Abstract
Accurate physical activity monitoring is essential to understand the impact of physical activity on one’s physical health and overall well-being. However, advances in human activity recognition algorithms have been constrained by the limited availability of large labelled datasets. This study aims to leverage recent advances in self-supervised learning to exploit the large-scale UK Biobank accelerometer dataset—a 700,000 person-days unlabelled dataset—in order to build models with vastly improved generalisability and accuracy. Our resulting models consistently outperform strong baselines across eight benchmark datasets, with an F1 relative improvement of 2.5–130.9% (median 24.4%). More importantly, in contrast to previous reports, our results generalise across external datasets, cohorts, living environments, and sensor devices. Our open-sourced pre-trained models will be valuable in domains with limited labelled data or where good sampling coverage (across devices, populations, and activities) is hard to achieve.
Similar content being viewed by others
Introduction
Cost-effective wearable sensors have gained increasing interest for their potential to revolutionise healthcare owing to their wide range of applications, including fitness and wellness tracking, remote patient monitoring1,2, early disease detection3,4, real-time clinical trials5,6,7, large-scale population health studies8,9,10,11, and personalised medicine12. Consumer-grade devices allow users to obtain summary movement and behaviour metrics such as sleep quality, sedentary time, pace, and step counts. Critical to their effectiveness is the use of reliable algorithms to infer human activities from motion sensor data. However, methodological progress in human activity recognition has been constrained by the limited availability of large representative labelled datasets.
Contrary to fields that have benefited from an explosion of data and subsequent methodological leaps, such as computer vision13,14,15,16,17,18 and natural language processing19,20,21,22, wearables-based human activity recognition research still relies on very small datasets, the majority of which are collected in an artificial setting (e.g., participants following a predefined script in a lab environment and under supervision). Further, this small-data limitation confounds research findings involving data-hungry deep learning methods; for example, there exist empirical studies23,24, suggesting that deep learning methods such as DeepConvLSTM25 did not significantly improve upon more conventional methods relying on simple statistics of the sensor signal.
In this paper, we leverage the UK Biobank accelerometer dataset to realise the full potential of deep learning methods for activity recognition. The UK Biobank is a unique large-scale study that recruited roughly half a million participants, of which more than 100,000 wore a wrist accelerometer for 7 days in their usual environments (as opposed to lab settings), amounting to over 700,000 person-days (and many terabytes) of free-living, 24/7 human motion data.
In order to make use of this unlabelled dataset, we build upon recent advances in self-supervised learning, which have shown great results in this regard, with popular examples such as GPT26. A suite of self-supervised learning methods have been explored for wearable sensor data with success, including multi-task self-supervision27, masked reconstruction28, contrastive learning18,29,30, and bootstraping31,32. A recent benchmark provided a comprehensive assessment of existing self-supervised approaches for human activity recognition and concluded that multi-task self-supervision could learn the most generic features applicable to different downstream tasks33. Existing methods either used the same data for pre-training and fine-tuning or were only trained on datasets with a small size (n = 100), a limiting factor for the generalisability of the pre-trained models. By applying multi-task self-supervision on a large unlabelled dataset with three simple tasks, arrow of time, permutation, and time warping27,34, we showed for the first time that a pre-trained model that could generalise to a wide range of downstream activity recognition datasets important for clinical and health applications.
Our main contributions are:
-
We demonstrate the application of multi-task self-supervised learning on a tera-scale wearables dataset to realise the full potential of deep learning in building state-of-the-art activity recognition models. We discuss engineering challenges in training on large and high-dimensional sensor data and other technical considerations.
-
In contrast to previous works, we conduct a more realistic evaluation of the utility of self-supervised human activity recognition by factoring in common issues seen in practical use cases of pre-trained models such as domain shift and task shift35. In particular, our models show consistent outperformance on external datasets.
-
We release our pre-trained models to enable the digital health research community to build high-performing models for their own use cases. Our models will be especially useful in domains with limited data.
Results
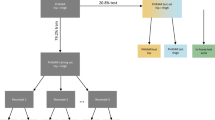
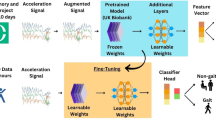
Figure 1 provides a schematic overview of our paper: first, we applied multi-task self-supervised learning to pre-train a deep convolutional neural network on 700,000 person-days of free-living accelerometer data from the UK Biobank; second, the pre-trained network is evaluated via transfer learning on eight benchmark datasets to assess representation quality on various activity types and populations.
Step 1 involves multi-task self-supervised learning on 700,000 person-days of data from the UK Biobank. In step 2, we evaluate the utility of the pre-trained network in eight benchmark human activity recognition baselines via transfer learning. Reproduced and modified with permissions from ref. 37.
