Abstract
Though advanced cancers generally display complex molecular portfolios, there is a subset of patients whose malignancies possess only one genomic alteration or alterations in one oncogenic pathway. We assess how N-of-One therapeutic strategies impact outcomes in these patients. From 12/2012 to 9/2018, 429 therapy-evaluable patients with diverse treatment-refractory cancers were presented at Molecular Tumor Boards at Moores Cancer Center at UC San Diego. The clinical benefit rate, defined by RECIST1.1, was assessed for patients with solid tumors who underwent next-generation sequencing (NGS) profiling revealing one genomic or pathway alteration, subsequently managed with N-of-One therapies. Nine of 429 patients (2.1%) met evaluation criteria. Using matched therapy indicated by NGS, the clinical benefit rate (stable disease ≥ 6 months/partial/complete response) was 66.7%. Median progression-free survival was 11.3 months (95% CI: 3.4–not evaluable). Thus, a small subset of diverse cancers has single pathway alterations on NGS testing. These patients may benefit from customized therapeutic matching.
Similar content being viewed by others
Recent advances in precision medicine have quickly transformed treatment strategies for patients with advanced cancers. Currently, genomic testing allows physicians to identify mutated genes and tailor treatment to precisely target the alterations in their malignancies. This approach has been therapeutically beneficial for several cancer types with particular genomic alterations, including BCR-ABL kinase inhibition for chronic myeloid leukemia and KIT inhibition for gastrointestinal stromal tumor (GIST), as well as BRAF and Her2 inhibition for multiple tumor types1,2. Furthermore, immunotherapies, such as anti-PD-1/PD-L1 blockades, have shown durable responses among patients with high tumor mutational burden (TMB), microsatellite instability-high (MSI-high), and PD-L1 overexpression/amplification3,4,5.
Despite the identification of numerous biomarkers for targeted therapy, oncology medications have significantly higher drug development attrition rates than medications for non-oncology indications6. This is partly because drugs that fail to elicit a durable response in a significant subgroup of patients are frequently abandoned, even if the drug exhibits significant activity in a small proportion of people7. Two of the major obstacles to treatment are intra-tumor heterogeneity, in which multiple genomic clonal populations exist within a neoplasm, and inter-tumor heterogeneity, in which multiple tumors in the same patient possess different co-occurring genomic alterations. In fact, patients with advanced cancer harbor a median of five unique oncogenic alterations, suggesting that therapeutics should be individualized and, if indicated, utilize a combination approach8. Still, when multiple co-driver alterations exist, it seems likely that patients will exhibit primary or secondary resistance to targeted therapeutic strategies.
Accordingly, we hypothesized that patients whose advanced cancers harbored a single alteration or alterations in a single genomic pathway on interrogation with next-generation sequencing (NGS) would respond especially well to cognate targeted therapy. Herein, we show that such individuals, while uncommon, can often achieve objective and durable responses when administered agents that are well matched to their molecular alteration(s) across a spectrum of cancer types and genomic abnormalities.
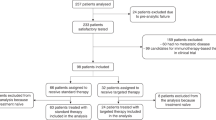
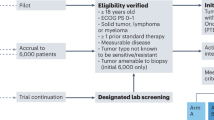
Overall, 715 distinct patients with advanced cancer were discussed at face-to-face Molecular Tumor Board (MTB) meetings. Among 429 patients who were subsequently treated and evaluable for outcome analysis, nine patients had a single genomic alteration or alterations in one molecular pathway that were treated with matched targeted therapy (Fig. 1). All nine patients had NGS performed on tissue by Foundation Medicine (FoundationOne™, Cambridge, Massachusetts, http://www.foundationmedicine.com) (Clinical Laboratory Improvement Amendments (CLIA)-certified). The FoundationOne™ tissue assay utilized during the study period interrogated between 182 and 324 cancer-related genes. Median patient age was 41 years (range, 14–72 years). They received a median of two lines of therapy, including matched therapy indicated by NGS results (range, 1–5).
Only solid tumors were included. Patients who had only one alteration on an initial profiling test but subsequently had more genomic profiling after MTB presentation and demonstrated additional alterations were excluded. All included patients had NGS profiling. Patients were treated within six months of MTB. Patients who received immunotherapy based on MSI-high or TMB-high were not included in the current analysis9.
Eight patients received small molecule targeted agent(s), while one patient with PDL1 amplification received immune checkpoint blockade. Among the nine patients, three patients achieved partial response (PR) and an additional three patients achieved stable disease (SD) ≥ 6 months. The remaining three patients had progressive disease (PD). Altogether, the clinical benefit rate was 66.7% (i.e., 6/9 patients) (Table 1). Median progression-free survival (PFS) was 11.3 months (95% CI: 3.4–not evaluable), while median overall survival (OS) was not reached (95% confidence interval (CI): 8.4–not evaluable) (Fig. 2). Only Patient #5467 (Table 1), who received immunotherapy, experienced a serious adverse event (SAE), namely Grade 3 pancreatitis. No other Grade 3–4 adverse events were observed, according to Common Terminology Criteria for Adverse Events.
