Abstract
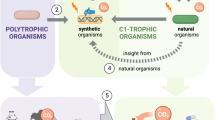
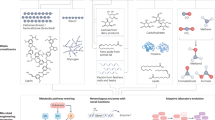
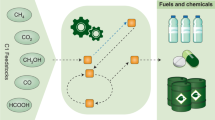
Production of industrial chemicals using renewable biomass feedstock is becoming increasingly important to address limited fossil resources, climate change and other environmental problems. To develop high-performance microbial cell factories, equivalent to chemical plants, microorganisms undergo systematic metabolic engineering to efficiently convert biomass-derived carbon sources into target chemicals. Over the past two decades, many engineered microorganisms capable of producing natural and non-natural chemicals have been developed. This Review details the current status of representative industrial chemicals that are produced through biological and/or chemical reactions. We present a comprehensive bio-based chemicals map that highlights the strategies and pathways of single or multiple biological reactions, chemical reactions and combinations thereof towards production of particular chemicals of interest. Future challenges are also discussed to enable production of even more diverse chemicals and more efficient production of chemicals from renewable feedstocks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
10 September 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
A European Strategy for Plastics in a Circular Economy (European Commission, 2018).
The future of plastic. Nat. Commun. 9, 2157 (2018).
Yang, D., Cho, J. S., Choi, K. R., Kim, H. U. & Lee, S. Y. Systems metabolic engineering as an enabling technology in accomplishing sustainable development goals. Microb. Biotechnol. 10, 1254–1258 (2017).
Lee, J. W. et al. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat. Chem. Biol. 8, 536–546 (2012).
Chubukov, V., Mukhopadhyay, A., Petzold, C. J., Keasling, J. D. & Martin, H. G. Synthetic and systems biology for microbial production of commodity chemicals. NPJ Syst. Biol. Appl. 2, 16009 (2016).
Clomburg, J. M., Crumbley, A. M. & Gonzalez, R. Industrial biomanufacturing: the future of chemical production. Science 355, aag0804 (2017).
Lee, S. Y. & Kim, H. U. Systems strategies for developing industrial microbial strains. Nat. Biotechnol. 33, 1061–1072 (2015).
Lee, J. W., Kim, T. Y., Jang, Y. S., Choi, S. & Lee, S. Y. Systems metabolic engineering for chemicals and materials. Trends Biotechnol. 29, 370–378 (2011).
Jang, Y. S. et al. Bio-based production of C2–C6 platform chemicals. Biotechnol. Bioeng. 109, 2437–2459 (2012).
Sarria, S., Kruyer, N. S. & Peralta-Yahya, P. Microbial synthesis of medium-chain chemicals from renewables. Nat. Biotechnol. 35, 1158–1166 (2017).
Pfleger, B. F., Gossing, M. & Nielsen, J. Metabolic engineering strategies for microbial synthesis of oleochemicals. Metab. Eng. 29, 1–11 (2015).
Becker, J. & Wittmann, C. Advanced biotechnology: metabolically engineered cells for the bio-based production of chemicals and fuels, materials, and health-care products. Angew. Chem. Int. Ed. Engl. 54, 3328–3350 (2015).
Rudroff, F. et al. Opportunities and challenges for combining chemo- and biocatalysis. Nat. Catal. 1, 12–22 (2018).
Zhang, M. M., Wang, Y., Ang, E. L. & Zhao, H. Engineering microbial hosts for production of bacterial natural products. Nat. Prod. Rep. 33, 963–987 (2016).
Krivoruchko, A., Zhang, Y., Siewers, V., Chen, Y. & Nielsen, J. Microbial acetyl-CoA metabolism and metabolic engineering. Metab. Eng. 28, 28–42 (2015).
Long, M. R., Ong, W. K. & Reed, J. L. Computational methods in metabolic engineering for strain design. Curr. Opin. Biotechnol. 34, 135–141 (2015).
Copeland, W. B. et al. Computational tools for metabolic engineering. Metab. Eng. 14, 270–280 (2012).
King, Z. A., Lloyd, C. J., Feist, A. M. & Palsson, B. O. Next-generation genome-scale models for metabolic engineering. Curr. Opin. Biotechnol. 35, 23–29 (2015).
Chae, T. U., Choi, S. Y., Kim, J. W., Ko, Y. S. & Lee, S. Y. Recent advances in systems metabolic engineering tools and strategies. Curr. Opin. Biotechnol. 47, 67–82 (2017).
Choi, K. R. & Lee, S. Y. CRISPR technologies for bacterial systems: current achievements and future directions. Biotechnol. Adv. 34, 1180–1209 (2016).
Jensen, M. K. & Keasling, J. D. Recent applications of synthetic biology tools for yeast metabolic engineering. FEMS Yeast Res. 15, 1–10 (2015).
Smanski, M. J. et al. Synthetic biology to access and expand nature’s chemical diversity. Nat. Rev. Microbiol. 14, 135–149 (2016).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes. Nucleic Acids Res. 46, 633–639 (2018).
Hadadi, N., Hafner, J., Shajkofci, A., Zisaki, A. & Hatzimanikatis, V. ATLAS of Biochemistry: A repository of all possible biochemical reactions for synthetic biology and metabolic engineering studies. ACS Synth. Biol. 5, 1155–1166 (2016).
