Abstract
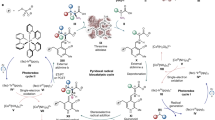
Polyketides found in nature originate from backbones synthesized through iterative decarboxylative Claisen condensations catalysed by polyketide synthases (PKSs). However, PKSs suffer from complicated architecture, energy inefficiencies, complex regulation, and competition with essential metabolic pathways for extender unit malonyl-CoA, all combining to limit the flux of polyketide biosynthesis. Here we show that certain thiolases, which we term polyketoacyl-CoA thiolases (PKTs), catalyse polyketide backbone formation via iterative non-decarboxylative Claisen condensations, hence offering a synthetic and efficient alternative to PKSs. We show that PKTs can synthesize polyketide backbones for representative lactone, alkylresorcinolic acid, alkylresorcinol, hydroxybenzoic acid and alkylphenol polyketide families, and elucidate the basic catalytic mechanism and structural features enabling this previously unknown activity. PKT-catalysed reactions offer a route to polyketide formation that leverages the simple architecture of thiolases to achieve higher ATP efficiencies and reduced competition with essential metabolic pathways, all of which circumvent intrinsic inefficiencies of PKSs for polyketide product synthesis.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that data supporting the findings of this study are available within the paper and its Supplementary Information. Supplementary Table 3 provides a list of the GenBank accession numbers of the 17 key enzymes used in this study. Data for Figs. 1–6 and Extended Data Fig. 6 are available as Source Data with this paper. All other data are available from the authors on reasonable request.
References
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. 48, 4688–4716 (2009).
Ma, S. M. et al. Complete reconstitution of a highly reducing iterative polyketide synthase. Science 326, 589–592 (2009).
Yu, D. Y., Xu, F. C., Zeng, J. & Zhan, J. X. Type III polyketide synthases in natural product biosynthesis. IUBMB Life 64, 285–295 (2012).
Bond, C., Tang, Y. & Li, L. Saccharomyces cerevisiae as a tool for mining, studying and engineering fungal polyketide synthases. Fungal Genet. Biol. 89, 52–61 (2016).
Staunton, J. & Weissman, K. J. Polyketide biosynthesis: a millennium review. Nat. Prod. Rep. 18, 380–416 (2001).
Shen, B. Polyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms. Curr. Opin. Chem. Biol. 7, 285–295 (2003).
Huo, L. et al. Heterologous expression of bacterial natural product biosynthetic pathways. Nat. Prod. Rep. 36, 1412–1436 (2019).
Pfeifer, B. A. & Khosla, C. Biosynthesis of polyketides in heterologous hosts. Microbiol. Mol. Biol. Rev. 65, 106–118 (2001).
Lowry, B. et al. In vitro reconstitution and analysis of the 6-deoxyerythronolide B synthase. J. Am. Chem. Soc. 135, 16809–16812 (2013).
Chan, Y. A., Podevels, A. M., Kevany, B. M. & Thomas, M. G. Biosynthesis of polyketide synthase extender units. Nat. Prod. Rep. 26, 90–114 (2009).
Brownsey, R., Boone, A., Elliott, J., Kulpa, J. & Lee, W. Regulation of acetyl-CoA carboxylase. Biochem. Soc. Trans. 34, 223–227 (2006).
Yan, D. et al. Repurposing type III polyketide synthase as a malonyl-CoA biosensor for metabolic engineering in bacteria. Proc. Natl Acad. Sci. USA 115, 9835–9844 (2018).
Keatinge-Clay, A. T. The structures of type I polyketide synthases. Nat. Prod. Rep. 29, 1050–1073 (2012).
Jiang, C., Kim, S. Y. & Suh, D. Y. Divergent evolution of the thiolase superfamily and chalcone synthase family. Mol. Phylogenetetics Evol. 49, 691–701 (2008).
Austin, M. B. & Noel, A. J. P. The chalcone synthase superfamily of type III polyketide synthases. Nat. Prod. Rep. 20, 79–110 (2003).
