Abstract
Circadian clocks are conserved time-keeper mechanisms in some prokaryotes and higher eukaryotes. Chromatin modification is emerging as key regulatory mechanism for refining core clock gene expression. Rhythmic changes in histone marks are closely associated to the TIMING OF CAB EXPRESSION 1 (TOC1) Arabidopsis clock gene. However, the chromatin-related modifiers responsible for these marks remain largely unknown. Here, we uncover that the chromatin modifier HISTONE DEACETYLASE 9 (HDA9) and the Evening complex (EC) component EARLY FLOWERING 3 (ELF3) directly interact to regulate the declining phase of TOC1 after its peak expression. We found that HDA9 specifically binds to the TOC1 promoter through the interaction with ELF3. The EC-HDA9 complex promotes H3 deacetylation at the TOC1 locus, contributing to suppressing TOC1 expression during the night, the time of EC function. Therefore, we have identified the mechanism by which the circadian clock intertwines with chromatin-related components to shape the circadian waveforms of gene expression in Arabidopsis.
Similar content being viewed by others
Introduction
Circadian clocks generate biological rhythms with a period of 24 h and control plant growth and development in synchronization with the environmental cycles. Multiple transcriptional feedback loops define the basic architecture of the plant circadian clock. In Arabidopsis, two single-MYB transcription factors, CIRCADIAN CLOCK–ASSOCIATED 1 (CCA1) and LATE ELONGATED HYPOCOTYL (LHY), repress transcription of the TOC1/PSEUDO RESPONSE REGULATOR 1 (PRR1) that in turn represses CCA1 and LHY expression, constituting the central loop1,2. The central loop is interlocked with additional transcriptional loops. In the morning, a transcriptional repressing wave includes additional members of the PRR family including PRR5, PRR7, and PRR9 and CCA1 and LHY3,4. Evening-expressed clock components such as TOC1 and the EC contribute to the repression of the morning-expressed genes5,6,7,8. Additional transcriptional regulation also underlies circadian frameworks9,10,11. Moreover, accumulating evidence suggests that multiple layers of regulation such as alternative splicing, controlled protein turnover, and posttranslational modification further contribute to a precise rhythmic oscillation12,13,14.
Chromatin conformation influences the accessibility of transcriptional regulator(s) to the associated DNA regions. Chemical modifications at histone N-terminal tails contribute to proper packing of chromatin15. In particular, histone acetylation is implicated primarily in transcriptional activation16. Histone acetyltransferases (HATs) catalyze the addition of acetyl groups to lysine residues at histones, neutralizing positive charges and thus decreasing the affinity of histones to DNA17. This facilitates the accessibility of transcriptional regulators and other chromatin modifiers18. In contrast, histone deacetylases (HDACs) antagonize the action of HATs by removing the acetyl groups from histone19.
The first study relating chromatin modification and the Arabidopsis circadian clock identified the circadian changes in Histone 3 acetylation (H3ac) at the TOC1 promoter. The antagonistic action of CCA1 at dawn and another single MYB clock-related transcription factor REVEILLE8/LHY-CCA1-LIKE5 (RVE8/LCL5) was found to define the hypo-acetylated and hyper-acetylated states of H3 at the TOC1 promoter during the day10. Indeed, CCA1 facilitates a repressive chromatin conformation at dawn either by interfering with HAT accessibility or by recruiting HDAC activity20. During the day, RVE8/LCL5 antagonizes the CCA1 repressive function, favoring H3ac10. These counteracting functions precisely shape the waveform of TOC1 expression10. The rhythmic accumulation of different histone marks was also observed at the promoters of other core clock components, including CCA1, LHY, LUX ARRHYTHMO (LUX), and PRRs, and correlates with their transcript accumulation21.
Despite the clear rhythms in histone acetylation at the TOC1 locus, the chromatin remodeling factor(s) contributing to the rhythms of histone acetylation and its inner working mechanism remain elusive. Here, we report that HDA9, a member of the reduced potassium dependency 3 (RPD3)/HDA1 family class I HDAC22,23, is involved in the circadian regulation of TOC1 by suppressing its expression. HDA9 specifically interacts with an EC component, ELF3, and the physical interaction enables HDA9 to bind to the TOC1 promoter. HDA9 might facilitate a closed chromatin structure at the TOC1 promoter, contributing to its declining phase of expression during the night period. These results provide an insight into how temporal regulation of histone acetylation is achieved in order to stably maintain circadian activity.
Results
Altered circadian oscillation in hda9 mutant plants
Previous studies have shown that histone deacetylation is important for the rhythmic oscillation in histone acetylation at the core clock promoters10,21,24. As pharmacological inhibition of the RPD3/HDA1 family class I HDAC activities25 with TSA (Trichostatin A) affects the circadian oscillation20,25, we investigated their possible function within the circadian clock. Among the four members of this family (HDA6, HDA7, HDA9, and HDA19)26, we focused on two members not previously studied, HDA7 and HDA9.