Hadadi, N. & Hatzimanikatis, V. Design of computational retrobiosynthesis tools for the design of de novo synthetic pathways. Curr. Opin. Chem. Biol. 28, 99–104 (2015).
Kumar, A., Wang, L., Ng, C. Y. & Maranas, C. D. Pathway design using de novo steps through uncharted biochemical spaces. Nat. Commun. 9, 184 (2018).
Shin, J. H., Kim, H. U., Kim, D. I. & Lee, S. Y. Production of bulk chemicals via novel metabolic pathways in microorganisms. Biotechnol. Adv. 31, 925–935 (2013).
Feher, T. et al. Validation of RetroPath, a computer-aided design tool for metabolic pathway engineering. Biotechnol. J. 9, 1446–1457 (2014).
Kan, S. B., Lewis, R. D., Chen, K. & Arnold, F. H. Directed evolution of cytochrome c for carbon-silicon bond formation: Bringing silicon to life. Science 354, 1048–1051 (2016).
Kan, S. B. J., Huang, X., Gumulya, Y., Chen, K. & Arnold, F. H. Genetically programmed chiral organoborane synthesis. Nature 552, 132–136 (2017).
Choi, S., Song, C. W., Shin, J. H. & Lee, S. Y. Biorefineries for the production of top building block chemicals and their derivatives. Metab. Eng. 28, 223–239 (2015).
Pereira, B. et al. Efficient utilization of pentoses for bioproduction of the renewable two-carbon compounds ethylene glycol and glycolate. Metab. Eng. 34, 80–87 (2016).
Chae, T. U., Choi, S. Y., Ryu, J. Y. & Lee, S. Y. Production of ethylene glycol from xylose by metabolically engineered Escherichia coli. AIChE J. (2018).
Sousa, A. F. et al. Biobased polyesters and other polymers from 2,5-furandicarboxylic acid: a tribute to furan excellency. Polym. Chem. 6, 5961–5983 (2015).
Luo, Z. W., Kim, W. J. & Lee, S. Y. Metabolic engineering of Escherichia coli for efficient production of 2-pyrone-4,6-dicarboxylic acid from glucose. ACS Synth. Biol. 7, 2296–2307 (2018).
Masuno, M. N. et al. Methods of producing para-xylene and terephthalic acid. US patent 2013/0245316 A1 (2013).
Luo, Z. W. & Lee, S. Y. Biotransformation of p-xylene into terephthalic acid by engineered Escherichia coli. Nat. Commun. 8, 15689 (2017).
Tomas, R. A., Bordado, J. C. & Gomes, J. F. p-Xylene oxidation to terephthalic acid: a literature review oriented toward process optimization and development. Chem. Rev. 113, 7421–7469 (2013).
Sangeetha, V. H., Deka, H., Varghese, T. O. & Nayak, S. K. State of the art and future prospectives of poly(lactic acid) based blends and composites. Polym. Compos. 39, 81–101 (2018).
Sauer, M., Porro, D., Mattanovich, D. & Branduardi, P. 16 years research on lactic acid production with yeast - ready for the market? Biotechnol. Genet. Eng. Rev. 27, 229–256 (2010).
Jung, Y. K., Kim, T. Y., Park, S. J. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of polylactic acid and its copolymers. Biotechnol. Bioeng. 105, 161–171 (2010).
Paddon, C. J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Burk, M. J., Burgard, A. P., Osterhout, R. E. & Pharkya, P. Microorganisms and methods for the biosynthesis of adipate, hexamethylenediamine and 6-aminocaproic acid. US patent 2010/0317069 A1 (2010).
Cordova, L. T. & Alper, H. S. Central metabolic nodes for diverse biochemical production. Curr. Opin. Chem. Biol. 35, 37–42 (2016).
Sánchez-Riera, F., Cameron, D. C. & Cooney, C. L. Influence of environmental factors in the production of R(−)-1, 2-propanediol by Clostridium thermosaccharolyticum. Biotechnol. Lett. 9, 449–454 (1987).
Siebert, D. & Wendisch, V. F. Metabolic pathway engineering for production of 1,2-propanediol and 1-propanol by Corynebacterium glutamicum. Biotechnol. Biofuels 8, 91 (2015).
Nakamura, C. E. & Whited, G. M. Metabolic engineering for the microbial production of 1,3-propanediol. Curr. Opin. Biotechnol. 14, 454–459 (2003).
Chen, Z. et al. Metabolic engineering of Corynebacterium glutamicum for the production of 3-hydroxypropionic acid from glucose and xylose. Metab. Eng. 39, 151–158 (2017).
Li, Y., Wang, X., Ge, X. & Tian, P. High production of 3-hydroxypropionic acid in Klebsiella pneumoniae by systematic optimization of glycerol metabolism. Sci. Rep. 6, 26932 (2016).
Chu, H. S. et al. Direct fermentation route for the production of acrylic acid. Metab. Eng. 32, 23–29 (2015).
Matsubara, M. et al. Fermentative production of 1-propanol from d-glucose, l-rhamnose and glycerol using recombinant Escherichia coli. J. Biosci. Bioeng. 122, 421–426 (2016).