Weighted single-task training
When training individual pretext tasks, we found that without weighted sampling, all the tasks had worse convergence behaviour (Fig. 2). The performance degradation was most pronounced for the AoT and permutation. The test performance for the AoT stayed at the random chance level, and the test performance for permutation dropped ~10% points without weighted sampling.
Multi-task self-supervised learning
To investigate how different self-supervision configurations perform in three downstream datasets, we picked one large (Capture-24), medium (Rowlands), and small (Opportunity) dataset for evaluation. We trained different tasks both individually and jointly using 1000 subjects from the UK Biobank, then we fine-tuned the models on the subsequent human activity recognition benchmarks (Table 1).
The differences between different self-supervision combinations on large datasets (Capture-24 and Rowlands) was smaller than that of the smaller dataset (Opportunity). There was no clear best-performing configuration, and thus, for ease of comparison, we chose to use all tasks in pre-training for the remaining experiments. In addition, training more tasks together might yield the most general representation for different downstream datasets.
Downstream performance—human activity recognition
Table 2 summarises the F1 and Kappa scores for eight human activity recognition datasets. The random forest models outperformed the deep learning models trained from scratch for all except the Capture-24 dataset, which is the largest labelled dataset in our evaluations (Table 4). The performance gap between random forest and training from scratch was the largest in smaller datasets. Meanwhile, pre-trained models outperformed the models trained from scratch and random forest in all eight datasets. Fine-tuning all layers was better than fine-tuning just the fully connected layers after the ConV layers.
The most significant improvement using pre-training was seen on the small datasets. Conversely, the benefit of self-supervised learning was more modest for larger datasets. In Capture-24, the F1 improvement was 2.5% when comparing the model with and without self-supervised pre-training. Nonetheless, with self-supervised pre-training, the median relative F1 improvement was 18.4% when compared to the same network trained from scratch and 8.5% when compared to the random forest model.
Transfer learning using labelled pre-training
Even though supervised pre-training can boost the learning outcome substantially more than training from scratch (Table 2 vs Table 3), self-supervised pre-training without labels could outperform supervised pre-training when using Rowlands and Capture-24 as the source data.
Ablation studies
Varying labelled data in the downstream
We observed that pre-trained models did well regardless of the number of labelled subjects in two downstream datasets (Fig. 3a). However, fully supervised and random forest models were more susceptible to the number of labelled subjects. The performance gain for having more labelled subjects was roughly linear with respect to the number of subjects included with a greater increase when we had fewer labelled subjects.
Varying unlabelled data in the pre-training
We also found that the downstream human activity recognition performance appeared to increase linearly with respect to the number of unlabelled subjects on a log scale (Fig. 3b, left). The self-supervision performance boost with more unlabelled subjects pre-training was most significant in the smallest dataset, Opportunity. Furthermore, if the number of participants is fixed at 10,000 in pre-training, the data ratio included per subject did not significantly influence the downstream performance. Notably, the downstream performance did not degrade more than 10% in F1 even when we reduced the the amount of data per subject from 100 to 25%. (Fig. 3b, right).
Understanding the representation
Cluster analysis
We used UMAP36 with default parameters for low-dimensional projections for visualisation. This was applied to the raw inputs, untrained features, and self-supervision-derived features without fine-tuning. Results for two of the downstream datasets are shown in Fig. 4, and the remaining results can be found in Supplementary Fig. 2. Across all datasets, we observed that the self-supervision-derived features were better at clustering similar activities (e.g., walking, stair climbing vs sitting, writing, typing) as well as their intensities (e.g., lying down, sitting, standing vs jogging, sports), exhibiting better intra-class compactness and inter-class separability.
Feature interpretation
Next, we visualised two exemplary pretext self-supervised task predictions in the presence of repetitive low- and high-intensity activities: shaking hands (Supplementary Fig. 4a) and playing tennis (Supplementary Fig. 4b). During tennis playing, a repetitive high, intensity activity, relevance scores tended to highlight the moments around the natural movements of swinging and hitting the tennis ball (Supplementary Figs. 4b and 5). When performing a repetitive low-intensity activity experiment, for example, shaking hands (Supplementary Fig. 4a), layer-wise relevance propagation appeared to also identify the intensity and natural signal periodicity as indicative of the original activity. In contrast, for augmented signals, our model attributed more during periods of visually unrealistic motion dynamics, such as unnatural fragmentation in activity frequency or synchronisation mismatches between sensor axes. Interestingly, stationary movement periods were not relevant for detecting the pretext tasks.
