA, gene effect: F(1,12) = 2.70, p = 0.13; day effect: F(7,84) = 175.84, p < 0.0001; interaction: F(7,84) = 3.16, p = 0.005] (Fig. 4e, f). In contrast, both L-PGDS−/− and wild-type mice showed similar synchronization when the light cycle was delayed by 8 h [two-way repeated measures ANOVA, gene effect: F(1,12) = 0.11, p = 0.75; day effect: F(7,84) = 121.54, p < 0.0001; interaction: F(7,84) = 1.01, p = 0.43] (Fig. 4g, h).
L-PGDS−/− mice show normal light-induced c-Fos expression
Light-induced c-Fos-expression was examined in the SCN in L-PGDS−/− mice (Supplementary Fig. 4). Light stimulation at CT 21 markedly increased the number of c-Fos-immunoreactive cells in the SCN in both L-PGDS−/− and wild-type mice. There was no difference in the distribution and number of c-Fos-immunoreactive cells under light stimulation between genotypes [L-PGDS−/−, 411.9 ± 39.07 cells; wild type, 453.6 ± 32.2 cells; two-way ANOVA, not significant].
L-PGDS−/− mice show a normal nonvisual light responses
To examine the integrity of visual pathways, we examined the pupillary light reflex in L-PGDS−/− mice. There were no differences in pupil sizes under scotopic conditions between genotypes (L-PGDS−/−, 1.87 ± 0.06 mm2; wild type, 1.99 ± 0.08 mm2; Student’s t test, not significant) (Supplementary Fig. 5a, b). Light-induced (100 and 600 lx) light reflexes were not significantly different in L-PGDS−/− and wild-type mice (Supplementary Fig. 5c, d).
Negative masking responses to light (nocturnal animals are normally passive in a high-illumination milieu18) were normal in L-PGDS−/− mice (Supplementary Fig. 5e–g). The amount of activity in mice exposed to a 2-h light pulse (100 and 400 lx) during the early night (Zeitgeber time 13–15) was similarly suppressed in L-PGDS−/− and wild-type mice [two-way ANOVA, 100 lx, gene effect: F(1,26) = 1.03, p = 0.32; light effect: F(1,26) = 17.14, p = 0.003; interaction: F(1,26) = 0.21, p = 0.65; 400 lx, gene effect: F(1,23) = 2.48, p = 0.13; light effect: F(1,23) = 121.41, p < 0.0001; interaction: F(1,23) = 0.13, p = 0.72].
DP2/CRTH2 mediates light-induced phase advance
To determine which subtype of PGD2 receptors, i.e., DP1 or DP2/CRTH2, is responsible for light-induced phase advance, we examined circadian entrainment of locomotor activity in DP1−/− and CRTH2−/− mice. A light pulse (20 or 100 lx, 30 min) in the late subjective night (CT 21) induced phase advance of locomotor activity rhythms at significantly lower levels in CRTH2−/− mice than in wild-type mice, although a stronger light pulse (600 lx, 30 min) at CT 21 induced similar levels of phase advance in CRTH2−/− and wild-type mice [two-way ANOVA, gene effect: F(1,58) = 1.03, p < 0.0001; light effect: F(2,58) = 16.97, p < 0.0001; interaction: F(2,58) = 0.60, p = 0.55] (Fig. 5c, f and Supplementary Fig. 3b). CRTH2−/− mice showed normal levels of phase delay compared with wild-type mice (Fig. 5d). DP1−/− mice showed normal levels of phase advance and delay even with exposure to very dim light (20 lx, 30 min) compared with wild-type mice (Fig. 5a, b).
Phase shift induced by light stimulation at CT 15 or CT 21 in DP1−/− (a, b) and CRTH2−/− (c, d) mice and wild-type mice of the respective genetic backgrounds. a, c Representative double-plotted actograms. Quantification of the phase shift induced by the indicated light stimulation. The values are expressed as the mean ± SEM (n = 3–6 (b), n = 9–14 (d) per group). e, f Phase shift induced by light stimulation at CT 15 or CT 21 in CD-1 wild-type mice administered the CRTH2 blocker CAY10471 or a vehicle (Riger’s solution) 30 min before light stimulation. e Representative double-plotted actograms. f Quantification of the phase shift. The values are expressed as the mean ± SEM (n = 10 per group). *p < 0.05; **p < 0.01. Statistically significant differences were assessed using two-way ANOVA followed by Tukey–Kramer tests. Yellow arrowheads indicate light stimulation (20 lx, 30 min). Paired red lines represent the onset of activity.
