Abstract
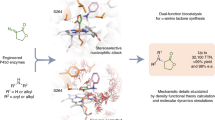
Ketones are crucial intermediates in synthesis and frequent moieties in many products. The direct regioselective synthesis of ketones from internal alkenes could simplify synthetic routes and solve a long-standing challenge in catalysis. Here we report the laboratory evolution of a cytochrome P450 enzyme for the direct oxidation of internal arylalkenes to ketones with several thousand turnovers. This evolved ketone synthase benefits from 15 crucial mutations, most of them distal to the active site. Computational analysis revealed that all these mutations collaborate to generate and tame a highly reactive carbocation intermediate. This is achieved through a confined, rigid, and geometrically and electrostatically preorganized active site. The engineered enzyme exploits a metal–oxo species for ketone synthesis and enables various challenging alkene functionalization reactions. This includes the catalytic, enantioselective oxidation of internal alkenes to ketones and formal asymmetric hydrofunctionalizations of internal alkenes in combination with other biocatalysts.

Similar content being viewed by others
Main
Ketones are essential structural motifs in pharmaceuticals, agrochemicals, materials and natural products. They are also versatile reactants in a wide range of reactions, for example, in the synthesis of chiral alcohols and amines1. Thus, efficient catalytic methods to produce ketones have long been sought. Selective ketone synthesis via the simple aerobic oxidation of internal alkenes would represent a powerful addition to synthetic organic chemistry. This is because internal alkenes are easily accessible from petroleum and renewable resources as well as from well-established reactions such as olefin metathesis2 and carbonyl olefination3. Due to the lack of efficient and selective catalysts, the hydroboration–oxidation method using toxic and expensive boranes is still widely used4,5. In addition to methods using stoichiometric reagents, various catalytic strategies to access ketones from internal alkenes have been developed. In particular, attempts have been made to expand the well-studied Wacker–Tsuji oxidation from terminal to internal alkenes6,7,8,9. However, internal alkenes are less reactive in such palladium-catalysed Wacker-type oxidations, resulting in high catalyst loadings and consequently low total turnover numbers (<20). In addition to activity, selectivity is also a major challenge in catalyst development (Fig. 1a). High regioselectivities in Wacker-type oxidation reactions depend on specific internal alkenes bearing directing groups that favour the formation of one ketone over the other. Many of these protocols are not aerobic oxidations or depend on stoichiometric reagents such as peroxides and phenylsilanes10,11. A recently published dual catalytic approach used water as the terminal oxidant and produced hydrogen gas as a by-product12. In general, efficient and regioselective aerobic oxidation of internal alkenes to ketones is not only a considerable challenge in catalyst development, but is also highly desired as it has the potential to streamline the synthesis of many important molecules.
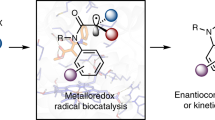
a, Regioselectivity is a challenge in the catalytic oxidation of internal alkenes to ketones. b, Design of a catalytic process to convert internal alkenes into ketones using high-valent metal–oxo species as the catalytic oxidant. High-valent metal–oxo species typically react with alkenes in epoxidation reactions. The ketone product is accessible by stabilizing a carbocation intermediate, and the reaction proceeds via a coupled electron–hydride transfer. c, Given that enzymatic alkene to ketone oxidation can be accessed, many challenging functionalization reactions of internal alkenes can be realized in combination with established biocatalysts such as ketoreductases39,40 (i), ω-transaminases39 (ii) and imine reductases41,42 (iii).
We have recently started to explore the potential of enzymes to directly oxidize alkenes to carbonyl compounds13,14. In an initial study, we applied directed evolution to engineer an enzyme that oxidizes terminal alkenes to the corresponding aldehydes, attaining opposite selectivity to the widely used Wacker–Tsuji oxidation reaction. The evolved iron haem-based biocatalyst achieved aldehyde formation by controlling an oxo transfer reaction using a high-valent metal–oxo complex as the catalytic oxidant (Fig. 1b). Many catalysts comprising such metal–oxo complexes, including cytochrome P450s15,16, peroxygenases17 and Jacobsen’s catalyst18,19, are used to efficiently epoxidize alkenes via highly concerted reaction pathways. The key to harnessing oxo transfer for carbonyl-selective alkene oxidation is to create a catalyst that is able to intercept the epoxidation and couple oxo transfer with an electron–hydride transfer process (Fig. 1b). This can be achieved by accessing a highly reactive carbocation intermediate and making it available for organic synthesis. While this is currently out of reach for small-molecule catalysts, our recent study13 showed that reactive centres in enzymes can be optimized with high precision to fully access and exploit such reactive carbocation intermediates. Recent mechanistic studies on this oxo transfer reaction revealed that alkene epoxidation is a dynamically favoured process and that carbonyl formation can be accomplished by controlling the accessible conformations of the radical and carbocation intermediates14. In addition, electrostatic preorganization of the enzyme active site favours the formation of this key carbocation intermediate14.
Here we report a generalizable approach to develop enzymes for the aerobic oxidation of alkenes to ketones. In particular, we have expanded the enzymatic alkene-to-carbonyl oxidation from terminal to internal arylalkenes. This generates ketones with high activity and regioselectivity in a direct aerobic oxidation reaction. Key aspects of this study are the directed enzyme evolution of a ketone synthase as well as computational studies to rationalize and propose how more than a dozen beneficial mutations collaborate to access this catalytic cycle. Furthermore, we applied the ketone synthase in synthesis as a stand-alone catalyst to generate various benzyl ketones as well as in combination with established biocatalysts in cascade reactions. The latter application enabled the formal asymmetric hydration and hydroamination (Fig. 1c) of internal alkenes, reactions that are particularly sought after with only limited catalytic solutions (Supplementary Figs. 2 and 3)20,21,22,23,24,25.
Results
Directed enzyme evolution
To explore whether the enzyme-controlled metal–oxo-mediated mechanism can be harnessed for efficient and selective ketone synthesis, we set out to evolve a promiscuous enzyme. We first aimed to identify a starting point for directed evolution by determining the activity and selectivity of various cytochrome P450 monooxygenases in the oxidation of internal alkenes. In particular, we focused on an in-house mutant library of a cytochrome P450 monooxygenase from Labrenzia agreggata (P450LA1), which has been recently evolved to oxidize terminal alkenes to aldehydes13. trans-β-Alkylstyrenes have been chosen as model substrates because such internal arylalkenes are particularly challenging substrates in Wacker-type oxidation, yielding mixtures of ketone products7,26. Indeed, we recently confirmed the initial activity with this substrate type13, demonstrating that the coupled electron–hydride transfer process (Fig. 1b) is accessible for ketone synthesis.
The variant P7E (P450LA1-T121A-V123I-N201K-H206W-N209S-I326V-Y385H-E418G) was selected from the initial screening as it showed reasonable selectivity and the highest level of product formation under the screening conditions (Supplementary Fig. 4). Using trans-β-methylstyrene as the substrate, phenylacetone was produced with a total turnover number (TTN) of 225; however, the performance of the catalyst was limited by the low selectivity of the reaction (Fig. 2a). In addition to phenylacetone (31%), epoxidation (67%) and allylic oxidation (2%) were also observed. It is worth noting that variant P7E did not produce the regioisomeric ketone propiophenone (Supplementary Fig. 5).
a, Activity and selectivity as a function of evolution starting with wild-type P450LA1 (Wt). Biotransformations were performed with 0.625 µM P450 enzyme variant, 5 mM trans-β-methylstyrene (1), 5 mM NADH cofactor and 1 vol% isopropanol in reaction buffer. The reactions were shaken at room temperature for 2 h before UHPLC analysis. The TTN was calculated by dividing the concentration of the ketone produced by the enzyme concentration used. TTN data are shown as a bar graph as a mean of six experiments (experimental triplicates of biological duplicates, n = 6). The individual data points are shown as white circles. The ketone selectivity was determined as the proportion of ketone in relation to all oxidation products formed (including epoxides and allylic oxidation products) and the data points are shown as black dots. b, Trajectory of the directed evolution experiment, showing the amino acid substitutions in each round. Inset: key data regarding the entire directed evolution experiment starting from Wt. c, Epoxides are not intermediates in the reaction as they are not converted by the ketone synthase via isomerization into the corresponding ketones. d, Homology model of P7E with the mutations introduced to generate the ketone synthase shown as purple spheres. The haem cofactor is shown as black sticks.
