Abstract
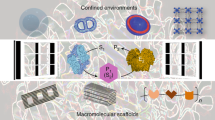
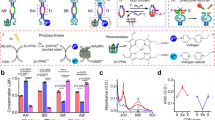
Cellular transformations, such as gene expression or temporal protein activities, are controlled by complex stimuli-responsive network circuitries regulated by enzymes, metabolites or transcription factors. Inspired by nature, extensive research efforts are directed to mimic these processes by in vitro chemical systems. Here we report on the assembly of constitutional dynamic networks (CDNs), composed of nucleic acid–enzyme conjugates, that act as modules for triggered, network-driven, biocatalytic cascades and for the intercommunication of network-guided biocatalytic cascades. Two CDNs were assembled—one network includes a constituent module functionalized with glucose oxidase and horseradish peroxidase in spatially close positions, and the second CDN includes a constituent module modified at sterically intimate positions with nicotinamide adenine dinucleotide and alcohol dehydrogenase. Biocatalytic cascades proceed in the two networks and, on the triggered reconfiguration of the CDNs, controlled and switchable biocatalytic cascades in the CDNs are demonstrated. The two CDNs are coupled, and the triggered feedback-driven intercommunication of the networks is realized.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Fell, D. A. & Wagner, A. The small world of metabolism. Nat. Biotechnol. 18, 1121–1122 (2000).
Guelzim, N., Bottani, S., Bourgine, P. & Képès, F. Topological and causal structure of the yeast transcriptional regulatory network. Nat. Genet. 31, 60–63 (2002).
Davidson, E. H. et al. A genomic regulatory network for development. Science 295, 1669–1678 (2002).
Barabási, A. L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Oliveri, P. & Davidson, E. H. Gene regulatory network controlling embryonic specification in the sea urchin. Curr. Opin. Genet. Dev. 14, 351–360 (2004).
Levine, M. & Davidson, E. H. Gene regulatory networks for development. Proc. Natl Acad. Sci. USA 102, 4936–4942 (2005).
Peter, I. S. & Davidson, E. H. Assessing regulatory information in developmental gene regulatory networks. Proc. Natl Acad. Sci. USA 114, 5862–5869 (2017).
Montagne, K., Plasson, R., Sakai, Y., Fujii, T. & Rondelez, Y. Programming an in vitro DNA oscillator using a molecular networking strategy. Mol. Syst. Biol. 7, 466 (2011).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Semenov, S. N. et al. Rational design of functional and tunable oscillating enzymatic networks. Nat. Chem. 7, 160–165 (2015).
Schaffter, S. W. & Schulman, R. Building in vitro transcriptional regulatory networks by successively integrating multiple functional circuit modules. Nat. Chem. 11, 829–838 (2019).
Kim, J., White, K. S. & Winfree, E. Construction of an in vitro bistable circuit from synthetic transcriptional switches. Mol. Syst. Biol. 2, 68 (2006).
Wang, F., Liu, X. & Willner, I. DNA switches: from principles to applications. Angew. Chem. Int. Ed. 54, 1098–1129 (2015).
Lu, C. H. et al. Switchable catalytic acrylamide hydrogels cross-linked by hemin/G-quadruplexes. Nano Lett. 13, 1298–1302 (2013).
Liu, D. & Balasubramanian, S. A proton-fuelled DNA nanomachine. Angew. Chem. Int. Ed. 42, 5734–5736 (2003).
Hu, Y., Cecconello, A., Idili, A., Ricci, F. & Willner, I. Triplex DNA nanostructures: from basic properties to applications. Angew. Chem. Int. Ed. 56, 15210–15233 (2017).
Liang, X., Mochizuki, T. & Asanuma, H. A supra-photoswitch involving sandwiched DNA base pairs and azobenzenes for light-driven nanostructures and nanodevices. Small 5, 1761–1768 (2009).
Asanuma, H. et al. Enantioselective incorporation of azobenzenes into oligodeoxyribonucleotide for effective photoregulation of duplex formation. Angew. Chem. Int. Ed. 40, 2671–2673 (2001).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003).
