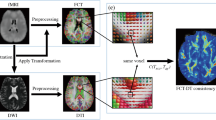
Abstract
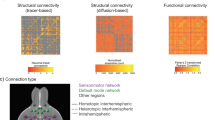
The brain is constituted of multiple networks of functionally correlated brain areas, out of which the default-mode network (DMN) is the largest. Most existing research into the DMN has taken a corticocentric approach. Despite its resemblance with the unitary model of the limbic system, the contribution of subcortical structures to the DMN may be underappreciated. Here, we propose a more comprehensive neuroanatomical model of the DMN including subcortical structures such as the basal forebrain, cholinergic nuclei, anterior and mediodorsal thalamic nuclei. Additionally, tractography of diffusion-weighted imaging was employed to explore the structural connectivity, which revealed that the thalamus and basal forebrain are of central importance for the functioning of the DMN. The contribution of these neurochemically diverse brain nuclei reconciles previous neuroimaging with neuropathological findings in diseased brains and offers the potential for identifying a conserved homologue of the DMN in other mammalian species.
Similar content being viewed by others
Introduction
In 1979, for the first time to our knowledge, David Ingvar used Xenon clearance to investigate resting wakefulness1. When aligned by scalp and skull markers, the 11 brains examined indicated an evident increase of the blood flow levels in the frontal lobe interpreted as a surrogate for undirected, spontaneous, conscious mental activity. Later, positron emission tomography (PET) was used to map more systematically task-related activation in the brain, often with resting wakefulness as a control task. The contrast between task-related and resting wakefulness led to the description of deactivation (i.e., active at rest more than during the task) in a set of regions, including retrosplenial cortex, inferior parietal cortex, dorsolateral frontal cortex inferior frontal cortex, left inferior temporal gyrus, medial frontal regions and amygdala2,3 that quickly bore the name of default-mode network (DMN)4. In these studies, skull landmarks or structural magnetic resonance imaging (MRI) were used to align PET images in Talairach stereotaxic or in Montreal Neurological Institute (MNI) templates2,3. The advent of functional magnetic resonance imaging (fMRI), particularly of methods for analysing functional connectivity, led to the allocation of new structures to this network, such as the hippocampal formation5,6,7.
Today, the DMN has largely been a cortically defined set of network nodes. Consisting of distinct regions/nodes distributed across the ventromedial and lateral prefrontal, posteromedial and inferior parietal, as well as the lateral and medial temporal cortex, the DMN is considered a backbone of cortical integration8,9,10,11. Its subcortical components are, however, less well characterised. Studies of whole-brain network organisation reveal subregions of the cerebellum12,13 and striatum14 that are functionally connected with the cortical regions of the DMN. Seed-based functional connectivity studies further demonstrate additional DMN-specific connectivity to several subcortical structures, including the amygdala15,16 and striatum17. The thalamus has also been shown to be structurally and functionally connected to DMN regions18,19. These studies are important, as a cleaner characterisation of the anatomy of the DMN is an essential step towards understanding its functional role and its involvement in brain diseases. Particularly, an increased activity characterises the regions that compose DMN during tasks involving autobiographical, episodic and semantic memory, mind wandering, perspective-taking or future thinking20,21; Bendetowicz et al.22. Conversely, DMN regions show a decreased neural activity during attention-demanding and externally oriented tasks21,23. Finally, altered connectivity in the DMN has been observed in a large variety of brain diseases, including Alzheimer’s disease, Parkinson’s disease, schizophrenia, depression, temporal lobe epilepsy, attention deficit and hyperactivity disorder, drug addiction, among others24,25,26,27,28,29. Hence, while prior research provides first hints towards a broader definition of the DMN system, further research is necessary to articulate the anatomical extent of specific subcortical contributions, and to understand the independent contribution of these structures in DMN function and pathologies.
Yet, since the DMN has repeatedly been characterised as a cohesive functional network5, an average of brain images relying exclusively on anatomical references and landmarks may be suboptimal30,31 whether the method employed is a surface-based or volume-based registration30,32. Small structures of the brain may be particularly susceptible to this misalignment, especially when MRI lacks contrast. Besides morphology, cytoarchitecture and function are poorly overlapping, especially in the DMN33,34. Consequently, functional areas present in every subject may not overlap after averaging all structurally aligned brain images in a group analysis30,35. This biological misalignment can be particularly problematic for revealing significant small regions of the DMN (Fig. 1). A better alignment is also essential for the subcortical structures of the brain, as their variability is still considerable36,37,38,39. Specifically, cytoarchitectonic studies have shown that only one-quarter of the volume of cholinergic nuclei overlaps in at least half of the individuals studied39. Similarly, structures such as mammillary bodies, nucleus basalis of Meynert, or anterior thalamic nuclei can vary in size, morphology and locations, and are particularly prone to misalignment with the current methods of structural registration32,40,41,42. Functional alignment methods have already been used to overcome the high interindividual variability of the morphology of some areas of the heteromodal association cortex and led to a more accurate mapping of resting-state functional connectivity43,44,45. They also have led to better predictions of task activation patterns in group analysis when compared with morphological alignment methods and have recently enabled much improved models of cortical parcellation43,46.
Illustration of intersubject alignment of brain images and unitary model of the limbic system. Blue and green rectangles represent the same functional area in two subjects, while yellow rectangles illustrate the overlap of the two individual areas after alignment. With structural alignment, there can be a complete misalignment of small functional areas (a) or partial misalignment of large functional areas (b) due to functional–anatomical variability or poor anatomical contrasts in MRI imaging. If an additional step of functional alignment is performed, an optimised overlap of functional areas is obtained. c The limbic system as originally depicted by Papez52 and (d) diagram of the unitary model of the limbic system49,50,51,165,166,167,168
To address several of these challenges that may impede proper group analysis if relying exclusively on MRI structural landmarks, we propose to revisit the anatomical scaffold of the DMN using a coregistration based on functional alignment. We hypothesised that using a functional alignment will reveal structures of basal forebrain and the Papez’s circuits, namely anterior and mediodorsal thalamic nuclei and mammillary bodies, as constituent nodes of the DMN for several reasons. First, all these regions are highly interconnected which suggest they belong to the same functional system47,48. Second, the current conceptualisation of DMN anatomy resembles the unitary model of the limbic system which, through the coordination of its subregions, subserves the elaboration of emotion, memories and behaviour49,50,51,52. Third, the basal forebrain comprises a group of neurochemically diverse nuclei, involved in dopaminergic, cholinergic and serotoninergic pathways, that are crucial in the pathophysiology of the aforementioned diseases that affect the DMN connectivity. Finally, recent electrophysiological evidence has shown that in rats the basal forebrain exhibits the same pattern of gamma oscillations than DMN and that it influences the activity of the anterior cingulate cortex53.
Therefore, in this study, we used a functional alignment of resting-state functional MRI (rs-fMRI)-based individual DMN maps to build a more comprehensive DMN model that includes the contribution of subcortical structures. To provide a complete window into the anatomy of the DMN, we explored the structural connectivity of our new model of the DMN using tractography imaging techniques and revealed that the thalamus and basal forebrain had high importance in term of values of node degree and centrality for the DMN.
Results
Comparison between structural and functional alignment
DMN connectivity maps obtained from structural and functional alignmentsare displayed in Fig. 2a–c. In both maps, classical areas of the DMN were observed, namely: posterior cingulate cortex and retrosplenial cortex; ventromedial, anteromedial and dorsal prefrontal cortex; temporal pole; middle temporal gyrus; hippocampus and parahippocampal cortex; amygdala and the posterior parietal cortex.
Maps of the DMN structurally or functionally aligned. a 3D view of the two DMN left panel corresponds to the structural space alignment, right panel to the functional space alignment, b brain sections of the structurally aligned DMN, c brain sections of the functionally aligned DMN, d subtraction of the structurally and the functionally aligned DMN maps, e statistical comparison (paired t test) between the two methods of alignment —structurally and functionally aligned DMN—in the three hypothesised regions, with colours indicating statistically significant differences at two levels of significance: <0.05 and <0.01, family-wise error (FWE) corrected p-values (higher in the functional space). DPFC dorsal prefrontal cortex, PPC posterior parietal cortex, VLPFC ventrolateral prefrontal cortex, MTG middle temporal gyrus, PCC posterior cingulate cortex, C caudate, DPFC dorsal prefrontal cortex, AMPFC antero-median prefrontal cortex, VMPFC ventro-median prefrontal cortex, TP temporal pole, BF basal forebrain, T thalamus, PH parahippocampal. n = 20 participants
Figure 2d illustrates the simple voxel-based subtraction between the DMN connectivity maps obtained from structural and functional alignments. Higher average connectivity was achieved in the functionally aligned DMN map in large areas, such as the medial prefrontal cortex and posterior cingulate cortex, mostly in the border zones. In fact, the highest differences in connectivity between structural and functional aligned DMN were at the level of the basal forebrain and thalamus. These areas were poorly or even not represented with alignment in the MNI152 space, but were visible after the functional alignment. DMN connectivity was also visible in the medial mesencephalic region, as well in inferior regions in the caudate nuclei, ventrolateral prefrontal cortex, cerebellar tonsils and cerebellar hemispheres. As expected, a difference was found between the two maps bilaterally in the thalamus and in the basal forebrain, and in a peripheral zone of the left mammillary body (Fig. 2e, Table 1). Unthresholded statistical maps of the comparison between the two methods of alignment—structurally and functionally aligned DMN—are available at neurovault.org (https://neurovault.org/collections/OCAMCQFK/). Two representative cases of individual maps in the MNI152 space and in the functional space are presented in the Supplementary Fig. 1.