We next aimed to optimize the activity and selectivity of P7E by directed evolution. Eleven rounds of evolution uncovered twelve beneficial mutations to convert P7E into an efficient ketone synthase (Fig. 2a,b). The directed evolution experiment involved mainly site-saturation mutagenesis using the 22c approach27 as well as combinatorial mutagenesis libraries (Supplementary Tables 1 and 2). In each round of directed evolution, usually six to eight amino acid positions of the haem domain were randomized individually, and high-throughput ultrahigh-performance liquid chromatography (UHPLC) analysis was used to identify the beneficial mutations (Supplementary Fig. 6). Interestingly, the hit rate was rather low. Typically, we could identify only one or two beneficial mutations per round of evolution that showed enhanced activity as well as selectivity. While the introduced mutations often altered enzyme activity, mutations that changed the oxo-transfer mechanism towards ketone formation were more difficult to identify. Thus, during the directed evolution, we moved from randomizing amino acids that build the first shell of the active site towards amino acids in the second and third shells. We stopped the directed evolution campaign after 12 rounds of evolution, having addressed a total of 78 amino acid positions at least once, which corresponds to 18% of the amino acid sequence of the haem domain. The final variant was named ketone synthase (KS) and produced phenylacetone with high activity (2,630 TTN) and chemoselectivity (75%) as well as with complete regiocontrol. The allylic oxidation of trans-β-methylstyrene was a further competing reaction pathway in the course of directed evolution, however, the epoxide was the only by-product observed using the final KS variant (Supplementary Figs. 7 and 8). Control experiments confirmed that epoxides were not isomerized by KS (Fig. 2c and Supplementary Fig. 9), highlighting that epoxides are not intermediates in the reaction and supporting that catalysis proceeds via the proposed coupled electron–hydride transfer mechanism. The evolved KS is 12-fold more active than the parent P7E variant and two orders of magnitude more efficient than previously reported catalysts (Supplementary Fig. 1).
The KS variant carries 18 mutations compared with the wild-type enzyme and 12 compared with P7E. Careful analysis revealed that 15 out of these 18 mutations are crucial to favour the ketone-forming pathway (Fig. 2a and Supplementary Fig. 4). Interestingly, the beneficial mutations are not limited to the active site. Many selectivity-determining mutations were introduced in the second and third shells, and their individual contributions add up to facilitate access to the ketone product (Fig. 2d). In addition to additive mutations, we observed cooperative effects that can be well visualized by following position 210, which changed during the evolution from threonine to valine to isoleucine and back to valine, always contributing to increases in selectivity and activity (Fig. 2b)28. Overall, there was no single mutation that changed the chemoselectivity in the reaction by more than 8%.
Mechanistic insights from computational modelling
To unravel the molecular basis of the evolved KS, we performed computational modelling. In particular, we aimed to describe the enzymatic reaction mechanism and explain how the multiple distal mutations contribute to fully enable the carbonyl-selective reaction pathway.
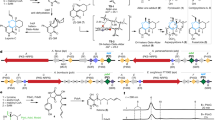
We first used density functional theory (DFT) to analyse the intrinsic reaction mechanism and geometric requirements using a computational truncated model (Fig. 3a; see Supplementary Methods and Supplementary Computational Part I for complete mechanistic descriptions). The calculations indicated that, as previously found for styrene and the formation of the corresponding aldehyde13,14, the epoxidation and ketone formation from substrate 1 share a common first step in which the first C–O bond is formed (TS1; Fig. 3a and Supplementary Fig. 19). In the doublet electronic state (TS1d, Gibbs free energy difference between reactant and transition state (ΔG‡) is 8.3 kcal mol−1), epoxide 3 is formed directly from TS1 via the formation of a second C–O bond; in the quartet electronic state (TS1q, ΔG‡ = 9.2 kcal mol−1, relative Gibbs free energy (ΔΔG) is 0.9 kcal mol−1 higher than TS1d), a covalent radical intermediate is formed (Int1q, Gibbs free energy difference (ΔG) is −13.7 kcal mol−1). This radical intermediate Int1q (Fig. 3a) can follow the epoxidation pathway, forming the second C–O bond through a low-energy TS (TS2q, ΔG‡ = 1.6 kcal mol−1), or form a covalent carbocation intermediate via intramolecular electron transfer from the substrate moiety to the porphyrin Fe (Int2q and Int2d, ΔG = 1.7 and −2.2 kcal mol−1, respectively). Int2d can be formed via a minimum-energy crossing point (MECP) from Int2q (Fig. 3a and Supplementary Fig. 23). In this truncated model, the formation of carbocation Int2 is triggered by a conformational change, namely the rotation of the benzyl group of the substrate (Fig. 3b and Supplementary Fig. 21). This allows optimal stabilization of the benzylic carbocation by maximizing hyperconjugation with the neighbouring σ(C–H) and σ(C–C) bonds as well as resonance with the aromatic ring, and by n→p interactions between the oxygen lone pairs and the empty carbocation p orbital (Fig. 3b and Supplementary Fig. 22)14. In this geometry, an intramolecular hydrogen bond between the ortho hydrogen of the phenyl ring and the oxygen atom is also established. Once formed, the carbocation within this conformation has its empty p orbital well aligned with the H atom, which can undergo a barrierless 1,2-hydride migration to form the final ketone product 2 in both the doublet and quartet electronic states. DFT calculations indicated that the carbonyl pathway can energetically compete with the epoxidation pathway as both are very similar in energy (TS1d versus TS1q), and that catalytic access to the ketone product depends on stabilizing the reactive intermediates in a specific conformation.
a, Calculated energy profile (ΔG and electronic energy (ΔE)) for the competing epoxidation and ketone formation pathways for trans-β-methylstyrene (1) using a computational truncated model. The energies of the doublet and quartet electronic states are reported in kcal mol−1. b, DFT-optimized structures for the radical (Int1) and carbocation (Int2) intermediates, and their ΔΔG. The key distances, dihedral angle (highlighted in orange) and spin densities (ρ) are given in ångstroms (Å), degrees (°) and atomic units (a.u.), respectively.
We previously found that intrinsic dynamic effects have an important influence on the reaction pathway after transition state TS1 and the formation of the first C–O bond14. The covalent radical intermediate Int1q formed from TS1 is kinetically activated, possessing a momentum vector that matches that of the epoxide-forming transition state TS2 (Supplementary Fig. 24). The intrinsic strong preference for epoxide formation is proposed to be a consequence of the dynamic behaviour of the radical intermediate, which is formed with an excess of energy that is not statistically distributed before further reacting14. This strong dynamic match favours alkene epoxidation over carbonyl formation, which requires a specific carbocation geometry to promote the 1,2-hydride migration (Fig. 3b). Consequently, we hypothesized that the carbonyl selectivity achieved by the evolved KS derives from conformational control over the intermediates due to geometric restrictions in the enzyme active site14.
