Abstract
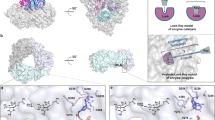
Artificial metalloenzymes (ArMs), which contain non-native, typically synthetic, metal cofactors, are a flourishing class of biocatalyst for unnatural reactions. Although the number of these reactions is rapidly increasing, multi-faceted mechanistic studies of ArMs comprising structural, kinetic, computational and cofactor binding data to reveal detailed mechanistic information on the effects of the protein scaffold on the structure and reactivity of ArMs are more limited. Here we report the structure of an unnatural P450 analogue using X-ray diffraction. We also report the kinetic analysis of its reaction, catalyst activation during an induction period, and the origins of the stereoselectivity for the cyclopropanation of a terpene catalysed by the iridium-containing P450 variant (Ir(Me)–CYP119). Our data reveal a mechanism initiated by the flip of the cofactor from an inactive to an active conformation. This change in conformation is followed by thousands of turnovers occurring by rate-determining formation of an iridium–carbene intermediate, thereby highlighting the influence of cofactor dynamics within a single active site on an ArM-catalysed reaction.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Atomic coordinates and structure factors for Ir(Me)–CYP119 have been deposited in the PDB with the PDB code 7UOR. The data on gas evolution that support the findings of this study are accessible through Dryad (https://doi.org/10.6078/D10Q6S). The other relevant kinetics data supporting the findings of this study are available within the Supplementary Information.
References
Davis, H. J. & Ward, T. R. Artificial metalloenzymes: challenges and opportunities. ACS Cent. Sci. 5, 1120–1136 (2019).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Dydio, P. et al. An artificial metalloenzyme with the kinetics of native enzymes. Science 354, 102–106 (2016).
Hyster Todd, K., Knörr, L., Ward Thomas, R. & Rovis, T. Biotinylated Rh(III) complexes in engineered streptavidin for accelerated asymmetric C–H activation. Science 338, 500–503 (2012).
Jeschek, M. et al. Directed evolution of artificial metalloenzymes for in vivo metathesis. Nature 537, 661–665 (2016).
Oohora, K. et al. Catalytic cyclopropanation by myoglobin reconstituted with iron porphycene: acceleration of catalysis due to rapid formation of the carbene species. J. Am. Chem. Soc. 139, 17265–17268 (2017).
Wolf, M. W., Vargas, D. A. & Lehnert, N. Engineering of RuMb: toward a green catalyst for carbene insertion reactions. Inorg. Chem. 56, 5623–5635 (2017).
Alonso-Cotchico, L. et al. Integrated computational study of the Cu-catalyzed hydration of alkenes in water solvent and into the context of an artificial metallohydratase. ACS Catal. 9, 4616–4626 (2019).
Bhagi-Damodaran, A., Petrik, I. D., Marshall, N. M., Robinson, H. & Lu, Y. Systematic tuning of heme redox potentials and its effects on O2 reduction rates in a designed oxidase in myoglobin. J. Am. Chem. Soc. 136, 11882–11885 (2014).
Bhagi-Damodaran, A. et al. Heme redox potentials hold the key to reactivity differences between nitric oxide reductase and heme-copper oxidase. Proc. Natl Acad. Sci. USA 115, 6195–6200 (2018).
Christoffel, F. et al. Design and evolution of chimeric streptavidin for protein-enabled dual gold catalysis. Nat. Catal. 4, 643–653 (2021).
Collot, J., Humbert, N., Skander, M., Klein, G. & Ward, T. R. Artificial metalloenzymes for enantioselective catalysis: the phenomenon of protein accelerated catalysis. J. Organomet. Chem. 689, 4868–4871 (2004).
Dürrenberger, M. et al. Artificial transfer hydrogenases for the enantioselective reduction of cyclic imines. Angew. Chem. Int. Ed. 50, 3026–3029 (2011).
Ke, Z., Abe, S., Ueno, T. & Morokuma, K. Catalytic mechanism in artificial metalloenzyme: QM/MM study of phenylacetylene polymerization by rhodium complex encapsulated in apo-ferritin. J. Am. Chem. Soc. 134, 15418–15429 (2012).
Muñoz Robles, V. et al. Toward the computational design of artificial metalloenzymes: from protein–ligand docking to multiscale approaches. ACS Catal. 5, 2469–2480 (2015).
Robles, V. M. et al. Structural, kinetic, and docking studies of artificial imine reductases based on biotin–streptavidin technology: an induced lock-and-key hypothesis. J. Am. Chem. Soc. 136, 15676–15683 (2014).
Stein, A. et al. A dual anchoring strategy for the directed evolution of improved artificial transfer hydrogenases based on carbonic anhydrase. ACS Cent. Sci. 7, 1874–1884 (2021).