Finally, we empirically compared the faithfulness of the explainable AI algorithms investigated and the combination of various layer-wise relevance propagation parameters, using sample-masking experiments for a random subset of 1000 (out-of-sample) subjects in the UK Biobank. Most explainable AI models consistently demonstrated the ability to identify relevant patterns for discriminating transformed samples from the original raw data when compared against a random model.
Discussion
Our work has shown that self-supervised pre-training consistently improved downstream human activity recognition, especially in small datasets, reducing the need for labelled data. The self-supervised representations generalise well across a range of external datasets, tasks, devices, health statuses, and populations, a key aspect in activity monitoring for clinical use. Our work represents the most robust human activity recognition foundational model to date, as it is trained on a much larger and more diverse dataset than previous efforts in this space. The scale of UK Biobank is several orders of magnitude greater than the largest datasets used by the current state-of-the-art such as the Fenland data9 (n = 160,000). In addition, not just big in size, but UK Biobank is much more diverse as it contains hundreds, if not thousands, of natural human activities—a crucial aspect regarding the training data and feature generalisation. Indeed, the obtained pre-trained model has already been used with success in enhancing digital monitoring for a clinical population with motor impairment37 and epidemiological research38,39.
With our pre-trained models, one can obtain a highly competitive activity recognition model with a small amount of labelled data, a feature important for clinical studies where labelled data is expensive to acquire. In contrast, previous studies applied self-supervised training and fine-tuning only on the same data sources24,27,40, making it necessary to pre-train on every new dataset in practical applications. A recent attempt has been made to systematically evaluate the effect of self-supervised techniques by pre-training on Capture-24, providing a good baseline for the performance evaluation in human activity recognition33. However, pre-training on Capture-24 with roughly 100 participants will not be able to characterise the impact of self-supervised pre-trained models. The representation quality is superior if the pre-training is done on datasets with richer population characteristics. Our pre-trained network can serve as a foundational human activity recognition model that removes the need to pre-train on unseen datasets.
We found that the representation quality from self-supervision was always better than that of supervised learning in an apple-to-apple comparison when using Rowlands as the source of pre-training. Self-supervised learning with other modalities has also found that self-supervised pre-training can outperform supervised pre-training18,41. Pre-training on the UK Biobank yields the most significant improvement among all the self-supervised representations, as the human activity recognition performance is uniformly better than other pre-trained baselines (Table 3). The performance boost could be attributed to the large data volume, diverse activity classes and rich population characteristics of the UK Biobank over alternative pre-training datasets. Recent investigations on large language models have profiled the trade-off between model size and data volume when the compute budget is fixed42, suggesting model size is another important dimension for pre-training worthy of further investigation for human sensing data.
This study highlights new questions to prioritise in future. Due to a current lack of raw accelerometer datasets in different regions of the world, a limitation of our work is that the pre-training data (UK Biobank) consists mostly of Caucasians from the UK. A multi-modal representation that also includes electrocardiogram and other time-series wearable sensor data sources will also be important to consider when such datasets are collected in the future. Lastly, future work could also investigate the representation quality using more recent self-supervised learning approaches on free-living UK Biobank accelerometers28,32,33,40,43, in addition to multi-task learning. We attempted to use Autoencoder and contrastive learning for pre-training. However, we could not obtain high-quality representation using the UK Biobank43. We suspect this was mainly due to the difference between free-living and lab-based activity data, which can be further analysed to compare the performance of different self-supervised methods.
We have developed and evaluated a self-supervised deep neural network on large-scale activity tracking data. The features obtained improved on prior state-of-the-art performance across eight benchmark human activity recognition datasets. Our open-sourced model represents a foundation model that others can build upon for state-of-the-art human activity recognition applications. The improved physical activity measurement will help to understand better the influence of physical activity on different disease outcomes, especially for populations that have been under-represented in previous studies.
Methods
We used tri-axial accelerometer data from wrist-worn activity trackers, which record acceleration on three orthogonal axes at a high sampling rate (e.g., 100 Hz). The main benefit of wrist-worn activity trackers is their high user compliance, resulting in days, if not weeks, of continuous recordings. Following ref. 44, we split the signals into windows of equal duration, effectively treating them as independent inputs to the human activity recognition models. We can then label each window with an activity class. Throughout this study, we linearly resampled all data to 30 Hz resolution and used ten-second-long windows to compare the downstream benchmarks fairly. The 30 Hz sampling rate was used because most human activities have a frequency less than 10 Hz. We used a sampling rate that is higher than the presumed Nyquist rate (20 Hz) to ensure that we did not lose any useful signal.