We next examined the effect of intracerebroventricular administration of the DP2/CRTH2 antagonist CAY10471 or DP1 antagonist BW A868C on light-induced phase advance in wild-type mice in the CD-1 genetic background. CAY10471 significantly diminished light-induced (20 lx, 30 min) phase advance at CT 21 [two-way ANOVA, treat effect: F(1,36) = 2.18, p = 0.15; light time effect: F(1,36) = 116.20, p < 0.0001; interaction: F(1,36) = 7.60, p = 0.009] (Fig. 5e, f). In contrast, BW A868C-treated mice showed normal levels of phase advance compared with vehicle-treated mice at each light pulse (20 or 600 lx) (Supplementary Fig. 6).
Discussion
PACAP is cotransmitted with glutamate in melanopsin-containing retinal ganglion cells, which monosynaptically innervate the SCN, and mediates nonvisual photoreception-regulated light-induced circadian entrainment, negative masking of locomotor activity, and the pupillary light reflex12,19,20. Since PACAP−/− mice show impaired light-induced circadian entrainment and negative masking and unusually early onset of activities during the light-to-dark transition period (an “early-bird” phenotype)9,10, in the present study, we conducted transcriptome analysis of laser capture-microdissected SCNs from PACAP−/− and wild-type mice with or without light stimulation in the late subjective night. We identified that L-Pgds showed the second largest change between the illuminated mutant mice and basal wild-type mice (Table 1). It is intriguing that L-Pgds expression was increased by light specifically at CT 21. We found that the individual intensity of L-Pgds signals in each cell was significantly increased by light at CT 21 in the SCN in wild-type mice (Fig. 2e), suggesting that the increased L-Pgds levels (Fig. 1c) are likely attributable to the increased individual intensity of L-Pgds signals in each cell in the SCN. Fujimori et al. demonstrated that the promoter of L-Pgds has an E-box motif using a luciferase reporter assay21, suggesting that L-Pgds may show changes in circadian expression under light stimulation. The blunted induction of L-Pgds in PACAP−/− mice by light at CT 21 might arise from abnormal basal expression of L-Pgds in the SCN in PACAP−/− mice (Supplementary Fig. 2a). Purb showed the largest change between the illuminated mutant mice and basal wild-type mice (Table 1). Recent studies have shown that PURB is a single-stranded nucleic acid-binding protein that is involved in the regulation of DNA replication and transcription; however, little has been reported on its function in the central nervous system. Furthermore, Purb-deficient mice are not available. Therefore, although it is important to understand the role of PURB in the circadian system, we assume that it would take considerable effort to convincingly determine how PURB is involved in the phase advance of the circadian rhythm in response to the light pulse. The role of Purb and the other genes differentially expressed in the current study should be addressed in future research.
L-PGDS−/− mice showed impaired phase advance under light at CT 21 but normal phase delay under light at CT 15. These results may implicate L-PGDS in phase advance-selective re-entrainment. PACAP−/− mice have been shown to convey parametric light information depending on intensity and duration. PACAP and L-PGDS may thus be involved in a mechanism for directional asymmetry in circadian entrainment (where phase advance is impaired while phase delay is normal in PACAP−/− and L-PGDS−/− mice). Previous studies have found that L-Pgds gene expression was upregulated by protein kinase C through derepression of notch-HES signaling and augmentation of AP-2β in human TE671 (medulloblastoma of cerebellum) cells22. Moreover, a key prostanoid enzyme, cyclooxygenase-2, and PGE2 biosynthesis were induced by interleukin 1 beta (IL-1β) via protein kinase C activation and mitogen-activated protein kinases cascade in the glial cells23. PACAP has been shown to stimulate extracellular signal-regulated kinase 1/2 activity24. Future studies into the functional relationships between PACAP- and L-PGDS-mediated signaling pathways will reveal the precise role of PACAP in light-induced circadian entrainment.
In the present study, the light-induced c-Fos expression was normally observed in L-PGDS−/− mice light stimulated at CT 21 (Supplementary Fig. 4), which is consistent with our previous result for PACAP−/− mice9. These results suggest that c-Fos is not critically involved in the PACAP- and L-PGDS-mediated phase advance of the central circadian clock.
Considering our current results, DP2/CRTH2 is also not involved in a mechanism of light-induced phase advance under 600 lx light (Fig. 5c, d and Supplementary Fig. 3b). Thus, the impaired light-induced phase advance under 600 lx in L-PGDS−/− mice may be explained by the PGD2 signaling-independent function of L-PGDS; however, the molecular mechanism underlying the function of L-PGDS is currently unclear. L-PGDS is reported to act as an extracellular transporter of various lipophilic small molecules as well as a PGD2-synthesizing enzyme14. L-PGDS binds to all-trans-retinoic acids that have been identified as potential circadian entrainment factors25,26. Moreover, Lee et al. reported a novel nonenzymatic function of L-PGDS, i.e., regulation of glial cell migration and morphology by binding to the MARCKS heat shock protein27. Alternatively, either one of the two PGD2 receptor subtypes may be sufficient to mediate PGD2 signaling for the light-induced phase advance under 600 lx light. Further analysis, e.g., using DP1 and DP2/CRTH2 double-deficient mice, is needed to precisely understand the function of L-PGDS and PGD2 receptors signaling pathway in the light-induced circadian entrainment.