Haapalainen, A. M., Merilainen, G. & Wierenga, R. K. The thiolase superfamily: condensing enzymes with diverse reaction specificities. Trends Biochem. Sci. 31, 64–71 (2006).
Tang, S. Y. et al. Screening for enhanced triacetic acid lactone production by recombinant Escherichia coli expressing a designed triacetic acid lactone reporter. J. Am. Chem. Soc. 135, 10099–10103 (2013).
Campobasso, N. et al. Staphylococcus aureus 3-hydroxy-3-methylglutaryl-CoA synthase: crystal structure and mechanism. J. Biol. Chem. 279, 44883–44888 (2004).
Duncombe, G. R. & Frerman, F. E. Molecular and catalytic properties of the acetoacetyl-coenzyme A thiolase of Escherichia coli. Arch. Biochem. Biophys. 176, 159–170 (1976).
Yang, S. Y., Yang, X. Y. H., Healylouie, G., Schulz, H. & Elzinga, M. Nucleotide-sequence of the fadA Gene—primary structure of 3-ketoacyl-coenzyme-A thiolase from Escherichia coli and the structural organization of the fadAB operon. J. Biol. Chem. 265, 10424–10429 (1990).
Teufel, R. et al. Bacterial phenylalanine and phenylacetate catabolic pathway revealed. Proc. Natl Acad. Sci. USA 107, 14390–14395 (2010).
Harwood, C. S., Nichols, N. N., Kim, M. K., Ditty, J. L. & Parales, R. E. Identification of the pcaRKF gene cluster from Pseudomonas putida: involvement in chemotaxis, biodegradation, and transport of 4-hydroxybenzoate. J. Bacteriol. 176, 6479–6488 (1994).
Cheong, S., Clomburg, J. M. & Gonzalez, R. Energy- and carbon-efficient synthesis of functionalized small molecules in bacteria using non-decarboxylative Claisen condensation reactions. Nat. Biotechnol. 34, 556–561 (2016).
Parke, D., Garcia, M. A. & Ornston, L. N. Cloning and genetic characterization of dca genes required for β-oxidation of straight-chain dicarboxylic acids in Acinetobacter sp. strain ADP1. Appl. Environ. Microbiol. 67, 4817–4827 (2001).
Slater, S. et al. Multiple β-ketothiolases mediate poly(β-hydroxyalkanoate) copolymer synthesis in Ralstonia eutropha. J. Bacteriol. 180, 1979–1987 (1998).
Shanks, B. H. & Keeling, P. L. Bioprivileged molecules: creating value from biomass. Green. Chem. 19, 3177–3185 (2017).
Noor, E. et al. Pathway thermodynamics highlights kinetic obstacles in central metabolism. PLoS Comput. Biol. 10, e1003483 (2014).
Xie, D. et al. Microbial synthesis of triacetic acid lactone. Biotechnol. Bioeng. 93, 727–736 (2006).
Johnson, A. O. et al. Design and application of genetically-encoded malonyl-CoA biosensors for metabolic engineering of microbial cell factories. Metab. Eng. 44, 253–264 (2017).
Qian, S., Clomburg, J. M. & Gonzalez, R. Engineering Escherichia coli as a platform for the in vivo synthesis of prenylated aromatics. Biotechnol. Bioeng. 116, 1116–1127 (2019).
Collie, N. & Myers, W. V. I. I. The formation of orcinol and other condensation products from dehydracetic acid. J. Chem. Soc. Trans. 63, 122–128 (1893).
Gagne, S. J. et al. Identification of olivetolic acid cyclase from Cannabis sativa reveals a unique catalytic route to plant polyketides. Proc. Natl Acad. Sci. USA 109, 12811–12816 (2012).
Tan, Z., Clomburg, J. M. & Gonzalez, R. Synthetic pathway for the production of olivetolic acid in Escherichia coli. ACS Synth. Biol. 7, 1886–1896 (2018).
Lim, Y. P., Go, M. K. & Yew, W. S. Exploiting the biosynthetic potential of type III polyketide synthases. Molecules 21, 806 (2016).