Lu, C. H., Willner, B. & Willner, I. DNA nanotechnology: from sensing and DNA machines to drug-delivery systems. ACS Nano 7, 8320–8332 (2013).
Teller, C. & Willner, I. Functional nucleic acid nanostructures and DNA machines. Curr. Opin. Biotechnol. 21, 376–391 (2010).
Bath, J. & Turberfield, A. J. DNA nanomachines. Nat. Nanotechnol. 2, 275–284 (2007).
Seeman, N. C. From genes to machines: DNA nanomechanical devices. Trends Biochem. Sci. 30, 119–125 (2012).
Wang, S. et al. Controlling the catalytic functions of DNAzymes within constitutional dynamic networks of DNA nanostructures. J. Am. Chem. Soc. 139, 9662–9671 (2017).
Wang, S. et al. Light-induced reversible reconfiguration of DNA-based constitutional dynamic networks: application to switchable catalysis. Angew. Chem. Int. Ed. 57, 8105–8109 (2018).
Yue, L., Wang, S. & Willner, I. Triggered reversible substitution of adaptive constitutional dynamic networks dictates programmed catalytic functions. Sci. Adv. 5, eaav5564 (2019).
Zhou, Z., Yue, L., Wang, S., Lehn, J. M. & Willner, I. DNA-based multiconstituent dynamic networks: hierarchical adaptive control over the composition and cooperative catalytic functions of the systems. J. Am. Chem. Soc. 140, 12077–12089 (2018).
Yue, L. et al. Consecutive feedback-driven constitutional dynamic networks. Proc. Natl Acad. Sci. USA 116, 2843–2848 (2019).
Yue, L. et al. Intercommunication of DNA-based constitutional dynamic networks. J. Am. Chem. Soc. 140, 8721–8731 (2018).
Yue, L., Wang, S. & Willner, I. Three-dimensional nucleic-acid-based constitutional dynamic networks: enhancing diversity through complexity of the systems. J. Am. Chem. Soc. 141, 16461–16470 (2019).
Jitrapakdee, S. et al. Structure, mechanism and regulation of pyruvate carboxylase. Biochem. J. 413, 369–387 (2008).
Vázquez-González, M., Wang, C. & Willner, I. Biocatalytic cascades operating on macromolecular scaffolds and in confined environments. Nat. Catal. 3, 256–273 (2020).
Elani, Y., Law, R. V. & Ces, O. Vesicle-based artificial cells as chemical microreactors with spatially segregated reaction pathways. Nat. Commun. 5, 5305 (2014).
Vriezema, D. M. et al. Positional assembly of enzymes in polymersome nanoreactors for cascade reactions. Angew. Chem. Int. Ed. 46, 7378–7382 (2007).
Liang, J. et al. Peptide-induced super-assembly of biocatalytic metal–organic frameworks for programmed enzyme cascades. Chem. Sci. 10, 7852–7858 (2019).
Chen, W. H., Vázquez-González, M., Zoabi, A., Abu-Reziq, R. & Willner, I. Biocatalytic cascades driven by enzymes encapsulated in metal–organic framework nanoparticles. Nat. Catal. 1, 689–695 (2018).
Wilner, O. I. et al. Enzyme cascades activated on topologically programmed DNA scaffolds. Nat. Nanotechnol. 4, 249–254 (2009).
Fu, J. et al. Multi-enzyme complexes on DNA scaffolds capable of substrate channelling with an artificial swinging arm. Nat. Nanotechnol. 9, 531–536 (2014).
Ngo, T. A., Nakata, E., Saimura, M. & Morii, T. Spatially organized enzymes drive cofactor-coupled cascade reactions. J. Am. Chem. Soc. 138, 3012–3021 (2016).
Xin, L., Zhou, C., Yang, Z. & Liu, D. Regulation of an enzyme cascade reaction by a DNA machine. Small 9, 3088–3091 (2013).