Functional connectivity of the DMN in the functional space
Twenty-four regions of interest were defined based on the DMN we obtained in the functional space and concordant with the previous anatomical models of the DMN5,8. Nine additional regions were defined as being solely revealed after the functional alignment. These regions included the left and right thalamus, the left and right basal forebrain, the midbrain, the left and right ventral lateral prefrontal cortex (VLPFC) and the left and right caudate nucleus (inferior regions of the nuclei), resulting in a total of 33 regions of interest (Supplementary Fig. 2). The association strength determined by Pearson’s correlation between the rs-fMRI time series of the regions of interest (i.e., functional connectivity) were higher with alignment in the functional space, compared with structural space, in all pairs of regions (Fig. 3a). The difference was statistically significant in 18% of pairs after Bonferroni correction for multiple comparisons (p-value < 0.0001). Table 2 represents the MNI coordinates of the centres of gravity of all the regions of interest in the DMN map. The alternative correlation matrices using cortical DMN areas according to Gordon et al.54 are presented in the Supplementary Fig. 3.
Functional connectivity. a Matrices of the Pearson’s correlations between rs-fMRI time series of the regions of interest in the structural and in the functional space. b Graph representation of the partial correlations between regions of interest in the functional space (connections with partial correlation above 0.2 are depicted; darker grey tones represent stronger connections). Statistically significant partial correlations have a dashed borderline (one-sample t test; p < 0.0001, which corresponds to the Bonferroni corrected level of significance). The left side structures are not represented, for a clearer visualisation. DPFC dorsal prefrontal cortex, PPC posterior parietal cortex, VLPFC ventrolateral prefrontal cortex, Rsp retrosplenial cortex, MTG middle temporal gyrus, PCC posterior cingulate cortex, C caudate, DPFC dorsal prefrontal cortex, AMPFC antero-median prefrontal cortex, VMPFC ventro-median prefrontal cortex, TP temporal pole, BF basal forebrain, T thalamus, PH parahippocampal region, CbH cerebellar hemisphere, CbT cerebellar tonsil, Amy amygdala, MidB midbrain. n = 20 participants
Regarding the hypothesised areas, the left basal forebrain demonstrated higher correlations with the right antero-median prefrontal cortex, the right posterior parietal cortex and the midbrain area, while the right basal forebrain had higher correlations with the right temporal pole and with the left cerebellar hemisphere (Supplementary Data 1). The left thalamus had higher correlations with the left parahippocampal region, the left temporal pole, the right and the left antero-median prefrontal cortex, the left and the right ventrolateral prefrontal cortex and the right posterior parietal cortex, while the right thalamus had higher correlations with the right and the left antero-median prefrontal cortex, with the left temporal pole and with the left cerebellar tonsil (Supplementary Data 1). No significant difference was found for partial correlations. Tables presenting the median, range and interquartile range of partial correlations between all regions of interests, the statistical comparison of partial correlations between the two methods of alignment and the partial correlations of two representative cases are also shown in the Supplementary Data 2, 3, 4 and 5, respectively.
Anatomical validation in thalamic, basal forebrain and mesencephalic areas
Figure 4 illustrates the intersection of the new DMN map after its translation to the MNI group space using individual inverse transformation matrices specific for each individual. All subjects’ DMN spatially overlapped with the templates of the left anterior thalamic nucleus, mediodorsal thalamic nuclei, medial septal nuclei and left nucleus accumbens (Table 3)39,55,56,57,58. The number of subjects with an intersection with the right anterior thalamic nucleus, right nucleus accumbens and ventral tegmental area was also very high (95, 95 and 90%, respectively), while the intersection with the other basal forebrain nuclei occurred in approximately half of the subjects, possibly due to their very small size. Maps of the mean and standard deviation Pearson’s correlation of the new DMN map after registration to the MNI space are available in the Supplementary Figs. 4 and 5, respectively.
Density maps of the functionally aligned individual DMN networks superimposed in the MNI152 space. Colour bar represents the percentage of individuals with a significant correlation in each voxel. This map is freely available at https://neurovault.org/collections/CTTXXAYJ/. n = 20 participants
Tractography
We explored the structural connectivity of our new model of the DMN using tractography imaging techniques. The regions of interest were the same used for functional connectivity analysis (Supplementary Fig. 2). Figure 5 represents the structural connectivity of the network.
The results indicated that both anterior and posterior portions of the cingulum, as well as inferior longitudinal fasciculus, the second branch of the superior longitudinal fasciculus, the posterior segment of arcuate fasciculus, the uncinate fasciculus and some fibres of the frontal orbito-polar tract (Fig. 5, upper panel) connected the different nodes of the DMN. In addition, the anatomical connectivity of the basal forebrain and the thalamus with other regions of interest included: the anterior thalamic projections, connecting thalamus with medial prefrontal cortex; the cingulum, connecting basal forebrain with medial prefrontal cortex and posterior cingulate cortex; the fornix, connecting basal forebrain (specifically the region correspondent to the medial septal nuclei) to the hippocampus and fibres connecting basal forebrain and thalamus, some of the most medial possibly corresponding to the bundle of Vicq D’Azyr (Fig. 5, lower panel). Statistical maps of the tractography analysis are presented in the Supplementary Fig. 6.
Graph theory analysis
Figure 6 represents the analysis of the DMN structural network with a graph theory approach using the 33 regions of interest defined from the DMN in the functional space.
Graph theory analysis of structural connectivity. The node size represents node degree and the node colour illustrates node betweenness centrality. The edges denote presence of structural connection. DPFC dorsal prefrontal cortex, PPC posterior parietal cortex, VLPFC ventrolateral prefrontal cortex, Rsp retrosplenial cortex, MTG middle temporal gyrus, PCC posterior cingulate cortex, C caudate, DPFC dorsal prefrontal cortex, AMPFC antero-median prefrontal cortex, VMPFC ventro-median prefrontal cortex, TP temporal pole, BF basal forebrain, T thalamus, PH parahippocampal region, CbH cerebellar hemisphere, CbT cerebellar tonsil, Amy amygdala, MidB midbrain. n = 20 participants
The results indicate that high degrees and high betweenness centrality in the network were obtained for the basal forebrain and thalamic regions, alongside the medial prefrontal cortex and the posterior cingulate–retrosplenial cortex, as well as in regions which were previously considered as hubs in the DMN.
More precisely, the [maximum–minimum] range of distribution of node degrees was [12–1] and the median [interquartile range] was 5 [8–2]. The node degrees of the left and right thalamus were 9 and 7, and of the left and right basal forebrain were 8 and 7, respectively (Supplementary Table 1). Therefore, thalamus and basal forebrain are among the structures in the network that have connections with a high number of nodes. For betweenness centrality, the [maximum–minimum] range of distribution was [0.104–0] and the median [interquartile range] was 0.004 [0.03–0.001]. The betweenness centrality of the left and right thalamus was 0.03, and of the left and right basal forebrain were 0.03 and 0.02, respectively (Supplementary Table 1). Hence, thalamus and basal forebrain make part of a high fraction of shortest paths in the network, that is, the shortest connections between two nodes.
A table showing the values for node degree and betweenness centrality at the individual level in two representative cases is shown in the Supplementary Table 2.
Discussion
In this study, we revisited the constituent elements of the DMN using an optimised method of coregistration in a functional space, besides the conventional structural alignment. Three main findings emerge from this research in healthy humans. First, higher functional connectivity correlation and sharper anatomical details were achieved when registering the DMN maps in a functional space. Second, we confirmed the hypothesis that structures of basal forebrain and anterior and mediodorsal thalamic nuclei belong to the DMN. Lastly, we characterised in detail the structural connectivity underlying functional connectivity. Based on these findings, we provided a more comprehensive neurobiological model of the DMN that bridges the gap between local differences in subcortical structures and global differences in the DMN reported in clinical studies.
