Abstract
Anatomically modern humans reached East Asia more than 40,000 years ago. However, key questions still remain unanswered with regard to the route(s) and the number of wave(s) in the dispersal into East Eurasia. Ancient genomes at the edge of the region may elucidate a more detailed picture of the peopling of East Eurasia. Here, we analyze the whole-genome sequence of a 2,500-year-old individual (IK002) from the main-island of Japan that is characterized with a typical Jomon culture. The phylogenetic analyses support multiple waves of migration, with IK002 forming a basal lineage to the East and Northeast Asian genomes examined, likely representing some of the earliest-wave migrants who went north from Southeast Asia to East Asia. Furthermore, IK002 shows strong genetic affinity with the indigenous Taiwan aborigines, which may support a coastal route of the Jomon-ancestry migration. This study highlights the power of ancient genomics to provide new insights into the complex history of human migration into East Eurasia.
Similar content being viewed by others
Introduction
After the major Out-of-Africa dispersal of Homo sapiens around 60,000 years ago (60 kya), modern humans rapidly expanded across the vast landscapes of Eurasia1. Both fossil and ancient genomic evidence suggest that groups ancestrally related to present-day East Asians were present in eastern China by as early as 40 kya2. Two major routes for these dispersals have been proposed, either from the northern or southern parts of the Himalaya mountains1,3,4,5,6. Population genomic studies on present-day humans7,8 have exclusively supported the southern route origin of East Asian populations. On the other hand, the archaeological record provides strong support for the northern route as the origin of human activity, particularly for the arrival at the Japanese archipelago located at the east end of Eurasian continent. The oldest use of Upper Paleolithic stone tools goes back 38,000 years, and microblades, likely originated from an area around Lake Baikal in Central Siberia, are found in the northern island (i.e., Hokkaido; ~25 kya) and main-island (i.e., Honshu; ~20 kya) of the Japanese archipelago9. However, few human remains were found from the Upper Paleolithic sites in the archipelago. The Jomon culture started >16 kya, characterized by a hunter-fisher-gathering lifestyle with the earliest use of pottery in the world10. This Jomon culture lasted until a start of rice cultivation which brought by people migrated from the Eurasian continent, plausibly through the Korean peninsula, to northern parts of Kyushu island in the Japanese archipelago 3 kya. Several lines of archaeological evidence support the cultural continuity from the Upper-Paleolithic to the Jomon period, providing a hypothesis that the Jomon people are direct descendants of Upper-Paleolithic people who likely remained isolated in the archipelago until the end of Last Glacial Maximum9,11,12. Therefore, ancient genomics of the Jomon can provide new insights into the origin and migration history of East Asians.
A critical challenge for ancient genomics with samples from the Japanese archipelago is the inherent nature of warm and humid climate conditions except for the most north island, Hokkaido, and the soils indicating strong acidity because of the volcanic islands, which generally result in poor DNA preservation13,14. Though whole-genome sequences of two Hokkaido Jomon individuals dated to be 3500–3800-year-old were recently published with sufficient coverage15, a partial genome of a 3000-year-old Jomon individual from the east-north part of Honshu Japan was reported, with very limited coverage (~0.03-fold) due to the poor preservation16. To identify the origin of the Jomon people, we sequenced the genome of a 2500-year-old Jomon individual (IK002) excavated from the central part of Honshu to 1.85-fold genomic coverage. Comparing this IK002 genome with ancient Southeast Asians, we previously reported genetic affinity between IK002 and the 8000 years old Hòabìnhian hunter-gatherer17. This direct evidence on the link between the Jomon and Southeast Asians, thus, suggests the southern route origin of the Jomon lineage. Nevertheless, key questions still remain as to (1) whether the Jomon were the direct descendant of the Upper Paleolithic people who were the first migrants into the Japanese archipelago and (2) whether the Jomon, as well as present-day East Asians, retain ancestral relationships with people who took the northern route.
Here, we test the deep divergence of the Jomon lineage and the impacts of southern- versus northern-route ancestry on the genetic makeup of the Jomon. The Jomon forms a lineage basal to both ancient and present-day East Asians; this deep origin supports the hypothesis that the Jomon were direct descendants of the Upper Paleolithic people. Furthermore, the Jomon has strong genetic affinities with the indigenous Taiwan aborigines. Our study shows that the Jomon-related ancestry is one of the earliest-wave migrants who might have taken a coastal route on the way from Southeast Asia toward East Asia.
Results
Ancient DNA sequencing of Jomon specimens
Initially, we extracted DNA of 12 individuals from three sites (Supplementary Fig. 1), which were accompanied by remains associated with the Jomon culture and had no evidence for influence from the following culture called Yayoi. The endogenous DNA contents for ten out of the twelve individuals were <1.0% due to poor DNA preservation, while those of two individuals, IK002 and HG02, were more than 1.0% (Supplementary Data 1). From those remaining two individuals, only the ~2500 years old IK002 exhibited typical patterns of DNA damage expected for ancient remains18,19,20 (Supplementary Fig. 2), which was then selected for whole-genome sequencing. A total of 29 double-stranded sequencing libraries were generated using DNA extracted from the teeth and petrous bone of IK002, yielding endogenous DNA contents ranging from 1.14 to 19.09% (Supplementary Data 2). The libraries were sequenced to average coverages of 1.85-fold for autosomal genome and 146-fold for mitochondrial genome, with low estimated contamination rate of 0.5% (95% CI: 0.01–2.2%, Supplementary Fig. 3)21. We found IK002 to be assigned to mitochondrial haplogroup N9b1, which is rare among present-day Japanese people (<2.0%), but common in the Jomon22,23. We determined the genetic sex with the method based on the ratio of Y chromosome to X chromosome; the estimate of <0.3 supports that IK002 is female, which is consistent with morphological assessments.
The origin of IK002
To make inferences on the genetic relationship of IK002 with geographically diverse human populations, we merged the IK002 genomic data with a diverse panel of previously published ancient genomes24,25,26,27,28,29,30, as well as 300 high-coverage present-day genomes from the Simons Genome Diversity Project31. We extracted genotypes for a set of 2,043,687 SNP sites included in the “2240K” capture panel32. First, we characterized IK002 in the context of worldwide populations using principal component analysis (PCA)33,34. We found that IK002 sat in between present-day East Asians and a cluster of ancient Hòabìnhian hunter-gatherers and the Upper-Paleolithic (40 kya) individual from Tiányuán Cave17,30,35 (Fig. 1a). Second, the Honshu Jomon, IK002, closely clusters with the two Hokkaido Jomon, F23 and F5; we confirm that IK002 and the Hokkaido Jomon form a clade to the exclusion of other populations using f4-statistics (Supplementary Data 3). Henceforth, we will use IK002 as the representative of the Jomon people in this paper. Third, when using a smaller number of SNPs (41,264 SNPs) including the present-day Ainu36 from Hokkaido (Supplementary Fig. 1), IK002 clusters with the Hokkaido Ainu (Supplementary Fig. 4), supporting previous findings that the Hokkaido Ainu are direct descendants of the Jomon people16,36,37,38,39,40,41,42,43. Outgroup-f3 statistics support those PCA clustering, with IK002 sharing most genetic drift with the Hokkaido Jomon, followed by the Ainu (Supplementary Fig. 5 & Supplementary Data 4). Thus, our results indicate that IK002 is genetically distinct from present-day people in East Eurasia or even in Japan, with the exception of the Hokkaido Ainu.