It has been shown that PACAP, L-PGDS, DP1, and DP2/CRTH2 are expressed in SCN11,12; PACAP is expressed in neurons and astrocytes28; L-PGDS is expressed in leptomeningeal cells, neurons, and oligodendrocytes29; DP1 is expressed in neurons and microglia30; and DP2/CRTH2 is expressed mainly in astrocytes29. These data suggest that L-PGDS/PGD2 and downstream DP2/CRTH2 signaling may represent a mechanism that is linked to neural and astrocytic signaling cascade involved in light-induced phase advance of the central circadian clock. Future studies, e.g., cell type-specific single-cell transcriptome analysis, may reveal the precise mechanism.
In the brain, PGD2 acts as the most potent endogenous sleep-promoting substance reported thus far16 and is involved in PGE2-induced neuropathic pain31. DP1 receptor signaling is involved in these PGD2 actions. In a genetic demyelination model of twitcher mice, L-PGDS has been implicated in neural protection32, while hematopoietic PGD synthase (H-PGDS), which is responsible for the production of PGD2 in inflammatory responses, plays a role in the progression of neural inflammation29. The PGD2 concentration in rat cerebrospinal fluid shows circadian rhythmicity in parallel with the sleep–wake cycle33, and L-PGDS levels in human serum also show circadian rhythmicity, with a nocturnal increase34. In the cultured peripheral fibroblasts, 15-deoxy Δ[12, 14] PG J2 (15d-PGJ2) and PGJ2, a derivative of PGD2, were reported to reset the peripheral circadian clock26,35. However, the function of L-PGDS as well as the two PGD2 receptor subtypes DP1 and DP2/CRTH2 in the circadian clock remains unknown. To our knowledge, the present results provide the first evidence that L-PGDS-derived PGD2 and the downstream DP2/CRTH2 specifically mediate light-induced phase advance of the central circadian clock.
Although DP1 was originally identified as a homolog of other PG receptors36, DP2/CRTH2 is a member of the G protein-coupled leukocyte chemoattractant receptor family, which is selectively expressed in Th2 but not Th1 lineage cells, and is thereby named CRTH2 (chemoattractant receptor-homologous molecule expressed on Th2 cells)37. In contrast to the functions of DP1, the functions of DP2/CRTH2 in the brain are not well understood. Mohri et al. have reported that both DP1 and DP2/CRTH2 are expressed in cultured astrocytes, stimulation of which leads to enhanced GFAP production, suggesting that PGD2 plays an important role in microglia/astrocyte interactions29. Previously, we showed that 15d-PGJ2 enhances nerve growth factor-induced neurite outgrowth in vitro, through activation of DP2/CRTH238. More recently, we showed that DP2/CRTH2 is critically involved in impairments of emotional aspects induced by lipopolysaccharide or tumor (colon 26) inoculation39,40 and cognitive dysfunction induced by the N-methyl-d-aspartate receptor antagonist MK-80141. These results suggest that DP2/CRTH2 antagonism has potential as a therapeutic target for behavioral symptoms. Because the circadian clock is closely associated with the sleep–wake cycle1,2,5, the present results may provide an additional mechanism for the somnogenic effect of PGD2 in that DP2/CRTH2-mediated PGD2 signaling regulates light-induced the central circadian entrainment.
Although it remains unclear how PGD2-DP2/CRTH2 signaling induces circadian entrainment, in NIH3T3 cells, 15d-PGJ2, a putative ligand of DP2/CRTH2, has been shown to trigger rhythmic endogenous clock gene expression and transiently upregulate Cry1, Cry2, and Rorα expressions26. Similar regulatory mechanism of clock gene expression may mediate circadian entrainment in which PGD2-DP2/CRTH2 signaling is involved.
In this study, we used three different mouse strains, PACAP−/− in a CD-1 background, L-PGDS−/− and CRTH2−/− in a BALB/c background, and DP1−/− in a C57BL/6 background. PACAP−/− mice in the C57BL/6 background showed extremely high postnatal mortality42. Therefore, we backcrossed the null mutation onto the CD-1 mouse background. The difference between average time of phase shift at 20 lx in wild-type mice shown Figs. 4 and 5 might arise from the different between mouse strains (e.g., BALB/c, Fig. 4, vs C57BL/6, Fig. 5a, b). Schwartz reported that the two strains of BALB/c and C57BL/6 mice showed a large difference in the free-running period and phase advance during the late subjective night to early subjective day43. It is known that mouse strain differences may reflect a wide range of sensitivities to light, as eye color, thickness of outer nuclear layer in retina, melatonin deficiency and locomotor activity differ among mouse strains44. Although all mouse strains examined in this study showed phase advance during the late subjective night and we performed all experiments using wild-type and mutant mice with the same mouse background, the difference in the mouse backgrounds must be carefully taken into account to interpret the data obtained from different strains. Therefore, the present results observed in DP1−/− mice must be carefully interpreted and scrutinized in future research.