The difference between alignment in the functional space and the structural space was characterised by an increase in the connectivity strength across the brain, as well as in many subcortical areas, classically not considered to be constituent nodes of the DMN. As previously reported, this confirmed that registration in the functional space provides a more accurate interindividual anatomical description and is recommended when doing functional connectivity analyses43,44.
The maps of the DMN registered in the functional space revealed previously underappreciated parts of this network, such as basal forebrain and anterior and mediodorsal thalamic nuclei. Tractography analysis yielded structural connectivity of these new DMN regions to the other regions of the network. Mainly, the cingulum connected the basal forebrain with the medial prefrontal cortex, posterior cingulate region, retrosplenial cortex and hippocampus and parahippocampal regions59,60. The fornix linked basal forebrain, specifically the medial septal nuclei, with the hippocampus and parahippocampal regions49,61,62. The anterior thalamic projections connected the thalamus with medial and ventrolateral prefrontal regions63 and finally some of the most medial fibres connecting the basal forebrain with the thalamus, probably corresponded to the mammillothalamic tract of Vicq D’Azyr64,65. The graph theory approach66 applied to the measures of structural connectivity revealed a high node degree of the basal forebrain and the thalamus in the network, as well as high betweenness centrality67. These results indicate that basal forebrain and thalamus have high centrality within the network, and therefore can have an important role for network integration and resilience66,67,68, along with the classically defined hubs, such as the medial prefrontal region as well as posterior cingulate and retrosplenial region69.
The involvement of basal forebrain and anterior and mediodorsal thalamus in the DMN has theoretical and functional repercussions beyond the purely anatomical level. The involvement of the anterior and mediodorsal thalamic nuclei as well as the basal forebrain are concordant with the role of the DMN in memory processes8,70, as all these regions are relays of the unitary model of the limbic system49,50,51. Previous reports of engagement of the mediodorsal thalamic nucleus and the DMN during memory tasks and the memory deficits provoked by lesions of the anterior and the mediodorsal thalamic nuclei also support this claim71,72,73,74. At a neurochemical level, the basal forebrain is also a principal actor in the production of acetylcholine75. Acetylcholine has a physiological and a neuropharmacological effect on memory processes. For instance, cholinergic system mediates rhythmic oscillation in the hippocampus that facilitates encoding39,76. The basal forebrain also contains GABAergic and glutamatergic neurons that mediate hippocampal theta synchronisation through an indirect septo-hippocampal pathway77. By providing evidence of the involvement of medial septal cholinergic nucleus and its structural connection to the hippocampus in our DMN model, the present work indicates a match between connectivity, neurochemistry and cognition.
The same correspondence between connectivity, neurochemistry and cognition applies to the relation between DMN and emotional modulation78,79,80,81,82,83. The nucleus accumbens is a central output for the dopaminergic projections and is involved in emotion regulation and affect integration84,85. The nucleus accumbens also receives glutamatergic inputs from the hippocampus and the prefrontal cortex86 belonging to the DMN. Surprisingly, our analysis also revealed the ventral tegmental area, which is also a dopaminergic nucleus with projections to the nucleus accumbens and the medial prefrontal cortex87. This association with the mesolimbic dopaminergic pathway is reinforced by our results of functional connectivity, since ventromedial prefrontal cortex and midbrain were among the structures with highest partial correlations with basal forebrain. Hence, combining present and previous findings, we speculate that the DMN, as defined by functional connectivity, might have a putative role in the integration of cholinergic and dopaminergic systems dedicated to memory and emotion.
The new DMN’s subcortical structures identified in the current work have cognitive and neurochemical roles that open a new window to the understanding of distinct brain pathologies affecting DMN connectivity (Table 4). Indeed, functional connectivity in each area of the DMN is an estimate of the global coherence of the DMN. Since we demonstrated that limbic thalamus and basal forebrain are nodes with high degree and high centrality in the DMN, damage in these structures should lead to a drastic decrease of functional connectivity in the whole DMN88. For instance, Alzheimer’s disease is associated with degeneration of the cholinergic system, including the medial septal nuclei, even in the earliest clinical stages of Mild Cognitive Impairment89 and apparently related to decreased functional coherence and deactivation in hub regions of DMN90,91. The high centrality of the basal forebrain in the DMN network may explain this early link between DMN and Alzheimer’s disease. In schizophrenia, also associated with decreased DMN connectivity and activation92,93, neuropathological evidence suggests an abnormal glutamatergic–dopaminergic interaction at the level of nucleus accumbens94. In addition, the ventral tegmental area is connected to the nucleus accumbens through the mesolimbic system, the classical dopaminergic pathway associated with schizophrenia, and functional data show a decrease of connectivity between VTA and several brain regions, including the thalamus, in unmedicated schizophrenic patients95. The pathophysiology of others diseases, such as drug addiction, depression, temporal lobe epilepsy and attention-deficit and hyperactivity disorder involve modifications in the nucleus accumbens, medial septal nuclei or the thalamic nuclei that connect limbic regions as well as dysfunctional connectivity of the DMN (specified in Table 427,96,97,98,99,100,101,102,103). Hence, the involvement of the basal forebrain and the thalamic nuclei in the DMN appears to bridge the gap between the subcortical anatomical differences and the global differences in the DMN previously reported.
Our results is also concordant with recent findings. For instance, Kernbach et al. recently demonstrated that grey matter variability across the DMN can well predict population variation in the microstructural properties of core white matter tracts of the limbic system—the anterior thalamic radiation and the fornix in 10,000 people9. As another example, Margulies et al. studied functional gradients along cortical surface and found that DMN areas were at the opposite end of primary motor/sensory areas in a spectrum of connectivity differentiation and that DMN areas exhibit the most considerable geodesic distance at the cortical level, being equidistant to the unimodal cortical areas11. These investigators suggested that the DMN acts as a neural relay for transmodal information. We speculate that the thalamus and basal forebrain may follow the same model at a subcortical level, integrating functional networks related to primary functions and brainstem inputs to the associative areas104,105.
Interestingly, the thalamus and the basal forebrain are phylogenetically older than many cortical structures, and especially those that compose the DMN106,107,108. The inclusion of these structures in the anatomical model of the DMN can open a window to the exploration of DMN in other mammalian species as well109,110,111,112. The medial thalamus has already been shown to be part of the mouse’s DMN113, as found in rs-fMRI studies114,115,116. Our results are also concordant with recent neurophysiological evidence in rats about the influence of basal forebrain in the regulation of the DMN53. Namely, it has been shown that gamma-band local field potentials in the basal forebrain exerts influence on one of the hub regions of rat’s DMN, the anterior cingulate cortex53. The cellular basis for the association of these two areas has been characterised with retrograde tracing studies117,118. In humans, it has been demonstrated that basal forebrain has a major functional coupling with the anterior cingulate cortex and with ventromedial prefrontal cortex119. We speculate that the putative differences in the basal forebrain projections may be one explanation for the more diffuse DMN activation at the midline level in rats when compared with humans112.
One limitation of this work is that the overlap between the DMN map and the nuclei studied is based on comparison with templates or variability maps, and not with the individual location of the nucleus in the explored subjects. The limited capacity of structural MRI to differentiate these small nuclei does not allow such comparison. Besides, the reverse transformation of the DMN from the functional space to the MNI space may not be exact, due to inherent limitations of inverse transformations. Although these limitations may decrease the accuracy of the intersection quantification with discrete nuclei, they did not alter the apparent overlap with basal forebrain and with the thalamus. Furthermore, demonstrating that the additional step of functional alignment results in higher correlation values is somewhat circular. However, our purpose was to demonstrate that this increase in correlation revealed subcortical structures that were previously neglected in the literature. The intra-individual functional connectivity variation across time may also be a source of bias35. However, this factor and the lack of uniformity regarding the optimal functional alignment method have not precluded the achievement of higher accurate results in previous studies43,44,45,46. Finally, tractography analyses can produce inaccurate results120,121. In order to avoid these caveats, we employed methods that have previously demonstrated high anatomical reliability when compared with axonal tracing and with post-mortem dissections59,122,123,124.
In conclusion, this work demonstrates that the registration of individual DMN maps in a functional space improves the definition of the anatomy of DMN by including additional structures, such as the thalamus and basal forebrain. Future research should focus on the cascade of neurochemical and pathophysiological events that follow small subcortical lesions of the DMN.
Methods
The local ethics committee (Comité de Protection des Personnes “CPP Ile de France V”) approved the experiment, and all participants provided written informed consent.
Subjects and MRI acquisition
MRI images of subjects without neurological or psychiatric disease were obtained (age mean ± SD 29 ± 6 years, range 22–42 years; 11 females, 9 males) with a Siemens 3 Tesla Prisma system equipped with a 64-channel head coil.