Yang, P. et al. A new strategy for production of 5-aminolevulinic acid in recombinant Corynebacterium glutamicum with high yield. Appl. Environ. Microbiol. 82, 2709–2717 (2016).
Chu, H. S. et al. Metabolic engineering of 3-hydroxypropionic acid biosynthesis in Escherichia coli. Biotechnol. Bioeng. 112, 356–364 (2015).
Karp, E. M. et al. Renewable acrylonitrile production. Science 358, 1307–1310 (2017).
Craciun, L. et al. Preparation of acrylic acid derivatives from α- or β-hydroxy carboxylic acids. US patent US7538247B2 (2009).
Corma, A., Iborra, S. & Velty, A. Chemical routes for the transformation of biomass into chemicals. Chem. Rev. 107, 2411–2502 (2007).
Dishisha, T., Pyo, S. H. & Hatti-Kaul, R. Bio-based 3-hydroxypropionic- and acrylic acid production from biodiesel glycerol via integrated microbial and chemical catalysis. Microb. Cell Fact. 14, 200 (2015).
Guo, Z. et al. Dehydration of lactic acid to acrylic acid over lanthanum phosphate catalysts: the role of Lewis acid sites. Phys. Chem. Chem. Phys. 18, 23746–23754 (2016).
Meng, Y., Xue, Y., Yu, B., Gao, C. & Ma, Y. Efficient production of l-lactic acid with high optical purity by alkaliphilic Bacillus sp. WL-S20. Bioresour. Technol. 116, 334–339 (2012).
Thomas, K. C. & Ingledew, W. M. Production of 21% (v/v) ethanol by fermentation of very high gravity (VHG) wheat mashes. J. Ind. Microbiol. Biotechnol. 10, 61–68 (1992).
Ma, C. et al. Enhanced 2,3-butanediol production by Klebsiella pneumoniae SDM. Appl. Microbiol. Biotechnol. 82, 49–57 (2009).
Kim, J. W. et al. Enhanced production of 2,3-butanediol by engineered Saccharomyces cerevisiae through fine-tuning of pyruvate decarboxylase and NADH oxidase activities. Biotechnol. Biofuels 9, 265 (2016).
Atsumi, S., Hanai, T. & Liao, J. C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451, 86–89 (2008).
Abdel-Rahman, M. A., Tashiro, Y. & Sonomoto, K. Recent advances in lactic acid production by microbial fermentation processes. Biotechnol. Adv. 31, 877–902 (2013).
Kwon, S., Yoo, I. K., Lee, W. G., Chang, H. N. & Chang, Y. K. High-rate continuous production of lactic acid by Lactobacillus rhamnosus in a two-stage membrane cell-recycle bioreactor. Biotechnol. Bioeng. 73, 25–34 (2001).
Carlos Serrano-Ruiz, J. & Dumesic, J. A. Catalytic upgrading of lactic acid to fuels and chemicals by dehydration/hydrogenation and C–C coupling reactions. Green Chem. 11, 1101–1104 (2009).
Carlson, T. L. & Peters, J., E. M. Low PH lactic acid fermentation. US patent US6475759B1 (2002).
Choi, S. Y. et al. One-step fermentative production of poly(lactate-co-glycolate) from carbohydrates in Escherichia coli. Nat. Biotechnol. 34, 435–440 (2016).
Choi, S. Y. et al. Engineering the xylose-catabolizing Dahms pathway for production of poly(d-lactateco-glycolate) and poly(d-lactate-co-glycolate-co-d-2-hydroxybutyrate) in. Escherichia coli. Microb. Biotechnol. 10, 1353–1364 (2017).
Shi, D. J., Wang, C. L. & Wang, K. M. Genome shuffling to improve thermotolerance, ethanol tolerance and ethanol productivity of Saccharomyces cerevisiae. J. Ind. Microbiol. Biotechnol. 36, 139–147 (2009).
Alper, H., Moxley, J., Nevoigt, E., Fink, G. R. & Stephanopoulos, G. Engineering yeast transcription machinery for improved ethanol tolerance and production. Science 314, 1565–1568 (2006).
Fan, D., Dai, D. J. & Wu, H. S. Ethylene formation by catalytic dehydration of ethanol with industrial considerations. Materials 6, 101–115 (2012).
Song, D. Kinetic model development for dehydration of 2,3-butanediol to 1,3-butadiene and methyl ethyl ketone over an amorphous calcium phosphate catalyst. Ind. Eng. Chem. Res. 55, 11664–11671 (2016).
Vecchini, N., Galeotti, A. & Pisano, A. Process for the production of 1,3 butadiene from 1,3 butanediol. US patent 20170313633 A1 (2014).
Vecchini, N., Galeotti, A. & Pisano, A. Process for the production of 1,3-butandiene from 1,4-butanediol via tetrahydrofuran. WO patent 2016092517 (2016).
Kataoka, N., Vangnai, A. S., Tajima, T., Nakashimada, Y. & Kato, J. Improvement of (R)-1,3-butanediol production by engineered Escherichia coli. J. Biosci. Bioeng. 115, 475–480 (2013).
Burgard, A. P., Burk, M. J. & Pharkya, P. Methods and organisms for converting synthesis gas or other gaseous carbon sources and methanol to 1,3-butanediol. US patent 9284581 B2 (2009).