To study the enzymatic reaction, homology models for the parent P7E and evolved KS variants were first constructed (see Supplementary Methods and Supplementary Computational Part II for modelling details). Extensive molecular dynamics (MD) simulations (five independent replicas of 1,000 ns each, 5,000 ns in total for each system) revealed important differences between the overall structures of KS and the parent enzyme P7E that affect the shape of the active-site cavity (Supplementary Fig. 29). Visual inspection of the active site in the evolved KS reveals a more confined substrate binding pocket compared with the parent P7E (Fig. 4a and Supplementary Fig. 30). This reshaped active site is expected to improve the binding of substrate 1 in a reactive conformation and might also be important to control the conformations of the subsequently formed reactive intermediates.
a, Arrangement of the active site in variants P7E and KS characterized from MD simulations in their holo states. The cavity surface is shown in purple. b, Representative structure obtained from restrained MD simulations of trans-β-methylstyrene (1) bound in a reactive NAC in the KS active site. c, Analysis of active-site residues from accumulated MD trajectories. Top: increased rigidification from holo to substrate-bound states. ΔRMSF describes the root-mean-square fluctuation (RMSF) measured for active-site residues along NAC substrate-bound MD simulations compared with holo state simulations (RMSF(substrate bound) – RMSF(holo)). The more negative the value of ΔRMSF, the more rigid (that is less mobile) the residue becomes after binding the substrate in the simulated NAC. Bottom: interaction energies of individual residues and substrate bound in a NAC conformation. MM-GBSA substrate–residue pair interaction energies describe the strength of the interactions between trans-β-methylstyrene and surrounding active-site residues along the NAC substrate-bound MD simulations. Orange boxes highlight major changes in the flexibility of residues (ΔRMSF) and relevant interactions occurring between substrate and active site residues (MM-GBSA). d, Calculated QM/MM energy profile (ΔG and ΔE; the KS-wat model includes an explicit water molecule in the QM region, see Supplementary Computational Part II for details) for the competing epoxidation and ketone formation pathways for trans-β-methylstyrene (1) catalysed by KS (see also Supplementary Fig. 41). The energies of the doublet and quartet electronic states are reported in kcal mol−1. e, QM/MM optimized geometries for the key radical intermediate (KS-wat-Intlq) and carbocation intermediate (KS-wat-Int2q) formed in the KS active site. ΔΔG and key distances are given in kcal mol−1 and ångstroms (Å), respectively.
Next, we aimed to characterize the reactive binding conformations of the substrate in the P7E and KS active sites. We used substrate 1 in docking calculations, which were refined by extensive restrained MD simulations (five replicas of 500 ns, 2,500 ns in total) to characterize reactive near-attack conformations (NACs) leading to the key TS1 (see Supplementary Methods and Supplementary Computational Part II for modelling details). The simulations revealed that the KS shows improved substrate binding in relevant reactive NAC conformations due to a more packed active site compared with P7E (Fig. 4b and Supplementary Figs. 31 and 32). This increased packing is reflected in two distinct observations. First, a more pronounced rigidity was observed for the KS active-site residues upon substrate binding in the NAC conformation (Fig. 4c, ΔRMSF). This is especially highlighted by residues 121–124 of the B′C loop region and residues 273–284 in the I helix (Fig. 4c and Supplementary Fig. 33). Second, stronger interactions occur in the KS active site between the substrate and surrounding residues (Fig. 4c and Supplementary Fig. 34). The substrate bound in a reactive conformation shows stronger interactions with active-site residues L97, A121V, W211, A275 and I276 for KS compared with P7E. All these residues are located around the phenyl ring of the substrate and consequently restrict its accessible conformations (Fig. 4b).
The enhanced active-site packing also plays a crucial role in controlling the radical intermediate formed from TS1, as supported by quantum mechanics/molecular mechanics (QM/MM) calculations carried out on the evolved KS (Fig. 4d and Supplementary Fig. 41). First, tight interactions with surrounding residues help the intermediate to dissipate its excess vibrational energy and achieve thermal equilibration once formed14. The QM/MM-optimized radical intermediate formed in the KS active site (KS-wat-Int1q) has the phenyl group tightly packed between the same amino acids of the B′C loop region and I helix as identified by NAC analysis of the substrate (Fig. 4e). Second, active-site tightening and steric interactions between the radical intermediate and the active-site residues facilitate access to the geometry required for effective carbocation formation (KS-wat-Int2q) Fig. 4e and Supplementary Fig. 42 for an analysis of the required conformational change). This geometry allows stabilization of the carbocation intermediate due to stereoelectronic effects and aligns the cis-β-hydrogen atom with the empty carbon p orbital for a fast and selective 1,2-hydride migration. These steric interactions also increase the barrier for the second C–O bond formation (KS-wat-TS2), disfavouring the epoxidation pathway (Fig. 4e and Supplementary Fig. 41). These interactions mainly involve hydrophobic contacts between the substrate and active-site residues A275 and V278 (I helix), A121 and I123 (B′C loop), L97 (BB′ loop), and W329 (β1 region), which contribute to maintaining a tight packing of the substrate and intermediates along the reaction coordinate.
In addition to the geometric and steric effects exerted by the enzyme active site, the role of electronic and electrostatic effects was also analysed. Our calculations indicated that carbocation formation is also favoured by electronic effects due to polar interactions in the active site and electrostatic preorganization of the catalytic pocket. Computational modelling revealed the presence of an ordered water molecule in the active site, which, in the presence of the substrate, interacts via hydrogen bonds with T283 and the oxygen atom of the Fe–oxo complex (Supplementary Fig. 36). This ordered water molecule is reminiscent of the activation of molecular oxygen to generate the iron–oxo active species29,30. Computations showed that the presence of this hydrogen-bonded water molecule can promote the formation of these radical and carbocation intermediates (Supplementary Figs. 27 and 28)14. Additionally, computational modelling indicated that the electrostatic preorganization of the active site also contributes to promote carbocation formation and its stabilization relative to the radical intermediate in both electronic states (Supplementary Figs. 39 and 41), in line with our previous findings14. This is in part due to the local electric field that is generated in the enzyme active site, which has a major projection on the Fe−O axis and follows the Fe to O direction (Supplementary Figs. 25, and 26).
In summary, computational modelling suggests that the electrostatic preorganization of the active site, the polar interaction with an ordered water molecule and the strong conformational control imposed by the KS active site are responsible for ketone formation. This precise catalytic control promotes carbocation formation and destabilizes the epoxide-forming transition state compared with the truncated, enzyme-free, model DFT calculations.
Role of distal mutations on active site preorganization
To rationalize how numerous distal mutations (Fig. 2d) optimized the active site, we studied the enzyme conformational dynamics in variants P7E and KS using dynamic correlation networks. Dynamic networks describe connections between distal protein residues based on their correlated dynamic behaviour mediated by neighbouring residues (in sequence or three-dimensional space). It has been proposed that dynamic networks can modulate the transition between active and inactive conformations of enzymes31 and impact active-site preorganization by controlling enzyme conformational flexibility and reducing non-productive conformations32.