In summary, we obtained the following main findings: (1) L-PGDS-deficient mice showed impaired phase advance of circadian rhythm locomotor activity but normal phase delay when assessed using the effect of light pulses in DD conditions or with a jet lag model in which mice were exposed to an 8 h time shift (advance or delay), while (2) the mutant’s other nonvisual light responses, including light-induced locomotor suppression and pupillary light reflex, were normal. (3) DP2/CRTH2-deficient mice showed impaired light-induced phase advance but normal phase delay. Finally, (4) the selective DP2/CRTH2 antagonist CAY10471 impaired light-induced phase advance. These results show that L-PGDS-derived PGD2 and downstream DP2/CRTH2 signaling constitute a novel signaling cascade specifically involved in light-induced phase advance of the central circadian clock. The results also provide insights into the roles of prostanoids in the regulation of brain functions.
Methods
Mice
All animal care and handling procedures were approved by the Animal Care and Use Committee of the Graduate School of Pharmaceutical Sciences, Osaka University. The generation of PACAP–/−45, L-PGDS−/−31, DP1−/−46, and CRTH2−/−47 mice via gene targeting has been previously reported; these mice were backcrossed for at least 10 generations onto the CD-1 (PACAP−/−), C57BL/6 (DP1−/−), or BALB/c (L-PGDS−/− and CRTH2−/−) genetic background. Mice were kept under an LD cycle (light on from 8 a.m. to 8 p.m. unless otherwise specified) at a controlled room temperature. Pelleted food (CMF; Oriental Yeast, Osaka, Japan) and water were available ad libitum. Male knockout mice (PACAP−/−, L-PGDS−/−, DP1−/−, and CRTH2−/−) and wild-type mice of the respective genetic backgrounds were used for this study. All experiments were carried out on male mice at 6 to 12 weeks of age.
Microarray analysis
Mice synchronized to a normal LD cycle were introduced into DD conditions. At CT 21, the mice were illuminated at 20 lx for 30 min. One hour after illumination, the brains were removed, and 14-μm thick coronal brain sections, including the SCN, were adhered to noncoated glass slides, fixed by treatment with 75% ethanol for 30 s at −18 °C, and then dehydrated. The PixCell IIe Laser Capture Microdissection System (Arcturus) was used to isolate 30 SCN pieces (Supplementary Fig. 1a) from the 4 mouse groups: PACAP−/− and wild-type mice that were either light stimulated in the late subjective night (CT 21) or kept without light (n = 3 per group).
Total RNA was subsequently extracted using the RNeasy MinElute Cleanup Kit (Qiagen) according to the manufacturer’s protocols. RNA amplification and oligonucleotide microarrays were performed as described previously48,49, but with several modifications. For the first-round RNA amplification, 15 ng of total SCN RNA pooled from three mouse samples (5 ng per each mouse) was reverse transcribed using Superscript II reverse transcriptase (Invitrogen), and the double-stranded DNA was synthesized with DNA polymerase (Invitrogen). Using the double-stranded DNA as a template, antisense RNA was synthesized using the RiboAmp OA RNA Amplification Kit (Arcturus) according to the manufacturer’s instructions. The second-round amplification was performed similarly, except that the Enzo High Yield RNA Transcript Labeling Kit (Enzo Diagnostics) was used to prepare biotin-labeled antisense RNA. The resulting antisense RNA was hybridized to the GeneChip Mouse Genome 430A array (Affymetrix) using standard methods. Image files were processed using the Microarray Analysis Suite software (Affymetrix).
Microarray data analysis was performed as described previously48, with minor modifications. The robust multiarray analysis algorithm was used for background correction, normalization, and expression level summarization. Genes that showed 1.7-fold or greater absolute changes in signal intensity were extracted and classified using the k-means clustering algorithm. Functional enrichment analysis and cellular component analysis of the gene clusters was based on gene ontology pathway annotation terms using Data Mining Tool (Affymetrix) algorithm, with p values < 0.05 considered statistically significant. The ToppGene Suite (https://toppgene.cchmc.org/) was also used for gene ontology annotation-based functional classification of the genes in the clusters shown in Supplementary Fig. 1b. The gene ontology annotations were cut off at p < 0.05 and false discovery rate < 0.05. The gene annotation was limited for the analysis of biological processes (1000 ≤ n ≤ 10,000).