Datasets
Our multi-task self-supervised training relied on the unlabelledUK Biobank dataset, which contains roughly 700,000 person-days of free-living activity data (>100,000 participants, 7 days of wear). The free-living aspect is important because the data can contain all sorts of activities, as opposed to lab data which are constrained to scripted activities only. The UK Biobank data (project ref 21/NW/0157) is covered by ethical approval from the NHS National Research Ethics.
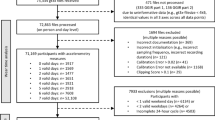
For the subsequent activity recognition benchmarks, we considered eight external labelled datasets that vary in size (600–600,000 samples), activity classes (4–18 classes), devices (5 different brands), device placements (4 configurations), populations (differ in age, sex, and health status), and collection protocol (free-living, scripted, and lab settings). See Table 4 and Supplementary Table 1 for detailed dataset characteristics. Three datasets had license information, and five datasets had explained informed consent information (Supplementary Table 2). We removed the classes that were not present in all the subjects in small datasets with less than ten individuals during data cleaning. The Michael J. Fox Foundation Levodopa Response (MJFF-LR) study was included to assess the generalisability of our model in a clinical population with motor impairment, Parkinson’s disease in our case. For the MJFF-LR study, we only included data collected in a lab because not all the participants had free-living data. Finger-to-nose and repeated-arm movement tasks were also removed from MJFF-LR as these two activities were performed using both arms in alternation, but we only used the data from one arm. We further merged three walking classes into one.
Even though we reused existing datasets, we made our best effort to enumerate the license and consent information for all the included datasets, as our data involved human subjects. We observed that many open benchmark datasets that we used did not have suitable licensing or consent information, possibly due to the lack of data governance awareness at the time of the study.
Multi-task self-supervised-learning
We considered three self-supervised tasks from ref. 27, which were first used in ref. 34 as data augmentation techniques. Eight transformations were included in the previous exploration of multi-task learning27. We chose arrow of time, permutation and time warping to maximise learning features related to human motion dynamics. Supplementary Methods Section explains why other transformations were not chosen.
Arrow of time (AoT) flips the signal along the time axis, effectively playing the signal in reverse. Permutation breaks the signal into chunks and shuffles them. We set the number of chunks to four and the minimum length of each chunk to at least ten timestamps. Time warping (TW) stretches and compresses arbitrary segments of the signal, effectively slowing down and speeding up the signal randomly.
Following ref. 27, we treated each of the tasks as a binary problem predicting whether a transformation has been applied. In the multi-task learning (MTL) setting, not all the tasks might benefit human activity recognition when trained jointly, so we assessed how different task combinations could influence the downstream performance. We computed the cross-entropy loss for each task and weighed all the tasks equally in the loss calculation.
Weighted sampling
Motion data collected in the real world contains large portions of low movement periods that are less informative (Supplementary Fig. 1), which is an issue for our self-supervised tasks as static signals remain virtually unchanged after the transformations. We found it crucial to perform weighted sampling for improved training stability and convergence: during training, we sample the data windows in proportion to their standard deviation so as to give more weight to high-movement periods.
Network training
We adapted a ResNet-V2 with 18 layers and 1D convolutions45 for the main trunk (feature extractor), totalling 10M parameters. The learned feature vector was of size 1024. All the tasks shared the same feature extractor. Then, we attached a softmax layer for each of the self-supervised tasks. In the downstream evaluation, we added a fully connected (FC) layer of size 512 in between the feature extractor and softmax readout. The network structure was fixed for all the downstream evaluations.
For self-supervised learning, we load up to four subjects from the UK Biobank at each iteration. For each subject, we first sampled one day out of the week-long data, from which we again sampled 1500 10-s windows to make up a training batch. Self-supervised transformations were then applied to the batch of data. Since the axis orientation differs between device manufacturers, we used random axis swaps and rotations to augment the training data to embed this invariance into our models. For optimisation, we used Adam46 with a learning rate of 1e-3. To account for large batch sizes, 1500 × 4 = 6000, we applied linear scaling for the learning rate with five epochs as burn-in47. We distributed the network training over four Tesla V100-SXM2 with 32 GB of memory. It took about 420 GPU hours to train the MTL model (about 20 epochs). We used an 8:2 ratio for the train/test split for all the self-supervised experiments. For fine-tuning, we used the same training setup as the pre-training where possible, except for the batch size, which was re-adjusted depending on the size of each dataset.
Evaluation—human activity recognition
To evaluate the downstream human activity recognition performance, we used held-one-subject-out cross-validation for the datasets that had <10 subjects. We additionally removed activity classes not done by all the subjects in these small datasets. For datasets with ≥10 subjects, we used five-fold subject-wise cross-validation instead. Each cross-validation had a 7:1:2 split ratio for train/validation/test sets. We used early-stopping with a patience of five to avoid over-fitting. For training runs that did not converge, we reported the best performance after using three different random seeds for weight initialisation.