a Principal component analysis (PCA) of ancient and present-day individuals from worldwide populations after the out-of-Africa expansion. Gray labels represent population codes showing coordinates for individuals. Colored circles indicate ancient individuals. b ADMIXTURE ancestry components (K = 15) for ancient and selected contemporary individuals. The color of light blue represents the component of IK002, which is shared with the present-day Japanese and Ulchi. c Maximum-likelihood phylogenetic tree (TreeMix) with bootstrap support of 100% unless indicated otherwise. The tree shows phylogenetic relationship among present-day Southeast/East Asians, Northeast Siberians, Native Americans, and ancient East Eurasians. Mbuti are the present-day Africans; Ust’Ishim is an Upper-Paleolithic individual (45 kya) from Western Siberia83; Mal′ta (MA-1)25 and Sunghir is Upper-Paleolithic individuals (24 kya and 34 kya)29, and Loschbour is a Mesolithic individual from West Eurasia88; La368 is a pre-Neolithic Hòabìnhian hunter-gatherer (8.0 kya) in Laos and Önge is the present-day hunter-gatherers in the Andaman island, both of who are from Southeast Asia17; Tiányuán is an Upper-Paleolithic individual (40 kya) in Beijing, China35; Kusunda are the present-day minority people in Nepal; Chokhopani is an Iron-age individual (3.0–2.4 kya) and Sherpa are the present-day minority people, both of who are in Tibet6; Han, Ami and main-island Japanese are the present-day East Asians31; Devils Gate Cave is a Neolithic individual (8.0 kya) in the Primorye region of Northeast Siberia, and Shamanka and Lokomotive are Early-Neolithic individuals (8.0 kya) from Central Siberia, respectively47; USR1 and Clovis are late-Paleolithic individuals (11.5 kya and 12.6 kya) in Alaska and Montana, respectively49,89. Colored arrows represent the migration pathways and signals of admixture among all datasets. The migration weight represents the fraction of ancestry derived from the migration edge.
Subsequently, we carried out model-based unsupervised clustering using ADMIXTURE44 (Supplementary Fig. 6). Assuming K = 15 ancestral clusters (Fig. 1b), an ancestral component unique to IK002 appears, which is the most prevalent in the Hokkaido Ainu (average 79.3%). This component is also shared with present-day Honshu Japanese as well as Ulchi (9.8% and 6.0%, respectively) (Fig. 1b). Those results also support the strong genetic affinity between IK002 and the Hokkaido Ainu.
We used ALDER45 in order to date the timing of admixture in populations with Jomon ancestry. Using IK002 and the Hokkaido Jomon as a merged source population representing Jomon ancestry, and present-day Han Chinese as the second source representing mainland East Asian ancestry, we estimated the admixture in present-day Honshu Japanese to be between 60 and 77 generations ago (~1700–2200 years ago assuming 29 years/generation), which is slightly earlier than previous estimates8 but more consistent with the archaeological record (Supplementary Data 5). This indicates the admixture started and continuously occurred after the Yayoi period. For the Ulchi we estimated a more recent timing (31–47 generations ago) consistent with the higher variance in the IK002 component observed in ADMIXTURE. Finally, we detected more recent (17–25 generations ago) admixture for the Hokkaido Ainu, likely a consequence of still ongoing gene flow between the Hokkaido Ainu and Honshu Japanese. The estimates of admixture timing are consistent when replacing Han with Korean, Ami or Devil′s Gate cave as mainland East Asian source population, and exponential curves from a single admixture event fit the observed LD curve well (Supplementary Fig. 7 & Supplementary Data 5).
To further explore the deep relationships between the Jomon and other Eurasian populations, we used TreeMix46 to reconstruct admixture graphs of IK002 and 18 ancient and present-day Eurasians and Native Americans (Fig. 1c & Supplementary Fig. 8). We found the IK002 lineage placed basal to the divergence between ancient and present-day Tibetans6,31 and to the common ancestor of the remaining ancient/present-day East Eurasians31,47,48 and Native Americans49,50. These genetic relationships are stable across different numbers of migration incorporated into the analysis. Major gene flow events recovered include the well-documented contribution of the Mal′ta individual (MA-1) to the ancestor of Native Americans25,49, as well as a contribution of IK002 to present-day mainland Japanese (m = 3–8; Supplementary Fig. 8). IK002 can be modeled as a basal lineage to East Asians, Northeast Asians/East Siberians, and Native Americans, supporting a scenario in which their ancestors arrived through the southern route and migrated from Southeast Asia toward Northeast Asia7,17. However, regarding Native Americans, high genetic contributions (11.8–36.8%) were detected from the Upper Paleolithic individual, MA-1, which means that Native Americans were admixture between the southern and the northern routes as shown in Raghavan et al. (2014). The divergence of IK002 from the ancestors of continental East Asians therefore likely predates the split between East Asians and Native Americans, which has been previously estimated at 26 kya49. Thus, our TreeMix results support the hypothesis that IK002 is a direct descendant of the people who brought the Upper Paleolithic stone tools 38,000 years ago into the Japanese archipelago.
The impacts of the Northern route migration into East Asia
Taking advantage of the earliest divergence of the IK002 (Fig. 1c & Supplementary Fig. 8), we address a question if the Upper-Paleolithic people who took the northern route of the Himalayas mountains to arrive east Eurasia made genetic contribution to populations migrated from Southeast Asia. Under the assumption that MA-1 is a descendant of a northern route wave, we tested gene flow from MA-1 to IK002, as well as to the other ancient and present-day Southeast/East Asians and Northeast Asians/East Siberians by three different forms of D statistics: D(Mbuti, MA-1; X, Ami), D(Mbuti, X; MA-1, Ami), and D(Mbuti, Ami; X, MA-1).
The first D statistics (shown as red in Fig. 2) provides results consistent with previous findings on the prevalence of MA-1 ancestry in the present-day Northeast Asians/East Siberians (Z < −3; p < 0.003, Supplementary Data 6)25, while none of Southeast/East Asians, except for Oroqen, shows a significant deviation from zero. The tree relationships observed in Fig. 1c are confirmed from the other two different forms between Ami and all of the tested populations with some variation that is mostly explained by the MA-1 gene flow (cyan and green in Fig. 2, Supplementary Data 7 & Supplementary Data 8). Therefore, there was no detectable signature of gene flow from MA-1 to the ancient/present-day Southeast/East Asians including IK002.