Baez, A., Cho, K. M. & Liao, J. C. High-flux isobutanol production using engineered Escherichia coli: a bioreactor study with in situ product removal. Appl. Microbiol. Biotechnol. 90, 1681–1690 (2011).
van Leeuwen, B. N., van der Wulp, A. M., Duijnstee, I., van Maris, A. J. & Straathof, A. J. Fermentative production of isobutene. Appl. Microbiol. Biotechnol. 93, 1377–1387 (2012).
Peters, M. W., Taylor, J. D., Jenni, M., Manzer, L. E. & Henton, D. E. Integrated process to selectively convert renewable isobutanol to p-xylene. US patent 2011/0087000 A1 (2011).
Moon, H. G. et al. One hundred years of clostridial butanol fermentation. FEMS Microbiol. Lett. 363, fnw001 (2016).
Patakova, P. et al. Comparative analysis of high butanol tolerance and production in clostridia. Biotechnol. Adv. 36, 721–738 (2018).
Jimenez-Bonilla, P. & Wang, Y. In situ biobutanol recovery from clostridial fermentations: a critical review. Crit. Rev. Biotechnol. 38, 469–482 (2018).
Lee, J. et al. Metabolic engineering of Clostridium acetobutylicum ATCC 824 for isopropanol-butanol-ethanol fermentation. Appl. Environ. Microbiol. 78, 1416–1423 (2012).
Anbarasan, P. et al. Integration of chemical catalysis with extractive fermentation to produce fuels. Nature 491, 235–239 (2012).
Beller, H. R., Lee, T. S. & Katz, L. Natural products as biofuels and bio-based chemicals: fatty acids and isoprenoids. Nat. Prod. Rep. 32, 1508–1526 (2015).
Peralta-Yahya, P. P., Zhang, F., del Cardayre, S. B. & Keasling, J. D. Microbial engineering for the production of advanced biofuels. Nature 488, 320–328 (2012).
Fairley, P. Introduction: Next generation biofuels. Nature 474, S2–5 (2011).
Xu, P. et al. Modular optimization of multi-gene pathways for fatty acids production in E. coli. Nat. Commun. 4, 1409 (2013).
Gajewski, J., Pavlovic, R., Fischer, M., Boles, E. & Grininger, M. Engineering fungal de novo fatty acid synthesis for short chain fatty acid production. Nat. Commun. 8, 14650 (2017).
Liao, J. C., Mi, L., Pontrelli, S. & Luo, S. Fuelling the future: microbial engineering for the production of sustainable biofuels. Nat. Rev. Microbiol. 14, 288–304 (2016).
Dellomonaco, C., Clomburg, J. M., Miller, E. N. & Gonzalez, R. Engineered reversal of the beta-oxidation cycle for the synthesis of fuels and chemicals. Nature 476, 355–359 (2011).
Sheppard, M. J., Kunjapur, A. M. & Prather, K. L. J. Modular and selective biosynthesis of gasoline-range alkanes. Metab. Eng. 33, 28–40 (2016).
Choi, Y. J. & Lee, S. Y. Microbial production of short-chain alkanes. Nature 502, 571–574 (2013).
Cheon, S., Kim, H. M., Gustavsson, M. & Lee, S. Y. Recent trends in metabolic engineering of microorganisms for the production of advanced biofuels. Curr. Opin. Chem. Biol. 35, 10–21 (2016).
Cao, Y. X. et al. Heterologous biosynthesis and manipulation of alkanes in Escherichia coli. Metab. Eng. 38, 19–28 (2016).
d’Espaux, L. et al. Engineering high-level production of fatty alcohols by Saccharomyces cerevisiae from lignocellulosic feedstocks. Metab. Eng. 42, 115–125 (2017).
Rodriguez, G. M., Tashiro, Y. & Atsumi, S. Expanding ester biosynthesis in Escherichia coli. Nat. Chem. Biol. 10, 259–265 (2014).
Qiao, K., Wasylenko, T. M., Zhou, K., Xu, P. & Stephanopoulos, G. Lipid production in Yarrowia lipolytica is maximized by engineering cytosolic redox metabolism. Nat. Biotechnol. 35, 173–177 (2017).
Xu, P., Qiao, K., Ahn, W. S. & Stephanopoulos, G. Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals. Proc. Natl Acad. Sci. USA 113, 10848–10853 (2016).
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 30, 354–359 (2012).
Zhou, Y. J., Buijs, N. A., Siewers, V. & Nielsen, J. Fatty acid-derived biofuels and chemicals production in Saccharomyces cerevisiae. Front. Bioeng. Biotechnol. 2, 32 (2014).
Zhou, Y. J. et al. Production of fatty acid-derived oleochemicals and biofuels by synthetic yeast cell factories. Nat. Commun. 7, 11709 (2016).
Kurosawa, K., Boccazzi, P., de Almeida, N. M. & Sinskey, A. J. High-cell-density batch fermentation of Rhodococcus opacus PD630 using a high glucose concentration for triacylglycerol production. J. Biotechnol. 147, 212–218 (2010).
Li, Y., Zhao, Z. & Bai, F. High-density cultivation of oleaginous yeast Rhodosporidium toruloides Y4 in fed-batch culture. Enzyme Microb. Technol. 41, 312–317 (2007).