Real-time quantitative PCR was performed using amplified RNA from laser-captured microdissected SCNs, Superscript II reverse transcriptase (Invitrogen), and the DyNAmo SYBR Green qPCR Kit (Finnzymes). Primer sequences are shown in Supplementary Table 2. Gapdh was amplified as a control.
In situ hybridization
Brain sections (20 μm-thick) including the SCN were subjected to in situ hybridization performed as described previously45,50. A cDNA fragment of mouse L-Pgds (GenBank accession number NM_008963.1; nucleotides 2–471) was used as a template to synthesize 35S-CTP-labeled cRNA probes. The sections counterstained with cresyl violet clearly revealed morphologically distinct SCNs as a pair of densely aggregated neural cells. In situ hybridization signals of L-Pgds was detected as intense clusters of black grains under bright-field illumination as well as white grains under dark-field illumination. We counted the number of L-Pgds signals on individual cell bodies that were merged with Nissl-stained neurons in the SCN using ImageJ software (NIH) (Fig. 2b). To quantitatively determine the L-Pgds-expressing neurons in the whole SCN, five coronal SCN sections every four sections per mouse were used for statistical analysis.
Immunohistochemistry
Immunohistochemistry was performed as described previously9,10. After in situ hybridization, the sections were incubated with a mouse anti-NeuN antibody (1:1000; Santa Cruz, #sc-246957) or a rabbit anti-Olig2 antibody (1:100; IBL, #18953) a rabbit anti-c-Fos antibody (1:1000; Santa Cruz, #sc-52) overnight at 4 °C. The Vectastain Elite ABC Kit (Vector Lab) was used for immunostaining according to the manufacturer’s protocols. For double immunostaining, we used a 1:1000 dilution of a goat anti-L-PGDS antibody (Santa Cruz, sc-14825), a 1:2000 dilution of a rabbit anti-VIP antibody or a 1:1000 dilution of a rabbit anti-AVP antibody (both anti-VIP and anti-AVP antibodies were kindly provided by Dr. Buijs). After three washes with 0.2% Triton X-100 in PBS, the sections were incubated for 1 h at room temperature in a secondary antibody solution consisting of anti-goat IgG coupled to Alexa-594 (red, 1:200 dilution; #A-11058, Life Technologies) and anti-rabbit IgG coupled to Alexa-488 (green, 1:1000 dilution; #A-11008, Life Technologies) in 0.2% Triton X-100 in PBS with 3% bovine serum albumin. After three washes, the sections were mounted on glass slides with Fluoromount (Diagnostic BioSystems) and dried before imaging. Fluorescent images were captured using a BIO-REVO BZ-9000 fluorescence microscope (Keyence).
Behavioral study
Light-induced phase shifts in locomotor activity rhythms were examined as described previously9,10. Briefly, mice were transferred to DD conditions after being entrained to a 12L:12D cycle, with monitoring of their locomotor activity by far-infrared apparatus (Bio-Medica). After more than 8 days in DD conditions, animals were exposed to a white light pulse of the indicated intensities at CT 15 or CT 21, and their behavioral rhythms were further recorded. The double-plot actograms were produced and the phase shifts were calculated based on the distance between two regression lines drawn from the onset of activity before and after the light pulse using MATLAB (The MathWorks) and the method of Daan and Pittendrigh9,51.
Resynchronization to phase shifts of the LD schedule was examined using the jet lag model as follows. After mice were entrained to a 12L:12D cycle with monitoring of their locomotor activity by running wheels, the LD cycle was advanced or delayed by 8 h, and the locomotor activity was further measured for 9 days. The phase shifts were determined as the onset time of activity before and after the shift of the LD cycle.
Intracerebroventricular injections
Intracerebroventricular injections were performed as described previously39. CD-1 mice were anesthetized and placed in a stereotaxic instrument (Narishige). A G-4 cannula (Eicom) was implanted, −0.4 mm posterior, 1.0 mm lateral, and 2.3 mm ventral from the bregma. After cannula implantation, each mouse was given 1 mg/kg buprenorphine (Sigma-Aldrich) to relieve the pain and housed individually for at least 1 week before performing experiments. Thirty minutes before light stimulation, a DP2/CRTH2-selective antagonist, CAY10471, and a DP1 selective antagonist, BW A868C (Cayman Chemical), were diluted with Ringer’s solution (1:100, Fuso Pharmaceutical Industries) and injected at a volume of 2 and 4.6 μl, respectively, at an infusion rate of 1 μl/min, using a microinjection pump (KD Scientific). We previously examined the effect of CAY10471 on DP2/CRTH2 and found that CAY10471 pretreatment before 30 min is sufficient to block DP2/CRTH2 signaling39. Zhao et al. previously demonstrated that BW A868C pretreatment for 20 min was sufficient to block DP1 signaling52. The day after behavioral experiments, each mouse was intracerebroventricularly injected with 3 μl of 1% (w/v) Evans blue solution (Sigma-Aldrich), and a coronal section of the brain was prepared. The intracerebroventricular injection was judged to be successful if the third ventricle was stained by Evans blue.