Remnant of the Jomon-related ancestry in the coastal region
The old divergence of IK002 (Fig. 1c) implies a negligible contribution to later ancient and present-day mainland East Asian groups. We further tested this implication by using f4-statistics with the form of (Mbuti, IK002; X, Chokhopani). If IK002 was a true outgroup to later East Asian groups, this statistic is expected to be zero for any test population X. However, we find that together with Japanese, present-day Taiwan aborigines (i.e., Ami and Atayal), as well as populations from the Okhotsk-Primorye region (i.e., Ulchi and Nivhk), also showed a significant (Z < −3; p < 0.003) excess of allele sharing with IK002. Populations in the inland of the eastern part of the Eurasian continent on the other hand were consistent with forming a clade with Chokhopani (Fig. 3), which suggests the presence of Ikawazu Jomon-related ancestry in the present-day coastal populations in East Asia. The signal is also present in the Neolithic individuals from Devil′s Gate Cave in the Primorye region (Z < −3; p < 0.003; Supplementary Fig. 9), suggesting that, at 8 kya, populations who had IK002-related ancestry in the region had already been largely but not completely replaced by later migrations. Interestingly, the genetic affinity to IK002 was found only in the coastal region but not in the inland for both ancient and present-day populations (Fig. 3).
Heatmaps of f4(Mbuti, IK002;X, Chokhopani), where X are the present-day East Eurasian populations. The color and size represents the value of f4-statistics. The shape represents statistical significances of genetic affinities based on Z-score. Triangle label means statistical significance with |Z| > 3 (P < 0.01), inverted triangle means weak significance with |Z| = 2–3 and circle means non-significance with |Z| < 2 (P > 0.05).
We carried out admixture graph modeling to further characterize the contributions of IK002-related ancestry to other East Asian populations. To that end, we first fit a backbone graph including ancient genomes representative of major divergences among East Asian lineages: IK002 (early dispersal); Chokhopani (later dispersal, East Asia), and Shamanka (later dispersal, Siberia). Test populations of interest were then modeled as three-way mixtures of early (IK002) and later (Chokhopani, Shamanka) dispersal lineages, using a grid search of admixture proportions within qpGraph. Consistent with the results from the f4-statistics, we find that models without contribution from IK002 result in poor fit scores for Japanese, Devil′s Gate Cave and Ami, as opposed to inland groups such as Han which do not require IK002-related ancestry (Supplementary Fig. 10). The range of admixture fractions with good model fit is generally quite wide, with best fit models showing IK002-related contributions of 8%, 4 and 41% into Japanese, Devil′s Gate Cave and Ami, respectively (Supplementary Fig. 10 & Supplementary Fig. 11). While the substantial contribution into Ami seems at odds with the lower f4-statistics compared with Japanese, the lineage admixing with Ami shares only a very short branch with IK002, suggesting a contribution from a distinct group with an early divergence from the IK002 lineage. We note that this backbone graph fitting assumed an unadmixed Jomon lineage, as opposed to a previously suggested dual-ancestry model where Jomon is admixed between Önge- and Ami-related ancestry. This alternative base model provides an equivalent admixture graph fit, however we find no evidence for shared genetic drift between the Önge and the ancestral Jomon lineage in qpGraph (Supplementary Fig. 12a & Supplementary Fig. 12b), or using direct f4-statistics (Supplementary Data 9 & Supplementary Data 10). Additional sampling of early East Asian human remains will be needed to further resolve the relationships among these deep lineages, but nevertheless either model supports the deep origins of Jomon.
Discussion
This study takes advantage of whole-genome sequence data from the 2500 years old Jomon individual, IK002, dissecting the origins of present-day East Asians. IK002 is modeled as a basal lineage to East Asians, Northeast Asians/East Siberians, and Native Americans (basal East Eurasians, bEE) after the divergence between Tiányuán and the ancestor of hunters-gatherers in Southeast Asia (Fig. 4)7,17. We clearly show the early divergence of IK002 from the common ancestor of the other ancient and present-day East Eurasian and Native Americans (Fig. 1c). Given that the split between the East Asian lineage and the Northeast Asians/East Siberian and Native American lineage was estimated to be 26 kya49, the divergence of the lineage leading to IK002 is likely to have occurred before this time but after 40 kya when the Tiányuán appeared (Fig. 4). Therefore, our results support the archaeological evidence based on lithic industry that the Jomon are direct descendants of the Upper-Paleolithic people who started living in the Japanese archipelago 38 kya (Supplementary Discussion).
The basal East Eurasians (bEE) are an ancient population that had no divergence among the ancestors of East Asians, Northeast Asians/East Siberian, and Native Americans. NA-ES-NA presents another ancient population that had no split between the ancestors of Northeast Asians/East Siberian and Native Americans.
We use the MA-1 ancestry as a proxy for ancestral populations who took the northern route of Himalaya mountains to come to East Eurasia. The fine stone tool, i.e., microblade, is a representative technology that was originally developed around Lake Baikal in Central Siberia during the Upper-Paleolithic period9. This microblade culture was also reached the Hokkaido island ~25 kya and the main island of the Japanese archipelago ~20 kya. If this culture was brought by demic diffusion, IK002 should still retain the MA-1 ancestry. However, we find no evidence on the genetic affinity of MA-1 with ancient and present-day Southeast/East Asians including Devil′s Gate Cave (8.0 kya), Chokhopani (3.0–2.4 kya), and IK002 (2.5 kya) (Fig. 2). Therefore, we conclude that MA-1 gene flow occurred after the divergence between the ancestral populations of Northeast Asians/East Siberians/Native Americans (NA-ES-NA) and East Asians (Fig. 4): namely, East Asians originated in Southeast Asia without any detectable genetic influence from the ancestor who took the northern route. There are two hypothetical possibilities to explain the contradiction between genome data and archaeological records. The first possibility is that MA-1 may not be a direct ancestor who invented the microblade culture. The second is that, if the assumption is correct, then the northern-route culture represented by microblade was brought to the Japanese archipelago by the NA-ES-NA population who must have had substantial gene flow from MA-1, which was a typical “cultural diffusion.” The first and second possibilities can be examined by obtaining genome data of Upper-Paleolithic specimens hopefully accompanying with microblade newly excavated from around Lake Baikal and from the Primorye region, respectively.