Yang, Y. et al. Programming rotary motions with a hexagonal DNA nanomachine. Chem. Eur. J. 25, 5158–5162 (2019).
Qu, J. et al. Synthetic multienzyme complexes, catalytic nanomachineries for cascade biosynthesis in vivo. ACS Nano 13, 9895–9906 (2019).
You, C., Myung, S. & Zhang, Y. H. P. Facilitated substrate channeling in a self-assembled trifunctional enzyme complex. Angew. Chem. Int. Ed. 51, 8787–8790 (2012).
Lin, J. L., Palomec, L. & Wheeldon, I. Design and analysis of enhanced catalysis in scaffolded multienzyme cascade reactions. ACS Catal. 4, 505–511 (2014).
Zhang, Y. & Hess, H. Toward rational design of high-efficiency enzyme cascades. ACS Catal. 7, 6018–6027 (2017).
Lancaster, L., Abdallah, W., Banta, S. & Wheeldon, I. Engineering enzyme microenvironments for enhanced biocatalysis. Chem. Soc. Rev. 47, 5177–5186 (2018).
Lin, J. L. & Wheeldon, I. Kinetic enhancements in DNA–enzyme nanostructures mimic the Sabatier principle. ACS Catal. 3, 560–564 (2013).
Idan, O. & Hess, H. Origins of activity enhancement in enzyme cascades on scaffolds. ACS Nano 7, 8658–8665 (2013).
Ellis, G. A. et al. Artificial multienzyme scaffolds: pursuing in vitro substrate channeling with an overview of current progress. ACS Catal. 9, 10812–10869 (2019).
Tigges, M., Marquez-Lago, T. T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009).
Fung, E. et al. A synthetic gene–metabolic oscillator. Nature 435, 118–122 (2005).
Wu, Q. et al. Cascade enzymes within self-assembled hybrid nanogel mimicked neutrophil lysosomes for singlet oxygen elevated cancer therapy. Nat. Commun. 10, 240 (2019).
Acknowledgements
Our research is supported by the Israel Science Foundation.
Author information
Authors and Affiliations
Contributions
C.W. performed the experiments, analysed the results and participated in writing the paper. L.Y. participated in the design of CDNs and analysis of the results. I.W. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Figs. 1–33, discussion, Tables 1–4, Notes 1–4 and references.
Source data
Source Data Fig. 1
Time-dependent fluorescence changes generated by the DNAzyme reporter units associated with the constituents in CDNs and time-dependent absorbance changes of ABTS∙– in CDNs.
Source Data Fig. 2
Time-dependent fluorescence changes generated by the DNAzyme reporter units associated with the constituents in CDNs and time-dependent absorbance changes induced by ADH/NAD+/MB+ cascade and ADH/NAD+/pyruvic acid cascade.
Source Data Fig. 3
Time-dependent absorbance changes induced by two biocatalytic cascades guided by intercommunicating CDNs.
Source Data Fig. 4
Time-dependent absorbance changes of two biocatalytic cascades guided by feedback-driven intercommunication of coupled CDNs.
Rights and permissions
About this article
Cite this article
Wang, C., Yue, L. & Willner, I. Controlling biocatalytic cascades with enzyme–DNA dynamic networks. Nat Catal 3, 941–950 (2020). https://doi.org/10.1038/s41929-020-00524-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-020-00524-7
This article is cited by
-
Analytical device miniaturization for the detection of circulating biomarkers
Nature Reviews Bioengineering (2023)
-
Spatiotemporal control for integrated catalysis
Nature Reviews Methods Primers (2023)
-
Hierarchically encapsulating enzymes with multi-shelled metal-organic frameworks for tandem biocatalytic reactions
Nature Communications (2022)
-
Electrostatically cooperative host-in-host of metal cluster ⊂ ionic organic cages in nanopores for enhanced catalysis
Nature Communications (2022)
-
Cascaded dissipative DNAzyme-driven layered networks guide transient replication of coded-strands as gene models
Nature Communications (2022)