Levering, J., Broddrick, J. & Zengler, K. Engineering of oleaginous organisms for lipid production. Curr. Opin. Biotechnol. 36, 32–39 (2015).
Ajjawi, I. et al. Lipid production in Nannochloropsis gaditana is doubled by decreasing expression of a single transcriptional regulator. Nat. Biotechnol. 35, 647–652 (2017).
Daboussi, F. et al. Genome engineering empowers the diatom Phaeodactylum tricornutum for biotechnology. Nat. Commun. 5, 3831 (2014).
Miao, S., Wang, P., Su, Z. & Zhang, S. Vegetable-oil-based polymers as future polymeric biomaterials. Acta Biomater. 10, 1692–1704 (2014).
Zhu, Y., Romain, C. & Williams, C. K. Sustainable polymers from renewable resources. Nature 540, 354–362 (2016).
Park, S. Y., Yang, D., Ha, S. H. & Lee, S. Y. Metabolic engineering of microorganisms for the production of natural compounds. Adv. Biosys. 2, 1700190 (2018).
Whited, G. M. et al. Development of a gas-phase bioprocess for isoprene-monomer production using metabolic pathway engineering. Ind. Biotechnol. 6, 152–163 (2010).
Yang, J. et al. Metabolic engineering of Escherichia coli for the biosynthesis of alpha-pinene. Biotechnol. Biofuels 6, 60 (2013).
Sarria, S., Wong, B., Garcia Martin, H., Keasling, J. D. & Peralta-Yahya, P. Microbial synthesis of pinene. ACS Synth. Biol. 3, 466–475 (2014).
Meadows, A. L. et al. Rewriting yeast central carbon metabolism for industrial isoprenoid production. Nature 537, 694–697 (2016).
Ozaydin, B., Burd, H., Lee, T. S. & Keasling, J. D. Carotenoid-based phenotypic screen of the yeast deletion collection reveals new genes with roles in isoprenoid production. Metab. Eng. 15, 174–183 (2013).
Kim, H. U., Charusanti, P., Lee, S. Y. & Weber, T. Metabolic engineering with systems biology tools to optimize production of prokaryotic secondary metabolites. Nat. Prod. Rep. 33, 933–941 (2016).
Curran, S. C. et al. Probing the flexibility of an iterative modular polyketide synthase with non-native substrates in vitro. ACS Chem. Biol. 13, 2261–2268 (2018).
Liu, Q. et al. Engineering an iterative polyketide pathway in Escherichia coli results in single-form alkene and alkane overproduction. Metab. Eng. 28, 82–90 (2015).
Yuzawa, S. et al. Comprehensive in vitro analysis of acyltransferase domain exchanges in modular polyketide synthases and its application for short-chain ketone production. ACS Synth. Biol. 6, 139–147 (2017).
Hagen, A. et al. Engineering a polyketide synthase for in vitro production of adipic acid. ACS Synth. Biol. 5, 21–27 (2016).
Yuzawa, S., Keasling, J. D. & Katz, L. Insights into polyketide biosynthesis gained from repurposing antibiotic-producing polyketide synthases to produce fuels and chemicals. J. Antibiot. 69, 494–499 (2016).
Averesch, N. J. H. & Kromer, J. O. Metabolic engineering of the shikimate pathway for production of aromatics and derived compounds-Present and future strain construction strategies. Front. Bioeng. Biotechnol. 6, 32 (2018).
Fischer-Romero, C., Tindall, B. J. & Juttner, F. Tolumonas auensis gen. nov., sp. nov., a toluene-producing bacterium from anoxic sediments of a freshwater lake. Int. J. Syst. Bacteriol. 46, 183–188 (1996).
Kim, B., Park, H., Na, D. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of phenol from glucose. Biotechnol. J. 9, 621–629 (2014).
Balderas-Hernandez, V. E. et al. Metabolic engineering for improving anthranilate synthesis from glucose in Escherichia coli. Microb. Cell Fact. 8, 19 (2009).
Balderas-Hernandez, V. E. et al. Catechol biosynthesis from glucose in Escherichia coli anthranilate-overproducer strains by heterologous expression of anthranilate 1,2-dioxygenase from Pseudomonas aeruginosa PAO1. Microb. Cell Fact. 13, 136 (2014).
Kim, B., Binkley, R., Kim, H. U. & Lee, S. Y. Metabolic engineering of Escherichia coli for the enhanced production of l-tyrosine. Biotechnol. Bioeng. 115, 2554–2564 (2018).
Miao, L., Li, Q., Diao, A., Zhang, X. & Ma, Y. Construction of a novel phenol synthetic pathway in Escherichia coli through 4-hydroxybenzoate decarboxylation. Appl. Microbiol. Biotechnol. 99, 5163–5173 (2015).
Li, M. et al. De novo production of resveratrol from glucose or ethanol by engineered Saccharomyces cerevisiae. Metab. Eng. 32, 1–11 (2015).
Li, M., Schneider, K., Kristensen, M., Borodina, I. & Nielsen, J. Engineering yeast for high-level production of stilbenoid antioxidants. Sci. Rep. 6, 36827 (2016).
Yang, J. E. et al. One-step fermentative production of aromatic polyesters from glucose by metabolically engineered Escherichia coli strains. Nat. Commun. 9, 79 (2018).