The genome of the Ikawazu Jomon (IK002) strongly supports the dual structure model of the origins of modern Japanese37. The PCA plot and phylogenetic tree showed that the present-day Japanese fell in the cluster of present-day East Asians (e.g., Han Chinese) but not clustered with IK002 (Fig. 1a & c), while a signal of gene flow was detected from IK002 to present-day Japanese (Fig. 1c & Supplementary Fig. 8). The PCA and ADMIXTURE showed the close relationship between IK002 and the Hokkaido Ainu even in the genome-wide structure reflected by linkage blocks (Supplementary Fig. 4 & Supplementary Fig. 6). These results fit the hypothesis that the Ainu and the Jomon share the common ancestor: the present-day Honshu Japanese are the hybrid between the Jomon and migrants from the East Eurasian continent, and the Hokkaido Ainu have less influence of genetic contribution of the migrants37,42.
IK002 gave new insights into the migration route from south to north in East Eurasia. The f4-statistics suggest that both the ancient and the present-day East Asians are closer to IK002 than Chokhopani (ancient Tibetans, 3.0–2.4 kya) in the coastal region but not in the inland region (Fig. 3 & Supplementary Fig. 9). Recent archaeological evidence shows little cultural influence from ancestral Austronesians to the Ryukyu islands (i.e., southernmost islands in the Japanese archipelago) in the last 10,000 years51. Given this cultural continuity, it is unlikely that this extra genetic affinity to IK002 was formed due to recent gene flow as shown in a previous study52 (see Supplementary Discussion). Here, we provide two explanations for this signal: (1) the earliest-wave of migration from south to north occurred through the coastal region, and/or (2) the migration occurred in both the coastal and inland regions, but the genetic components of the earliest-wave were drowned out by back-migration(s) from north to south occurred in the inland region. In the early migration of anatomically modern humans, the route along the coast has been primarily thought to be important3,53,54,55,56. The use of water craft could support such explanation for the expansions through the islands and the coastal region3, which supports the first explanation. There could be, however, potential criticisms: such archaeological evidence of craft boat is hardly found. Ulchi and Nivkh show significantly negative values of f4-statistics (Z = −4.541 and −10.148, respectively. p < 0.000006). This could be an influence of the Hokkaido Ainu who are likely to be direct descendants of the Jomon people. The ancestor of the Ainu people could have admixed with the Okhotsk people57 who were genetically as well as morphologically close to Ulchi and Nivkh in the the Primorye region58,59,60,61,62. The second explanation is that the track of the earliest-wave was erased in the inland but left over in the coastal region. Taiwan aborigines (Ami and Atayal) and Igorot are the Austronesian minorities. Taiwan aborigines are thought to have come from the East Eurasian continent 13.2 ± 3.8 kya63, though the origin of Igorot (which can be derogatory, and could be best referred as Kankanaey) is not well-known.
The coastal-migration hypothesis would be supported by morphological traits of the Jomon individuals excavated from the Hobi shell-mound site, which is geographically very close to the Ikawazu shell-mound site in the Atsumi peninsula. Kaifu and Masuyama (2018) examined spatiotemporal variation in humeral shaft thickness using a sample of 1003 prehistoric individuals from various sites in the Japanese Archipelago, including 797 Jomon individuals64. The results showed that the Jomon humeri were thicker in coastal populations than in inland populations. Kaifu and Masuyama (2018) hypothesized that this was caused by not only fishing in the outer sea, but also active marine transportation by rowing boats. This previous study suggests the Hobi Jomon individuals in the Atsumi peninsula were the people who adapted to coastal environment.
However, there is still lack of ancient genome data in East Asia critical to understand the peopling history of East Eurasians. Although our data support the idea that IK002 was the direct descendant of the Upper-Paleolithic people, how/where those Upper-Paleolithic people migrated to the Japanese archipelago remains unanswered. It is required to analyze more ancient genome data in order to fill the gap and to prove the hypothetical migration routes.
Methods
Human remains from archaeological sites in Jomon periods
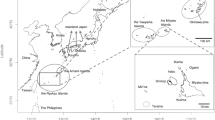
Twelve individuals from three archaeological sites (Ikawazu and Hobi Shell- mounds and Hegi Cave) were applied to this genomic study. The Ikawazu Shell-mound site locates in the Tahara city [34° 38′ 43″ north latitude; 137° 8′ 52″east longitude], Aichi Prefecture, where is in the central part of main island of the Japanese archipelago. The Hobi Shell-mounds, which are close to each other within ~5 km, locate on the center of main island (Honshu) (Supplementary Fig. 1). The Hegi Cave site locates on the northern part of Kyushu island. Based on conventional chronology of potteries, the individuals from the Hegi Cave site and the Hobi and Ikawazu Shell-mounds were assigned to the early to middle Jomon period (ca. 5000–8000 years ago) and the late to final Jomon periods (ca. 3500–2500 years ago), respectively (Supplementary Data 1).
Archaeological information of the Ikawazu remains
The Ikawazu Shell-mound was initially excavated in 191865: over 100 individuals were excavated from the site, accompanied with the Jomon potteries assigned to the late–final Jomon period (ca. 3500–2500 years ago) based on the pottery chronology. More recently, one of us (T.M.) and colleagues excavated the new section within the Ikawazu Shell-mound site in 2010 and found six buried individuals of complete skeletal remains. IK001 and IK002 were recovered from Pit No.4, and showed a better state of preservation than those of the other remains. IK001 and IK002 had morphologically typical Jomon characteristics. On the side of the IK002 head, a Jomon pottery, so called a Gokan-no-mori type Jomon ware-corded which is typical in the late Jomon period, was offered. IK001 was excavated together with IK002: the former was an infant, and the latter was a late-middle-age woman. Preliminary PCR-direct sequencing of mitochondrial DNA (mtDNA) showed IK001 and IK002 had different mt D-loop sequences, suggesting they were not a mother–child relationship.
We extracted the collagen from IK001 and IK002, and obtained the purified gelatins for radiocarbon dating that were carried out using a compact AMS at The University Museum in The University of Tokyo. The conventional radiocarbon ages were estimated to 2638 ± 16 BP and 2681 ± 16 BP, respectively. Given that the Ikawazu people built the large shell-mound, it is likely that IK001 and IK002 had applied to marine resources. To correct the marine reservoir effect depending on intake ratio of marine fish and shell, we measured stable isotope ratios of carbon and nitrogen from extracted gelatins of IK001 and IK002. Calculating contribution of amino acids from marine resources with the two end-points of terrestrial herbivore and marine fish, the 50% marine were estimated. These data were calibrated by OxCal 4.3 (calibration program based on the calibration curve of IntCal 13)66,67, and the calibrated ages of IK001 and IK002 showed 2699–2367 cal BP (95% CI) and 2720–2418 cal BP (95.4% CI), respectively. Because these ages were assigned to the Gokan-no-mori period which has no evidence of rice cultivation, we confirmed that IK001 and IK002 were individuals from the Jomon period accompanied with typical Jomon culture. We sampled teeth of IK001 (M1) and IK002 (M3), and fragments of the petrous bone of IK002.