To that end, we obtained HDA7 and HDA9-deficient mutants, hda7–2 and hda9–1, and analyzed rhythmic expression of the circadian oscillator gene, CCA1. Quantitative real-time RT-PCR (RT-qPCR) analysis revealed that the circadian expression of CCA1 was unaffected in hda7–2 mutant compared with wild type (Supplementary Fig. 1). However, the lack of HDA9 led to clear alterations in CCA1 circadian oscillation (Supplementary Figs. 1 and 2). CCA1 phase appeared advanced, which suggests a possible shortening of the circadian period in hda9–1 (Supplementary Fig. 1). Consistently, expression of circadian output genes, COLD, CIRCADIAN RHYTHM, AND RNA BINDING (CCR2) and CHLOROPHYLL A/B BINDING–PROTEIN 2 (CAB2), and several circadian oscillator genes displayed a similar pattern with a period shortening and advanced rhythmic phase (Fig. 1a, b; also see Supplementary Figs. 3–5). Thus, HDA9 is important for proper circadian oscillation.
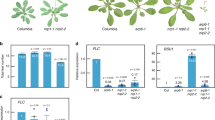
Mutation of HDA9 alters circadian oscillation. Two-week-old seedlings grown under neutral day conditions (ND) were transferred to continuous light conditions (LL) at ZT0. Whole seedlings (n > 15) were harvested from ZT48 to ZT116 to analyze transcript accumulation of CCR2 (a) and CAB2 (b). eIF4a was used as the normalization control. Two technical replicates were averaged. Bars indicate the standard deviation. The white and gray boxes indicate the subjective day and night, respectively
Binding of HDA9 to the TOC1 promoter
We next aimed to identify whether HDA9 regulation of circadian gene expression occurs through direct binding to a core clock locus. To that end, we performed chromatin immunoprecipitation (ChIP) assays using 35 S:MYC-HDA9 transgenic plants. ChIP enrichment was examined by quantitative PCR (qPCR) in the promoter regions of selected clock genes, which include key clock-related cis-elements, such as CCA1-binding site (CBS), evening element (EE), G-box, and/or LUX binding site (LBS) (Fig. 2a)5,7,27,28, in which H3ac levels are known to be rhythmically oscillating21. Our results showed no significant ChIP amplification except at the TOC1 promoter (Fig. 2b). HDA9 specifically associated with a promoter region at the TOC1 locus (Fig. 2b), but not within TOC1 coding region (Supplementary Fig. 6).
HDA9 associates with TOC1 promoter to catalyze H3 deacetylation. a Genomic structures of core clock genes. Exons are represented by black boxes. Underbars indicate the regions amplified by PCR after chromatin immunoprecipitation (ChIP). Red, blue, green, and yellow arrowheads represent CBS, EE, G-box, and LBS motifs, respectively. b Binding of HDA9 to clock gene promoters. Two-week-old plants entrained with ND cycles were subjected to LL. Plants were harvested at ZT16 for ChIP analysis with anti-MYC antibody. Enrichment of fragmented genomic regions was analyzed by ChIP-qPCR. Biological triplicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05; difference between Col-0 and 35 S:MYC-HDA9 plants). Bars indicate the standard error of the mean. c H3ac levels at the TOC1 locus in hda9–1. Two-week-old plants entrained with ND cycles were subjected to LL. Plants were harvested at ZT12 and ZT16 for ChIP analysis with anti-H3ac antibody. Biological triplicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05; difference between Col-0 and hda9–1 plants). Bars indicate the standard error of the mean. d Transient expression assays. The effector and reporter constructs were coexpressed into Arabidopsis protoplasts. The GUS activities were measured fluorimetrically. Biological triplicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05). Bars indicate the standard error of the mean. e Expression of TOC1 in hda9–1. Two-week-old seedlings grown under ND were transferred to LL at ZT0. Whole seedlings were harvested from ZT48 to ZT68. eIF4a was used as the normalization control. Two technical replicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05; difference between Col-0 and hda9–1 plants). Bars indicate the standard deviation. The white and gray boxes indicate the subjective day and night
To further explore the circadian function of HDA9, we examined its possible oscillatory expression, but found that the transcript accumulation of HDA9 did not significantly oscillate throughout a circadian cycle (Supplementary Fig. 7). Thus, HDA9 mRNA expression is not circadianly regulated. We nevertheless asked whether HDA9 might rhythmically bind to the TOC1 promoter. Notably, binding of HDA9 was observed at Zeitgeber Time 16 (ZT16) (Fig. 2b and Supplementary Fig. 8), when TOC1 expression is rather low1. No obvious amplification was observed at other time points examined (Supplementary Fig. 8).
HDA9 protein biochemically catalyzes the removal of acetyl groups from histone H3 proteins22,23. Since HDA9 binds to the TOC1 promoter (Fig. 2b), we hypothesized that HDA9 might contribute to changes in H3ac accumulation at the TOC1 locus. ChIP assays with an anti-H3ac antibody revealed that H3ac accumulation at a distal region of the TOC1 promoter declined from ZT12 to ZT16 in wild type (Fig. 2c), consistent with the declined expression of TOC11. However, the reduction of H3ac at the TOC1 promoter was impaired in hda9–1 mostly at ZT16 (Fig. 2c), when HDA9 binds to the TOC1 promoter (Fig. 2b and Supplementary Fig. 8). HDA9 might bind to the specific promoter regions and then catalyze H3 deacetylation of neighboring sites. The spatial separation was consistent with the molecular function of other chromatin modifiers29.