Zhou, W. et al. Biosynthesis of phlorisovalerophenone and 4-hydroxy-6-isobutyl-2-pyrone in Escherichia coli from glucose. Microb. Cell Fact. 15, 149 (2016).
Huang, L., Xue, Z. & Zhu, Q. Method for the production of resveratrol in a recombinant bacterial host cell. US patent WO2006124999A2 (2007).
Choi, O. et al. Biosynthesis of plant-specific phenylpropanoids by construction of an artificial biosynthetic pathway in Escherichia coli. J. Ind. Microbiol. Biotechnol. 38, 1657–1665 (2011).
Mizuuchi, Y. et al. Novel type III polyketide synthases from Aloe arborescens. FEBS J. 276, 2391–2401 (2009).
Fujii, I. Functional analysis of fungal polyketide biosynthesis genes. J. Antibiot. 63, 207–218 (2010).
Moriguchi, T., Kezuka, Y., Nonaka, T., Ebizuka, Y. & Fujii, I. Hidden function of catalytic domain in 6-methylsalicylic acid synthase for product release. J. Biol. Chem. 285, 15637–15643 (2010).
Sabatini, M. et al. Biochemical characterization of the minimal domains of an iterative eukaryotic polyketide synthase. FEBS J. 285, 4494–4511 (2018).
Crawford, J. M. et al. Structural basis for biosynthetic programming of fungal aromatic polyketide cyclization. Nature 461, 1139–1243 (2009).
Shen, B. & Hutchinson, C. R. Deciphering the mechanism for the assembly of aromatic polyketides by a bacterial polyketide synthase. Proc. Natl Acad. Sci. USA 93, 6600–6604 (1996).
Kim, E. J., Son, H. F., Kim, S., Ahn, J. W. & Kim, K. J. Crystal structure and biochemical characterization of β-keto thiolase B from polyhydroxyalkanoate-producing bacterium Ralstonia eutropha H16. Biochem. Biophys. Res. Commun. 444, 365–369 (2014).
Jez, J. M. et al. Structural control of polyketide formation in plant-specific polyketide synthases. Chem. Biol. 7, 919–930 (2000).
Blaisse, M. R., Fu, B. & Chang, M. C. Y. Structural and biochemical studies of substrate selectivity in Ascaris suum thiolases. Biochemistry 57, 3155–3166 (2018).
Clomburg, J. M. et al. Integrated engineering of β-oxidation reversal and ω-oxidation pathways for the synthesis of medium chain omega-functionalized carboxylic acids. Metab. Eng. 28, 202–212 (2015).
Dellomonaco, C., Clomburg, J. M., Miller, E. N. & Gonzalez, R. Engineered reversal of the β-oxidation cycle for the synthesis of fuels and chemicals. Nature 476, 355–359 (2011).
Kim, S., Clomburg, J. M. & Gonzalez, R. Synthesis of medium-chain length (C6-C10) fuels and chemicals via β-oxidation reversal in Escherichia coli. J. Ind. Microbiol. Biotechnol. 42, 465–475 (2015).
Krivoruchko, A., Zhang, Y., Siewers, V., Chen, Y. & Nielsen, J. Microbial acetyl-CoA metabolism and metabolic engineering. Metab. Eng. 28, 28–42 (2015).
Acknowledgements
We thank C. Pennington at Rice University for assistance with LC–MS analysis.
Author information
Authors and Affiliations
Contributions
R.G. designed the research. Z.T., S.C. and J.M.C. performed the research. S.Q. performed structure simulation and alignment. Z.T., J.M.C. and R.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a patent application (US patent application no. PCT/US2016/045037).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Reactions catalysed by members of the thiolase superfamily and structures of corresponding enzymes.
a, Reactions catalysed by members of the thiolase superfamily; (b) Three-dimensional structures of corresponding enzymes (monomer). The structures shown are those of Medicago sataiva chalcone synthase PKS III (1bi5), Streptomcyes albus ketosynthase domain of polyketide synthase KS/PKS I (4wky), Streptomyces coelicolor KS/PKS II (1tqy), E. coli β-ketoacyl-acyl-carrier-protein synthase KAS I (2vb8), E. coli KAS II (1kas), E. coli KAS III (1hn9), Arabidopsis thaliana KCS (modelling structure), human mitochondria thiolase I (4c2j), E. coli thiolase II (5f0v), and Bacillus subtilis HMGS (4yxq).