Sano, C. History of glutamate production. Am. J. Clin. Nutr. 90, 728S–732S (2009).
Shimizu, H. & Hirasawa, T. in Amino Acid Biosynthesis: Pathways, Regulation and Metabolic Engineering (ed. Wendisch, V. F.) 1–38 (Springer, Heidelberg, 2007).
Park, S. H. et al. Metabolic engineering of Corynebacterium glutamicum for l-arginine production. Nat. Commun. 5, 4618 (2014).
Cho, J. S. et al. CRISPR/Cas9-coupled recombineering for metabolic engineering of Corynebacterium glutamicum. Metab. Eng. 42, 157–167 (2017).
Kim, S. Y., Lee, J. & Lee, S. Y. Metabolic engineering of Corynebacterium glutamicum for the production of l-ornithine. Biotechnol. Bioeng. 112, 416–421 (2015).
Zelder, O. et al. Improved process for the production of gamma-aminobutyric acid (GABA). WO patent 2015/092599 A1 (2015).
Park, S. J. et al. Synthesis of nylon 4 from gamma-aminobutyrate (GABA) produced by recombinant Escherichia coli. Bioprocess Biosyst. Eng. 36, 885–892 (2013).
Chae, T. U., Ko, Y. S., Hwang, K. S. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of four-, five- and six-carbon lactams. Metab. Eng. 41, 82–91 (2017).
Zhang, J. et al. Metabolic engineering of Escherichia coli for the biosynthesis of 2-pyrrolidone. Metab. Eng. Commun. 3, 1–7 (2016).
Kinoshita, S., Nakayama, K. & Udaka, S. The fermentative production of l-ornithine preliminary report. J. Gen. Appl. Microbiol. 3, 276–277 (1957).
Qian, Z. G., Xia, X. X. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of putrescine: a four carbon diamine. Biotechnol. Bioeng. 104, 651–662 (2009).
Na, D. et al. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat. Biotechnol. 31, 170–174 (2013).
Noh, M., Yoo, S. M., Kim, W. J. & Lee, S. Y. Gene expression knockdown by modulating synthetic small RNA expression in Escherichia coli. Cell Syst. 5, 418–426 (2017). e414.
Guettler, M. V., Jain, M. K. & Rumler, D. Method for making succinic acid, bacterial variants for use in the process, and methods for obtaining variants. US patent 5573931 A (1996).
Lee, P. C., Lee, W. G., Lee, S. Y. & Chang, H. N. Succinic acid production with reduced by-product formation in the fermentation of Anaerobiospirillum succiniciproducens using glycerol as a carbon source. Biotechnol. Bioeng. 72, 41–48 (2001).
Okino, S. et al. An efficient succinic acid production process in a metabolically engineered Corynebacterium glutamicum strain. Appl. Microbiol. Biotechnol. 81, 459–464 (2008).
Lee, J. W. et al. Homo-succinic acid production by metabolically engineered Mannheimia succiniciproducens. Metab. Eng. 38, 409–417 (2016).
Lange, A. et al. Bio-based succinate from sucrose: High-resolution 13C metabolic flux analysis and metabolic engineering of the rumen bacterium Basfia succiniciproducens. Metab. Eng. 44, 198–212 (2017).
Rush, B. J. & Fosmer, A. M. Methods for succinate production. US patent application US20140363862A1 (2014).
Raab, A. M., Gebhardt, G., Bolotina, N., Weuster-Botz, D. & Lang, C. Metabolic engineering of Saccharomyces cerevisiae for the biotechnological production of succinic acid. Metab. Eng. 12, 518–525 (2010).
Gao, C. et al. Robust succinic acid production from crude glycerol using engineered Yarrowia lipolytica. Biotechnol. Biofuels 9, 179 (2016).
Ahn, J. H., Jang, Y. S. & Lee, S. Y. Production of succinic acid by metabolically engineered microorganisms. Curr. Opin. Biotechnol. 42, 54–66 (2016).
Hong, U. G. et al. Hydrogenation of succinic acid to 1,4-butanediol over rhenium catalyst supported on copper-containing mesoporous carbon. J. Nanosci. Nanotechnol. 13, 7448–7453 (2013).
Hong, U. G. et al. Hydrogenation of succinic acid to tetrahydrofuran (THF) over rhenium catalyst supported on H2SO4-treated mesoporous carbon. Appl. Catal. A Gen. 415, 141–148 (2012).
Hong, U. G., Hwang, S., Seo, J. G., Lee, J. & Song, I. K. Hydrogenation of succinic acid to γ-butyrolactone (GBL) over palladium catalyst supported on alumina xerogel: Effect of acid density of the catalyst. J. Ind. Eng. Chem. 17, 316–320 (2011).
Werpy, T., Frye, J., J. G., Wang, Y. & Zacher, A. H. Methods of making pyrrolidones. US patent US 6706893 B2 (2004).
Burgard, A., Burk, M. J., Osterhout, R., Van Dien, S. & Yim, H. Development of a commercial scale process for production of 1,4-butanediol from sugar. Curr. Opin. Biotechnol. 42, 118–125 (2016).
Ling, L. B. & Ng, T. K. Fermentation process for carboxylic acids. US patent 4877731 A (1989).