Given that H3ac is associated with gene activation16, HDA9 function may facilitate TOC1 repression. To test this hypothesis, we performed transient expression assays using Arabidopsis protoplasts. The TOC1 promoter was fused to the 35S minimal promoter. The recombinant reporter plasmid and the effector construct expressing HDA9 gene were co-transformed into Arabidopsis protoplasts. Co-expression of these constructs repressed the GUS activity by 40% (Fig. 2d). In addition, we also examined TOC1 expression in HDA9-misexpressed plants. In hda9–1 mutant plants, TOC1 expression was clearly advanced, resulting in higher expression during the day and just after subjective dusk (Fig. 2e and Supplementary Fig. 9). Taken together, we propose that HDA9 is recruited to the TOC1 promoter after its peak time and represses TOC1 expression by promoting H3 deacetylation.
Protein–protein interaction of HDA9 and ELF3
Our results show that the HDA9 protein is recruited to the TOC1 promoter. Given that HDA9 is not circadianly regulated, we hypothesized that additional molecular component(s) might contribute to the rhythmic function of HDA9. As HDA9 itself does not have binding selectivity on specific DNA regions, DNA-binding proteins may be required for specific HDA9 binding to the TOC1 promoter. To examine this hypothesis, we carried out yeast-two-hybrid assays with expression constructs containing core clock transcription factors. The HDA9-GAL4 DNA binding domain fusion construct was co-expressed with a construct expressing a clock gene fused in-frame to the 3’-end of GAL4 activation domain in yeast cells. Cell growth on the selective medium revealed that HDA9 specifically interacted with ELF3 (Fig. 3a), but not with other examined clock proteins (Fig. 3a).
HDA9 interacts with EC. a Yeast-two-hybrid assays. Yeast-two-hybrid assays were performed with the HDA9 proteins fused to the DNA-binding domain (BD) of GAL4 and clock components fused with the transcriptional activation domain (AD) of GAL4 for analysis of interactions. Interactions were examined by cell growth on selective media. -LWHA indicates Leu, Trp, His, and Ade drop-out plates. -LW indicates Leu and Trp drop-out plates. GAL4 was used as a positive control. b BiFC assays. Partial fragments of YFP protein were fused with HDA9 and ELF3 and co-expressed in Arabidopsis protoplasts. IDD14-RFP was used as a nucleus marker. Scale bar, 10 µm. c Coimmunoprecipitation assays. Agrobacterium tumefaciens cells containing 35S:HDA9-MYC and 35S:ELF3-GFP constructs were coinfiltrated to 3-week-old N. benthamiana leaves. Epitope-tagged proteins were detected immunologically using corresponding antibodies. Full blot images were shown in Supplementary Fig. 10
To support the interaction of HDA9 with ELF3 in vivo, we performed bimolecular fluorescent complementation (BiFC) analysis using Arabidopsis protoplasts. The HDA9 cDNA sequence was fused in-frame to the 5’-end of a gene sequence encoding the C-terminal half of YFP (cYFP), and the ELF3 gene was fused in-frame to the 5’-end of a sequence encoding the N-terminal half of YFP (nYFP). The fusion constructs were then transiently co-expressed in Arabidopsis protoplasts. The HDA9-ELF3 combination was able to visualize yellow fluorescence, which was exclusively detected in the nucleus (Fig. 3b). Moreover, in planta interactions of HDA9 and ELF3 were also verified (Fig. 3c and Supplementary Fig. 10). These results indicate that HDA9 associates with ELF3 to repress TOC1 expression.
The EC facilitates HDA9 binding to the TOC1 promoter
Since ELF3 is a key component of the EC30, HDA9 could be associated with the EC. This is in agreement with the temporal association of HDA9 to TOC1, because EC function is highly relevant at ZT1630. However, it is currently unknown whether the EC indeed associates with the TOC1 promoter. To test this possibility, we employed pELF3:ELF3-MYC/elf3–1, pELF4:ELF4-HA/elf4–2, and pLUX:LUX-GFP/lux-4 transgenic plants30. ChIP assays were performed with epitope-tagged transgenic plants immunoprecipitated with corresponding antibodies. DNA bound to epitope-tagged proteins was analyzed by qPCR assays. The ChIP-qPCR analysis showed that the TOC1 promoter was enriched by ELF3, ELF4, and LUX (Figs. 4a–c), also in the regions where HDA9 is recruited (Fig. 2a, b). Furthermore, binding of EC components to the TOC1 promoter occurred mainly at ZT16 (Fig. 4a–c), which is consistent with the timing of HDA9 binding (Fig. 2b).
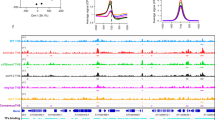
EC directly binds to the TOC1 promoter. a–c Binding of ELF3, ELF4, and LUX to the TOC1 locus. Two-week-old plants entrained with ND cycles were subjected to LL. Plants were harvested at ZT12 and ZT16 for ChIP analysis. Enrichment of fragmented genomic regions was analyzed by ChIP-qPCR (see Fig. 2a). Two biological replicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05, **P < 0.01). Bars indicate the standard error of the mean. d Effects of ELF3, ELF4, and LUX on TOC1 repression. The effector and reporter constructs were transiently coexpressed in Arabidopsis protoplasts. The GUS activities were measured fluorimetrically. Biological triplicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05). Bars indicate the standard error of the mean
Direct binding of EC to the TOC1 promoter most likely allows transcriptional repression of TOC1. To verify the possibility, we conducted transient expression assays with the TOC1 promoter fused to the 35S minimal promoter. The recombinant reporter plasmid and the effector construct expressing ELF3, ELF4, or LUX were co-transformed into Arabidopsis protoplasts. As expected, they bound to the TOC1 promoter and repressed its expression (Fig. 4d), although they had different transcriptional repressive activity. Given that EC components are known to have overlapped and separate biological functions, ELF3 and LUX may play a particular role in TOC1 regulation.