Analysis of the holo enzymes revealed the emergence of an expanded dynamic network in KS compared with P7E. Shortest path maps (SPMs)33, which describe communication between neighbouring residues, show that the new enlarged dynamic network in KS is centred on the substrate binding pocket and completely surrounds it (Fig. 5 and Supplementary Fig. 37). The expanded dynamic network includes residues in the I helix and the 116–124 B′C loop region that were not part of the original network in P7E, but are important for the conformational control of the substrate and intermediates (Fig. 4c), as described earlier. The enhancement and tuning of the protein dynamic network can be attributed to the newly introduced mutations, which is in line with previous studies31,32,33,34, including important epistatic effects, as in related cytochrome P450s that directly impact catalysis28. Although only four mutations appear as nodes in the SPMs in the KS, practically all of the introduced mutations, including the distal ones, appear at adjacent positions in three-dimensional space (Supplementary Figs. 37 and 38), thus contributing to the protein dynamic behaviour31,32,33,34. The enhanced dynamic network in the KS is not only maintained, but strengthened in the reactive substrate-bound state (Fig. 5 and Supplementary Fig. 38), similarly to what has been observed in other laboratory-evolved enzymes32,35. Taking all this into account, the observed reshaping and rigidification of the active site in the KS can be attributed to the changes in the dynamic network of the evolved enzyme due to distal mutations.
SPMs33 calculated from accumulated MD simulation time for variants P7E and KS in their holo and ‘substrate bound in a NAC conformation’ states. The sizes of the spheres (nodes of the network) and black lines (edges of the network) are indicative of their weight in the network. Altered amino acid residues during evolution are shown in stick format and highlighted in orange. Haem cofactor and substrate 1 are shown in stick format, in grey and cyan, respectively. Insets: active site with bound substrate.
Substrate scope and application in synthesis
To explore the potential applications of such a KS, we studied the substrate scope of this enzyme and performed reactions on a preparative scale. The KS with its confined active site accepted various structurally related trans-β-methylstyrene derivatives and produced the corresponding ketones with efficiencies of up to several thousand TTNs and selectivities of up to 79% (Fig. 6a). Various substitutions are tolerated, including substitutions in the ortho (11), meta (10) and para (4–8) positions of the aromatic ring as well as in the trans-β-alkenic position (9). We also tested various structurally more unrelated internal alkenes, but these were not accepted as substrates by the highly optimized active site (Supplementary Fig. 10). As an example, cis-β-methylstyrene was converted by the KS into epoxides but not phenylacetone (Supplementary Fig. 11). This agrees with recent mechanistic findings14 on the evolved cytochrome P450 that oxidized terminal alkenes to aldehydes13, highlighting that hydride migration during the catalytic cycle of these evolved enzymes based on P450LA1 proceeds selectively from the cis position. This also supports our conclusions that catalysts optimized for metal–oxo-mediated alkene-to-ketone oxidation require a highly confined and preorganized reactive centre.
a, Substrate scope of the KS reaction. Reactions were carried out using 0.625 µM KS, 5 mM of the corresponding substrate and 5 mM NADH cofactor. TTN values were determined after 2 h reaction time. aMaximum TTN was determined after 48 h biotransformation combined with a cofactor regeneration system. b, KS application on a preparative scale for the synthesis of ketones as well as in the formal asymmetric hydration and hydroamination of internal alkenes. See Supplementary Figs. 16 and 17 for more details. e.r., enantiomeric ratio; GDH, glucose dehydrogenase; PAR, phenylacetaldehyde reductase; LBv-ADH, alcohol dehydrogenase from Lactobacillus brevis; IRED, imine reductase pIR-23 (Cystobacter ferrugineus (CF)IRED).
Next, we explored substitutions at the α-position to study the enantioselectivity of the enzymatic oxidation reaction. Even though trans-α,β-dimethylstyrene (17) was not used as substrate in the screening, and the enzyme therefore was not selected for enantioselectivity during the directed evolution, the KS enabled the synthesis of the chiral ketone 12 with good enantioselectivity towards the (S) enantiomer (enantiomeric ratio, 87:13, S/R; Fig. 6a and Supplementary Fig. 13). Stereoselectivity is achieved by enantiofacial discrimination during the 1,2-hydride migration after the first C–O bond formation, in line with previous observations for a related engineered enzyme14. The enantioselectivity detected is also consistent with the selectivity model proposed on the basis of the computational modelling carried out on trans-β-methylstyrene (Supplementary Fig. 13). It is worth highlighting that catalytic asymmetric oxidation of internal alkenes to chiral ketones is currently unknown. Further engineering of this cytochrome P450 enzyme or other iron-dependent monooxygenases will reveal whether KSs can be generated with even higher enantioselectivity.
The activity of the KS could be further optimized by changing the reaction conditions. Lengthening the reaction time and application of a cofactor recycling system enabled the conversion of alkene 1 to ketone 2 with a TTN of up to 4,750 (Supplementary Fig. 14). To demonstrate that these reactions can be performed on a preparative scale (1.0 mmol), phenylacetone (2) was synthesized using a catalyst loading of 0.025 mol% KS (Fig. 6b). The product was isolated in 61% yield, with atmospheric oxygen and glucose as the only stoichiometric reagents.
Formal asymmetric hydrofunctionalization of internal alkenes
Laboratory evolution of non-natural enzyme function can provide access to new synthetic pathways and solve long-standing challenges in synthesis36,37,38. In this regard, we performed asymmetric redox hydrations and hydroaminations on internal arylalkenes by combining the KS with ketoreductases39,40 and imine reductases41,42 in cascade reactions on a preparative scale (1.0 mmol). Using this set-up, the unactivated internal arylalkene 1 was converted into chiral phenylethanols and phenylethylamine, which are important structural motifs in top-selling pharmaceuticals (Supplementary Fig. 15). The chiral alcohols 13 and 14 as well as amine 16 were produced with excellent enantioselectivity (enantiomeric ratio of up to >99:1) in isolated yields of 66, 69 and 39%, respectively (Fig. 6b). An important feature of these reactions is that (1) they only depend on simple stoichiometric reagents such as atmospheric oxygen and isopropanol (asymmetric hydration) or atmospheric oxygen and glucose (asymmetric hydroamination) and (2) simple unprotected amines (such as 15) can be used as amine donors. This approach can in principle be expanded to other internal arylalkenes (Fig. 6a) or other amines as amine donors42,43.
In addition, we aimed to generate two stereocentres using the trisubstituted alkene 17 as the starting material (Fig. 7). Alcohol 18 was obtained via ketone 12 with low activity but excellent selectivity, yielding a single stereoisomer. This approach combines the KS-catalysed enantioselective alkene-to-ketone oxidation via an asymmetric hydride migration with diastereofacial differentiation of the ketoreductase.
Alkene 17 was used as a substrate to generate two stereocentres. The enzymatic reaction generates a single stereoisomer of alcohol 18 via ketone 12 and combines enantioselective oxidation by the KS with diastereofacial discrimination by a ketoreductase (PAR). The enzymatic reaction is compared with a chemical synthesis that generates all possible stereoisomers (see Supplementary Fig. 18 for more information).
Such conversions of unactivated internal alkenes to chiral alcohols and amines are a particular challenge in catalysis (Supplementary Figs. 2 and 3) with only limited catalytic solutions20,21,22,23,24,25. There are natural enzymes that catalyse direct hydroaminations of internal alkenes, however, due to their underlying mechanism, they depend on activated internal alkenes such as α,β-unsaturated carboxylic acids44,45. As internal unactivated arylalkenes are easily accessible and ketones are substrates for multiple (bio)catalysts40,42,46,47,48,49,50,51, many new synthetic routes can be envisioned using evolved KSs.