Extended Data Fig. 2 Purified enzymes used in this study.
The sources of enzymes are as follows. PKS III 2-PS was from Gerbera hybrid; DcaF was from Acinetobacter sp. ADP1; FadAx was from Pseudomonas putida; PcaF was from Pseudomonas putida; BktB was from Ralstonia eutropha; AtoB was from E. coli; EcFadA was from E. coli; PaaJ was from E. coli; HMGS was from Staphylococcus aureus; FadB was from E. coli; PaaH was from E. coli; The expected molecular weights of enzymes with N terminal 6 x His-tag calculated by tool from ExPASy, (https://web.expasy.org/compute_pi/) are indicated in parentheses. BktB (5#, 11#, 16#), PaaJ (8#, 14#) and EcFadA (7#, 12#) appeared more than once as we initially employed them for in vitro TAL biosynthesis tests (Fig. 2), and then we further purified and used them for the in vitro assay of ketoacyl-CoA thiolase, 3-ketoadipyl-CoA thiolase and PKT activities (see Fig. 6a). For EcFadA WT (7#, 12#) and EcFadA_BktB (21#), the expected MWs from ExPASy calculations are 41.9 kDa and 43.2 kDa, while actual sizes on SDS-PAGE appear ~40 kDa and ~41 kDa.
Extended Data Fig. 3 Primary sequence alignment of thiolases and PKS III.
a, Primary sequence alignment of PKS III 2-PS (2-pyrone synthase from Gerbera hybrid) with AtoB (from E. coli). The sequence identity is only 10.0%; (b) Primary sequence alignment of PKS III 2-PS with FadAx (from Pseudomonas putida). The sequence identity is only 8.7%; (c) Primary sequence alignment of PKS III 2-PS with BktB (from Ralstonia eutropha). The sequence identity is only 13.6%. d, Primary sequence alignment of PKS III ORS (from Rhododendron dauricum) with BktB. The sequence identity is only 9.9%.
Extended Data Fig. 4 Max-min Driving Force (MDF) for the synthesis of specified polyketides from acetyl-CoA through the proposed PKT-based pathway.
MDF calculations assume minimum and maximum metabolite concentrations of 0.000001 and 0.01 M, respectively, and a fixed NADPH/NADP + ratio of 10.
Extended Data Fig. 5 Summary of SDS-PAGE analysis of samples in this study.
a, SDS-PAGE analysis of the soluble expression of BktB and 2-PS in JC01 (DE3) ΔatoB host strain. M: marker; 1: purified BktB; 2: purified 2-PS; 3: soluble fraction of JC01 (DE3) ΔatoB expressing pCDF-bktB, the arrow indicates the soluble BktB protein; 4: soluble fraction of JC01 (DE3) ΔatoB harbouring pCDF-2-ps, the arrow indicates the soluble 2-PS protein. b, SDS-PAGE analysis of the soluble expression of BktB WT and C90S, H350A, C380S mutants. M: marker; 1: soluble fraction of BL21 (DE3) expressing BktB WT; 2: soluble fraction of BL21 (DE3) expressing BktB C90S; 3: soluble fraction of BL21 (DE3) expressing BktB H350A; 4: soluble fraction of BL21 (DE3) expressing BktB C380S. c, SDS-PAGE analysis of the soluble expression of EcFadA and BktB mutants after segment swapping. M: marker; 1: soluble fraction of BL21 (DE3) expressing EcFadA WT; 2: soluble fraction of BL21 (DE3) expressing EcFadA_BktB; 3: soluble fraction of BL21 (DE3) expressing BktB WT; 4: soluble fraction of BL21 (DE3) expressing BktB_EcFadA.