DNA extraction
All the subsequent experiments of ancient DNA were performed in the clean room exclusively built for ancient DNA analyses installed in Department of Anatomy, Kitasato University School of Medicine. The bone and tooth pieces were cut by a sterile disc drill to separate crowns (enamel), roots (cementum and dentine) of teeth for all the samples, and the inner part of petrous bone only for IK002. DNA extraction was carried out with the protocol that is based on the previous studies24,68 and modified in this study.
The tooth samples were cut by a sterile and UV-irradiated disc drill to separate crown (enamel) and root (cementum and dentine). DNA extraction of the root was carried out by the Gamba method with our modification. The teeth were washed by 3% sodium hypochlorite solution (Sigma-Aldrich) for 15 min, in order to decrease the degree of modern DNA contamination. After washing the teeth with ultrapure water (Thermo Fisher Scientific) and 99% ethanol (Sigma-Aldrich), the teeth were dried on the clean bench in the clean room for 16 h. The washed samples were pulverized by freezer mill (ShakeMaster Auto ver 2.0, BioMedical Science Inc.), and fine powder was obtained. To release DNA molecules from the sample powder, 200 mg tooth powder was incubated for 24 h at 55 °C followed by 24 h at 37 °C in 2 ml DNA LoBind tube (Eppendorf) with 1 ml lysis buffer in final concentrations of 20 mM Tris HCl (pH 7.5), 0.7% N-lauroylsarcosine, 47.5 mM EDTA (pH 8), 0.65 U/ml Proteinase K, shaking at 900 rpm in a Thermomixer (Eppendorf). The samples were then centrifuged at 13,000 g for 10 min and the supernatants were discarded. Fresh lysis buffer (1 ml) was added to the pellet, vortexed, and the incubation and centrifugation steps were repeated. The second supernatants were then transferred to ultrafiltration tubes (Amicon® Ultra-4 Centrifugal Filter Unit 30K, Merck), diluted with 3 ml TE (pH 8.0) and centrifuged at 2,000 g until final concentrations of ~100 ml were obtained. These volumes were then transferred to silica column (MiniElute PCR Purification Kit, QIAGEN) and purified according to manufacture’s instructions, except for adding TWEEN 20 (at 0.05% final concentration) to 60 ul EB buffer pre-heated to 60 °C at the final step.
The petrous bone was cut by a sterile and UV-irradiated disc drill, and three pieces where Pinhasi et al. (2015) named as “C-part”69 were obtained (C1, C2, C3); the pieces were washed by ultrapure water (Thermo Fisher Scientific) and 99% ethanol (Sigma-Aldrich). After the dried pieces were drilled and homogenized, ~500 mg bone powder was obtained from the three pieces. The first powder of 150 mg was used to extract DNA molecules following the modified protocol mentioned above. The powders of C1, C2, C3 were rinsed by ultrapure water [Rinsed supernatant], then treated with pre-digestion buffer containing 20 mM Tris HCl (pH 7.5), 0.7% N-lauroylsarcosine, 0.4 M EDTA (pH 8), 0.65 U/ml recombinant Proteinase K for 30 min at shaking at 900 rpm in a Thermomixer (Eppendorf). The mixture was then centrifuged at 13,000 g for 10 min and the supernatant was transferred to a 2 ml tube DNA LoBind [Pre-digestion]. Fresh lysis buffer (1 ml) containing 20 mM Tris HCl (pH 7.5), 0.7% N-lauroylsarcosine, 47.5 mM EDTA (pH 8), 0.65 U/ml recombinant Proteinase K was added to the pellet. After vortexed and incubated for 24 h at 55 °C followed by shaking at 900 rpm for 24 h at 37 °C, the first extract was obtained [Extract 1]. This step was then repeated, and the second extract [Extract 2] was obtained. The residual pellet was pulverized by wet-grinding with shaking sterile beads in grinding cylinder. Fresh lysis buffer containing 20 mM Tris HCl (pH 7.5), 0.7% N-lauroylsarcosine, 0.4 M EDTA (pH 8), 0.65 U/ml recombinant Proteinase K was added into the pulverized pellet, and the pellet was incubated for 24 h at 55 °C followed by shaking at 900 rpm for 24 h at 37 °C in 2 ml tube, the third extract was obtained [Extract 3]. The five elutes (rinsed and pre-digestion supernatants and three extracts) were filtrated following the protocol mentioned in the paragraph of DNA extraction from tooth. Finally, we obtained five DNA extracts from each petrous bone piece (total 15 extracts).
Library construction
DNA extracts obtained from these samples were quantified and qualified by Qubit 3.0 (Thermo Fisher Scientific) and Bioanalyzer (Agilent). Twenty-two extracts were used to construct 34 NGS libraries for Illumina sequencing system in Kitasato University. The NGS sequencing was carried out using MiSeq (Illumina) in Kyushu University and HiSeq (Illumina) in National Institute of Genetics in Japan. For cross-check, we separately prepared another five extracts from IK002, and made the NGS libraries in Copenhagen University and sequenced them using HiSeq in the Danish National High-Throughput DNA Sequencing Centre in Copenhagen.
Concerning C1 and C3, we used only one elute, [Extract 2], to construct NGS libraries and used to run on NGSs only in Japan. Meanwhile, concerning C2, we used five elutes, [Rinsed supernatant], [Pre-digestion], [Extract 1], [Extract 2], [Extract 3], and for constructing NGS libraries of C2, two different protocols were used separately in two laboratories (Kitasato University in Japan, and Copenhagen University Geogenetics Laboratory in Denmark) for inter-laboratory crosschecking. In the Kitasato University, the bead-based size selection protocol with the NEBNext Ultra DNA library preparation kit (New England Biolabs: NEB) was used. To remove large DNA fragments that could be contaminants from modern organisms, we modified the NEB original protocol: we adjusted the mixing ratio of the Agencourt AMPure XP solution (Beckman Coulter), the Solid Phase Reversible Immobilization magnetic bead solution. In the Copenhagen University, the protocol shown in Allentoft et al. (2015)26 was used to make NGS libraries. Eventually, we constructed 6 libraries from tooth and 18 libraries from the petrous bone in the Kitasato University, and 5 libraries from the petrous bone in the Copenhagen University; totally we provided 29 libraries from IK002.