Statistics and reproducibility
Experimental data were analyzed using one-way or two-way analysis of variance (ANOVA). The Tukey–Kramer post hoc test was also performed after significant main effects for drug, time or luminescence intensity were observed. The criterion for statistical significance was p < 0.05. Statistical analyses were performed using Stat View software (version 5.0; SAS Institute). Each experiment was repeated at least three times, and sample sizes and numbers are indicated in detail in each figure legend.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Microarray data have been deposited to the DDBJ Genomic Expression Archive (GEA) and are available at the accession number E-GEAD-376 and A-GEOD-8299. Data that support the findings of this study are available from the corresponding authors upon reasonable request. Source data underlying plots shown in figures are provided in Supplementary Data 1.
Code availability
Details of publicly available software used in the study are given in the methods and the Nature Research Reporting Summary linked to this article.
References
Bass, J. & Takahashi, J. S. Circadian integration of metabolism and energetics. Science 330, 1349–1354 (2010).
Golombek, D. A. & Rosenstein, R. E. Physiology of circadian entrainment. Physiol. Rev. 90, 1063–1102 (2010).
Albrecht, U. Timing to perfection: the biology of central and peripheral circadian clocks. Neuron 74, 246–260 (2012).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Tahara, Y., Aoyama, S. & Shibata, S. The mammalian circadian clock and its entrainment by stress and exercise. J. Physiol. Sci. 67, 1–10 (2017).
Cha, H. K., Chung, S., Lim, H. Y., Jung, J. W. & Son, G. H. Small molecule modulators of the circadian molecular clock with implications for neuropsychiatric diseases. Front. Mol. Neurosci. 11, 496 (2018).
Vaudry, D. et al. Pituitary adenylate cyclase-activating polypeptide and its receptors: 20 years after the discovery. Pharmacol. Rev. 61, 283–357 (2009).
Zhang, L. & Eiden, L. E. Two ancient neuropeptides, PACAP and AVP, modulate motivated behavior at synapses in the extrahypothalamic brain: a study in contrast. Cell Tissue Res. 375, 103–122 (2019).
Kawaguchi, C. et al. Changes in light-induced phase shift of circadian rhythm in mice lacking PACAP. Biochem. Biophys. Res. Commun. 310, 169–175 (2003).
Kawaguchi, C. et al. PACAP-deficient mice exhibit light parameter-dependent abnormalities on nonvisual photoreception and early activity onset. PLoS ONE 5, e9286 (2010).
Hannibal, J., Moller, M., Ottersen, O. P. & Fahrenkrug, J. PACAP and glutamate are co-stored in the retinohypothalamic tract. J. Comp. Neurol. 418, 147–155 (2000).
Hannibal, J. Melanopsin is expressed in PACAP-containing retinal ganglion cells of the human retinohypothalamic tract. Investig. Ophthalmol. Vis. Sci. 45, 4202–4209 (2004).
Nagata, A. et al. Human brain prostaglandin D synthase has been evolutionarily differentiated from lipophilic-ligand carrier proteins. Proc. Natl. Acad. Sci. USA 88, 4020–4024 (1991).
Urade, Y. & Eguchi, N. Lipocalin-type and hematopoietic prostaglandin D synthases as a novel example of functional convergence. Prostaglandins Other Lipid Mediat. 68-69, 375–382 (2002).
Pinzar, E. et al. Prostaglandin D synthase gene is involved in the regulation of non-rapid eye movement sleep. Proc. Natl. Acad. Sci. USA 97, 4903–4907 (2000).
Urade, Y. & Hayaishi, O. Prostaglandin D2 and sleep/wake regulation. Sleep Med. Rev. 15, 411–418 (2011).
Reghunandanan, V. & Reghunandanan, R. Neurotransmitters of the suprachiasmatic nuclei. J. Circadian Rhythms 4, 2 (2006).
Redlin, U. Neural basis and biological function of masking by light in mammals: suppression of melatonin and locomotor activity. Chronobiol. Int. 18, 737–758 (2001).
Hannibal, J., Brabet, P. & Fahrenkrug, J. Mice lacking the PACAP type I receptor have impaired photic entrainment and negative masking. Am. J. Physiol. Regul. Integr. Comp. Physiol. 295, R2050–2058 (2008).