Conclusions
Our findings demonstrate that cytochrome P450 can be evolved to oxidize internal arylalkenes to ketones with high activity and selectivity. We used the evolved KS to directly convert internal arylalkenes into ketones, including a catalytic enantioselective example. In addition, we combined the KS with established biocatalysts to convert internal arylalkenes into chiral phenylethanols and phenylethylamines on a preparative scale with high activity, regio- and enantioselectivity. What stands out is that the KS enables many important, highly selective functionalization reactions of internal arylalkenes that have so far largely eluded efficient and selective catalysis. Similar to many other engineered enzymes that enable new catalytic reactions52,53, the substrate scope is initially limited. However, we envision that further protein engineering and laboratory evolution of the enzyme (or other cytochrome P450s) will expand the substrate scope to various alkenes, as has been demonstrated with many other enzymes that catalyse natural and non-natural chemical reactions.
Engineered enzyme active sites can exert an exceptional degree of control over reactive intermediates53. This is achieved in the KS evolution by generating a confined, rigid and preorganized active site through multiple mutations that enhance a pre-existing dynamic network. The evolved KS provides access to a highly reactive carbocation species and thus enables direct ketone production from high-valent metal–oxo species. We foresee that the high degree of confinement54 in enzyme active sites could be generally leveraged to enable other challenging catalytic cycles by precisely controlling the accessible conformations of reactive intermediates. Furthermore, we also envision that this coupled oxo–electron–hydride transfer process could be fully exploited with less reactive alkyl-substituted alkenes as substrates.
Methods
Cloning and library creation
pET22b(+) was used as a cloning and expression vector for all P450LA1-derived enzyme variants described in this study. Site-saturation mutagenesis libraries were generated using the ‘22c-trick’ method27. Primer sequences are available in Supplementary Data Table 1. The obtained PCR products were digested with DpnI, purified on agarose gel and ligated using Gibson Assembly Master Mix55. The DNA was further purified before transformation into electrocompetent Escherichia coli BL21(DE3) cells (E. cloni EXPRESS BL21(DE3), Lucigen). Typically, 90 transformants were analysed per 22c-trick library, resulting in significant oversampling. In addition, the quality of each variant library was examined by sequencing (quick quality control)27 and only libraries with nucleic acid distributions that reflect the 22 codons used were screened. For combinatorial mutagenesis libraries, primers were designed following the multichange isothermal mutagenesis protocol and the libraries were constructed as described above56.
Expression of cytochrome P450 variant libraries in 96-well format
An overnight culture plate (96-well format) was filled with Terrific Broth (TB) medium (with double the amount of glycerol, 100 μg ml−1 ampicillin, 500 μl well−1) and each well was inoculated with a single colony. The plate was incubated for 20 h at 37 °C and 250 rpm (25 mm orbital) using humidity control. The expression culture was then inoculated with 50 μl well−1 of the overnight culture in 610 μl well−1 TB medium (100 μg ml−1 ampicillin, 2× glycerol) in a fresh 96-well plate and incubated for 4 h at 37 °C and 250 rpm. After the plate had been cooled on ice water for 10 min, expression was induced with 40 μl well−1 induction master mix in TB medium (0.2 mM isopropyl-β-d-thiogalactopyranosid (IPTG), 0.5 mM 5-aminolevulinic acid, final concentration) and incubated for a further 20 h at 25 °C and 250 rpm. The cells were pelleted by centrifugation (3,220g, 10 min, 4 °C) and stored at −20 °C for at least 1 d.
UHPLC screening of cytochrome P450 variant libraries
Lysis buffer (0.1 M Na2HPO4, 0.15 M NaCl, 2% glycerol, pH 8, 1 mg ml−1 lysozyme, 0.2 mg ml−1 DNaseI, 200 μl well−1) was added to the pelleted and frozen cells for lysis. The plate was incubated for 4 h at 4 °C with occasional strong manual shaking (otherwise 400 rpm on a plate shaker). The plate was then centrifuged (3,220g, 10 min, 4 °C) and 150 µl well−1 of the lysate was transferred to another 96-well plate that already contained 10 µl well−1 substrate solution (trans-β-methylstyrene (1) dissolved in 1:1:0.55 dimethylsulfoxide–isopropanol–water, 15 mM final concentration) and 240 µl well−1 NADH buffer solution (5 mM final concentration). The plate was sealed with adhesive polystyrene foil and incubated for 2 h at 25 °C and 400 rpm. Acetonitrile was added (600 μl well−1) and the mixture homogenized before subsequent resting incubation for 30 min. Precipitated protein was pelleted by centrifugation (3,220g, 10 min, 25 °C) and 150 μl well−1 of the supernatant was transferred to a 96-well screening plate by centrifugal filtration (AcroPrep Advance 96 filter plate, 0.2 µm polytetrafluoroethylene, Pall). The screening plate was sealed with heat-sealing aluminium foil and samples were finally submitted for UHPLC analysis.
Small-scale expression of cytochrome P450 variants
The P450LA1-derived enzyme variants with elevated selectivity found in each round of evolution were streaked on LB-based agar medium and incubated overnight at 37 °C. An individual colony was inoculated into the overnight culture in 5 ml LB medium (100 µg ml−1 ampicillin, final concentration) at 37 °C and 180 rpm. Expression cultures were inoculated with 500 µl of preculture into 50 ml TB medium (100 µg ml−1 ampicillin, 2× glycerol, final concentration) in a 250-ml flask without baffles and incubated for 2–3 h at 37 °C and 180 rpm until an optical density at 600 nm (OD600) of 0.6–0.8 was reached. The flask was cooled on ice for 10 min before expression was induced (0.2 mM IPTG, 0.5 mM 5-aminolevulinic acid, final concentration). The induced cells were shaken for 20 h at 25 °C and 180 rpm. The cells were collected (3,220g, 10 min, 4 °C) and stored at −20 °C for at least 1 d.
Biotransformations
The pelleted and frozen cells were lysed in lysis buffer (3 ml per g cell wet weight, 0.1 M Na2HPO4, 0.15 M NaCl, 2% glycerol, pH 8, 1 mg ml−1 lysozyme, 0.2 mg ml−1 DNaseI) for 4 h on ice. After centrifugation (20,238g, 4 °C, 10 min), the enzyme concentration contained in the supernatant was determined by ferrous CO binding difference spectroscopy following an established method57. A defined volume of lysate (0.625 µM final enzyme concentration) was used in biotransformations. Thus, the lysate, NADH buffer solution (80 µl, 5 mM final concentration) and 16 µl substrate solution (trans-β-methylstyrene (1) or another internal alkene in isopropanol, 5 mM final concentration) were mixed in reaction buffer (0.1 M Na2HPO4, 0.15 M NaCl, 2% glycerol, pH 8, 800 µl final volume) in a sealable GC vial, and the mixture was incubated for 2 h at room temperature and 400 rpm (8 mm orbital shaker). An additional volume of acetonitrile (800 µl) was added and the homogenized sample was left to stand for 30 min. After the precipitate had been pelleted by centrifugation (20,238g, 4 °C, 10 min), the supernatant was submitted to UHPLC analysis. The TTN values after 2 or 48 h reaction time were calculated as the ratio of ketone product and cytochrome P450 concentration. The selectivity for the ketone product was calculated according to the following formula: concentration ketone/(concentration ketone + concentration epoxide + concentration cinnamyl alcohol).
Large-scale expression of KS
E. coli BL21(DE3) cells were transformed with plasmid DNA encoding for P450LA1-derived KS and grown overnight in 5 ml LB medium (100 µg ml−1 ampicillin, final concentration) at 37 °C and 180 rpm. Expression cultures were inoculated with 4 ml of preculture into 400 ml TB medium (100 µg ml−1 ampicillin, 2× glycerol, final concentration) in a 2-l flask with baffles and incubated for 2–3 h at 37 °C and 100 rpm until an OD600 of 0.6–0.8 was reached. The flask was cooled on ice for 10 min before expression was induced (0.2 mM IPTG, 0.5 mM 5-aminolevulinic acid, final concentration). Induced cells were shaken for 20 h at 20 °C and 100 rpm. The cells were collected (4,357g, 10 min, 4 °C) and stored at −20 °C for at least 1 d.