Upp, D. M. et al.Engineering dirhodium artificial metalloenzymes for diazo coupling cascade reactions. Angew. Chem. Int. Ed. 60, 23672–23677 (2021).
Villarino, L. et al. Cofactor binding dynamics influence the catalytic activity and selectivity of an artificial metalloenzyme. ACS Catal. 10, 11783–11790 (2020).
Yu, Y. et al. A designed metalloenzyme achieving the catalytic rate of a native enzyme. J. Am. Chem. Soc. 137, 11570–11573 (2015).
Zubi, Y. S., Liu, B., Gu, Y., Sahoo, D. & Lewis, J. C. Controlling the optical and catalytic properties of artificial metalloenzyme photocatalysts using chemogenetic engineering. Chem. Sci. 13, 1459–1468 (2022).
Hayashi, T. et al. Capture and characterization of a reactive haem–carbenoid complex in an artificial metalloenzyme. Nat. Catal. 1, 578–584 (2018).
Carminati, D. M., Moore, E. J. & Fasan, R. in Methods in Enzymology Vol. 644 (ed. Tawfik, D. S.) 35–61 (Academic Press, 2020).
Natoli, S. N. & Hartwig, J. F. Noble−metal substitution in hemoproteins: an emerging strategy for abiological catalysis. Acc. Chem. Res. 52, 326–335 (2019).
Zhang, R. K., Huang, X. & Arnold, F. H. Selective CH bond functionalization with engineered heme proteins: new tools to generate complexity. Curr. Opin. Chem. Biol. 49, 67–75 (2019).
Rabe, K. S., Kiko, K. & Niemeyer, C. M. Characterization of the peroxidase activity of CYP119, a thermostable P450 from Sulfolobus acidocaldarius. ChemBioChem 9, 420–425 (2008).
Hayashi, T., Sano, Y. & Onoda, A. Generation of new artificial metalloproteins by cofactor modification of native hemoproteins. Isr. J. Chem. 55, 76–84 (2015).
Moffat, K., Loe, R. S. & Hoffman, B. M. The structure of metmanganoglobin. J. Mol. Biol. 104, 669–685 (1976).
Bushnell, G. W., Louie, G. V. & Brayer, G. D. High-resolution three-dimensional structure of horse heart cytochrome c. J. Mol. Biol. 214, 585–595 (1990).
Pearson, A. R. et al. The crystal structure of cytochrome P460 of Nitrosomonas europaea reveals a novel cytochrome fold and heme−protein cross-link. Biochemistry 46, 8340–8349 (2007).
Cedervall, P., Hooper, A. B. & Wilmot, C. M. Structural studies of hydroxylamine oxidoreductase reveal a unique heme cofactor and a previously unidentified interaction partner. Biochemistry 52, 6211–6218 (2013).
Smith, M. A. & Lancaster, K. M. The eponymous cofactors in cytochrome P460s from ammonia-oxidizing bacteria are iron porphyrinoids whose macrocycles are dibasic. Biochemistry 57, 334–343 (2018).
Tran, A.-T. T., Kalish, H., Balch, A. L. & La Mar, G. N. Solution 1H NMR investigation of the seating and rotational “hopping” of centrosymmetric etioheme-I in myoglobin: effect of globin origin and its oxidation/spin state on heme dynamics. J. Biol. Inorg. Chem. 5, 624–633 (2000).
Ueno, T. et al. Crystal structures of artificial metalloproteins: tight binding of FeIII(Schiff-base) by mutation of Ala71 to Gly in apo-myoglobin. Inorg. Chem. 43, 2852–2858 (2004).
Abe, S. et al. Design and structure analysis of artificial metalloproteins: selective coordination of his64 to copper complexes with square-planar structure in the apo-myoglobin scaffold. Inorg. Chem. 46, 5137–5139 (2007).
Key, H. M. et al. Beyond iron: iridium-containing P450 enzymes for selective cyclopropanations of structurally diverse alkenes. ACS Cent. Sci. 3, 302–308 (2017).
Gu, Y., Natoli, S. N., Liu, Z., Clark, D. S. & Hartwig, J. F. Site-selective functionalization of (sp3)C−H bonds catalyzed by artificial metalloenzymes containing an iridium-porphyrin cofactor. Angew. Chem. Int. Ed. 58, 13954–13960 (2019).
Segel, I. H. Enzyme Kinetics: Behavior and Analysis of Rapid Equilibrium and Steady-State Enzyme Systems (Wiley, 1975).
Hargrove, M. S., Barrick, D. & Olson, J. S. The association rate constant for heme binding to globin is independent of protein structure. Biochemistry 35, 11293–11299 (1996).