Data output, processing, and authentication
The 29 libraries were sequenced on a flowcell using the Ilumina HiSeq 2500 and the HiSeq reagent kit of normal and rapid mode for 100 cycles in paired end in the National Institute of Genetics in Japan and the Danish National High-Throughput DNA Sequencing Centre in Denmark. After running HiSeq, the sequence reads were called by Illumina’s Real Time Analysis (RTA) or CASAVA ver. 1.8.2 (Illumina) base-calling software. The HiSeq output-data were processed using customizable NGS pipeline in the Geogenetics Laboratory and the Kitasato University. AdapterRemoval ver. 270 was used to trim adapters terminal N’s (–trimns), low quality bases (-trim qualities,–minquality 2) and short reads (–minlength 30), and filtered reads were checked with FastQC ver. 0.11.771. The filtered reads were mapped against hg19, human reference genome, by BWA ver. 0.5.9. Mapped reads with mapping quality below Phred score 30 and duplicates were removed using SAMtools72 and the MarkDuplicates tool of Picard Tools (http://broadinstitute.github.io/picard/). Read depth and coverage were determined using pysam (https://github.com/pysam-developers/pysam) and Bedtools (https://github.com/arq5x/bedtools2).
Misincorporation patterns were assessed using mapDamage273. The degree of modern DNA contamination was estimated by ContamMix21 focused on mitogenome sequences. The resulting sequence assembly and haplotype for mitochondrial genome was visualized using MitoSuite ver. 1.0.974.
Analysis panels for population genetic inference were obtained by merging the mapped reads of IK002 and relevant previously published ancient individuals with two reference datasets of modern individuals:
-
Panel “2240K”: Genotypes for 404 whole-genome sequenced modern individuals31,75,76,77,78,79,80, at 2,043,687 autosomal SNPs targeted for in-solution capture in previously published ancient DNA panels81,82,83.
-
Panel “HO Ainu”: Genotypes for 2119 modern individuals from the Human Origins panel84, as well as 36 Ainu individuals85, at 41,264 SNPs overlapping between the two panels.
For both panels, pseudo-haploid genotypes for ancient individuals were generated by randomly sampling an allele passing filters (mapping quality ≥ 30 and base quality ≥ 30) at the reference panel SNP positions.
Principal component analysis and ADMIXTURE
As a first assessment of the genetic affinities of the study individuals we carried out PCA, as previously described29,33,86. In particular, we projected the low coverage ancient individuals onto the PCs inferred from different sets of modern and high coverage ancient individuals, using the ‘lsqproject’ and ‘autoshrink’ options in smartpca from the EIGENSOFT package33, on both analysis panels. To explore shared genetic component between IK002, Ainu and the other populations, we ran ADMIXTURE ver. 1.3.044 on the “HO Ainu” panel. ADMIXTURE was run in ten replicates, for K values ranging from K = 2 and K = 20, and best runs were selected and aligned using pong87.
TreeMix
Maximum-likelihood trees and admixture graphs were inferred using TreeMix46. A subset of populations from the “2240K” panel were chosen to represent different ancestries of East Eurasians and Native Americans; IK002, East Asians (Han, Ami, Japanese and Devils Cave), Northeast Siberians (Lokomotiv and Shamanka, the ancient Siberians), Native Americans (Clovis and USR1, the ancestry of Native American), Himalayan (Sherpa, Kusunda and Chokhopani, the ancient highlander) and Southeast Asians (Önge and La368, the Hoabinhian). Furthermore, Tiányuán, Mal’ta (MA-1) and Ust’Ism were included as a landmark of divergence events happened in the Upper Paleolithic period. We used Mbuti as an outgroup and ran TreeMix with m = 0 to eight migration events to fit admixture graphs to the data. We only considered the SNP sites that are non-missing in all individuals included in this analysis and chose the tree under each condition that showed the highest likelihood among ten replicates with different random seeds.
f-statistics and D-statistics
We used the D-statistic framework and f4-statistical analyses to investigate patterns of admixture and shared ancestry in our data set. All D-statistics were calculated from allele frequencies using the estimators described previously26,29, with standard errors obtained from a block jackknife (the jackknife parameter = 0.050, the number of blocks = 714). Calculating of D-statistics was carried out by qpDstat in the AdmixTools ver. 4.184. The values of D-statistics were visualized and mapped by R software. f4-statistics was calculated by qpDstat with the f4 mode.
ALDER admixture dating
To infer the timing of admixture we used ALDER45 on the “HO Ainu” panel, for Japanese, Ainu and Ulchi as target populations. We used IK002 and the two Hokkaido Jomon individuals as a combined Jomon source, and Han, Ami, Korean or the ancient individuals from Devil’s Gate cave as mainland East Asian source populations.
qpGraph modeling
Admixture graph modeling with qpGraph84 was carried out on the “2240K” panel. First, a backbone graph including ancient genomes representative of major divergences among East Asian lineages was fit: IK002 (early dispersal); Chokhophani (later dispersal, East Asia), and Shamanka (later dispersal, Siberia). Test populations of interest were then modeled as three-way mixtures of early (IK002) and later (Chokhopani, Shamanka) dispersal lineages, using a grid search in 1% increments of the two independent admixture proportions (using the ‘lock’ function in qpGraph).
Statistics and reproducibility
Ancient genomic data were generated using multiple libraries, which ensure reproducibility. Contamination ratios were <1%. All statistics was done using available packages and reproducibility can be accomplished using our own parameters mentioned in Methods.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
This study has been evaluated by the Education Affairs in Tahara city in Aichi Prefecture, Japan. All genomic data (fastq formats) are available for download in the DNA DataBank of Japan (DDBJ) Sequence Read Archive (DRA. https://www.ddbj.nig.ac.jp/dra/index-e.html) under the accession numbers DRA005042 (Sample accession SAMD00058001) and at the European Nucleotide Archive (ENA) with accession number PRJEB26721 (Sample accession SAMEA4665869). The final bam datasets of mitochondrial- and nuclear-genome sequences, population genetic datasets of AdmixTools format (“.indv”, “.snp” and “.geno” files) and other source data are available on the server in University of Tokyo (https://drive.google.com/drive/folders/1IYZaq1WUbcP_ER2wWx224vP7rkX86NHw?usp=sharing). There is no any restrictions on data access in this study.
References
Bae, C. J., Douka, K. & Petraglia, M. D. On the origin of modern humans: Asian perspectives. Science 358, eaai9067 1–7 (2017).
Shang, H., Tong, H., Zhang, S., Chen, F. & Trinkaus, E. An early modern human from Tianyuan Cave, Zhoukoudian, China. Proc. Natl. Acad. Sci. USA 104, 6573–6578 (2007).
Pope, K. O. & Terrell, J. E. Environmental setting of human migrations in the circum-Pacific region. J. Biogeogr. 35, 1–21 (2008).
Kaifu, Y., Izuho, M. & Goebel, T. Modern human dispersal and behavior in Paleolithic Asia. in Emergence and diversity of modern human behavior in Paleolithic Asia (eds Kaifu, Y., Izuho, M., Goebel, T., Sato, H., & Ono, A.) 535–566 (Texas A & M University Press College Station, 2015).
Reyes-Centeno, H., Hubbe, M., Hanihara, T., Stringer, C. & Harvati, K. Testing modern human out-of-Africa dispersal models and implications for modern human origins. J. Hum. Evol. 87, 95–106 (2015).