An axial 3D T1-weighted imaging data set covering the whole head was acquired for each participant (286 slices, voxel resolution = 0.7 mm3, echo time (TE) = 2.17 ms, repetition time (TR) = 2400 ms, flip angle = 9°).
rs-fMRI images were obtained using T2*-weighted echo-planar imaging (EPI) with blood oxygenation level-dependent (BOLD) contrast. EPIs (TR/TE = 2050/25 ms) comprised 42 axial slices acquired with a multiband pulse125,126,127,128 covering the entire cerebrum (voxel size = 3 mm3) including 290 brain scan volumes in one run of 10 min.
A diffusion-weighted imaging (DWI) acquisition sequence, fully optimised for tractography, provided isotropic (1.7 × 1.7 × 1.7 mm) resolution and coverage of the whole head with a posterior–anterior phase of acquisition, with an echo time (TE) = 75 ms. A repetition time (TR) equivalent to 3500 ms was used. At each slice location, six images were acquired with no diffusion gradient applied (b-value of 0 s mm−2). In addition, 60 diffusion-weighted images were acquired, in which gradient directions were uniformly distributed on the hemisphere with electrostatic repulsion. The diffusion weighting was equal to a b-value of 2000 s mm−2. This sequence was fully repeated with reversed phase-encode blips. This provides us with two data sets with distortions going in opposite directions. From these pairs, the susceptibility-induced off-resonance field was estimated using a method similar to that described in ref. 129 and corrected on the whole diffusion-weighted data set using the tool TOPUP and EDDY as implemented in FSL130.
rs-fMRI analysis—overview
rs-fMRI images were corrected for artefacts in the Funcon-Preprocessing tool of the Brain Connectivity and Behaviour toolkit (http://toolkit.bcblab.com131,132,133). Then, they were registered to T1 high-resolution individual structural images and normalised to MNI152 standard space using Advanced Normalization Tools (ANTS; http://stnava.github.io/ANTs;134,135). Preprocessing steps in details:
-
a.
fMRI images were first motion corrected using MCFLIRT132, then corrected for slice timing, smoothed with a full half-width maximum equal to 1.5 times the largest voxel dimension and finally filtered for low temporal frequencies using a Gaussian-weighted local fit to a straight line. These steps are available in FEAT, as part of FSL package133.
-
b.
fMRI images were linearly registered to the T1 images, and subsequently to the MNI152 template (2 mm) using affine and diffeomorphic transformations136,137. Confounding signals were discarded from fMRI by regressing out a confound matrix from the functional data. The confound matrix included the estimated motion parameters obtained from the previously performed motion correction, the first eigenvariate of the white matter and cerebrospinal fluid, as well as their first derivative. Eigenvariates were extracted using fslmeants combined with the–eig option. White matter and cerebrospinal fluid eigenvariates were extracted using masks based on the T1-derived 3-classes segmentation thresholded to a probability value of 0.9, registered to the rs-fMRI images and binarised. Finally, the first derivative of the motion parameters, white matter and cerebrospinal fluid signal were calculated by linear convolution between their time course and a [−1 0 1] vector.
-
c.
Since the rs-fMRI signal can be heavily affected by motion, even following motion correction between temporally adjacent volumes138, we estimated the signal fluctuation associated with motion and regressed it out from the fMRI. To this aim, we employed a recently developed and validated procedure based on data-driven Independent Component Analysis (ICA), termed ICA-Aroma139. This method performs an ICA decomposition of the data and estimates which components reflect motion-related noise in the fMRI signal on the basis of a robust set of spatial and temporal features. This is made possible due the distinctiveness of the motion-related components isolated by ICA on the fMRI signal140. This approach outperforms other methods such as the regression of the motion parameter estimates, while limiting in the same time the loss in degrees of freedom139. Compared with spike removal methods such as scrubbing141, ICA-Aroma has the advantage of preserving the temporal structure of the fMRI signal.
Individual DMN maps in the structural space
Individual subject-tailored/fitted DMN maps were obtained by correlation with seed regions of interest of a functional parcellated brain template. The regions used for the seed-based functional connectivity analysis were those defined as DMN regions in the resting-state parcellation map by Gordon et al.54,142,143. Gordon et al. created these parcellations according to abrupt changes in resting-state’s time course profile, each parcel having a homogenous time course profile. This general-purpose atlas provides a total of 40 DMN nodes (20 in each hemisphere). The DMN nodes as defined by Gordon et al. are available in the Supplementary Fig. 7 and Supplementary Table 3, respectively). These 40 regions were used as seeds. The correlation map for each seed was obtained using the Funcon-Connectivity tool of the Brain Connectivity and Behaviour toolkit (http://toolkit.bcblab.com;131). This tool calculates the Pearson’s correlation between the mean rs-fMRI time course of the seed with the rest of the brain to generate the functional connectivity map for every seed131.
The DMN map of each participant was obtained by calculating the median of the 40 seed-based correlation maps of each individual using FSL144. We then calculated the median across the DMN maps of the 20 participants to obtain the median DMN map in the MNI152 space. The median was used, instead of the mean, because it is less sensitive to outliers, being a better centrality measure for small sample sizes145.
Individual DMN maps in the functional space
To achieve the proposed optimised map of the DMN, the same 20 individual maps were functionally aligned with each other in a new functional space using the following steps:
Individual DMN maps were aligned with each other using ANTs’ script buildtemplateparallel.sh, defining cross-correlation as the similarity measure and greedy SyN as the transformation model136,137,146. This approach consists in an iterative (n = 4) diffeomorphic transformation to a common space. The group map was obtained by calculating the median of all DMN maps after functional alignment.
The resulting maps correspond to alignment of the 20 individual DMN maps in a functional space (Fig. 2).
Functional connectivity comparison of the two DMN
We applied a 0.3 threshold on the functional connectivity strength for both the functional and structural-based DMN maps. This value was chosen because it corresponds to a medium effect size147. Time series of rs-fMRI of each individual in the different regions of interest identified in the DMN maps were extracted and the mean value of the voxels of each region was obtained, using the command fslstats (with the options –k and –M)144. Cortical regions of interest were defined according to the previous anatomical models of the DMN5,8. Subcortical regions of interest were defined manually based on the experienced judgement of the neuroanatomists among the authors combined with the careful comparison with previously published atlases148,149. Correlation and partial correlation coefficients were determined as measures of functional connectivity.
Two matrices, representing the median correlation values in the MNI152 space and in the functional space, were created using BrainNet Viewer150. A paired t test was calculated for each cell of the two connectivity matrices using Python’s Scipy package, version 0.19.1 (https://www.scipy.org; scipy.stats.ttest_rel). A p-value corrected for multiple comparisons at a threshold of <0.0001 (Bonferroni correction) was used. Circos software was used to illustrate functional connections in the functional space (http://circos.ca;151).
Two alternative correlation matrices using cortical DMN areas according to Gordon et al.54, instead of the ones defined in classical anatomical models of Andrews-Hanna et al.8 and Buckner et al.5, were calculated.
Anatomical validation in thalamic, basal forebrain and mesencephalic areas
Meynert nuclei, medial septal nuclei and diagonal band of Broca probabilistic maps were derived from the work of Zaborszky et al., the Harvard-Oxford probabilistic atlas was employed for the nucleus accumbens, Talairach atlas registered to the MNI152 space for mammillary bodies and thalamic nuclei, and Harvard Ascending Arousal Network Atlas for ventral tegmental area39,55,56,57,58. A percentage of volume overlap between the DMN map and each nucleus of interest was subsequently calculated for each subject. The same Pearson’s correlation threshold of 0.3 was applied for this analysis, as specified in 2.2.3.
Tractography analysis
Diffusion-Weighted Images were corrected for signal drift152, motion and eddy current artefacts using ExploreDTI (http://www.exploredti.com;153).
Whole-brain tractography was performed on the software StarTrack using a deterministic approach (https://www.mr-startrack.com). A damped Richardson–Lucy algorithm was applied for spherical deconvolutions154. A fixed fibre response corresponding to a shape factor of α = 1.5 × 10.3 mm2/s was adopted. The defined number of iterations was 150, and the geometric damping parameter was 8. The absolute threshold was defined as three times the spherical fibre orientation distribution (FOD) of a grey matter isotropic voxel and the relative threshold as 8% of the maximum amplitude of the FOD155. A modified Euler algorithm was used123. An angle threshold of 35°, a step size of 0.85 mm and a minimum length of 20 mm were chosen.
Diffusion tensor images were registered to the MNI152 standard space and, then, into the functional space applying the affine and diffeomorphic deformation generated in the previous sections, using the tool tractmath as part of the software package Tract Querier156.