After the network was trained on the UK Biobank using ~100,000 participants, we further fine-tuned the network on the eight labelled downstream datasets to perform human activity detection using two approaches: (1) fine-tuning all the layers (2) freezing the trunk (feature extractor) and fine-tuning only the FC layers in the end. We also report the model performance for a network of the same architecture but fully trained from scratch, and a strong random forest model with tried-and-tested time series features, which has often been neglected in baseline model comparisons8,48,49,50. See the Supplementary Methods Section for the list of features used.
In addition, a shared implementation was introduced for our network training, model evaluation and preprocessing. Differences in experiment setup such as training rates, regularisation and data augmentation can lead to inconsistent results51. A unified evaluation framework would ensure a fair comparison between different baseline models. Our evaluation framework contrasts with previous work, where there is no fixed evaluation protocol across the benchmark datasets, making it hard to compare model performance with the current state-of-the-art. The results produced in our paper would serve as the baseline for future human activity recognition research.
Transfer learning
Pre-training on a larger labelled dataset and fine-tuning on a smaller dataset is a common technique in practical application that has been under-reported as a baseline for self-supervision. The success of transfer learning, however, depends on how similar the source and target datasets are. Hence, we included experiments using the two largest labelled datasets, Capture-24 and Rowlands for pre-training, which were then fine-tuned on other labelled datasets.
The benefits of data volume
In the ablation studies, we investigated how the downstream performance differs on two axes, the amount of labelled data and the amount of unlabelled data. Concretely, we gradually increase the number of labelled subjects in both Capture-24 and Rowlands in the downstream evaluation to assess whether our pre-trained model can still do well in a limited-data regime. In terms of unlabelled data, we experimented with pre-training that had 100 to 100,0000 participants with one order of magnitude increment. We also varied the amount of unlabelled data per subject from 0.25 to 1 using 10,000 participants. A data ratio of 0.25 means that if one day of data per subject was used previously, then only 6 h of data per subject would now be used for training. Investigating how unlabelled data influences downstream performance will guide how much data one needs to have to obtain an effective self-supervised model for human activity recognition.
Understanding network representation
Contextualising layer-wise relevance propagation
We applied layer-wise relevance propagation (LRP) to visually investigate the signal characteristics relevant for detecting the pretext tasks52,53. It is inherently more difficult to visually interpret attribution heatmaps generated through Explainable AI (XAI) frameworks on time-series signals. To overcome this lack of visual ground truth, we devised a set of simple contextual experiments to evaluate our LRP attribution results. Using the same accelerometer as the UK Biobank, we recorded a participant performing two activities under video observation: (1) low-intensity scripted (hand-shaking) and (2) high-intensity unscripted (playing tennis). We acquired a ground truth (the context) for the accelerometer activity through the time-synced video observations, enabling a better visual interpretation of the sensor-based characteristics attributed as relevant for detecting different pretexts. Holistic interpretations were formed based on visualising the raw sensor signal, its analogues time-frequency representation through continuous wavelet transform (CWT) scalograms54, as well as the time- and pretext task-localised LRP relevance scores, all with respect to observing the concurrent video recordings. Details on the XAI contextual LRP (cLRP) framework are described in the Supplementary Methods Section.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The UK Biobank accelerometer dataset used for training can be requested by application (https://www.ukbiobank.ac.uk/enable-your-research/register). All the other evaluation datasets can be downloaded via https://github.com/OxWearables/ssl-wearables. The MJFF-LR study can be accessed via https://www.synapse.org/#!Synapse:syn20681023/wiki/594686 after registration. The Rowlands dataset can be requested by contacting Alex Rowlands directly.
Code availability
The underlying code for for this study is available in OxWearables/ssl-wearables on Github and can be accessed via this link https://github.com/OxWearables/ssl-wearables.
References
Seshadri, D. R. et al. Wearable sensors for covid-19: a call to action to harness our digital infrastructure for remote patient monitoring and virtual assessments. Front. Digital Health 2, 558695 (2020).
Small, S. R. et al. Current clinical utilisation of wearable motion sensors for the assessment of outcome following knee arthroplasty: a scoping review. BMJ Open 9, e033832 (2019).
Lubitz, S. A. et al. Detection of atrial fibrillation in a large population using wearable devices: the Fitbit heart study. Circulation 146, 1415–1424 (2022).
Cheong, S. H. R., Ng, Y. J. X., Lau, Y. & Lau, S. T. Wearable technology for early detection of covid-19: a systematic scoping review. Prevent. Med. 162, 107170 (2022).