Since the EC recruits HDA9 to repress gene expression through H3 deacetylation, we measured H3ac levels at the TOC1 promoter in elf3–8, elf4–101, and lux-6 mutants. ChIP-qPCR analysis revealed that H3 deacetylation at a distal region of the TOC1 promoter was impaired at ZT16 in all mutant examined (Fig. 5a–c), in a similar fashion to that observed for hda9–1 mutant (Fig. 2c). In addition, we also analyzed expression of TOC1 in elf3–8 and lux-6 and found that TOC1 expression was arrhythmic and elevated at some time points, including the subjective night (ZT64-ZT68) (Fig. 5d, e; also see Supplementary Figs. 11 and 12). Consistently, CCA1 gene expression was also arrhythmic and suppressed during the subjective day (Supplementary Fig. 13), which correlates with the elevated expression of its repressor TOC12. These results suggest that the EC and HDA9 act together in the control of histone deacetylation at the TOC1 promoter.
EC represses TOC1 expression by promoting H3 deacetylation. a–c Elevated H3ac levels at TOC1 locus in EC mutants. Two-week-old plants entrained with ND cycles were subjected to LL. Plants were harvested at ZT12 and ZT16 for ChIP analysis with anti-H3ac antibody. Two biological replicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05). Bars indicate the standard error of the mean. d and e Expression of TOC1 in elf3–8 and lux-6. Two-week-old seedlings grown under ND were transferred to LL at ZT0. Whole seedlings were harvested from ZT48 to ZT68. eIF4a was used as the normalization control. Two technical replicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05). Bars indicate the standard deviation. The white and gray boxes indicate the subjective day and night
The importance of EC function for HDA9 binding to the TOC1 promoter was tested using 35S:MYC-HDA9 transgenic plants crossed with the elf3–8 mutant. ChIP assays using an anti-MYC antibody showed that HDA9 binding to the TOC1 locus was reduced in the elf3–8 mutant background compared with the wild-type background (Fig. 6a), although HDA9 protein similarly accumulated in both backgrounds (Supplementary Fig. 14). Since HDA9 promotes H3 deacetylation at the cognate regions, we also measured H3ac levels in the same plants. H3ac levels were reduced in 35S:MYC-HDA9 plants compared with wild type, but H3 deacetylation was compromised in 35S:MYC-HDA9xelf3–8 plants, which was equivalent to elf3–8 (Fig. 6b). Further, 35S:MYC-HDA9 transgenic plants displayed robust circadian expression of TOC1, while the amplitude was reduced compared with wild type (Supplementary Fig. 15). However, arrhythmic circadian expression was observed in 35S:MYC-HDA9xelf3–8 (Fig. 6c), similar to EC mutants (Fig. 5d, e). Indeed, HDA9 function is most likely dependent on EC. Transient expression assays using Arabidopsis protoplasts revealed that HDA9 binds to the TOC1 promoter to inhibit expression (Fig. 2d). However, the HDA9 function disappeared in elf3–8 and lux-6 mutants (Fig. 6d). These results suggest that HDA9 requires the ELF3, possibly EC, for binding to the TOC1 promoter.
EC is required for HDA9 binding to the TOC1 promoter. a Binding of HDA9 to the TOC1 locus. b H3ac levels in 35S:MYC-HDA9xelf3–8. In a and b, two-week-old plants entrained with ND cycles were subjected to LL. Plants were harvested at ZT16 for ChIP analysis. Two biological replicates were averaged and statistically analyzed with Student’s t-test (*P < 0.05). Bars indicate the standard error of the mean. n.s., not significant. c Expression of TOC1 in 35S:MYC-HDA9xelf3–8. Two-week-old seedlings grown under ND were transferred to LL at ZT0. Whole seedlings were harvested from ZT48 to ZT68. eIF4a was used as the normalization control. Biological triplicates were averaged. Bars indicate the standard error of the mean. The white and gray boxes indicate the subjective day and night. d Transient expression assays. The GUS activities were measured fluorimetrically. Biological triplicates were averaged and statistically analyzed with Student’s t-test (***P < 0.001). Bars indicate the standard error of the mean
In summary, TOC1 expression is regulated by rhythmic changes in histone acetylation states. A-yet-unidentified HAT activity is required for the rising phase of TOC1 expression. After its peak expression, HDA9 is recruited to the TOC1 promoter through the interaction with ELF3, possibly the EC. The EC-HDA9 complex facilitates H3 deacetylation at the TOC1 locus to reach its basal expression during night period (Fig. 7). Diurnal oscillation of H3ac levels may be pervasive in core clock genes, ensuring circadian homeostasis.
Working diagram of the EC-HDA9 complex in circadian control. Transcript accumulation of TOC1 correlates with H3ac levels at the gene promoter. As-yet-unidentified HAT catalyzes H3ac at the TOC1 promoter during the daytime. After peak expression, the ELF3 protein, presumably evening complex (EC), recruits HDA9. The EC-HDA9 complex binds to the TOC1 promoter and removes H3ac at neighboring sites to decline its expression during nighttime
Discussion
Rhythmic expression of core clock genes is intimately associated with the levels of H3ac, especially H3K56ac and H3K9/14ac, and H3K4me3 at the gene promoters in Arabidopsis21,31. Despite the connection between chromatin modification and the circadian control, just a few responsible chromatin modifiers have been demonstrated to participate in circadian chromatin remodeling.
