Abstract
Zebrafish have proven to be a valuable model to study human cancer biology with the ultimate aim of developing new therapies. Danio rerio are amenable to in vivo imaging, high-throughput drug screening, mutagenesis, and transgenesis, and they share histological and genetic similarities with Homo sapiens. The significance of zebrafish in the field of precision oncology is rapidly emerging. Indeed, modeling cancer in zebrafish has already been used to identify tumor biomarkers, define therapeutic targets and provide an in vivo platform for drug discovery. New zebrafish studies are starting to pave the way to direct individualized clinical applications. Patient-derived cancer cell xenograft models have demonstrated the feasibility of using zebrafish as a real-time avatar of prognosis and drug response to identify the most ideal therapy for an individual patient. Genetic cancer modeling in zebrafish, now facilitated by rapidly evolving genome editing techniques, represents another innovative approach to recapitulate human oncogenesis and develop individualized treatments. Utilizing zebrafish to design customizable precision therapies will improve the clinical outcome of patients afflicted with cancer.
Similar content being viewed by others
Introduction
Precision medicine in oncology arises from recognition that patient-specific clinical, genetic, and molecular features dictate effectiveness of a given treatment. Therefore, precision oncology seeks to identify the most effective therapy for an individual patient, based on characterization of their cancer. The development of genomic technologies and molecular diagnostics enables detection of cancer biomarkers. These relevant abnormalities associated with specific cancers lead to the identification of actionable targets. Diagnostic (associated with the presence of a specific pathophysiological state), prognostic (associated with disease outcome), and predictive (associated with drug response) cancer biomarkers guide clinical treatment decisions and direct the use of drugs that modulate the activity of the specific actionable target.1,2
Zebrafish (Danio rerio) have rapidly emerged as a promising animal model of human cancer. Histological, molecular, and genetic similarities to Homo sapiens facilitate zebrafish studies of human malignancies. Zebrafish are amenable to in vivo fluorescent imaging, chemical and genetic screens, transgenesis, and high-throughput mutagenesis assays, which have brought zebrafish to the center stage of future advances in the field of precision oncology.3 A variety of attributes have contributed to the emergence of zebrafish as an attractive vertebrate model organism. Zebrafish are easy and inexpensive to maintain and breed with high fecundity, which facilitates large studies and high-throughput in vivo assays. Another advantage of working with zebrafish embryos is their conserved vertebrate features develop rapidly and genetic studies restricted to embryos can be completed in days to weeks rather than weeks to months as is often the case with mammalian models.3,4 However, it is important to note that zebrafish sexual maturation takes about three months, so generation studies (i.e., transgenics, knockouts, etc.) require a similar time frame as murine models. The small size, external development, and transparency of zebrafish embryos make them amenable to fluorescent live imaging to monitor physiological processes (e.g., development, morphogenesis, angiogenesis, etc.) and pathological phenomena (e.g., cancer initiation, tumorigenesis, metastasis, etc.). Taken together, the attractive features of the zebrafish model system underscore the reasons it has gained prominence in the study of cancer and serves as an excellent addition to other common oncology models and platforms.
This review aims to provide an overview of how current zebrafish cancer studies lay the groundwork for utilization of this model organism in precision oncology, highlighting specific studies oriented to the development of zebrafish-based patient-specific approaches for cancer treatment. The challenges and shortcomings of zebrafish cancer studies are presented as areas of the field requiring advancements and growth.
Zebrafish: From modeling classic cancer research to precision oncology
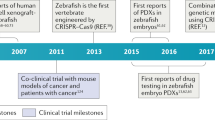
The application of the zebrafish model to precision oncology remains in its infancy, and there are not yet examples of direct use of zebrafish to guide patient-specific cancer treatments in the clinic. However, the field has matured enough to move toward this aim in the near future. Modeling cancer in zebrafish has provided important insights that contribute to the development of precision oncology as well as straightforward examples of advantages and feasibility of direct clinical utilization (Table 1).