DFT calculations
All DFT calculations were performed using the Gaussian 09 software package58. An enzyme-free truncated model was used [Fe=O(Por)(SCH3)(1)] that included the iron–oxo active species of compound I (Fe=O), a porphyrin pyrrole core (Por), a methanethiolate group to mimic the cysteine (Cys) axial ligand and the trans-β-methylstyrene substrate (1). The resulting model had a neutral total charge and two different energetically accessible electronic states, doublet and quartet, were considered. The unrestricted hybrid (U)B3LYP functional59,60,61 was used with an ultrafine integration grid and included the conductor-like polarizable continuum model (CPCM; dichloromethane, relative permittivity ε = 8.9) to estimate the dielectric permittivity in the enzyme active site. The 6-31G(d) basis set was used for all atoms except Fe, for which the SDD basis set and related SDD pseudopotential were employed. All optimized stationary points were characterized as minima using frequency calculations, including transition states that show a single imaginary frequency that describes the corresponding reaction coordinate. Intrinsic reaction coordinate calculations were performed to ensure that the optimized transition states connect the expected reactants and products. Enthalpies and entropies were obtained at 1 atm and 298.15 K. Enthalpy calculations were corrected using the harmonic oscillator approximation, by increasing all frequencies below 100 cm−1 to 100 cm−1 using Goodvibes (v.3.0.1) Python script62. Single-point energy calculations were carried out with the previously described functional ((U)B3LYP with an ultrafine grid and CPCM dichloromethane conductor model) and the Def2TZVP basis set on all atoms, and included empirical Grimme D3 dispersion corrections with Becke–Johnson (D3BJ) damping63.
MD simulations
MD simulations in explicit water were performed using the AMBER18 package64,65. Parameters for the trans-β-methylstyrene (1) substrate were generated within the Antechamber66 module in the AMBER18 package using the general AMBER force field (gaff2) with partial charges set to fit the electrostatic potential generated at the HF/6-31G(d) level by the restrained electrostatic potential model. Parameters for the haem compound I and the axial Cys were taken from ref. 67. The enzyme variants were solvated in a pre-equilibrated cubic box with a 10-Å buffer of transferable intermolecular potential water molecules using the AMBER18 leap module, resulting in the addition of ∼16,000 solvent molecules. Explicit counter ions (Na+ or Cl−) were introduced to neutralize the system. All subsequent calculations were performed using the Stony Brook modification of the Amber14 force field (ff14SB)68. A two-stage geometry optimization approach was used. In the first stage, the positions of solvent molecules and ions were minimized by imposing positional restraints on the solute by a harmonic potential with a force constant of 500 kcal mol−1 Å−2, and the second stage involved an unrestrained minimization of all the atoms in the simulation cell. The system was gently heated in six 50 ps steps, increasing the temperature by 50 K in each step (0–300 K) under constant volume and periodic boundary conditions. Water molecules were treated using the SHAKE algorithm, keeping the angle between the hydrogen atoms fixed. Long-range electrostatic effects were modelled using the particle-mesh Ewald method. An 8-Å cut-off was applied to Lennard-Jones and electrostatic interactions. Harmonic restraints of 30 kcal mol–1 were applied to the solute, and the Langevin scheme was used to control and equalize the temperature. The time step was kept at 1 fs during the heating stages. Each system was then equilibrated for 2 ns with a 2 fs time step at a constant pressure of 1 atm and temperature of 300 K without restraints. Once the systems had been equilibrated in the constant-temperature and constant-pressure ensemble, production trajectories were run under the constant-temperature and constant-volume ensemble and periodic boundary conditions. In particular, a total of 5,000 ns of simulations in the absence of substrate were accumulated for variants P7E and KS from five independent replicas of 1,000 ns for each system. The Cpptraj module from Ambertools utilities was used to process and analyse the trajectories, including cluster analyses. POVME 3.0 was used to analyse active-site volumes69.
Docking and substrate-bound MD simulations
The most representative structures from the previous holo state simulations were characterized by clustering of the accumulated simulation time, considering the protein backbone root mean square deviation. These structures were used for docking calculations with substrate 1, which were performed using AutoDock Vina70. The docking results were used as starting points for the restrained MD simulations, in which the distance between the centre of mass of the alkene (defined by atoms C1 and C2) of substrate 1 and the oxygen atom of haem compound I was kept restrained during the MD simulation (3.0–3.5 Å, using a 100 kcal mol−1 Å−2 force constant). This allowed the catalytically relevant binding poses of the substrate to be explored when it is in a NAC conformation to make the oxidation reaction happen, largely refining the docking predictions and preventing undesired unbinding events during the simulations. The same protocol previously described for the MD simulations was applied. A total of five independent replicas of 500 ns of production trajectory each were accumulated for each system, accumulating a total of 2.5 µs of restrained MD simulation time for each system. Substrate–residue interactions were calculated using the pairwise per-residue free-energy decomposition and molecular mechanics with generalized Born and surface area solvation (MM-GBSA) approach as implemented in the MMPBSA.py module71 from AmberTools18. The MM-GBSA energies of 500 structures (1 ns each) were calculated for each MD trajectory. The final reported MM-GBSA energy was the average of the 500 structures. SPMs for dynamic correlation network analysis were estimated using the module DynaComm.py (ref. 33).
QM/MM calculations
Initial structures for QM/MM modelling were selected from the substrate-bound restrained MD simulations of the KS. Representative snapshots were selected on the basis of the different binding poses explored by the substrate. All water molecules and counter ions beyond 3 Å from any residue of the protein, cofactors or substrates were removed. The QM region included the haem porphyrin pyrrole core, the Cys390 side chain, the iron centre and the whole substrate (1). The resulting QM region had a neutral charge, and both doublet and quartet low-lying electronic states were considered. All residues and water molecules outside a 12-Å shell from the QM region were kept frozen. QM/MM calculations were carried out using the ONIOM72 approach as implemented in the Gaussian 09 package58. Geometry optimizations were performed with the hybrid (U)B3LYP functional59,60,61 using an ultrafine integration grid and the 6-31G(d) basis set on all atoms except for iron, for which an SDD basis set and related SDD pseudopotential were used. The MM parameters and charges were identical to those used in the MD simulations. A two-step sequential optimization protocol using the QuadMacro optimization algorithm was used: a first optimization using a mechanical embedding scheme was initially performed, and once optimized, MM water molecules were kept frozen and a second optimization was performed within the electrostatic embedding scheme. Stationary points were verified as minima or saddle-point (transition-state) geometries after vibrational frequency analysis, and thermal corrections were obtained at 1 atm and 298.15 K. Single-point energy calculations on the optimized structures were performed at the (U)B3LYP/Def2TZVP level of theory within the electrostatic embedding scheme, and included empirical Grimme D3 dispersion corrections with Becke–Johnson (D3BJ) damping63.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The original materials and data that support the findings of this study are available within the paper and its Supplementary Information or can be obtained from the corresponding authors upon reasonable request.
References
Smith, M. B. & March, J. March’s Advanced Organic Chemistry (Wiley-VCH, 2007).
Grubbs, R. H. & Wenzel, A. G. Handbook of Metathesis 2nd edn (Wiley‐VCH, 2015).
Tekada, T. Modern Carbonyl Olefination: Methods and Applications (Wiley-VCH, 2006).