The SET DOMAIN GROUP 2 (SDG2)/ARABIDOPSIS TRITHORAX–RELATED 3 (ATXR3) protein establishes H3K4me3 mark at core clock gene loci to promote expression. The H3K4me3 mark inhibits clock repressor binding at the core clock promoters, facilitating transcriptional repression to target genes only at specific time-of-day21. Accordingly, the SDG2/ATXR3-deficient mutants exhibit a decrease in H3K4me3 accumulation, advanced clock repressor binding, and reduced amplitude of core clock gene expression21. In addition, the JmjC domain-containing histone demethylase JMJ30/JMJD5 is also involved in circadian homeostasis. The JMJ30/JMJD5 gene is circadian-regulated and peaks at dusk32,33. The central oscillators CCA1 and LHY directly bind to and repress JMJ30/JMJD5. Then, JMJ30/JMJD5 reciprocally regulates expression of CCA1 and LHY32.
A couple of HDACs have been shown to be implicated in the Arabidopsis circadian system. PRR5, PRR7, and PRR9 directly repress CCA1 and LHY34. Notably, PRRs interact with TOPLESS/TOPLESS RELATED PROTEINS (TPL/TPRs), as well as HDA6 and HDA1934. Consistently, inhibition of the TPL activity compromises transcriptional repression activities of PRR5, PRR7, and PRR9, lengthening circadian period34. The potent HDAC inhibitor TSA activates CCA1 expression, but this activation is impaired in the tpl-1 mutant, demonstrating that HDAC regulation is largely dependent on TPL34.
We here show that HDA9 regulates circadian oscillation by modulating TOC1 expression. HDA9 binds to the TOC1 promoter and stimulates H3 deacetylation to suppress TOC1 expression. HDA9 interacts with the EC component ELF3, and rhythmic binding of HDA9 to the TOC1 promoter is defined by the EC. Accordingly, although HDA9 has no obvious rhythmic expression pattern, the EC-HDA9 interactions facilitate temporal regulation of TOC1. Consistent with the fact that the EC is expressed highly during night period35, the EC-HDA9 complex is relevant in the declining phase of TOC1 expression. Nonetheless, many questions remain to be answered. Considering the variable alteration patterns of circadian gene expression in hda9–1 mutant, EC and HDA9 are extensively interconnected with other circadian and chromatin-related components, accounting for dynamic contribution to circadian oscillation. Moreover, it is sometimes observed the anti-phase of binding regions and histone modifications29,36. While HDA9 binds to a proximal region of the TOC1 promoter, changes in histone acetylation are drastically observed in a distal region. This may be due to propagation and expansion of epigenetic modification of chromatin. Chromatin remodeling components that regulate epigenetic contexts on chromatin would also be involved, and thus the responsible protein components should be further elucidated. Future studies will give a comprehensive view of circadian-chromatin networks. Altogether, these results exemplify a mechanism by which chromatin factors engage with clock components to rhythmically modulate chromatin conformation and hence transcriptional oscillation.
The HDA9 protein is implicated in plant developmental processes. HDA9 is highly expressed in dry seeds and represses seedling traits to maintain seed dormancy37. The hda9 mutants show reduced seed dormancy with higher expression of genes responsible for photosynthesis and photoautotroph, such as RuBisCO and RuBisCO activase (RCA)37. The HDA9 action is antagonistic to HDA6 and HDA19, which repress embryonic properties in germinating seeds37, balancing the proper transition from seed to autotropic seedling.
The HDA9-deficient mutants also exhibit early flowering phenotypes under short day conditions38,39. HDA9 binds to the AGAMOUS–LIKE 19 (AGL19) promoter and negatively regulates its transcription through the alterations in H3K9 and H3K27 acetylation levels at the AGL19 locus39. Consequently, the HDA9-AGL19 module mainly regulates FLOWERING LOCUS T (FT) expression, in parallel with photoperiod or autonomous pathways to prevent precocious flowering in short days39. The epigenetic regulation of AGL19 by HDA9 is also relevant, in part, during vernalization process. Given that HDA9 is functional when AGL19 is actively expressed, such in the adult stage or after vernalization39, this HDAC protein is able to associate with transcriptionally active genes to reset the chromatin state. Accumulating evidence further supports that the HDA9-POWERDRESS (PWR) complex stimulates H3 deacetylation at a genome-wide level and is implicated in additional developmental processes, such as floral stem cell fate, stress responses, and aging22,23.
Notably, HDA9 activity can be diurnally gated. HDACs often form a massive protein complex containing for instance Sin3, RPD3-type HDAC, SIN3-ASSOCIATED POLYPEPTIDE 18 (SAP18) and SAP3040. Homologs of each component are present in Arabidopsis: six SIN3-Like (SNL1-SNL6), four RPD3 HDACs, one SAP18, and two SAP30 function-relateds (AFR1 and AFR2)40. The Arabidopsis Sin3-HDAC complex is crucial for diurnal control of FT40. This complex accumulates at dusk and is recruited to FT promoter to ensure periodic histone deacetylation. Dynamic cycles of H3 acetylation and deacetylation ensure adequate levels of the gene transcription and thereby prevent precocious flowering in the premature inductive conditions40. This study further supports periodic functions of HDA9 in controlling circadian gene expression. H3ac states can be reversible and account for stably oscillating gene expression. Taken together, the HDA9 protein seems to integrate temporal information to regulate oscillator gene expression and a variety of physiological output processes in order to optimize plant growth and development.