Hoops, S. et al. COPASI—a COmplex PAthway SImulator. Bioinformatics 22, 3067–3074 (2006).
Deniau, C. et al. Thermodynamics of heme binding to the HasA(SM) hemophore: effect of mutations at three key residues for heme uptake. Biochemistry 42, 10627–10633 (2003).
Yukl, E. T. et al. Kinetic and spectroscopic studies of hemin acquisition in the hemophore HasAp from Pseudomonas aeruginosa. Biochemistry 49, 6646–6654 (2010).
Barik, A., Priyadarsini, K. I. & Mohan, H. Photophysical studies on binding of curcumin to bovine serum albumins. Photochem. Photobiol. 77, 597–603 (2003).
Feng, X. Z., Lin, Z., Yang, L. J., Wang, C. & Bai, C. L. Investigation of the interaction between acridine orange and bovine serum albumin. Talanta 47, 1223–1229 (1998).
Bhakta, M. N. & Wilks, A. The mechanism of heme transfer from the cytoplasmic heme binding protein PhuS to the ẟ-regioselective heme oxygenase of Pseudomonas aeruginosa. Biochemistry 45, 11642–11649 (2006).
Penning, T. M. Single-molecule enzymology of steroid transforming enzymes: transient kinetic studies and what they tell us. J. Steroid Biochem. Mol. Biol. 161, 5–12 (2016).
Owens, C. P., Du, J., Dawson, J. H. & Goulding, C. W. Characterization of heme ligation properties of Rv0203, a secreted heme binding protein involved in Mycobacterium tuberculosis heme uptake. Biochemistry 51, 1518–1531 (2012).
Wittwer, M. et al. Engineering and emerging applications of artificial metalloenzymes with whole cells. Nat. Catal. 4, 814–827 (2021).
Hollenberg, P. F. Mechanisms of cytochrome P450 and peroxidase-catalyzed xenobiotic metabolism. FASEB J. 6, 686–694 (1992).
Meunier, B., de Visser, S. P. & Shaik, S. Mechanism of oxidation reactions catalyzed by cytochrome P450 enzymes. Chem. Rev. 104, 3947–3980 (2004).
Hu, J., Allen, R., Rozinek, S. & Brancaleon, L. Experimental and computational characterization of photosensitized conformational effects mediated by protoporphyrin ligands on human serum albumin. Photochem. Photobiol. Sci. 16, 694–710 (2017).
Boens, N. et al. Rational design, synthesis, and spectroscopic and photophysical properties of a visible-light-excitable, ratiometric, fluorescent near-neutral pH indicator based on BODIPY. Chem 17, 10924–10934 (2011).
Acknowledgements
B.J.B., S.N.N. and J.F.H. acknowledge support from the Director, Office of Science, 271 US Department of Energy, under Contract No. DE-AC02-272 05CH1123. S.N.N. also thanks the NIH (F32-GM126652) and the Burroughs Wellcome fund (PDEP). K.N.H. thanks the National Institute of General Medical Sciences of the NIH (grant R01GM124480) for support. M.G.-B. acknowledges support from the Spanish Ministerio de Ciencia e Innovación (projects PID2019-111300GA-I00 and RyC2020-028628-I). The work conducted by the Joint BioEnergy Institute is supported by the US Department of Energy, Office of Science, Office of Biological and Environmental Research under contract no. DE-AC02-05CH11231 between LBNL and the US Department of Energy. We would like to thank R. G. Bergman, F. D. Toste, J. Y. Wang and E. D. Kalkman for valuable discussions and suggestions. We would like to thank A. Quest for helpful insight on figures.
Author information
Authors and Affiliations
Contributions
B.J.B. and J.F.H. designed and B.J.B. conducted the chemical kinetics and fluorimetry experiments and analysed the data. D.B.H. assisted with protein expression and purification. S.N.N. and J.H.P. designed and conducted the protein crystallography experiments. J.H.P. and P.D.A. analysed the crystallographic data. B.J.B., D.S.C. and J.F.H. interpreted the experimental data and prepared the experimental portion of the manuscript and Supplementary Information. M.G.-B. and K.N.H. designed the computational studies and interpreted the data. M.G.-B. conducted the computations, analysed the data and prepared the computational section of the manuscript and Supplemenary Information. All authors contributed to discussions, commented on and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Tables 1–10 and Figs. 1–23.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bloomer, B.J., Natoli, S.N., Garcia-Borràs, M. et al. Mechanistic and structural characterization of an iridium-containing cytochrome reveals kinetically relevant cofactor dynamics. Nat Catal 6, 39–51 (2023). https://doi.org/10.1038/s41929-022-00899-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-022-00899-9