Munos, B. et al. Mobile health: the power of wearables, sensors, and apps to transform clinical trials. Annal. NY Acad. Sci. 1375, 3–18 (2016).
Izmailova, E. S., Wagner, J. A. & Perakslis, E. D. Wearable devices in clinical trials: hype and hypothesis. Clin. Pharmacol. Ther. 104, 42–52 (2018).
Beauchamp, U. L., Pappot, H. & Holländer-Mieritz, C. The use of wearables in clinical trials during cancer treatment: systematic review. JMIR mHealth uHealth 8, e22006 (2020).
Willetts, M., Hollowell, S., Aslett, L., Holmes, C. & Doherty, A. Statistical machine learning of sleep and physical activity phenotypes from sensor data in 96,220 UK biobank participants. Sci. Rep. 8, 1–10 (2018).
Lindsay, T. et al. Descriptive epidemiology of physical activity energy expenditure in UK adults (the Fenland study). Int. J. Behav. Nutr. Phys. Activity 16, 1–13 (2019).
Straczkiewicz, M., James, P. & Onnela, J.-P. A systematic review of smartphone-based human activity recognition methods for health research. NPJ Digit. Med. 4, 148 (2021).
Khurshid, S. et al. Wearable accelerometer-derived physical activity and incident disease. NPJ Digit. Med. 5, 131 (2022).
Dunn, J., Runge, R. & Snyder, M. Wearables and the medical revolution. Pers. Med. 15, 429–448 (2018).
Doersch, C., Gupta, A. & Efros, A. A. Unsupervised visual representation learning by context prediction. In Proceedings of the IEEE International Conference on Computer Vision 1422–1430 (IEEE, 2015).
Zhang, R., Isola, P. & Efros, A. A. Colorful image colorization. In Computer Vision–ECCV 2016: 14th European Conference, Amsterdam, The Netherlands, October 11-14, 2016, Proceedings, Part III 14 649–666 (Springer, 2016).
Noroozi, M. & Favaro, P. Unsupervised learning of visual representations by solving jigsaw puzzles. In European Conference on Computer Vision, 69–84 (Springer, 2016).
Wei, D., Lim, J. J., Zisserman, A. & Freeman, W. T. Learning and using the arrow of time. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 8052–8060 (IEEE, 2018).
He, K., Fan, H., Wu, Y., Xie, S. & Girshick, R. Momentum contrast for unsupervised visual representation learning. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 9729–9738 (IEEE, 2020).
Chen, T., Kornblith, S., Norouzi, M. & Hinton, G. A simple framework for contrastive learning of visual representations. In International Conference on Machine Learning, 1597–1607 (PMLR, 2020).
Mikolov, T., Chen, K., Corrado, G. & Dean, J. Efficient estimation of word representations in vector space. Preprint at https://arxiv.org/abs/1301.3781 (2013).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. In Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Vol. 1 (Long and Short Papers) 4171–4186 (Association for Computational Linguistics, Minneapolis, Minnesota, 2019).
Radford, A., Narasimhan, K., Salimans, T. & Sutskever, I. Improving language understanding with unsupervised learning. (Name of the Blog, [Online], accessed 11 June 2018); Available from: https://openai.com/research/language-unsupervised
Radford, A. et al. Language models are unsupervised multitask learners. OpenAI Blog 1, 9 (2019).
Twomey, N. et al. A comprehensive study of activity recognition using accelerometers. In Informatics, Vol. 5, 27 (Multidisciplinary Digital Publishing Institute, 2018).
Haresamudram, H., Anderson, D. V. & Plötz, T. On the role of features in human activity recognition. In Proceedings of the 2019 ACM International Symposium on Wearable Computers (ISWC '19), 78–88 (Association for Computing Machinery, New York, NY, USA); https://doi.org/10.1145/3341163.3347727
Ordóñez, F. J. & Roggen, D. Deep convolutional and lstm recurrent neural networks for multimodal wearable activity recognition. Sensors 16, 115 (2016).
Brown, T. et al. Language models are few-shot learners. Adv. Neural Inf. Process. Syst. 33, 1877–1901 (2020).
Saeed, A., Ozcelebi, T. & Lukkien, J. Multi-task self-supervised learning for human activity detection. In Proceedings of the ACM on Interactive, Mobile, Wearable and Ubiquitous Technologies Vol. 3, 1–30 (ACM, 2019).