The same regions of interest used for functional connectivity calculation (section 2.2.3) were also used for the tractography analysis.
The command tckedit of MRtrix toolbox (http://www.mrtrix.org/157) was used to extract the tracts of interest. The –include arguments were composed by combinations of DMN regions of interest to guarantee that the selected streamlines crossed at least two DMN regions of interest.
In order to have a group-representative map for each tract, individual tracts were converted into maps using tckmap command of MRtrix toolbox (http://www.mrtrix.org;158). Each voxel was binarised in 1 or 0 according to being or not being intersected by a streamline159. Lastly, a one-sample t test was calculated using FSL randomise, with variance smoothing of 4 mm159. The resulting group-representative tract maps using this method were shown to have a good anatomical correspondence with histological atlas of white matter tracts159,160. BrainVisa was used to create the corresponding illustration of the tracts that reached significance161.
Graph theory analysis of structural connectivity
To explore whether the new putative regions of the DMN are essential structures and potential areas of vulnerability in the network, we investigated the hub properties of the network nodes using graph theory measures, namely node degree and betweenness centrality66. Node degree refers to the number of connections between a given node and the other nodes of the network. Betweenness centrality is the fraction of all shortest paths in the network that pass through a given node66.
For each participant, an anatomical connectome matrix of the DMN was built using the tck2connectome command of MRtrix (http://www.mrtrix.org;157). Each region of interest was defined as a node. Only the streamlines that ended in both regions of interest were considered162,163. The matrices were binarised depending on the existence or absence of streamlines connecting two regions of interest162,163. The defined threshold for the binarisation was 1 because the number of streamlines does not reflect the connectivity strength or the true number of axonal projections between two brain regions163,164, and previous evidence has shown that changing the streamline count threshold for binarisation (between 1 and 5) does not change the overall results of the network analysis. Brain Connectivity Toolbox for Python (https://pypi.python.org/pypi/bctpy) was used to obtain the network measures. The functions degrees_und and betweenness_bin were run, respectively, for each individual matrix67. The median values for each measure were obtained. The illustration of the network was made using Surf Ice (https://www.nitrc.org/projects/surfice/).
Statistics and reproducibility
In order to assess reproducibility, the effects of interest were also measured at the individual level (i.e., replicated for every subject).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The raw data used for this research are available on demand from the corresponding authors pedroascimentoalves@gmail.com and michel.thiebaut@gmail.com. The data for the statistical comparisons presented in the paper are available in Supplementary Data 1–5. Unthresholded statistical maps of the comparison between the two methods of alignment—structurally and functionally aligned DMN—are available at neurovault.org (https://neurovault.org/collections/OCAMCQFK/).
References
Ingvar, D. H. “Hyperfrontal” distribution of the cerebral grey matter flow in resting wakefulness; on the functional anatomy of the conscious state. Acta Neurol. Scand. 60, 12–25 (1979).
Mazoyer, B. et al. Cortical networks for working memory and executive functions sustain the conscious resting state in man. Brain Res Bull. 54, 287–298 (2001).
Shulman, G. et al. Top-down modulation of early sensory cortex. Cereb. cortex 7, 193–206 (1997a).
Raichle, M. E. et al. A default mode of brain function. Proc. Natl Acad. Sci. USA 98, 676–682 (2001).
Buckner, R. L., Andrews-Hanna, J. R. & Schacter, D. L. The brain’s default network: anatomy, function, and relevance to disease. Ann. N. Y. Acad. Sci. 1124, 1–38 (2008).
Greicius, M. D., Srivastava, G., Reiss, A. L. & Menon, V. Default-mode network activity distinguishes Alzheimer’ s disease from healthy aging: evidence from functional MRI. Proc. Natl Acad. Sci. USA 101, 4637–4642 (2004).
Vincent, J. L. et al. Coherent spontaneous activity identifies a hippocampal-parietal memory network. J. Neurophysiol. 96, 3517–3531 (2006).
Andrews-Hanna, J. R., Reidler, J. S., Sepulcre, J., Poulin, R. & Buckner, R. L. Functional-anatomic fractionation of the brain’s default network. Neuron 65, 550–562 (2010).
Kernbach, J. M. et al. Subspecialization within default mode nodes characterized in 10, 000 UK Biobank participants. Proc. Natl Acad. Sci. USA 115, 12295–12300 (2018).
Lopez-Persem, A., Verhagen, L., Amiez, C., Petrides, M. & Sallet, J. The human ventro medial prefrontal cortex sulcal morphology and its influence on its functional organization. bioRxiv 1–36, https://doi.org/10.1101/417824 (2018).
Margulies, D. S. et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc. Natl Acad. Sci. USA 113, 12574–12579 (2016).
Buckner, R. L., Krienen, F. M., Castellanos, A., Diaz, J. C. & Yeo, B. T. T. The organization of the human cerebellum estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 2322–2345 (2011).
Stoodley, C. J. & Schmahmann, J. D. NeuroImage Functional topography in the human cerebellum: a meta-analysis of neuroimaging studies. Neuroimage 44, 489–501 (2009).
Choi, E. Y., Yeo, B. T. T. & Buckner, R. L. The organization of the human striatum estimated by intrinsic functional connectivity. J. Neurophysiol. 108, 2242–2263 (2012).
Bzdok, D., Laird, A. R., Zilles, K. & Fox, P. T. An investigation of the structural, connectional, and functional subspecialization in the human amygdala. Hum. Brain Mapp. 34, 3247–3266 (2012).
Roy, A. K. et al. Functional connectivity of the human amygdala using resting state fMRI. Neuroimage 45, 614–626 (2009).
Di Martino, A. et al. Functional connectivity of human striatum: a resting state fMRI study. Cereb. Cortex 18, 2735–2747 (2008).
Cunningham, S. I., Tomasi, D. & Volkow, N. D. Structural and functional connectivity of the precuneus and thalamus to the default mode network. Hum. Brain Mapp. 38, 938–956 (2016).
Fransson, P. Spontaneous low-frequency BOLD signal fluctuations: an fMRI investigation of the resting-state default mode of brain function hypothesis. Hum. Brain Mapp. 26, 15–29 (2005).
Shapira-Lichter, I., Oren, N., Jacob, Y., Gruberger, M. & Hendler, T. Portraying the unique contribution of the default mode network to internally driven mnemonic processes. Proc. Natl Acad. Sci. USA 110, 4950–4955 (2013).
Shulman, G. et al. Common blood flow changes across visual tasks: II. Decreases in cerebral cortex. J. Cogn. Neurosci. 9, 648–663 (1997b).
Bendetowicz, D. et al. Two critical brain networks for generation and combination of remote associations. Brain 141, 217–233 (2018).
Spreng, R. N., Mar, R. A. & Kim, A. S. N. The Common neural basis of autobiographical memory, prospection, navigation, theory of mind, and the default mode: a quantitative meta-analysis. J. Cogn. Neurosci. 21, 489–510 (2009).
Broyd, S. J. et al. Default-mode brain dysfunction in mental disorders: a systematic review. Neurosci. Biobehav. Rev. 33, 279–296 (2009).
Geng, X. et al. Salience and default mode network dysregulation in chronic cocaine users predict treatment outcome. Brain 140, 1513–1524 (2017).
Tessitore, A. et al. Default-mode network connectivity in cognitively unimpaired patients with Parkinson disease. Neurology 79, 2226–2232 (2012).
Voets, N. L. et al. Structural substrates for resting network disruption in temporal lobe epilepsy. Brain 135, 2350–2357 (2012).
Whitfield-Gabrieli, S. & Ford, J. M. Default mode network activity and connectivity in psychopathology. Annu Rev. Clin. Psychol. 8, 49–76 (2012).
Zhu, X., Zhu, Q., Shen, H., Liao, W. & Yuan, F. Rumination and default mode network subsystems connectivity in first-episode, drug-naive young patients with major depressive disorder. Sci. Rep. 7, 43105 (2017).
Brett, M., Johnsrude, I. S. & Owen, A. M. The problem of functional localization in the human brain. Nat. Rev. Neurosci. 3, 243–249 (2002).
Thiebaut de Schotten, M. & Shallice, T. Identical, similar or different? Is a single brain model sufficient? Cortex 86, 172–175 (2017).
Despotovic, I., Goossens, B. & Philips, W. MRI segmentation of the human brain: challenges, methods, and applications. Comput Math. Methods Med. 2015, 1–23 (2015).
Bzdok, D. et al. Subspecialization in the human posterior medial cortex. Neuroimage 106, 55–71 (2015).