Methods
Plant materials and growth conditions
Arabidopsis thaliana (Columbia-0 ecotype) was used for all experiments unless otherwise specified. Plants were grown under neutral-day conditions (NDs; 12-h light/12-h dark cycles) with cool white fluorescent light (100 μmol photons m−2 s−1) at 22 °C. The elf3–8, elf4–101, hda7–2, and hda9–1 were previously reported39,41,42,43. The lux-6 mutant (SALK-022315) was obtained from Arabidopsis Biological Resource Center (ABRC). To produce transgenic plants overexpressing the HDA9 gene, a full-length cDNA was subcloned into the binary pBA002 vector under the control of the CaMV 35 S promoter. Agrobacterium tumefaciens-mediated Arabidopsis transformation was then performed.
Quantitative real-time RT-PCR analysis
Plants grown under conditions/ZT indicated were harvested and ground in liquid nitrogen. Total RNA was extracted by mixing the tissue powder with 700 μl of TRI reagent (TAKARA Bio, Singa, Japan). The extraction mixture was incubated at 65 °C for 5 min. After mixing with 700 μl of chloroform:isoamyl alcohol (24:1), the extraction mixture was centrifuged for 20 min at room temperature. The RNA in the liquid phase was precipitated at 4 °C in a final concentration of 2 M LiCl. RNA pellet was washed by 70% ethanol, briefly air dried, and suspended in distilled water. Reverse transcription (RT) was performed using Moloney Murine Leukemia Virus (M-MLV) reverse transcriptase (Dr. Protein, Seoul, South Korea) with oligo(dT18) to synthesize first-strand cDNA from 2 μg of total RNA. Total RNA samples were pretreated with an RNAse-free DNAse to remove DNA contamination.
Quantitative RT-PCR reactions were performed on the Step-One Plus Real-Time PCR System (Applied Biosystems). The PCR primers used are listed in Supplementary Table 1. The comparative CT method was used to determine the relative gene expression, with the expression of Eukaryotic translation initiation factor 4A1 (eIF4A) gene (At3g13920) as the internal control. All RT-qPCR reactions were performed with biological triplicates using total RNA samples extracted from three independent replicate samples.
Yeast two-hybrid assays
Yeast two-hybrid assays were performed using the BD Matchmaker system (Clontech, Mountain View, CA, USA). The pGADT7 vector was used for the GAL4 activation domain fusion, and the pGBKT7 vector was used for the GAL4 DNA binding domain fusion. The yeast strain AH109 harboring the LacZ and His reporter genes was used. PCR products were subcloned into the pGBKT7 and pGADT7 vectors. The expression constructs were co-transformed into yeast AH109 cells, and transformed cells were selected by growth on SD/-Leu/-Trp medium and SD/-Leu/-Trp/-His/-Ade.
Bimolecular fluorescence complementation (BiFC) assays
The HDA9 gene was fused in-frame to the 5′ end of a gene sequence encoding the C-terminal half of EYFP in the pSATN-cEYFP-C1 vector (E3082). The ELF3 cDNA sequences were fused in-frame to the 5′ end of a gene sequence encoding the N-terminal half of EYFP in the pSATN-nEYFP-C1 vector (E3081). Expression constructs were co-transformed into Arabidopsis protoplasts. Expression of the fusion constructs was monitored by fluorescence microscopy using a Zeiss LSM510 confocal microscope (Carl Zeiss, Jena, Germany).
Chromatin immunoprecipitation (ChIP) assays
The epitope-tagged transgenic plant samples were cross-linked with 1% formaldehyde, ground to powder in liquid nitrogen, and then sonicated (15-s ON/15-s OFF for seven times, each with 5 min). The sonicated chromatin complexes were bound with corresponding antibodies. Anti-MYC (05–724, Millipore, Billerica, USA), anti-H3ac (06–599, Millipore, Billerica, USA), anti-HA (ab9110, Abcam, Cambridge, USA), and salmon sperm DNA/protein A agarose beads (16–157, Millipore, Billerica, USA) were used for chromatin immunoprecipitation. The immunoprecipitated chromatin complexes were purified by incubating with 50 μl slurry of Protein-A Sepharose (GE, 17–5280–01) pre-equilibrated with 1 mg/ml salmon sperm DNA and 1 mg/ml BSA for 3 h at 4 °C. The affinity beads bound chromatin complexes were washed with 850 μl nuclei lysis buffer for three times, LNDET buffer (0.25 M LiCl, 1% NP40, 1% sodium deoxycholate, 1 mM EDTA, pH8.0) for three times and TE buffer (10 mM Tris–HCl, 1 mM EDTA, pH8.0) for three times. Chromatin complexes were eluted from the beads with use of 200 μl elution buffer (1% SDS, 0.1 M NaHCO3) followed by the digestion with 2.5 ml Proteinase-K (20 mg/ml, Invitrogen, Carlsbad, CA) for overnight at 65 °C. DNA was purified using phenol/chloroform/isoamyl alcohol and sodium acetate (pH 5.2). The level of precipitated DNA fragments was quantified by quantitative real-time PCR using specific primer sets (Supplementary Table 2). Values were normalized to an internal control (eIF4a). Values for control plants were set to 1 after normalization against eIF4a for quantitative PCR analysis.