Atkins, W. J., Burkhardt, E. R. & Matos, K. Safe handling of boranes at scale. Org. Process Res. Dev. 10, 1292–1295 (2006).
Burkhardt, E. R. & Matos, K. Boron reagents in process chemistry: excellent tools for selective reductions. Chem. Rev. 106, 2617–2650 (2006).
Morandi, B., Wickens, Z. K. & Grubbs, R. H. Regioselective Wacker oxidation of internal alkenes: rapid access to functionalized ketones facilitated by cross-metathesis. Angew. Chem. Int. Ed. 52, 9751–9754 (2013).
Morandi, B., Wickens, Z. K. & Grubbs, R. H. Practical and general palladium-catalyzed synthesis of ketones from internal olefins. Angew. Chem. Int. Ed. 52, 2944–2948 (2013).
Deluca, R. J. et al. Wacker-type oxidation of internal alkenes using Pd(Quinox) and TBHP. J. Org. Chem. 78, 1682–1686 (2013).
Mitsudome, T., Mizumoto, K., Mizugaki, T., Jitsukawa, K. & Kaneda, K. Wacker-type oxidation of internal olefins using a PdCl2/N,N-dimethylacetamide catalyst system under copper-free reaction conditions. Angew. Chem. Int. Ed. 49, 1238–1240 (2010).
Puls, F. & Knölker, H. Conversion of olefins into ketones by an iron-catalyzed Wacker-type oxidation using oxygen as the sole oxidant. Angew. Chem. Int. Ed. 57, 1222–1226 (2018).
Puls, F., Linke, P., Kataeva, O. & Knölker, H. Iron‐catalyzed Wacker‐type oxidation of olefins at room temperature with 1,3‐diketones or neocuproine as ligands. Angew. Chem. Int. Ed. 60, 14083–14090.
Zhang, G. et al. Anti-Markovnikov oxidation of β-alkyl styrenes with H2O as the terminal oxidant. J. Am. Chem. Soc. 138, 12037–12040 (2016).
Hammer, S. C. et al. Anti-Markovnikov alkene oxidation by metal-oxo-mediated enzyme catalysis. Science 358, 215–218 (2017).
Soler, J., Gergel, S., Klaus, C., Hammer, S. C. & Garcia-Borràs, M. Enzymatic control over reactive intermediates enables direct oxidation of alkenes to carbonyls by a P450 iron-oxo species. J. Am. Chem. Soc. 144, 15954–15968 (2022).
Guengerich, F. P. Mechanisms of cytochrome P450-catalyzed oxidations. ACS Catal. 8, 10964–10976 (2018).
Ortiz De Montellano, P. R. Cytochrome P450: Structure, Mechanism, and Biochemistry (Springer, 2005).
Wang, Y., Lan, D., Durrani, R. & Hollmann, F. Peroxygenases en route to becoming dream catalysts. What are the opportunities and challenges? Curr. Opin. Chem. Biol. 37, 1–9 (2017).
Zhang, W., Loebach, J. L., Wilson, S. R. & Jacobsen, E. N. Enantioselective epoxidation of olefins catalyzed by (salen)manganese complexes. J. Am. Chem. Soc. 112, 2801–2803 (1990).
Jacobsen, E. N., Zhang, W., Muci, A. R., Ecker, J. R. & Deng, L. Highly enantioselective epoxidation catalysts derived from 1,2-diaminocyclohexane. J. Am. Chem. Soc. 113, 7063–7064 (1991).
Hintermann, L. Recent developments in metal-catalyzed additions of oxygen nucleophiles to alkenes and alkynes. Top. Organomet. Chem. 31, 123–155 (2010).
Demming, R. M. et al. Asymmetric enzymatic hydration of unactivated, aliphatic alkenes. Angew. Chem. Int. Ed. 58, 173–177 (2019).
Musacchio, A. J. et al. Catalytic intermolecular hydroaminations of unactivated olefins with secondary alkyl amines. Science 730, 727–730 (2017).
Pirnot, M. T., Wang, Y.-M. & Buchwald, S. L. Copper hydride-catalyzed hydroamination of alkenes and alkynes. Angew. Chem. Int. Ed. 55, 48–57 (2015).
Xi, Y., Ma, S. & Hartwig, J. F. Catalytic asymmetric addition of an amine N–H bond across internal alkenes. Nature 588, 37–40 (2020).
Yang, Y., Shi, S.-L., Niu, D., Liu, P. & Buchwald, S. L. Catalytic asymmetric hydroamination of unactivated internal olefins to aliphatic amines. Science 349, 62–66 (2015).
Keinan, E., Seth, K. K. & Lamed, R. Organic synthesis with enzymes. 3. TBADH-catalyzed reduction of chloro ketones. Total synthesis of (+)-(S,S)-(cis-6-methyltetrahydropyran-2-yl)acetic acid: a civet constituent. J. Am. Chem. Soc. 108, 3474–3480 (1986).
Kille, S. et al. Reducing codon redundancy and screening effort of combinatorial protein libraries created by saturation mutagenesis. ACS Synth. Biol. 2, 83–92 (2013).
Acevedo-Rocha, C. G. et al. Pervasive cooperative mutational effects on multiple catalytic enzyme traits emerge via long-range conformational dynamics. Nat. Commun. 12, 1621 (2021).
Kalita, S., Shaik, S., Kisan, H. K. & Dubey, K. D. A paradigm shift in the catalytic cycle of P450: the preparatory choreography during O2 binding and origins of the necessity for two protonation pathways. ACS Catal. 10, 11481–11492 (2020).
Dubey, K. D. & Shaik, S. Cytochrome P450—the wonderful nanomachine revealed through dynamic simulations of the catalytic cycle. Acc. Chem. Res. 52, 389–399 (2019).
Osuna, S. The challenge of predicting distal active site mutations in computational enzyme design. Wiley Interdiscip. Rev. Comput Mol. Sci. 11, e1502 (2021).
Bunzel, H. A. et al. Evolution of dynamical networks enhances catalysis in a designer enzyme. Nat. Chem. 13, 1017–1022 (2021).
Romero-Rivera, A., Garcia-Borràs, M. & Osuna, S. Role of conformational dynamics in the evolution of retro-aldolase activity. ACS Catal. 7, 8524–8532 (2017).
Maria-Solano, M. A., Iglesias-Fernández, J. & Osuna, S. Deciphering the allosterically driven conformational ensemble in tryptophan synthase evolution. J. Am. Chem. Soc. 141, 13049–13056 (2019).
Calvó-Tusell, C., Maria-Solano, M. A., Osuna, S. & Feixas, F. Time evolution of the millisecond allosteric activation of imidazole glycerol phosphate synthase. J. Am. Chem. Soc. 144, 7146–7159 (2022).
Erb, T. J., Jones, P. R. & Bar-Even, A. Synthetic metabolism: metabolic engineering meets enzyme design. Curr. Opin. Chem. Biol. 37, 56–62 (2017).
Hammer, S. C., Knight, A. M. & Arnold, F. H. Design and evolution of enzymes for non-natural chemistry. Curr. Opin. Green Sustain. Chem. 7, 23–30 (2017).
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. Int. Ed. 57, 4143–4148 (2018).
Nestl, B. M., Hammer, S. C., Nebel, B. A. & Hauer, B. New generation of biocatalysts for organic synthesis. Angew. Chem. Int. Ed. 53, 3070–3095 (2014).
Wu, S., Snajdrova, R., Moore, J. C., Baldenius, K. & Bornscheuer, U. T. Biocatalysis: enzymatic synthesis for industrial applications. Angew. Chem. Int. Ed. 60, 88–119 (2021).