Recent literature on molecular profiling technologies has revealed that advanced cancer patients harbor a median of five molecular alterations8. Although rare, some patients may harbor only one molecular alteration after interrogation of several hundred oncogenic markers with NGS9. In this study, the administration of N-of-One treatments was retrospectively reviewed to assess how this approach impacted clinical benefit rates (i.e., PR + SD ≥ 6 months), PFS, and OS in patients harboring one alteration or alterations in one oncogenic pathway.
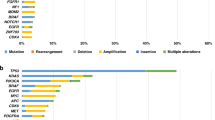
Overall, 2.1% (9/429) of evaluable patients had one gene/pathway alteration targeted with molecularly matched agents. Six of these nine patients achieved clinical benefit. The pathways successfully targeted included BRAF/MEK, CDK4/6, FGFR, KIT, and RET pathways, as well as PD-L1 amplification-associated immune suppression. Notably, the patient with a BRAF V600E mutant ovarian serous carcinoma tumor (i.e., not melanoma) was successfully treated with BRAF inhibitor dabrafenib and MEK inhibitor trametinib (PR ongoing at 10.7+ months). Furthermore, one of two patients with alterations expected to activate CDK4/6 was successfully treated with a CDK4/6 inhibitor as monotherapy (ovarian undifferentiated neuroendocrine cancer, PR lasting 8.4 months); this is in contrast to observations suggesting that matched CDK4/6 inhibitor monotherapy, such as palbociclib, is ineffective10, possibly for patients with multiple co-occurring mutations, unlike the patient discussed above. On the other hand, three of the nine patients did poorly. It is plausible that, despite a single pathway alteration on genomics, other important driver pathways were altered at the transcript or protein level in these patients. As such, in addition to tissue genomic profiling, more comprehensive analysis that includes cell-free DNA, transcriptomics, epigenetics, and immune profiling may be considered in the care of future patients.
Limitations of this paper include the retrospective nature of analysis, small sample size, and lack of controls. Despite these limitations, this study provides a window into the opportunity to leverage NGS to prescribe personalized matched therapies for patients with incurable malignancies whose cancers have not yet shown complicated genomic evolution. Trials such as NCI-MATCH11 and MSK-IMPACT12 have demonstrated the viability of deploying NGS to triage patients to targeted therapeutics, while I-PREDICT8 and WINTHER13 have evaluated the outcomes of patients receiving therapies that prioritize combination therapy matched to complex genomic and/or transcriptomic profiles. The current study suggests that there is a small subset of patients (~2%) with advanced metastatic disease whose tumors still demonstrate only single pathway alterations on NGS, and that such cancers remain amenable to focused pathway targeting.
Methods
Patient selection
We investigated the molecular profiling status (performed by CLIA-certified laboratories) and clinical outcomes of patients with advanced cancer presented at MTB meetings from December 2012 to September 2018, following guidelines of the institutional review board-approved Profile Related Evidence Determining Individualized Cancer Therapy (PREDICT) study (NCT02478931; ClinicalTrials.gov; Posted June 23, 2015) and any investigational therapies for which patients gave consent. Weekly in-person MTBs were held at the Moores Cancer Center at UC San Diego (UCSD) Health and followed protocols as previously described14. We studied patients with solid tumors harboring one genomic or pathway alteration managed with matched targeted therapy8,15. We excluded patients who received immunotherapy based on MSI-high or TMB-high. However, patients treated with checkpoint blockade were included if the agent targeted discrete alterations such as PD-L1 amplification4. Patients who had only one alteration on an initial profiling test but subsequently received additional NGS profiling that revealed further mutations after MTB discussions were excluded from this analysis.
Endpoints and statistics
In accordance with RECIST 1.1 criteria, all patients were assessed with the outcome endpoints of clinical benefit rate [i.e., stable disease (SD) ≥ 6 months, partial response (PR), or complete response (CR)] as determined by the treating physician. Median PFS and median OS were also evaluated by the Kaplan–Meier method. PFS was defined as the time from the start of therapy to disease progression or last follow-up date if progression-free (the latter being censored). OS was defined as the time from the start of therapy to death or last follow-up if alive (the latter being censored).
Declaration of ethical approval
This retrospective case series involves patients enrolled in the UCSD Study of Profile Related Evidence Determining Individualized Cancer Therapy (PREDICT). This study was performed in accordance with UCSD IRB guidelines, and for any investigational treatments for which patients gave consent. All patients underwent informed consent and signed consented forms in their native languages via licensed medical interpreters.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
De-identified data will be made available on reasonable request. Should qualified researchers contact the corresponding author for the de-identified dataset, they will not need to obtain ethical approval or sign a data usage agreement as patient confidentiality will be maintained.