Transient expression assays
For transient expression assays using Arabidopsis protoplasts, reporter and effector plasmids were constructed. The reporter plasmid contains a minimal 35S promoter sequence and the GUS-coding gene. The TOC1 promoter was inserted into the reporter plasmid. The coding regions of HDA9, ELF3, ELF4, and LUX were subcloned into the effector vector for transient overexpression under the control of the CaMV 35 S promoter. An amount of 2.5 × 105 protoplasts was co-transfected with 4 μg reporter construct, 4 μg effector construct, and 2 μg CaMV 35 S promoter-luciferase construct as the internal transfection control. Transfected cells were incubated at 22 °C overnight and harvested for LUC and GUS reporter assays.
Reporting Summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All relevant data are available from the corresponding author on request.
References
Alabadi, D. et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293, 880–883 (2001).
Gendron, J. M. et al. Arabidopsis circadian clock protein, TOC1, is a DNA-binding transcription factor. Proc. Natl Acad. Sci. USA 109, 3167–3172 (2012).
Nakamichi, N. et al. PSEUDO-RESPONSE REGULATORS 9, 7, and 5 are transcriptional repressors in the Arabidopsis circadian clock. Plant Cell 22, 594–605 (2010).
Farre, E. M., Harmer, S. L., Harmon, F. G., Yanovsky, M. J. & Kay, S. A. Overlapping and distinct roles of PRR7 and PRR9 in the Arabidopsis circadian clock. Curr. Biol. 15, 47–54 (2005).
Helfer, A. et al. LUX ARRHYTHMO encodes a nighttime repressor of circadian gene expression in the Arabidopsis core clock. Curr. Biol. 21, 126–133 (2011).
Chow, B. Y., Helfer, A., Nusinow, D. A. & Kay, S. A. ELF3 recruitment to the PRR9 promoter requires other evening complex members in the Arabidopsis circadian clock. Plant Signal. Behav. 7, 170–173 (2012).
Dixon, L. E. et al. Temporal repression of core circadian genes is mediated through EARLY FLOWERING 3 in. Arab. Curr. Biol. 21, 120–125 (2011).
Locke, J. C. et al. Experimental validation of a predicted feedback loop in the multi-oscillator clock of Arabidopsis thaliana. Mol. Syst. Biol. 2, 59 (2006).
Xie, Q. et al. LNK1 and LNK2 are transcriptional coactivators in the Arabidopsis circadian oscillator. Plant Cell 26, 2843–2857 (2014).
Farinas, B. & Mas, P. Functional implication of the MYB transcription factor RVE8/LCL5 in the circadian control of histone acetylation. Plant J. 66, 318–329 (2011).
Lau, O. S. et al. Interaction of Arabidopsis DET1 with CCA1 and LHY in mediating transcriptional repression in the plant circadian clock. Mol. Cell 43, 703–712 (2011).
Petrillo, E., Sanchez, S. E., Kornblihtt, A. R. & Yanovsky, M. J. Alternative splicing adds a new loop to the circadian clock. Commun. Integr. Biol. 4, 284–286 (2011).
Mas, P., Kim, W. Y., Somers, D. E. & Kay, S. A. Targeted degradation of TOC1 by ZTL modulates circadian function in Arabidopsis thaliana. Nature 426, 567–570 (2003).
Seo, P. J. & Mas, P. Multiple layers of posttranslational regulation refine circadian clock activity in Arabidopsis. Plant Cell 26, 79–87 (2014).
Shu, H., Wildhaber, T., Siretskiy, A., Gruissem, W. & Hennig, L. Distinct modes of DNA accessibility in plant chromatin. Nat. Commun. 3, 1281 (2012).
Eberharter, A. & Becker, P. B. Histone acetylation: a switch between repressive and permissive chromatin. Second in review series on chromatin dynamics. EMBO Rep. 3, 224–229 (2002).
Boycheva, I., Vassileva, V. & Iantcheva, A. Histone acetyltransferases in plant development and plasticity. Curr. Genom. 15, 28–37 (2014).
Gorisch, S. M., Wachsmuth, M., Toth, K. F., Lichter, P. & Rippe, K. Histone acetylation increases chromatin accessibility. J. Cell. Sci. 118, 5825–5834 (2005).
Yang, X. J. & Seto, E. HATs and HDACs: from structure, function and regulation to novel strategies for therapy and prevention. Oncogene 26, 5310–5318 (2007).
Perales, M. & Mas, P. A functional link between rhythmic changes in chromatin structure and the Arabidopsis biological clock. Plant Cell 19, 2111–2123 (2007).
Malapeira, J., Khaitova, L. C. & Mas, P. Ordered changes in histone modifications at the core of the Arabidopsis circadian clock. Proc. Natl Acad. Sci. USA 109, 21540–21545 (2012).
Kim, Y. J. et al. POWERDRESS and HDA9 interact and promote histone H3 deacetylation at specific genomic sites in. Arab. Proc. Natl Acad. Sci. USA 113, 14858–14863 (2016).
Chen, X. et al. POWERDRESS interacts with HISTONE DEACETYLASE 9 to promote aging in Arabidopsis. eLife 5, https://doi.org/10.7554/eLife.17214 (2016).
Hung, F. Y. et al. The Arabidopsis LDL1/2-HDA6 histone modification complex is functionally associated with CCA1/LHY in regulation of circadian clock genes. Nucleic Acids Res. https://doi.org/10.1093/nar/gky749 (2018).