Haresamudram, H. et al. Masked reconstruction based self-supervision for human activity recognition. In Proceedings of the 2020 ACM International Symposium on Wearable Computers (ISWC '20), 45–49 (Association for Computing Machinery, New York, NY, USA, 2020); https://doi.org/10.1145/3410531.3414306
Tang, C. I., Perez-Pozuelo, I., Spathis, D. & Mascolo, C. Exploring contrastive learning in human activity recognition for healthcare. Preprint at https://arxiv.org/abs/2011.11542 (2020).
Haresamudram, H., Essa, I. & Plötz, T. Contrastive predictive coding for human activity recognition. In Proceedings of the ACM on Interactive, Mobile, Wearable and Ubiquitous Technologies Vol. 5, 1–26 (ACM, 2021).
Grill, J.-B. et al. Bootstrap your own latent-a new approach to self-supervised learning. Adv. Neural Inf. Process. Syst. 33, 21271–21284 (2020).
Shah, K., Spathis, D., Tang, C. I. & Mascolo, C. Evaluating contrastive learning on wearable timeseries for downstream clinical outcomes. Preprint at https://arxiv.org/abs/2111.07089 (2021).
Haresamudram, H., Essa, I. & Plötz, T. Assessing the state of self-supervised human activity recognition using wearables. In Proceedings of the ACM on Interactive, Mobile, Wearable and Ubiquitous Technologies Vol. 6, 1–47 (ACM, 2022).
Um, T. T. et al. Data augmentation of wearable sensor data for Parkinson’s disease monitoring using convolutional neural networks. In Proceedings of the 19th ACM International Conference on Multimodal Interaction, ICMI 2017, 216–220 (ACM, 2017).
Quiñonero-Candela, J., Sugiyama, M., Schwaighofer, A. & Lawrence, N. D. Dataset Shift in Machine Learning (MIT Press, 2008).
McInnes, L., Healy, J. & Melville, J. Umap: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/pdf/1802.03426.pdf(2018).
Creagh, A. C. et al. Digital health technologies and machine learning augment patient reported outcomes to remotely characterise rheumatoid arthritis. NPJ Digit. Med. 7, 33 (2022).
Small, S. R. et al. Development and validation of a machine learning wrist-worn step detection algorithm with deployment in the UK biobank. Preprint at https://doi.org/10.1101/2023.02.20.23285750 (2023).
Yuan, H. et al. Self-supervised learning of accelerometer data provides new insights for sleep and its association with mortality. Preprint at https://www.medrxiv.org/content/10.1101/2023.07.07.23292251v1, https://doi.org/10.1038/s41746-024-01065-0 (2023).
Haresamudram, H., Essa, I. & Plötz, T. Contrastive Predictive Coding for Human Activity Recognition. Proc. ACM Interact. Mob. Wearable Ubiquitous Technol. 5, 26 (2021).
Tomasev, N. et al. Pushing the limits of self-supervised resnets: can we outperform supervised learning without labels on imagenet? Preprint at https://arxiv.org/abs/2201.05119 (2022).
Hoffmann, J. et al. Training compute-optimal large language models. Preprint at https://arxiv.org/abs/2203.15556 (2022).
Tang, C. I. et al. SelfHAR: Improving Human Activity Recognition through Self-training with Unlabeled Data. Proc. ACM Interact. Mob. Wearable Ubiquitous Technol. 5, 30 (2021).
Bulling, A., Blanke, U. & Schiele, B. A tutorial on human activity recognition using body-worn inertial sensors. ACM Comput. Surv. (CSUR) 46, 1–33 (2014).
He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks. In ECCV, 630–645 (Springer, 2016).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Goyal, P. et al. Accurate, large minibatch sgd: Training imagenet in 1 hour. Preprint at https://arxiv.org/abs/1706.02677 (2017).
Zhang, S. et al. Physical activity classification using the GENEA wrist-worn accelerometer. Med. Sci. Sports Exerc. 44, 742–748 (2012).
Mannini, A., Intille, S. S., Rosenberger, M., Sabatini, A. M. & Haskell, W. Activity recognition using a single accelerometer placed at the wrist or ankle. Med. Sci. Sports Exerc. 45, 2193 (2013).
Ellis, K., Kerr, J., Godbole, S., Staudenmayer, J. & Lanckriet, G. Hip and wrist accelerometer algorithms for free-living behavior classification. Med. Sci. Sports Exerc. 48, 933 (2016).
Oliver, A., Odena, A., Raffel, C. A., Cubuk, E. D. & Goodfellow, I. J. Realistic evaluation of deep semi-supervised learning algorithms. In Proceedings of the 32nd International Conference on Neural Information Processing Systems (NIPS'18), Vol. 31, 3239–3250 (Curran Associates Inc., Red Hook, NY, USA, 2018).