Eickhoff, S. B., Laird, A. R., Fox, P. T., Bzdok, D. & Hensel, L. Functional segregation of the human dorsomedial prefrontal cortex. Cereb. Cortex 26, 304–321 (2016).
Braga, R. M. & Buckner, R. L. Parallel interdigitated distributed networks within the individual estimated by intrinsic functional connectivity. Neuron 95, 457–471.e5 (2017).
Amunts, K. et al. Cytoarchitectonic mapping of the human amygdala, hippocampal region and entorhinal cortex: Intersubject variability and probability maps. Anat. Embryol. (Berl.). 210, 343–352 (2005).
Amunts, K. et al. Broca’s region revisited: cytoarchitecture and intersubject variability. J. Comp. Neurol. 412, 319–341 (1999).
Croxson, P.L., Forkel, S.J., Cerliani, L. & Thiebaut de Schotten, M. Structural variability across the primate brain: a cross-species comparison. Cereb. Cortex 1–13. https://doi.org/10.1093/cercor/bhx244 (2017).
Zaborszky, L. et al. Stereotaxic probabilistic maps of the magnocellular cell groups in human basal forebrain. Neuroimage 42, 1127–1141 (2008).
Liu, A. K. L., Chang, R. C. C., Pearce, R. K. B. & Gentleman, S. M. Nucleus basalis of Meynert revisited: anatomy, history and differential involvement in Alzheimer’s and Parkinson’s disease. Acta Neuropathol. 129, 527–540 (2015).
Möttönen, T. et al. Defining the anterior nucleus of the thalamus (ANT) as a deep brain stimulation target in refractory epilepsy: delineation using 3 T MRI and intraoperative microelectrode recording. NeuroImage Clin. 7, 823–829 (2015).
Tagliamonte, M., Sestieri, C., Romani, G. L., Gallucci, M. & Caulo, M. MRI anatomical variants of mammillary bodies. Brain Struct. Funct. 220, 85–90 (2013).
Langs, G., Golland, P. & Ghosh, S. Predicting activation across individuals with resting-state functional connectivity based multi-atlas label fusion. Med. Image Comput. Comput. Assist. Inter. 9350, 313–320 (2015).
Mueller, S. et al. Individual variability in functional connectivity architecture of the human brain. Neuron 77, 586–595 (2013).
Robinson, E. C. et al. MSM: a new flexible framework for multimodal surface matching. Neuroimage 100, 414–426 (2014).
Glasser, M. F. et al. A multi-modal parcellation of human cerebral cortex. Nature 536, 171–178 (2016).
Yakovlev, P. I. Motility, behaviour and the brain: stereodynamic organization and neural correlates of behaviour. J. Nerve Men. Dis. 107, 313–335 (1948).
Yakovlev, P. I. & Locke, S. Limbic nuclei of thalamus and connections of limbic cortex III. Corticocortical connections of the anterior cingulate gyrus, the cingulum, and the subcallosal bundle in monkey. Arch. Neurol. 5, 364–400 (1961).
Catani, M., Dell’Acqua, F. & Thiebaut de Schotten, M. A revised limbic system model for memory, emotion and behaviour. Neurosci. Biobehav. Rev. 37, 1724–1737 (2013).
MacLean, P. Some psychiatric implications of physiological studies on frontotemporal portion of limbic system (Visceral brain). Electroencephalogr. Clin. Neurophysiol. 4, 407–418 (1952).
MacLean, P. Psychosomatic disease and the “visceral brain.” Recent developments bearing on the Papez theory of emotion. Psychosom. Med. 11, 338–353 (1949).
Papez, J. A proposed mechanism of emotion. Arch. Neurol. Psychiatry 258, 725–743 (1937).
Nair, J. et al. Basal forebrain contributes to default mode network regulation. Proc. Natl Acad. Sci. USA 2018, 201712431 (2018).
Gordon, E. M. et al. Generation and evaluation of a cortical area parcellation from resting-state correlations. Cereb. Cortex 26, 288–303 (2016).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Edlow, B. L. et al. Neuroanatomic connectivity of the human ascending arousal system critical to consciousness and its disorders. J. Neuropathol. Exp. Neurol. 71, 531–546 (2012).
Lancaster, J. L. et al. Bias between MNI and talairach coordinates analyzed using the ICBM-152 brain template. Hum. Brain Mapp. 28, 1194–1205 (2007).
Talairach, J. & Tournoux, P. Co-planar Stereotaxic Atlas of the Human Brain. (Thieme, New York, 1988).
Catani, M. et al. Short frontal lobe connections of the human brain. Cortex 48, 273–291 (2012).
Catani, M., Howard, R. J., Pajevic, S. & Jones, D. K. Virtual in vivo interactive dissection of white matter fasciculi in the human brain. Neuroimage 17, 77–94 (2002).
Aggleton, J. The Amygdala: A Functional Analysis. 2nd edn (Oxford University Press, New York, 2000).
Saunders, R. & Aggleton, J. Origin and topography of fibers contributing to the fornix in macaque monkeys. Hippocampus 17, 396–411 (2007).
Behrens, T. E. J. et al. Non-invasive mapping of connections between human thalamus and cortex using diffusion imaging. Nat. Neurosci. 6, 750–757 (2003).
Balak, N. et al. Mammillothalamic and mammillotegmental tracts as new targets for dementia and epilepsy treatment. World Neurosurg. 110, 133–144 (2018).
Vicq D’Azyr, F. Traité d’anatomie et de physiologie, avec des planches coloriées représentant au naturel les divers organes de l’Homme et des Animaux. (François-Ambroise Didot, Paris, 1786).
Bullmore, E. & Sporns, O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009).
Rubinov, M. & Sporns, O. Complex network measures of brain connectivity: uses and interpretations. Neuroimage 52, 1059–1069 (2010).
Hagmann, P. et al. Mapping the structural core of human cerebral cortex. PLoS Biol. 6, 1479–1493 (2008).
van Oort, E. S. B., van Cappellen van Walsum, A. M. & Norris, D. G. An investigation into the functional and structural connectivity of the Default Mode Network. Neuroimage 90, 381–389 (2014).
Schacter, D. L., Addis, D. R. & Buckner, R. L. Remembering the past to imagine the future: the prospective brain. Nat. Rev. Neurosci. 8, 657–661 (2007).
Rabin, J. S., Gilboa, A., Stuss, D. T., Mar, R. A. & Rosenbaum, R. S. Common and unique neural correlates of autobiographical memory and theory of mind. J. Cogn. Neurosci. 22, 1095–1111 (2010).
Spreng, R. N. & Grady, C. L. Patterns of brain activity supporting autobiographical memory, prospection, and theory of mind, and their relationship to the default mode network. J. Cogn. Neurosci. 22, 1112–1123 (2010).
Child, N. D. & Benarroch, E. E. Anterior nucleus of the thalamus: functional organization and clinical implications. Neurology 81, 1869–1876 (2013).
Danet, L. et al. Thalamic amnesia after infarct: the role of the mammillothalamic tract and mediodorsal nucleus. Neurology 85, 2107–2115 (2015).
Zaborszky, L. et al. Neurons in the basal forebrain project to the cortex in a complex topographic organization that reflects corticocortical connectivity patterns: an experimental study based on retrograde tracing and 3D reconstruction. Cereb. Cortex 25, 118–137 (2015).
Hasselmo, M. E. The role of acetylcholine in learning and memory. Curr. Opin. Neurobiol. 16, 710–715 (2006).
Dannenberg, X. H. et al. Synergy of direct and indirect cholinergic septo-hippocampal pathways coordinates firing in hippocampal networks. J. Neurosci. 35, 8394–8410 (2015).
Raichle, M. E. The brain’s default mode network. Annu. Rev. Neurosci. 38, 433–447 (2015).
Spies, M. et al. Default mode network deactivation during emotion processing predicts early antidepressant response. Transl. Psychiatry 7, e1008 (2017).
Zhao, J., Tomasi, D., Wiers, C.E., Shokri-kojori, E. & Şükrü, B. Correlation between traits of emotion-based impulsivity and intrinsic default-mode network activity. Neural Plast., https://doi.org/10.1155/2017/9297621 (2017).
Bzdok, D. et al. Segregation of the human medial prefrontal cortex in social cognition. Front. Hum. Neurosci. 7, 1–17 (2013).
Alcalá-López, D. et al. Computing the social brain connectome across systems and states. Cereb. Cortex 1–26. https://doi.org/10.1093/cercor/bhx121 (2017).
Mars, R. B. et al. On the relationship between the “default mode network” and the “social brain. Front. Hum. Neurosci. 6, 1–9 (2012).
Floresco, S. B. The nucleus accumbens: an interface between cognition, emotion, and action. Annu. Rev. Psychol. 66, 25–52 (2015).