Engelund, A., Fahrenkrug, J., Harrison, A., Luuk, H. & Hannibal, J. Altered pupillary light reflex in PACAP receptor 1-deficient mice. Brain Res. 1453, 17–25 (2012).
Fujimori, K., Fujitani, Y., Kadoyama, K., Kumanogoh, H., Ishikawa, K. & Urade, Y. Regulation of lipocalin-type prostaglandin D synthase gene expression by Hes-1 through E-box and interleukin-1 beta via two NF-kappa B elements in rat leptomeningeal cells. J. Biol. Chem. 278, 6018–6026 (2003).
Fujimori, K., Kadoyama, K. & Urade, Y. Protein kinase C activates human lipocalin-type prostaglandin D synthase gene expression through de-repression of notch-HES signaling and enhancement of AP-2 beta function in brain-derived TE671 cells. J. Biol. Chem. 280, 18452–18461 (2005).
Molina-Holgado, E., Ortiz, S., Molina-Holgado, F. & Guaza, C. Induction of COX-2 and PGE(2) biosynthesis by IL-1beta is mediated by PKC and mitogen-activated protein kinases in murine astrocytes. Br. J. Pharmacol. 131, 152–159 (2000).
Barrie, A. P., Clohessy, A. M., Buensuceso, C. S., Rogers, M. V. & Allen, J. M. Pituitary adenylyl cyclase-activating peptide stimulates extracellular signal-regulated kinase 1 or 2 (ERK1/2) activity in a Ras-independent, mitogen-activated protein Kinase/ERK kinase 1 or 2-dependent manner in PC12 cells. J. Biol. Chem. 272, 19666–19671 (1997).
Tanaka, T., Urade, Y., Kimura, H., Eguchi, N., Hayaishi, O. & Lipocalin-type Prostaglandin, D. Synthase (Beta-Trace) Is a newly recognized type of retinoid transporter. J. Biol. Chem. 272, 15789–15795 (1997).
Nakahata, Y., Akashi, M., Trcka, D., Yasuda, A. & Takumi, T. The in vitro real-time oscillation monitoring system identifies potential entrainment factors for circadian clocks. BMC Mol. Biol. 7, 5 (2006).
Lee, S., Jang, E., Kim, J. H., Lee, W. H. & Suk, K. Lipocalin-type prostaglandin D2 synthase protein regulates glial cell migration and morphology through myristoylated alanine-rich C-kinase substrate: prostaglandin D2-independent effects. J. Biol. Chem. 287, 9414–9428 (2012).
Grimaldi, M. & Cavallaro, S. Functional and molecular diversity of PACAP/VIP receptors in cortical neurons and type I astrocytes. Eur. J. Neurosci. 8, 2767–2772 (1999).
Mohri, I. et al. Prostaglandin D2-mediated microglia/astrocyte interaction enhances astrogliosis and demyelination in twitcher. J. Neurosci. 26, 4383–4393 (2006).
Mohri, I. et al. Hematopoietic prostaglandin D synthase and DP1 receptor are selectively upregulated in microglia and astrocytes within senile plaques from human patients and in a mouse model of Alzheimer disease. J. Neuropathol. Exp. Neurol. 66, 469–480 (2007).
Eguchi, N. et al. Lack of tactile pain (allodynia) in lipocalin-type prostaglandin D synthase-deficient mice. Proc. Natl. Acad. Sci. USA 96, 726–730 (1999).
Taniike, M. et al. Perineuronal oligodendrocytes protect against neuronal apoptosis through the production of lipocalin-type prostaglandin D synthase in a genetic demyelinating model. J. Neurosci. 22, 4885–4896 (2002).
Pandey, H. P., Ram, A., Matsumura, H. & Hayaishi, O. Concentration of prostaglandin D2 in cerebrospinal fluid exhibits a circadian alteration in conscious rats. Biochem. Mol. Biol. Int. 37, 431–437 (1995).
Jordan, W. et al. Prostaglandin D synthase (beta-trace) in healthy human sleep. Sleep 27, 867–874 (2004).
Koinuma, S. et al. The resetting of the circadian rhythm by prostaglandin J2 is distinctly phase-dependent. FEBS Lett. 583, 413–418 (2009).
Hirata, M., Kakizuka, A., Aizawa, M., Ushikubi, F. & Narumiya, S. Molecular characterization of a mouse prostaglandin D receptor and functional expression of the cloned gene. Proc. Natl. Acad. Sci. USA 91, 11192–11196 (1994).
Nagata, K. et al. CRTH2, an orphan receptor of T-helper-2-cells, is expressed on basophils and eosinophils and responds to mast cell-derived factor(s). FEBS Lett. 459, 195–199 (1999).