Montavon, G., Binder, A., Lapuschkin, S., Samek, W. & Müller, K.-R. Layer-Wise Relevance Propagation: An Overview. In Explainable AI: Interpreting, Explaining and Visualizing Deep Learning. Lecture Notes in Computer Science (eds Samek, W., Montavon, G., Vedaldi, A., Hansen, L. & Müller, K.-R.) vol. 11700, 193–209 (Springer, Cham, 2019); https://doi.org/10.1007/978-3-030-28954-6_10
Creagh, A. P., Lipsmeier, F., Lindemann, M. & Vos, M. D. Interpretable deep learning for the remote characterisation of ambulation in multiple sclerosis using smartphones. Sci. Rep. 11, 14301 (2021).
Addison, P. S., Walker, J. & Guido, R. C. Time–frequency analysis of biosignals. IEEE Eng. Med. Biol. Magaz. 28, 14–29 (2009).
Doherty, A. et al. Large scale population assessment of physical activity using wrist worn accelerometers: the UK biobank study. PLoS ONE 12, e0169649 (2017).
Esliger, D. et al. Validation of the GENEA accelerometer. Med. Sci. Sports Exerc. 43, 1085–1093 (2011).
Weiss, G. M., Yoneda, K. & Hayajneh, T. Smartphone and smartwatch-based biometrics using activities of daily living. IEEE Access 7, 133190–133202 (2019).
Daneault, J.-F. et al. Accelerometer data collected with a minimum set of wearable sensors from subjects with Parkinson’s disease. Sci. Data 8, 48 (2021).
Sztyler, T. & Stuckenschmidt, H. On-body localization of wearable devices: an investigation of position-aware activity recognition. In 2016 IEEE International Conference on Pervasive Computing and Communications (PerCom), 1–9 (IEEE, 2016).
Roggen, D. et al. Collecting complex activity datasets in highly rich networked sensor environments. In 2010 Seventh International Conference on Networked Sensing Systems (INSS), 233–240 (IEEE, 2010).
Reiss, A. & Stricker, D. Introducing a new benchmarked dataset for activity monitoring. In 2012 16th International Symposium on Wearable Computers, 108–109 (IEEE, 2012).
Bruno, B., Mastrogiovanni, F., Sgorbissa, A., Vernazza, T. & Zaccaria, R. Analysis of human behavior recognition algorithms based on acceleration data. In 2013 IEEE International Conference on Robotics and Automation, 1602–1607 (IEEE, 2013).
Acknowledgements
The authors would like to thank all the helpful discussions and feedback we recevied from Gert Mertes, Henrique Aguiar, Andres Tamm, and Korsuk Sirinukunwattana. This research has been conducted using the UK Biobank Resource under Application Number 59070. This work is supported by: Novo Nordisk (H.Y., S.C. and A.D.); the Wellcome Trust [223100/Z/21/Z] (AD); GlaxoSmithKline (A.C., A.A. and D.C.); the British Heart Foundation Centre of Research Excellence [RE/18/3/34214] (A.D.); the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (A.D. and D.C.); and Health Data Research UK, an initiative funded by UK Research and Innovation, Department of Health and Social Care (England) and the devolved administrations, and leading medical research charities. It is also supported by the UK’s Engineering and Physical Sciences Research Council (EPSRC) with grants EP/S001530/1 (the MOA project) and EP/R018677/1 (the OPERA project); and the European Research Council (ERC) via the REDIAL project (Grant Agreement ID: 805194), and industrial funding from Samsung AI. D.A.C. was also supported by the Pandemic Sciences Institute at the University of Oxford; the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (BRC); an NIHR Research Professorship; a Royal Academy of Engineering Research Chair; and the InnoHK Hong Kong Centre for Centre for Cerebro-cardiovascular Engineering (COCHE). For the purpose of open access, the author has applied a CC-BY public copyright licence to any author-accepted manuscript version arising from this submission. The funding for the Levodopa Response Trial Data is from the Michael J. Fox Foundation. The authors would also like to thank Alex Rowlands and Mike Catt, who kindly shared their activity dataset with us. Their project was funded by a grant from Unilever Discover to the School of Sports and Health Sciences, University of Exeter. No funding bodies had any role in the analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
H.Y. and S.C.—conceptualisation, data processing, model development, and writing. A.P.C. and C.T.—model development and writing. D.A.C.—reviewing and supervision. A.D.—conceptualisation, model development, reviewing, and supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, H., Chan, S., Creagh, A.P. et al. Self-supervised learning for human activity recognition using 700,000 person-days of wearable data. npj Digit. Med. 7, 91 (2024). https://doi.org/10.1038/s41746-024-01062-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-024-01062-3