Laviolette, S. R. Dopamine modulation of emotional processing in cortical and subcortical neural circuits: Evidence for a final common pathway in schizophrenia? Schizophr. Bull. 33, 971–981 (2007).
Britt, J. P. et al. Synaptic and behavioral profile of multiple glutamatergic inputs to the nucleus accumbens. Neuron 76, 790–803 (2012).
Morales, M. & Margolis, E. B. Ventral tegmental area: cellular heterogeneity, connectivity and behaviour. Nat. Rev. Neurosci. 18, 73–85 (2017).
van den Heuvel, M. P. & Sporns, O. Network hubs in the human brain. Trends Cogn. Sci. 17, 683–696 (2013).
Grothe, M. et al. Reduction of basal forebrain cholinergic system parallels cognitive impairment in patients at high risk of developing alzheimer’s disease. Cereb. Cortex 20, 1685–1695 (2010).
Persson, J. et al. Altered deactivation in individuals with genetic risk for Alzheimer’s disease. Neuropsychologia 46, 1679–1687 (2008).
He, Y. et al. Regional coherence changes in the early stages of Alzheimer’s disease: a combined structural and resting-state functional MRI study. Neuroimage 35, 488–500 (2007).
Pomarol-Clotet, E. et al. Failure to deactivate in the prefrontal cortex in schizophrenia: dysfunction of the default mode network? Psychol. Med. 38, 1185–1193 (2008).
Bluhm, R. L. et al. Spontaneous low-frequency fluctuations in the BOLD signal in schizophrenic patients: Anomalies in the default network. Schizophr. Bull. 33, 1004–1012 (2007).
McCollum, L. A. & Roberts, R. C. Uncovering the role of the nucleus accumbens in schizophrenia: a postmortem analysis of tyrosine hydroxylase and vesicular glutamate transporters. Schizophr. Res. 169, 369–373 (2015).
Hadley, J. A. et al. Ventral tegmental area/midbrain functional connectivity and response to antipsychotic medication in schizophrenia. Neuropsychopharmacology 39, 1020–1030 (2014).
Butler, T. et al. Septal nuclei enlargement in human temporal lobe epilepsy without mesial temporal sclerosis. Neurology 80, 487–491 (2013).
Dinkelacker, V. et al. Hippocampal-thalamic wiring in medial temporal lobe epilepsy: enhanced connectivity per hippocampal voxel. Epilepsia 56, 1217–1226 (2015).
Ivanov, I. et al. Morphological abnormalities of the thalamus in youths with attention deficit hyperactivity disorder. Image Process. 397–408, https://doi.org/10.1176/appi.ajp.2009.09030398 (2010).
Scofield, M. D. et al. The nucleus accumbens: mechanisms of addiction across drug classes reflect the importance of glutamate homeostasis. Pharmacol. Rev. 68, 816–871 (2016).
Vialou, V. et al. DeltaFosB in brain reward circuits mediates resilience to stress and antidepressant responses. Nat. Neurosci. 13, 745–752 (2010).
Volkow, N. D. et al. Motivation deficit in ADHD is associated with dysfunction of the dopamine reward pathway. Mol. Psychiatry 16, 1147–1154 (2011).
Yamamura, T. et al. Association of thalamic hyperactivity with treatment-resistant depression and poor response in early treatment for major depression: a resting-state fMRI study using fractional amplitude of low-frequency fluctuations. Transl. Psychiatry 6, e754 (2016).
Zhu, Y., Wienecke, C. F. R., Nachtrab, G. & Chen, X. A thalamic input to the nucleus accumbens mediates opiate dependence. Nature 530, 219–222 (2016).
Mease, R. A., Metz, M. & Groh, A. Cortical sensory responses are enhanced by the higher-order thalamus. Cell Rep. 14, 208–215 (2016).
Dringenberg, H. C. & Olmstead, M. C. Integrated contributions of basal forebrain and thalamus to neocortical activation elicited by pedunculopontine tegmental stimulation in urethane-anesthetized rats. Neuroscience 119, 839–853 (2003).
Yamamoto, K., Solal, B. & Philippe, V. New perspective on the regionalization of the anterior forebrain in Osteichthyes. Dev. Growth Regen. 59, 175–187 (2017).
Butler, A. B. Evolution of the thalamus: a morphological and functional review. Thalamus Relat. Syst. 4, 35–58 (2008).
Karten, H. J. Vertebrate brains and evolutionary connectomics: on the origins of the mammalian “neocortex.” Phil. Trans. R. Soc. B 370, 20150060 (2015).
Rilling, J. K. et al. A comparison of resting-state brain activity in humans and chimpanzees. Proc. Natl Acad. Sci. USA 104, 17146–17151 (2007).
Vincent, J. L. et al. Intrinsic functional architecture in the anaesthetized monkey brain. Nature 447, 83–88 (2007).
Buckner, R. L. & Margulies, D. S. Macroscale cortical organization and a default-like transmodal apex network in the marmoset monkey. bioRxiv. https://doi.org/10.1101/415141 (2018).
Lu, H. et al. Rat brains also have a default mode network. Proc. Natl Acad. Sci. USA 109, 3979–3984 (2012).
Gozzi, A. & Schwarz, A. J. Large-scale functional connectivity networks in the rodent brain. Neuroimage 15, 496–509 (2016).
Sforazzini, F., Schwarz, A. J., Galbusera, A., Bifone, A. & Gozzi, A. Distributed BOLD and CBV-weighted resting-state networks in the mouse brain. Neuroimage 87, 403–415 (2014).
Bertero, A. et al. Autism-associated 16p11.2 microdeletion impairs prefrontal functional connectivity in mouse and human. Brain 141, 2055–2065 (2018).
Liska, A., Galbusera, A., Schwarz, A. J. & Gozzi, A. Functional connectivity hubs of the mouse brain. Neuroimage 115, 281–291 (2015).
Chandler, D. & Waterhouse, B. D. Evidence for broad versus segregated projections from cholinergic and noradrenergic nuclei to functionally and anatomically discrete subregions of prefrontal cortex. Front Behav. Neurosci. 6, 1–9 (2012).
Chandler, D. J., Lamperski, C. S. & Waterhouse, B. D. Identification and distribution of projections from monoaminergic and cholinergic nuclei to functionally differentiated subregions of prefrontal cortex. Brain Res. 1522, 38–58 (2013).
Markello, R. D., Spreng, R. N., Luh, W., Anderson, A. K. & Rosa, E. De Segregation of the human basal forebrain using resting state functional MRI. Neuroimage 173, 287–297 (2018).
Jones, D. K. & Cercignani, M. Twenty-five pitfalls in the analysis of diffusion MRI Data. NMR Biomed. 23, 803–820 (2010).
Maier-Hein, K. H., Neher, P. F. & Descoteaux, M. The challenge of mapping the human connectome based on diffusion tractography. Nat. Commun. 8, 1349–1361 (2017).
Thiebaut de Schotten, M. et al. A lateralized brain network for visuospatial attention. Nat. Neurosci. 14, 1245–1247 (2011a).
Dell’Acqua, F., Simmons, A., Williams, S. C. R. & Catani, M. Can spherical deconvolution provide more information than fiber orientations? Hindrance modulated orientational anisotropy, a true-tract specific index to characterize white matter diffusion. Hum. Brain Mapp. 34, 2464–2483 (2013).
Karolis, V. R., Corbetta, M. & Thiebaut de Schotten, M. The architecture of functional lateralisation and its relationship to callosal connectivity in the human brain. Nat. Commun. 10, 1417–1424 (2019).
Moeller, S. et al. Multiband multislice GE-EPI at 7 tesla, with 16-fold acceleration using partial parallel imaging with application to high spatial and temporal whole-brain FMRI. Magn. Reson. Med. 63, 1144–1153 (2010).
Setsompop, K. et al. Blipped-controlled aliasing in parallel imaging for simultaneous multislice echo planar imaging with reduced g-factor penalty. Magn. Reson. Med. 67, 1210–1224 (2012).
Xu, J. et al. Evaluation of slice accelerations using multiband echo planar imaging at 3T. Neuroimage 83, 991–1001 (2013).
Feinberg, D. A. et al. Multiplexed echo planar imaging for sub-second whole brain fmri and fast diffusion imaging. PLoS One 5, https://doi.org/10.1371/journal.pone.0015710 (2010).
Andersson, J. L. R., Skare, S. & Ashburner, J. How to correct susceptibility distortions in spin-echo echo-planar images: application to diffusion tensor imaging. Neuroimage 20, 870–888 (2003).
Smith, S. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage 23, 208–219 (2004).
Foulon, C. et al. Advanced lesion symptom mapping analyses and implementation as BCBtoolkit. Gigascience 7, 1–17 (2018).