Hatanaka, M. et al. 15d-Prostaglandin J2 enhancement of nerve growth factor–induced neurite outgrowth is blocked by the chemoattractant receptor-homologous molecule expressed on T-helper type 2 cells (CRTH2) antagonist CAY10471 in PC12 cells. J. Pharmacol. Sci. 113, 89–93 (2010).
Haba, R. et al. Central CRTH2, a second prostaglandin D2 receptor, mediates emotional impairment in the lipopolysaccharide and tumor-induced sickness behavior model. J. Neurosci. 34, 2514–2523 (2014).
Onaka, Y. et al. CRTH2, a prostaglandin D2 receptor, mediates depression-related behavior in mice. Behav. Brain Res. 284, 131–137 (2015).
Onaka, Y. et al. Prostaglandin D2 signaling mediated by the CRTH2 receptor is involved in MK-801-induced cognitive dysfunction. Behav. Brain. Res. 314, 77–86 (2016).
Gray, S. L., Cummings, K. J., Jirik, F. R. & Sherwood, N. M. Targeted disruption of the pituitary adenylate cyclase-activating polypeptide gene results in early postnatal death associated with dysfunction of lipid and carbohydrate metabolism. Mol. Endcrinol. 15, 1739–1747 (2001).
Schwartz, W. J. & Zimmerman, P. Circadian timekeeping in BALB/c and C57BL/6 inbred mouse strains. J. Neurosci. 11, 3685–3694 (1990).
Peirson, S. N., Brown, L. A., Pothecary, C. A., Benson, L. A. & Fisk, A. S. Light and the laboratory mouse. J. Neurosci. Methods 300, 26–36 (2018).
Hashimoto, H. et al. Altered psychomotor behaviors in mice lacking pituitary adenylate cyclase-activating polypeptide (PACAP). Proc. Natl. Acad. Sci. USA 98, 13355–13360 (2001).
Matsuoka, T. et al. Prostaglandin D2 as a mediator of allergic asthma. Science 287, 2013–2017 (2000).
Satoh, T. et al. Prostaglandin D2 plays an essential role in chronic allergic inflammation of the skin via CRTH2 receptor. J. Immunol. 177, 2621–2629 (2006).
Sugimoto, Y. et al. Microarray evaluation of EP4 receptor-mediated prostaglandin E2 suppression of 3T3-L1 adipocyte differentiation. Biochem. Biophys. Res. Commun. 322, 911–917 (2004).
Tsuchiya, S. et al. Characterization of gene expression profiles for different types of mast cells pooled from mouse stomach subregions by an RNA amplification method. BMC Genom. 10, 35 (2009).
Hashimoto, H. et al. Distribution of the mRNA for a pituitary adenylate cyclase-activating polypeptide receptor in the rat brain: an in situ hybridization study. J. Comp. Neurol. 371, 567–577 (1996).
Daan, S. & Pittendrigh, C. S. A functional analysis of circadian pacemakers in nocturnal rodents. II. variability phase response curves. J. Comp. Physiol. 106, 253–266 (1976).
Zhao, H., Ohinata, K. & Yoshikawa, M. Central prostaglandin D2 exhibits anxiolytic-like activity via the DP1 receptor in mice. Prostaglandins Other Lipid Mediat. 88, 68–72 (2009).
Acknowledgements
This work was supported in part by JSPS KAKENHI, grant numbers JP17H03989 (H.H.), JP17K19488 (H.H.), JP20H00492 (H.H.), JP19K07121 (A.H.-T.), JP16H01881 (Y.U.), JP21790077 (N.S.), JP18790053 (N.S.); MEXT KAKENHI, grant number JP18H05416 (H.H.); AMED, grant number JP19dm0107122 (H.H.), JP19dm0207061 (H.H.), JP20am0101084 (H.H.); and a grant from the Takeda Science Foundation (H.H.).
Author information
Authors and Affiliations
Contributions
C.K., A.H.-T., M.H., A.K., R.N., Y.Y., and Y.S. carried out the experimental work. K.N. contributed to the design of experiments regarding the actograms of mouse locomotor activity and light exposure. S.T., Y.S., A.I., and Y.O. performed the RNA sequence analysis. Y.S., A.I., Y.U., H.H., K.N., M.N., and S.N. contributed to resource acquisition and provided unpublished reagents/analytic tools. Y.S., A.I, K.T., and A.B. advised on study design and helped conceive the study. A.K. and Y.A. helped with interpretation of the data. N.S., A.H.-T., T.N., and H.H. conceived, designed and coordinated the entire study. C.K., A.H.-T., T.N., and H.H. wrote the paper. All authors reviewed and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kawaguchi, C., Shintani, N., Hayata-Takano, A. et al. Lipocalin-type prostaglandin D synthase regulates light-induced phase advance of the central circadian rhythm in mice. Commun Biol 3, 557 (2020). https://doi.org/10.1038/s42003-020-01281-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01281-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.