Song, C. W., Kim, D. I., Choi, S., Jang, J. W. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of fumaric acid. Biotechnol. Bioeng. 110, 2025–2034 (2013).
Xu, G., Liu, L. & Chen, J. Reconstruction of cytosolic fumaric acid biosynthetic pathways in Saccharomyces cerevisiae. Microb. Cell Fact. 11, 24 (2012).
Li, N. et al. Engineering Escherichia coli for fumaric acid production from glycerol. Bioresour. Technol. 174, 81–87 (2014).
Battat, E., Peleg, Y., Bercovitz, A., Rokem, J. S. & Goldberg, I. Optimization of l-malic acid production by Aspergillus flavus in a stirred fermentor. Biotechnol. Bioeng. 37, 1108–1116 (1991).
Zambanini, T. et al. Efficient malic acid production from glycerol with Ustilago trichophora TZ1. Biotechnol. Biofuels 9, 67 (2016).
Zhang, X., Wang, X., Shanmugam, K. T. & Ingram, L. O. l-Malate production by metabolically engineered Escherichia coli. Appl. Environ. Microbiol. 77, 427–434 (2011).
Zelle, R. M. et al. Malic acid production by Saccharomyces cerevisiae: engineering of pyruvate carboxylation, oxaloacetate reduction, and malate export. Appl. Environ. Microbiol. 74, 2766–2777 (2008).
Moharregh-Khiabani, D., Linker, R. A., Gold, R. & Stangel, M. Fumaric acid and its esters: an emerging treatment for multiple sclerosis. Curr. Neuropharmacol. 7, 60–64 (2009).
Vert, M. Chemical routes to poly(beta-malic acid) and potential applications of this water-soluble bioresorbable poly(beta-hydroxy alkanoate). Polym. Degradation Stab. 59, 169–175 (1998).
Li, X., Cai, Z., Li, Y. & Zhang, Y. Design and construction of a non-natural malate to 1,2,4-butanetriol pathway creates possibility to produce 1,2,4-butanetriol from glucose. Sci. Rep. 4, 5541 (2014).
Cheong, S., Clomburg, J. M. & Gonzalez, R. Energy- and carbon-efficient synthesis of functionalized small molecules in bacteria using non-decarboxylative Claisen condensation reactions. Nat. Biotechnol. 34, 556–561 (2016).
Zhao, M. et al. Metabolic engineering of Escherichia coli for producing adipic acid through the reverse adipate-degradation pathway. Metab. Eng. 47, 254–262 (2018).
Sato, S., Takahashi, R., Sodesawa, T. & Yamamoto, N. Dehydration of 1,4-butanediol into 3-buten-1-ol catalyzed by ceria. Catal. Commun. 5, 397–400 (2004).
Hunter, S. E., Ehrenberger, C. E. & Savage, P. E. Kinetics and mechanism of tetrahydrofuran synthesis via 1,4-butanediol dehydration in high-temperature water. J. Org. Chem. 71, 6229–6239 (2006).
Zhao, J. & Hartwig, J. F. Acceptorless, neat, ruthenium-catalyzed dehydrogenative cyclization of diols to lactones. Organometallics 24, 2441–2446 (2005).
Subba Rao, Y. V., Kulkarni, S. J., Subrahmanyam, M. & Ramo Rao, A. V. Modified ZSM-5 catalysts for the synthesis of five- and six-membered heterocyclic compounds. J. Org. Chem. 59, 3998–4000 (1994).
Clomburg, J. M. et al. Integrated engineering of beta-oxidation reversal and omega-oxidation pathways for the synthesis of medium chain omega-functionalized carboxylic acids. Metab. Eng. 28, 202–212 (2015).
Yu, J. L., Xia, X. X., Zhong, J. J. & Qian, Z. G. Direct biosynthesis of adipic acid from a synthetic pathway in recombinant Escherichia coli. Biotechnol. Bioeng. 111, 2580–2586 (2014).
Raj, K. et al. Biocatalytic production of adipic acid from glucose using engineered Saccharomyces cerevisiae. Metab. Eng. Commun. 6, 28–32 (2018).
Choi, Y. J., Park, J. H., Kim, T. Y. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of 1-propanol. Metab. Eng. 14, 477–486 (2012).
Zhang, K., Sawaya, M. R., Eisenberg, D. S. & Liao, J. C. Expanding metabolism for biosynthesis of nonnatural alcohols. Proc. Natl Acad. Sci. USA 105, 20653–20658 (2008).
Song, C. W., Lee, J., Ko, Y. S. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of 3-aminopropionic acid. Metab. Eng. 30, 121–129 (2015).
Song, C. W., Kim, J. W., Cho, I. J. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of 3-hydroxypropionic acid and malonic acid through beta-alanine route. ACS Synth. Biol. 5, 1256–1263 (2016).
Chae, T. U., Kim, W. J., Choi, S., Park, S. J. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of 1,3-diaminopropane, a three carbon diamine. Sci. Rep. 5, 13040 (2015).
Mimitsuka, T., Sawai, H., Hatsu, M. & Yamada, K. Metabolic engineering of Corynebacterium glutamicum for cadaverine fermentation. Biosci. Biotechnol. Biochem. 71, 2130–2135 (2007).