Jeong, C. et al. Long-term genetic stability and a high-altitude East Asian origin for the peoples of the high valleys of the Himalayan arc. Proc. Natl. Acad. Sci. USA 113, 7485–7490 (2016).
HUGO Pan-Asian SNP Consortium et al. Mapping human genetic diversity in Asia. Science 326, 1541–1545 (2009).
Jeong, C., Nakagome, S. & Di Rienzo, A. Deep history of East Asian populations revealed through genetic analysis of the Ainu. Genetics 202, 261–272 (2016).
Kaifu, Y., Izuho, M., Goebel, T., Sato, H. & Ono, A. Emergence and Diversity of Modern Human Behavior in Paleolithic Asia. (Texas A&M University Press, 2015).
Habu, J. Ancient Jomon of Japan. (Cambridge University Press, 2004).
Lambeck, K., Yokoyama, Y. & Purcell, T. Into and out of the Last Glacial Maximum: sea-level change during Oxygen Isotope Stages 3 and 2. Quat. Sci. Rev. 21, 343–360 (2002).
Nakagawa, T. et al. Pollen/event stratigraphy of the varved sediment of Lake Suigetsu, central Japan from 15,701 to 10,217 SG vyr BP (Suigetsu varve years before present): description, interpretation, and correlation with other regions. Quat. Sci. Rev. 24, 1691–1701 (2005).
Pinhasi, R. et al. Optimal ancient DNA yields from the inner ear part of the human petrous bone. PLoS ONE 10, e0129102 (2015).
Hofreiter, M. Ancient DNA. Oxford Bibliographies Online Datasets. https://doi.org/10.1093/obo/9780199941728-0036 (2014).
Kanzawa-Kiriyama, H. et al. Late Jomon male and female genome sequences from the Funadomari site in Hokkaido, Japan. Anthropol. Sci. 127, 83–108 (2019). 2019.
Kanzawa-Kiriyama, H. et al. A partial nuclear genome of the Jomons who lived 3000 years ago in Fukushima, Japan. J. Hum. Genet. 62, 213–221 (2017).
McColl, H. et al. The prehistoric peopling of Southeast Asia. Science 361, 88–92 (2018).
Briggs, A. W. et al. Patterns of damage in genomic DNA sequences from a Neandertal. Proc. Natl. Acad. Sci. USA 104, 14616–14621 (2007).
Orlando, L. et al. True single-molecule DNA sequencing of a pleistocene horse bone. Genome Res. 21, 1705–1719 (2011).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil–DNA–glycosylase treatment for screening of ancient DNA. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, 20130624 (2015).
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013).
Adachi, N., Shinoda, K.-I., Umetsu, K. & Matsumura, H. Mitochondrial DNA analysis of Jomon skeletons from the Funadomari site, Hokkaido, and its implication for the origins of Native American. Am. J. Phys. Anthropol. 138, 255–265 (2009).
Kanzawa-Kiriyama, H., Saso, A., Suwa, G. & Saitou, N. Ancient mitochondrial DNA sequences of Jomon teeth samples from Sanganji, Tohoku district, Japan. Anthropol. Sci. 121, 89–103 (2013).
Gamba, C. et al. Genome flux and stasis in a five millennium transect of European prehistory. Nat. Commun. 5, 5257 (2014).
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014).
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Jones, E. R. et al. Upper Palaeolithic genomes reveal deep roots of modern Eurasians. Nat. Commun. 6, 8912 (2015).
Sikora, M. et al. Ancient genomes show social and reproductive behavior of early Upper Paleolithic foragers. Science 358, 659–662 (2017).
Yang, M. A. et al. 40,000-year-old individual from Asia provides insight into early population structure in Eurasia. Curr. Biol. 27, 3202–3208.e9 (2017).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Fu, Q. et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl. Acad. Sci. U. S. A. 110, 2223–2227 (2013).
Jinam, T. et al. The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations. J. Hum. Genet. 57, 787–795 (2012).
Hanihara, K. Dual Structure Model for the Population History of the Japanese. Japan Rev. 2, 1–33 (1991).
Hanihara, K. Reanalysis of Local Variations in the Ainu Crania. Anthropol. Sci. 106, 1–15 (1998).
Shigematsu, M., Ishida, H., Goto, M. & Hanihara, T. Morphological affinities between Jomon and Ainu: reassessment based on nonmetric cranial traits. Anthropol. Sci. 112, 161–172 (2004).
Hammer, M. F. et al. Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes. J. Hum. Genet. 51, 47–58 (2006).
Ishida, H., Hanihara, T., Kondo, O. & Fukumine, T. Craniometric divergence history of the Japanese populations. Anthropol. Sci. 117, 147–156 (2009).
Nakagome, S. et al. Model-based verification of hypotheses on the origin of modern Japanese revisited by Bayesian inference based on genome-wide SNP data. Mol. Biol. Evol. 32, 1533–1543 (2015).
Yuasa, I. et al. Investigation of Japanese-specific alleles: most are of Jomon lineage. Leg. Med. 17, 52–55 (2015).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Loh, P.-R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254 (2013).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Damgaard, P. de B. et al. 137 ancient human genomes from across the Eurasian steppes. Nature 557, 369–374 (2018).
Siska, V. et al. Genome-wide data from two early Neolithic East Asian individuals dating to 7700 years ago. Sci. Adv. 3, e1601877 (2017).
Moreno-Mayar, J. V. et al. Terminal Pleistocene Alaskan genome reveals first founding population of Native Americans. Nature 553, 203–207 (2018).
Rasmussen, M. et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225–229 (2014).
Yamagiwa, K. et al. A possible new oldest pottery group in the Southern Ryukyu Islands, Japan: comparative analysis of elemental components of potsherds from the Shiraho-Saonetabaru Cave Site. J. Archaeological Sci.: Rep. 26, 101879 (2019).
Sato, T. et al. Genome-wide SNP analysis reveals population structure and demographic history of the ryukyu islanders in the southern part of the Japanese archipelago. Mol. Biol. Evol. 31, 2929–2940 (2014).
Stringer, C. Palaeoanthropology. Coasting out of Africa. Nature 405, 24–5 (2000). 27.
Cordaux, R. et al. Mitochondrial DNA analysis reveals diverse histories of tribal populations from India. Eur. J. Hum. Genet. 11, 253–264 (2003).
Cordaux, R. & Stoneking, M. South Asia, the Andamanese, and the genetic evidence for an ‘early’ human dispersal out of Africa. Am. J. Hum. Genet. 72, 1586–1590 (2003).
Underhill, P. A. A synopsis of extant Y chromosome diversity in East Asia and Oceania. THE PEOPLING OF EAST ASIA, (eds Sagart, L., Blench, R. and Sanchez-Mazas, A.) 297–214 (2005).