Sharma, M., Mangas-Sánchez, J., Turner, N. J. & Grogan, G. NAD(P)H-dependent dehydrogenases for the asymmetric reductive amination of ketones: structure, mechanism, evolution and application. Adv. Synth. Catal. 359, 2011–2025 (2017).
Ducrot, L., Bennett, M., Grogan, G. & Vergne-Vaxelaire, C. NAD(P)H-dependent enzymes for reductive amination: active site description and carbonyl-containing compound spectrum. Adv. Synth. Catal. 363, 328–351 (2021).
Guo, F. & Berglund, P. Transaminase biocatalysis: optimization and application. Green Chem. 19, 333–360 (2016).
Fu, H. et al. Chemoenzymatic asymmetric synthesis of the metallo-β-lactamase inhibitor aspergillomarasmine A and related aminocarboxylic acids. Nat. Catal. 1, 186–191 (2018).
Cui, Y. et al. Development of a versatile and efficient C–N lyase platform for asymmetric hydroamination via computational enzyme redesign. Nat. Catal. 4, 364–373 (2021).
Roddan, R., Ward, J. M., Keep, N. H. & Hailes, H. C. Pictet–Spenglerases in alkaloid biosynthesis: future applications in biocatalysis. Curr. Opin. Chem. Biol. 55, 69–76 (2020).
Brovetto, M., Gamenara, D., Méndez, P. S. & Seoane, G. A. C–C bond-forming lyases in organic synthesis. Chem. Rev. 111, 4346–4403 (2011).
Leisch, H., Morley, K. & Lau, P. C. K. Baeyer–Villiger monooxygenases: more than just green chemistry. Chem. Rev. 111, 4165–4222 (2011).
Knaus, T., Böhmer, W. & Mutti, F. G. Amine dehydrogenases: efficient biocatalysts for the reductive amination of carbonyl compounds. Green Chem. 19, 453–463 (2017).
Rudroff, F. et al. Opportunities and challenges for combining chemo- and biocatalysis. Nat. Catal. 1, 12–22 (2018).
Köhler, V. & Turner, N. J. Artificial concurrent catalytic processes involving enzymes. Chem. Commun. 51, 450–464 (2015).
Chen, K. & Arnold, F. H. Engineering new catalytic activities in enzymes. Nat. Catal. 3, 203–213 (2020).
Klaus, C. & Hammer, S. C. New catalytic reactions by enzyme engineering. Trends Chem. 4, 363–366 (2022).
Mitschke, B., Turberg, M. & List, B. Confinement as a unifying element in selective catalysis. Chem 6, 2515–2532 (2020).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Mitchell, L. A. et al. Multichange isothermal mutagenesis: a new strategy for multiple site-directed mutations in plasmid DNA. ACS Synth. Biol. 2, 473–477 (2013).
Guengerich, F. P., Martin, M. V., Sohl, C. D. & Cheng, Q. Measurement of cytochrome P450 and NADPH–cytochrome P450 reductase. Nat. Protoc. 4, 1245–1251 (2009).
Frisch, M. J. et al. Gaussian 09, Revision D.01 (Gaussian, 2009).
Lee, C., Yang, W. & Parr, R. G. Development of the Colle–Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37, 785–789 (1988).
Becke, A. D. Density‐functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Becke, A. D. Density-functional exchange-energy approximation with correct asymptotic behavior. Phys. Rev. A 38, 3098–3100 (1988).
Luchini, G. et al. bobbypaton/GoodVibes: GoodVibes v3.0.0 (v3.0.0). Zenodo https://doi.org/10.5281/zenodo.3346166 (2019).
Grimme, S., Ehrlich, S. & Goerigk, L. Effect of the damping function in dispersion corrected density functional theory. J. Comput. Chem. 32, 1456–1465 (2011).
Case, D. A. et al. AMBER18 (Univ. California, 2018).
Cruzeiro, V. W. D., Amaral, M. S. & Roitberg, A. E. Redox potential replica exchange molecular dynamics at constant pH in AMBER: implementation and validation. J. Chem. Phys. 149, 072338 (2018).
Li, P. & Merz, K. M. MCPB.py: a Python based metal center parameter builder. J. Chem. Inf. Model. 56, 599–604 (2016).
Shahrokh, K., Orendt, A., Yost, G. S. & Cheatham, T. E. Quantum mechanically derived AMBER-compatible heme parameters for various states of the cytochrome P450 catalytic cycle. J. Comput. Chem. 33, 119–133 (2012).
Maier, J. A. et al. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Wagner, J. R. et al. POVME 3.0: software for mapping binding pocket flexibility. J. Chem. Theory Comput. 13, 4584–4592 (2017).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Miller, B. R. et al. MMPBSA.py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Dapprich, S., Komáromi, I., Byun, K. S., Morokuma, K. & Frisch, M. J. A new ONIOM implementation in Gaussian98. Part I. The calculation of energies, gradients, vibrational frequencies and electric field derivatives. Theochem 461-462, 1–21 (1999).
Acknowledgements
We thank M. Gajdoš, C. Klaus, B. Nestl (Innophore), F. Ospina and the reviewers for fruitful discussion and comments to improve the manuscript. We further acknowledge M. Breuer (BASF) for disclosing sequence information for a ketoreductase, as well as H. Gröger (Bielefeld University) and N. Turner (University of Manchester) for providing plasmid DNA coding for glucose dehydrogenases and imine reductases, respectively. This work was supported by the Deutsche Forschungsgemeinschaft via the Emmy Noether programme (fellowship no. 420112577, to S.C.H.), the Generalitat de Catalunya AGAUR via a Beatriu de Pinós H2020 MSCA-Cofund 2018-BP-00204 project (to M.G.-B.), and the Spanish MICINN (Ministerio de Ciencia e Innovación) via the PID2019-111300GA-I00 project and the Ramón y Cajal programme via an RYC 2020-028628-I fellowship (to M.G.-B.). J.S. thanks the Spanish MIU (Ministerio de Universidades) for a predoctoral FPU fellowship (FPU18/02380). The computer resources at the Barcelona Supercomputing Center (BSC-RES) are acknowledged (RES-QSB-2019-3-0009, RES-QSB-2020-2-0016 and RES-QH-2021-2-0013).
Funding
Open access funding provided by Universität Bielefeld.
Author information
Authors and Affiliations
Contributions
S.C.H. led the project. S.G. and S.C.H. designed the wet-lab experiments. S.G. performed experiments. S.G. and S.C.H. analysed data. M.G.-B. designed and supervised the computational modelling. J.S. and M.G.-B. performed the computational work and analysed the data. A.K. and K.H.S. contributed to compound synthesis and the substrate scope studies. S.G., S.C.H., M.G.-B. and J.S. co-wrote the paper and all authors agreed on the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Figs. 1–44, Tables 1–15 and Notes 1–6.
Supplementary Data Table 1
Sequences of oligonucleotides.
Supplementary Data 1
Coordinates from MD simulations (five replicas) for the KS variant in the holo state.
Supplementary Data 2
Coordinates from restrained MD simulations (five replicas) for the KS variant with substrate bound.
Supplementary Data 3
Coordinates from MD simulations (five replicas) for the P7E parent variant in the holo state.
Supplementary Data 4
Coordinates from restrained MD simulations (five replicas) for the P7E variant with substrate bound.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gergel, S., Soler, J., Klein, A. et al. Engineered cytochrome P450 for direct arylalkene-to-ketone oxidation via highly reactive carbocation intermediates. Nat Catal 6, 606–617 (2023). https://doi.org/10.1038/s41929-023-00979-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-023-00979-4