Li, H. et al. The histone deacetylase inhibitor trichostatin a promotes totipotency in the male gametophyte. Plant Cell 26, 195–209, https://doi.org/10.1105/tpc.113.116491 (2014).
Liu, X. et al. Transcriptional repression by histone deacetylases in plants. Mol. Plant 7, 764–772 (2014).
Ezer, D. et al. The G-Box transcriptional regulatory code in. Arab. Plant Physiol. 175, 628–640 (2017).
Michael, T. P. & McClung, C. R. Phase-specific circadian clock regulatory elements in Arabidopsis. Plant Physiol. 130, 627–638 (2002).
Ko, J. H. et al. Growth habit determination by the balance of histone methylation activities in Arabidopsis. EMBO J. 29, 3208–3215 (2010).
Ezer, D. et al. The evening complex coordinates environmental and endogenous signals in Arabidopsis. Nat. Plants 3, 17087 (2017).
Song, H. R. & Noh, Y. S. Rhythmic oscillation of histone acetylation and methylation at the Arabidopsis central clock loci. Mol. Cells 34, 279–287 (2012).
Lu, S. X. et al. The Jumonji C domain-containing protein JMJ30 regulates period length in the Arabidopsis circadian clock. Plant Physiol. 155, 906–915 (2011).
Jones, M. A. et al. Jumonji domain protein JMJD5 functions in both the plant and human circadian systems. Proc. Natl Acad. Sci. USA 107, 21623–21628 (2010).
Wang, L., Kim, J. & Somers, D. E. Transcriptional corepressor TOPLESS complexes with pseudoresponse regulator proteins and histone deacetylases to regulate circadian transcription. Proc. Natl Acad. Sci. USA 110, 761–766 (2013).
Huang, H. & Nusinow, D. A. Into the evening: complex interactions in the Arabidopsis circadian clock. Trends Genet. 32, 674–686 (2016).
Yu, C. W. et al. HISTONE DEACETYLASE6 interacts with FLOWERING LOCUS D and regulates flowering in Arabidopsis. Plant Physiol. 156, 173–184 (2011).
van Zanten, M. et al. HISTONE DEACETYLASE 9 represses seedling traits in Arabidopsis thaliana dry seeds. Plant J. 80, 475–488 (2014).
Kim, W., Latrasse, D., Servet, C. & Zhou, D. X. Arabidopsis histone deacetylase HDA9 regulates flowering time through repression of AGL19. Biochem. Biophys. Res. Commun. 432, 394–398 (2013).
Kang, M. J., Jin, H. S., Noh, Y. S. & Noh, B. Repression of flowering under a noninductive photoperiod by the HDA9-AGL19-FT module in. Arab. New Phytol. 206, 281–294 (2015).
Gu, X., Wang, Y. & He, Y. Photoperiodic regulation of flowering time through periodic histone deacetylation of the florigen gene. Ft. PLoS Biol. 11, e1001649 (2013).
Hicks, K. A., Albertson, T. M. & Wagner, D. R. EARLY FLOWERING3 encodes a novel protein that regulates circadian clock function and flowering in Arabidopsis. Plant Cell 13, 1281–1292 (2001).
Kikis, E. A., Khanna, R. & Quail, P. H. ELF4 is a phytochrome-regulated component of a negative-feedback loop involving the central oscillator components CCA1 and LHY. Plant J. 44, 300–313 (2005).
Cigliano, R. A. et al. Histone deacetylase AtHDA7 is required for female gametophyte and embryo development in. Arab. Plant Physiol. 163, 431–440 (2013).
Acknowledgements
We would like to thank Dr. Clara Conicella for providing hda7–2 mutant. This work was supported by the Basic Science Research (NRF-2016R1D1A1B03931139) and Basic Research Laboratory (NRF-2017R1A4A1015620) programs provided by the National Research Foundation of Korea and by the Next-Generation BioGreen 21 Program (PJ01314501) provided by the Rural Development Administration, by research grants from the Spanish Ministry of Economy and Competitiveness, from the Generalitat de Catalunya (AGAUR), from the Global Research Network of the National Research Foundation of Korea, from the European Commission Marie Curie Research Training Network (ChIP-ET) to P.M. and by the CERCA Programme/Generalitat de Catalunya. We acknowledge financial support from the Spanish Ministry of Economy and Competitiveness, through the Severo Ochoa Programme for Centers of Excellence in R&D 2016–2019 (SEV‐2015‐0533).
Author information
Authors and Affiliations
Contributions
P.J.S. and P.M. conceived and designed the experiments. P.J.S. and P.M. wrote the paper. K.L. conducted experiments and contributed to the study design under the supervision of P.J.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, K., Mas, P. & Seo, P.J. The EC-HDA9 complex rhythmically regulates histone acetylation at the TOC1 promoter in Arabidopsis. Commun Biol 2, 143 (2019). https://doi.org/10.1038/s42003-019-0377-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-019-0377-7
This article is cited by
-
Diurnal RNAPII-tethered chromatin interactions are associated with rhythmic gene expression in rice
Genome Biology (2022)
-
Aschoff’s rule on circadian rhythms orchestrated by blue light sensor CRY2 and clock component PRR9
Nature Communications (2022)
-
Modulation of evening complex activity enables north-to-south adaptation of soybean
Science China Life Sciences (2021)
-
Histone acetylation dynamics regulating plant development and stress responses
Cellular and Molecular Life Sciences (2021)
-
Functions and mechanisms of plant histone deacetylases
Science China Life Sciences (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.