Shin, J. H. et al. Metabolic engineering of Corynebacterium glutamicum for enhanced production of 5-aminovaleric acid. Microb. Cell Fact. 15, 174 (2016).
Zhang, J. et al. Application of an acyl-CoA ligase from Streptomyces aizunensis for lactam biosynthesis. ACS Synth. Biol. 6, 884–890 (2017).
Cann, A. F. & Liao, J. C. Production of 2-methyl-1-butanol in engineered Escherichia coli. Appl. Microbiol. Biotechnol. 81, 89–98 (2008).
Lepore, A. W. et al. Catalytic dehydration of biomass derived 1-propanol to propene over M-ZSM-5 (M = H, V, Cu, or Zn). Ind. Eng. Chem. Res. 56, 4302–4308 (2017).
Borodina, I. et al. Establishing a synthetic pathway for high-level production of 3-hydroxypropionic acid in Saccharomyces cerevisiae via beta-alanine. Metab. Eng. 27, 57–64 (2015).
Park, S. J. et al. Metabolic engineering of Escherichia coli for the production of 5-aminovalerate and glutarate as C5 platform chemicals. Metab. Eng. 16, 42–47 (2013).
Adkins, J., Jordan, J. & Nielsen, D. R. Engineering Escherichia coli for renewable production of the 5-carbon polyamide building-blocks 5-aminovalerate and glutarate. Biotechnol. Bioeng. 110, 1726–1734 (2013).
Rohles, C. M., Giesselmann, G., Kohlstedt, M., Wittmann, C. & Becker, J. Systems metabolic engineering of Corynebacterium glutamicum for the production of the carbon-5 platform chemicals 5-aminovalerate and glutarate. Microb. Cell Fact. 15, 154 (2016).
Joo, J. C. et al. Production of 5-aminovaleric acid in recombinant Corynebacterium glutamicum strains from a Miscanthus hydrolysate solution prepared by a newly developed Miscanthus hydrolysis process. Bioresour. Technol. 245, 1692–1700 (2017).
Rohles, C. M. et al. A bio-based route to the carbon-5 chemical glutaric acid and to bionylon-6,5 using metabolically engineered Corynebacterium glutamicum. Green Chem. 20, 4662–4674 (2018).
Qian, Z. G., Xia, X. X. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of cadaverine: a five carbon diamine. Biotechnol. Bioeng. 108, 93–103 (2011).
Buschke, N. et al. Systems metabolic engineering of xylose-utilizing Corynebacterium glutamicum for production of 1,5-diaminopentane. Biotechnol. J. 8, 557–570 (2013).
Kim, H. T. et al. Metabolic engineering of Corynebacterium glutamicum for the high-level production of cadaverine that can be used for the synthesis of biopolyamide 510. ACS Sustain. Chem. Eng. 6, 5296–5305 (2018).
Pronk, J. T. et al. How to set up collaborations between academia and industrial biotech companies. Nat. Biotechnol. 33, 237–240 (2015).
Segler, M. H. S., Preuss, M. & Waller, M. P. Planning chemical syntheses with deep neural networks and symbolic AI. Nature 555, 604–610 (2018).
Kim, W. J., Kim, H. U. & Lee, S. Y. Current state and applications of microbial genome-scale metabolic models. Curr. Opin. Syst. Biol. 2, 9–17 (2017).
Chen, Z., Wilmanns, M. & Zeng, A. P. Structural synthetic biotechnology: from molecular structure to predictable design for industrial strain development. Trends Biotechnol. 28, 534–542 (2010).
Durre, P. & Eikmanns, B. J. C1-carbon sources for chemical and fuel production by microbial gas fermentation. Curr. Opin. Biotechnol. 35, 63–72 (2015).
de Lorenzo, V. et al. The power of synthetic biology for bioproduction, remediation and pollution control: the UN’s Sustainable Development Goals will inevitably require the application of molecular biology and biotechnology on a global scale. EMBO Rep. 19, e45658 (2018).
Acknowledgements
This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556 and NRF-2012M1A2A2026557) from the Ministry of Science and ICT through the National Research Foundation of Korea.
Author information
Authors and Affiliations
Contributions
S.Y.L. conceived the project and designed the study. All authors analysed literature, compiled data, planned the content and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Data 1
Bio-based chemicals map in poster format
Supplementary Information
Supplementary Note 1–2, Supplementary Figure 1, Supplementary Tables 1–3, Supplementary References
Rights and permissions
About this article
Cite this article
Lee, S.Y., Kim, H.U., Chae, T.U. et al. A comprehensive metabolic map for production of bio-based chemicals. Nat Catal 2, 18–33 (2019). https://doi.org/10.1038/s41929-018-0212-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-018-0212-4
This article is cited by
-
The forefront of chemical engineering research
Nature Chemical Engineering (2024)
-
Synthetic microbiology in sustainability applications
Nature Reviews Microbiology (2024)
-
A synthetic methylotrophic Escherichia coli as a chassis for bioproduction from methanol
Nature Catalysis (2024)
-
Systems metabolic engineering of Escherichia coli for hyper-production of 5‑aminolevulinic acid
Biotechnology for Biofuels and Bioproducts (2023)
-
Growth-coupled anaerobic production of isobutanol from glucose in minimal medium with Escherichia coli
Biotechnology for Biofuels and Bioproducts (2023)