References
Padma, V. V. An overview of targeted cancer therapy. BioMedicine 5, 19 (2015).
Kato, S., Subbiah, V. & Kurzrock, R. Counterpoint: successes in the pursuit of precision medicine: biomarkers take credit. J. Natl Compr. Cancer Netw. 15, 863–866 (2017).
Goodman, A. M. et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Therapeutics 16, 2598–2608 (2017).
Goodman, A. M. et al. Prevalence of PDL1 amplification and preliminary response to immune checkpoint blockade in solid tumors. JAMA Oncol. 4, 1237 (2018).
Marcus, L., Lemery, S. J., Keegan, P. & Pazdur, R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin. Cancer Res. 25, 3753–3758 (2019).
Hutchinson, L. & Kirk, R. High drug attrition rates—where are we going wrong? Nat. Rev. Clin. Oncol. 8, 189–190 (2011).
Iyer, G. et al. Genome sequencing identifies a basis for everolimus sensitivity. Science 338, 221–221 (2012).
Sicklick, J. K. et al. Molecular profiling of cancer patients enables personalized combination therapy: the I-PREDICT study. Nat. Med. 25, 744–750 (2019).
Kato, S. et al. Prognostic implications of RAS alterations in diverse malignancies and impact of targeted therapies. Int. J. Cancer 146, 3450–3460 (2020).
Edelman, M. J. et al. SWOG S1400C (NCT02154490)—a phase II study of palbociclib for previously treated cell cycle gene alteration–positive patients with stage IV squamous cell lung cancer (Lung-MAP Substudy). J. Thorac. Oncol. 14, 1853–1859 (2019).
Flaherty, K. T. et al. Molecular landscape and actionable alterations in a genomically guided cancer clinical trial: national cancer institute molecular analysis for therapy choice (NCI-MATCH). J. Clin. Oncol. 38, 3883–3894 (2020).
Cheng, D. T. et al. Memorial sloan kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT). J. Mol. Diagnostics 17, 251–264 (2015).
Rodon, J. et al. Genomic and transcriptomic profiling expands precision cancer medicine: the WINTHER trial. Nat. Med. 25, 751–758 (2019).
Kato, S. et al. Real-world data from a molecular tumor board demonstrates improved outcomes with a precision N-of-One strategy. Nat. Commun. 11, 4965 (2020).
Wheler, J. J. et al. Cancer therapy directed by comprehensive genomic profiling: a single center study. Cancer Res. 76, 3690–3701 (2016).
Acknowledgements
Funded in part by the Joan and Irwin Jacobs Fund and by National Cancer Institute grants P30 CA023100 (RK, JKS). The authors also acknowledge the support of NIH K08CA168999 and R21CA192072, as well as Pedal the Cause, David Foundation, and Kristen Ann Carr Fund (JKS).
Author information
Authors and Affiliations
Contributions
Study conception and design: D.G., R.K., S.K. Acquisition of the data: S.L., R.O., H.J.L., K.H.K. Data analysis and interpretation: D.G., R.K., J.K.S., S.K. Writing—original draft preparation: D.G., R.K., S.K. Writing—review and edits: S.L., R.O., H.J.L., K.H.K., J.K.S. Created visualizations: D.G. Project administration: R.K., S.K. All authors approve this submitted version, agree to be accountable for their contributions, and ensure that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. D.G. and R.K. are listed as co-first authors. The method used in assigning the authorship order among co-first authors D.G. and R.K. was based on the individual who contributed most to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing non-financial interests but the following competing financial interests: D.G., S.L., R.O., H.J.L., and K.H.K. have nothing to disclose. R.K. has received research funding from Incyte, Genentech, Merck Serono, Pfizer, Sequenom, Foundation Medicine, Guardant Health, and Konica Minolta, as well as consultant fees from LOXO, X-Biotech, Actuate Therapeutics, Roche, and NeoMed. She receives speaker fees from Roche, owns stock in IDbyDNA, and has an ownership interest in CureMatch, Inc. J.K.S. receives research funding from Novartis Pharmaceuticals, Amgen Pharmaceuticals, and Foundation Medicine; consultant fees from Grand Rounds, Loxo, and Deciphera; and speaker’s fees from Roche and Deciphera. He also owns stocks in Personalis. S.K. serves as a consultant for Foundation Medicine and receives speaker’s fees from Roche.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gupta, D., Kurzrock, R., Lee, S. et al. Case series of outcomes in advanced cancer patients with single pathway alterations receiving N-of-One therapies. npj Precis. Onc. 6, 18 (2022). https://doi.org/10.1038/s41698-022-00259-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-022-00259-7
This article is cited by
-
Nonsense-mediated RNA decay: an emerging modulator of malignancy
Nature Reviews Cancer (2022)