Jinam, T. A., Kanzawa-Kiriyama, H. & Saitou, N. Human genetic diversity in the Japanese Archipelago: dual structure and beyond. Genes Genet. Syst. 90, 147–152 (2015).
Ishida, H. Metric and nonmetric cranial variation of the Prehistoric Okhotsk people. Anthropol. Sci. 104, 233–258 (1996).
Komesu, A. et al. Nonmetric cranial variation in human skeletal remains associated with Okhotsk culture. Anthropol. Sci. 116, 33–47 (2008).
Kaburagi, M., Ishida, H., Goto, M. & Hanihara, T. Comparative studies of the Ainu, their ancestors, and neighbors: assessment based on metric and nonmetric dental data. Anthropol. Sci. 118, 95–106 (2010).
Sato, T. et al. Origins and genetic features of the Okhotsk people, revealed by ancient mitochondrial DNA analysis. J. Hum. Genet. 52, 618–627 (2007).
Sato, T. et al. Mitochondrial DNA haplogrouping of the Okhotsk people based on analysis of ancient DNA: an intermediate of gene flow from the continental Sakhalin people to the Ainu. Anthropol. Sci. 117, 171–180 (2009).
Trejaut, J. A. et al. Traces of archaic mitochondrial lineages persist in Austronesian-speaking Formosan populations. PLoS Biol. 3, e247 (2005).
Kaifu, Y. & Masuyama, T. Why humeri of the Jomon people are so thick?: Imprications from its inter-site variation. Anthropological Sci. (Jpn. Ser.) 126, 133–155 (2018).
Koganei, Y. On the ritual ablation of upper canine in the stone age people of Japan. Anthropol. Sci. 33, 31–36 (1918).
Ramsey, B. C. Bayesian analysis of radiocarbon dates. Radiocarbon 51, 337–360 (2009).
Reimer, P. J. et al. IntCal13 and Marine13 radiocarbon age calibration curves 0-50,000 years cal BP. Radiocarbon 55, 1869–1887 (2013).
Kemp, B. M. & Smith, D. G. Use of bleach to eliminate contaminating DNA from the surface of bones and teeth. Forensic Sci. Int. 154, 53–61 (2005).
Pinhasi, R. et al. Optimal ancient DNA yields from the inner ear part of the human petrous bone. PLoS ONE 10, e0129102 (2015).
Lindgreen, S. AdapterRemoval: easy cleaning of next generation sequencing reads. BMC Res. Notes 5, 337 (2012).
Andrews, S. & Others. FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Jónsson, H., Ginolhac, A., Schubert, M., Johnson, P. L. F. & Orlando, L. mapDamage2.0: fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics 29, 1682–1684 (2013).
Ishiya, K. & Ueda, S. MitoSuite: a graphical tool for human mitochondrial genome profiling in massive parallel sequencing. PeerJ 5, e3406 (2017).
Ayub, Q. et al. The Kalash genetic isolate: ancient divergence, drift, and selection. Am. J. Hum. Genet. 96, 775–783 (2015).
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014).
Raghavan, M. et al. The genetic prehistory of the New World Arctic. Science 345, 1255832 (2014).
Mondal, M. et al. Genomic analysis of Andamanese provides insights into ancient human migration into Asia and adaptation. Nat. Genet. 48, 1066–1070 (2016).
Raghavan, M. et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science 349, aab3884 (2015).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Jinam, T. et al. The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations. J. Hum. Genet. 57, 787–795 (2012).
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012).
Behr, A. A., Liu, K. Z., Liu-Fang, G., Nakka, P. & Ramachandran, S. pong: fast analysis and visualization of latent clusters in population genetic data. Bioinformatics 32, 2817–2823 (2016).
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
Rasmussen, M. et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225–229 (2014).
Acknowledgements
The excavation of the Ikawazu Jomon individual was supported by Grant-in-Aid for Scientific Research (B) (25284157) to Y.Y. The Ikawazu Jomon genome project was organized by H.I., and T.H. & H.O. who were supported by MEXT KAKENHI Grant Numbers 16H06408 and 17H05132, by Grant-in-Aid for Scientific Research on Innovative Areas (Cultural History of Paleoasia), and by Grant-in-Aid for Challenging Exploratory Research (23657167) and for Scientific Research (B) (17H03738). The Ikawazu Jomon genome sequencing was supported by JSPS KAKENHI Grant Number 16H06279 to A.To., and partly supported in the CHOZEN project in Kanazawa University, and in the Cooperative Research Project Program of the Medical Institute of Bioregulation, Kyushu University. Computations for the Ikawazu Jomon genome were partially performed on the NIG supercomputer at ROIS National Institute of Genetics.
Author information
Authors and Affiliations
Contributions
H.O. initiated and led the study. T.G., S.N., R.K., Y.Y., H.I., E.W., M.S., and H.O. designed the study. H.I., T.H., and N.S. supervised morphological aspects of the project, and Y.Y. and H.St. supervised archaeological aspects of the project. H.O. and M.S. supervised the overall project. Y.Y., Y.M., H.St., S.M., O.K., and N.S. excavated, curated and described the skeletal remains from the Ikawazu and the Hobi shell-mound sites, and T.T., T.W., and H.I. did those from the Hegi cave site. T.G., R.S., and H.O. performed the sampling for DNA analysis, and M.Y. conducted radiocarbon dating. T.G., T.S., K.K., S.R., R.S., and M.A. performed the DNA laboratory work. H.Sb., A.Ta., and A.To., performed deep sequencing with high-throughput sequencers. T.G., S.N., M.S., S.R., T.K., B.N., H.M., and T.S. analyzed or assisted in the analysis of data. T.G., S.N., M.S., and H.O. interpreted the results with considerable input from M.Y., Y.Y. and H.I., and wrote the paper. All authors discussed the results and contributed to the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gakuhari, T., Nakagome, S., Rasmussen, S. et al. Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations. Commun Biol 3, 437 (2020). https://doi.org/10.1038/s42003-020-01162-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01162-2
This article is cited by
-
Demographic history of Ryukyu islanders at the southern part of the Japanese Archipelago inferred from whole-genome resequencing data
Journal of Human Genetics (2023)
-
Spatial and temporal diversity of positive selection on shared haplotypes at the PSCA locus among worldwide human populations
Heredity (2023)
-
Parallel signatures of Mycobacterium tuberculosis and human Y-chromosome phylogeography support the Two Layer model of East Asian population history
Communications Biology (2023)
-
The genomic history of southwestern Chinese populations demonstrated massive population migration and admixture among proto-Hmong–Mien speakers and incoming migrants
Molecular Genetics and Genomics (2022)
-
New insights into the fine-scale history of western–eastern admixture of the northwestern Chinese population in the Hexi Corridor via genome-wide genetic legacy
Molecular Genetics and Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.