Extended Data Fig. 6 Assessment of malonyl-CoA availability through the flaviolin biosynthesis pathway.
Upper, flaviolin biosynthesis pathway: 5 molecules of malonyl-CoA are condensed by RppA to form flaviolin, which has a specific absorbance at 340 nm. Bottom, RppA was expressed with different MCS/ACC enzymes for characterization of malonyl-CoA availability in strain JC01 (DE3) ΔatoB. Engineered E. coli strains were cultured in LB-like MOPS medium + 2% (wt/v) glycerol. For strains harbouring MCS, 12 mM malonate sodium was also included. Error bars represent s.d. calculated from at least three biological replicates.
Extended Data Fig. 7 Mass spectrometry (MS) information of the samples.
a, MS information of orcinol standard and orcinol sample extracted from in vivo samples; (b) MS information of ORA standard and ORA sample extracted from in vivo samples; (c) MS of 6-MSA standard and 6-MSA sample extracted from in vitro samples. d, MS of m-cresol standard and m-cresol sample extracted from in vitro samples.
Extended Data Fig. 8 The proposed PKT-based platforms for aloesone, octaketide SEK4, norsolorinate and tetracenomycin F1 biosynthesis.
a, The proposed PKT-based platform for aloesone biosynthesis. b, The proposed PKT-based platform for SEK4 biosynthesis. c, The proposed PKT-based platform for norsolorinate biosynthesis. d, The proposed PKT-based platform for tetracenomycin F1 biosynthesis.
Extended Data Fig. 9 Catalytic mechanism of polyketoacyl-CoA thiolase (PKT) and representative PKS III 2-PS.
Cys/His/Cys is the catalytic triad of PKT activity, enabling a two-step, ping-pong mechanism for the thiolytic condensation reaction using substrates with multiple keto-groups (for example acetoacetyl-CoA). Notably, this catalytic mechanism is distinct from that of PKS III, where the core catalytic triad is Cys/His/Asn, with Asn (asparagine) playing a critical role in catalysing malonyl-CoA decarboxylation and stabilizing the condensation transition state.
Extended Data Fig. 10 Exploiting PKT to synthesize additional PK backbones and corresponding PKs (R = groups other than CH3).
Synthesis of polyketide backbones for representative lactone, alkylresorcinolic acid, and alkylresorcinol polyketide families through up to 3 rounds of iterative non-decarboxylative Claisen condensations catalyzed by PKTs.
Supplementary information
Supplementary Information
Supplementary Tables 1–4.
Source data
Source Data Fig. 1
All amino acid sequences of thiolases superfamily used in Fig. 1b.
Source Data Fig. 2
Statistical Source Data for Figs. 2b and 2d.
Source Data Fig. 3
Statistical Source Data for Figs. 3b and 3d.
Source Data Fig. 4
Statistical Source Data for Fig. 4c.
Source Data Fig. 5
Statistical Source Data for Fig. 5c.
Source Data Fig. 6
Statistical Source Data for Figs. 6a and 6d.
Source Data Extended Data Fig. 6
Statistical Source Data for Extended Fig. 6.
Rights and permissions
About this article
Cite this article
Tan, Z., Clomburg, J.M., Cheong, S. et al. A polyketoacyl-CoA thiolase-dependent pathway for the synthesis of polyketide backbones. Nat Catal 3, 593–603 (2020). https://doi.org/10.1038/s41929-020-0471-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-020-0471-8
This article is cited by
-
Acetyl-CoA-independent malonyl-CoA biosynthesis
Nature Catalysis (2024)
-
Biorenewable and circular polydiketoenamine plastics
Nature Sustainability (2023)
-
Peroxisomal KAT2 (3-ketoacyl-CoA thiolase 2) gene has a key role in gingerol biosynthesis in ginger (Zingiber officinale Rosc.)
Journal of Plant Biochemistry and Biotechnology (2023)
-
Thioester-mediated biocatalytic amide bond synthesis with in situ thiol recycling
Nature Catalysis (2022)
-
An artificial pathway for polyketide biosynthesis
Nature Catalysis (2020)