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17, 825–841 (2002).
Woolrich, M. W. et al. Bayesian analysis of neuroimaging data in FSL. Neuroimage 45, S173–S186 (2009).
Avants, B. & Gee, J. C. Geodesic estimation for large deformation anatomical shape averaging and interpolation. Neuroimage 23, 139–150 (2004).
Avants, B. B. et al. The optimal template effect in hippocampus studies of diseased populations. Neuroimage 49, 2457–2466 (2010).
Avants, B. B. et al. A reproducible evaluation of ANTs similarity metric performance in brain image registration. Neuroimage 54, 2033–2044 (2011).
Klein, A. et al. Evaluation of 14 nonlinear deformation algorithms applied to human brain MRI registration. Neuroimage 46, 786–802 (2009).
van Dijk, K. R. A., Sabuncu, M. R. & Buckner, R. L. The influence of head motion on intrinsic functional connectivity MRI. Neuroimage 59, 431–438 (2012).
Pruim, R. H. R., Mennes, M., Buitelaar, J. K. & Beckmann, C. F. Evaluation of ICA-AROMA and alternative strategies for motion artifact removal in resting state fMRI. Neuroimage 112, 278–287 (2015).
Salimi-khorshidi, G. et al. Automatic denoising of functional MRI data: combining independent component analysis and hierarchical fusion of classifiers. Neuroimage 90, 449–468 (2014).
Power, J. D., Barnes, K. A., Snyder, A. Z., Schlaggar, B. L. & Petersen, S. E. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142–2154 (2012).
Shine, J. M. et al. The dynamics of functional brain networks: integrated network states during cognitive task performance. Neuron 92, 544–554 (2016).
Tomasi, D. & Volkow, N. D. Association between brain activation and functional connectivity. Cereb Cortex 1–13, https://doi.org/10.1093/cercor/bhy077 (2018).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. Fsl. Neuroimage 62, 782–790 (2012).
Kenney, J. Mathematics of Statistics. (Chapman & Hall, London, 1939).
Avants, B. B., Epstein, C. L., Grossman, M. & Gee, J. C. Symmetric diffeomorphic image registration with cross-correlation: evaluating automated labeling of elderly and neurodegenerative brain. Med. Image Anal. 12, 26–41 (2008).
Cohen, J. A power prime. Psychol. Bull. 112, 155–159 (1992).
Nieuwenhuys, R., Voogd, J. & van Huijzen, C. The Human Central Nervous System: A Synopsis and Atlas, 4th edn. (Steinkopff, 2008).
Catani, M. & Thiebaut de Schotten, M. Atlas of Human Brain Connections, 1st edn. (Oxford University Press, 2012).
Xia, M., Wang, J. & He, Y. BrainNet Viewer: a network visualization tool for human brain connectomics. PLoS One 8. https://doi.org/10.1371/journal.pone.0068910 (2013).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Vos, S. B. et al. The importance of correcting for signal drift in diffusion MRI. Magn. Reson. Med. 77, 285–299 (2017).
Leemans, A., Jeurissen, B., Sijbers, J. & Jones, D. ExploreDTI: a graphical toolbox for processing, analyzing, and visualizing diffusion MR data. In 17th Scientific Meeting, International Society for Magnetic Resonance in Medicine. 3537 (Hawaii, 2009).
Dell’Acqua, F. et al. A modified damped Richardson-Lucy algorithm to reduce isotropic background effects in spherical deconvolution. Neuroimage 49, 1446–1458 (2010).
Thiebaut de Schotten, M. et al. Damage to white matter pathways in subacute and chronic spatial neglect: a group study and 2 single-case studies with complete virtual “in vivo” tractography dissection. Cereb. Cortex 24, 691–706 (2014).
Wassermann, D. et al. The white matter query language: a novel approach for describing human white matter anatomy. Brain Struct. Funct. 221, 4705–4721 (2016).
Tournier, J. D., Calamante, F. & Connelly, A. MRtrix: diffusion tractography in crossing fiber regions. Int. J. Imaging Syst. Technol. 22, 53–66 (2012).
Calamante, F., Tournier, J. D., Jackson, G. D. & Connelly, A. Track-density imaging (TDI): super-resolution white matter imaging using whole-brain track-density mapping. Neuroimage 53, 1233–1243 (2010).
Thiebaut de Schotten, M. et al. Atlasing location, asymmetry and inter-subject variability of white matter tracts in the human brain with MR diffusion tractography. Neuroimage 54, 49–59 (2011b).
Bürgel, U. et al. White matter fiber tracts of the human brain: three-dimensional mapping at microscopic resolution, topography and intersubject variability. Neuroimage 29, 1092–1105 (2006).
Rivière, D., Geffroy, D., Denghien, I., Souedet, N. & Cointepas, Y. Anatomist: a python framework for interactive 3D visualization of neuroimaging data. In Python in Neuroscience Workshop (2011).
Shu, N. et al. Diffusion tensor tractography reveals disrupted topological efficiency in white matter structural networks in multiple sclerosis. Cereb. Cortex 21, 2565–2577 (2011).
Gong, G. et al. Mapping anatomical connectivity patterns of human cerebral cortex using in vivo diffusion tensor imaging tractography. Cereb. Cortex 19, 524–536 (2009).
Jones, D. K., Knösche, T. R. & Turner, R. White matter integrity, fiber count, and other fallacies: the do’s and don’ts of diffusion MRI. Neuroimage 73, 239–254 (2013).
Aggleton, J. P. Understanding anterograde amnesia: disconnections and hidden lesions. Q. J. Exp. Psychol. 61, 1441–1471 (2008).
Haber, S. N., Kunishio, K. & Mizobuchi, M. The orbital and medial prefrontal basal ganglia circuit through the primate. Neuroscience 39, 323–338 (1995).
Haber, S. N., Wolfe, D. P. & Groenewegen, H. J. The relationship between ventral striatal efferent fibers and the distribution of peptide-positive woolly fibers in the forebrain of the rhesus monkey. Neuroscience 39, 323–338 (1990).
Kunishio, K. & Haber, S. N. Primate cingulostriatal projection: limbic striatal versus sensorimotor striatal input. J. Comp. Neurol. 350, 337–356 (1994).
Aggleton, J. P., Pralus, A., Nelson, A. J. D. & Hornberger, M. Thalamic pathology and memory loss in early Alzheimer’s disease: moving the focus from the medial temporal lobe to Papez circuit. Brain 139, 1877–1890 (2016).
Osorio, I., Overman, J., Giftakis, J. & Wilkinson, S. B. High frequency thalamic stimulation for inoperable mesial temporal epilepsy. Epilepsia 48, 1561–1571 (2007).
Salanova, V. et al. Long-term efficacy and safety of thalamic stimulation for drug-resistant partial epilepsy. Neurology 84, 1017–1025 (2015).
Acknowledgements
P.N.A. work was financially supported by the Research Experience Fellowship grant of the European Academy of Neurology and by “PRÉMIO JOÃO LOBO ANTUNES” - SCML. The research leading to these results received funding from the “Agence Nationale de la Recherche” [grant number ANR-13- JSV4-0001-01]. This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement No. 818521) and from the Fondation pour la Recherche Médicale (FRM DEQ20150331725).
Author information
Authors and Affiliations
Contributions
P.N.A. implemented part of the methods, performed the analyses and wrote the manuscript. C.F. implemented part of the methods and edited the paper, Vyacheslav Karolis contributed to the methods and edited the paper, D.B., D.S.M. and E.V. helped conceptually and edited the paper, M.T.S. conceived and coordinated the study, reviewed the neuroimaging data, wrote the paper and provided funding.
Corresponding authors
Ethics declarations
Competing interests
Michel Thiebaut de Schotten is an Editorial Board Member for Communications Biology, but was neither involved in the editorial review of nor the decision to publish this article. The authors declare no other competing financial or non-financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alves, P.N., Foulon, C., Karolis, V. et al. An improved neuroanatomical model of the default-mode network reconciles previous neuroimaging and neuropathological findings. Commun Biol 2, 370 (2019). https://doi.org/10.1038/s42003-019-0611-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-019-0611-3
This article is cited by
-
Multimodal neural correlates of dispositional resilience among healthy individuals
Scientific Reports (2024)
-
Language evolution and computational capabilities: conceptualization of the first language units
International Journal of Anthropology and Ethnology (2023)
-
Functional connectivity of cognition-related brain networks in adults with fetal alcohol syndrome
BMC Medicine (2023)
-
Effect of Tai Chi on Young Adults with Subthreshold Depression via a Stress–Reward Complex: A Randomized Controlled Trial
Sports Medicine - Open (2023)
-
The default network dominates neural responses to evolving movie stories
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.