Classic cancer modeling via mutagenesis, transgenesis, and xenotransplantation has contributed in numerous ways to precision oncology (Fig. 1, left). Zebrafish cancer models have facilitated (i) the identification and in vivo validation of molecular players in tumorigenesis and metastasis, (ii) the definition of actionable alterations and therapeutic targets, and (iii) the discovery of tumor biomarkers and genetic signatures as potential diagnostic and prognostic indicators. Moreover, several studies have exemplified the potential of zebrafish models to contribute more significantly and directly to precision oncology through (iv) identifying and testing drugs for targeted inhibition of specific pathways/alterations by utilizing zebrafish as an in vivo drug screening platform. A number of small molecules that might represent new targeted drugs for individualized medicine have been identified through this approach. Notably, the rapidly increasing number of patient-derived cancer cell xenografts5,6,7,8,9,10 places zebrafish on the road toward its clinical application for the treatment of individual cancer patients. Various studies have demonstrated the applicability of these models in (v) evaluating patient prognosis in vivo and (vi) directing individualized treatments in real-time based on responses to drugs of patient cancer cell xenografts (Fig. 1, right). Taken together, modeling cancer in zebrafish has evolved to the extent that precision oncology applications are emerging.
Applications of the zebrafish model in precision oncology. Classic cancer research using zebrafish has contributed to precision oncology through the establishment of numerous cancer models, leading not only to significant advancements in cancer biology, but also to the definition of targeted drugs suitable for personalized cancer treatments (blue, left). Possible applications of zebrafish in the clinic to drive personalized therapies for specific patients have also been shown. The feasibility of this approach has been demonstrated through the use of patient-derived zebrafish xenografts and generation of transgenic zebrafish modeling mutations or translocations defining a specific patient’s tumor (red, right)
Genetic models of cancer
To date, innumerable zebrafish genetic models of cancer have been generated, and the number continues to rapidly increase. Genetic cancer models have been developed using various strategies, including transient, stable, and double transgenesis and various inducers of mutagenesis (Table 2). Their use in precision oncology is gaining momentum. Here, we will discuss the most significant reports exemplifying this evolution.
Melanoma
Melanoma research offers many concrete examples of genetic zebrafish models used for the definition of new therapeutic targets and as an in vivo platform for drug screening. Melanoma accounts for the death of over 70% of skin cancer patients and only 14% of patients with metastatic disease survive for five years. Unlike many other tumor types, new cases and mortality of melanoma are still rising.11,12 While some oncogenic driver mutations, such as BRAF and NRAS, have been identified in melanoma, the efficacy of therapies is limited and the prognosis of metastatic melanoma patients remains poor.13 Many spontaneous, oncogene-driven zebrafish models of melanoma exist. In 2005, Patton et al.14 expressed BRAFV600E in melanocytes using the microphtalmia-associated transcription factor a (mitfa) promoter. These fish developed nevi, but required a p53M214K mutant zebrafish background for melanoma development in ~5% of zebrafish by four months.14 A crestin:EGFP reporter, recapitulating the embryonic neural crest expression patter of crestin, showed that a fate change occurs at melanoma initiation in this model, as a single melanocyte reactivates the neural crest progenitor state.15 Similarly, human oncogenic NRASQ61K expression under the control of mitfa promoter resulted in a transgenic fish that required p53 loss of function for the genesis of melanoma.16 The first p53 mutation-independent model was developed through expression of human oncogenic HRASG12V driven by the same mitfa promoter fragment. In this model, however, melanoma does not arise at a high frequency and takes several months to develop.17Instead, when HRASG12V expression is driven in melanocyte progenitor cells by the kita (c-kit in humans) promoter, melanoma occurs spontaneously by 1–3 months in ~20% of fish.18 Recently, a novel zebrafish transgenic model of uveal melanoma was created by expressing oncogenic GNAQQ209P in the melanocyte lineage using again the mitfa promoter. The corresponding p53 inactivation was also required for the malignant progression in this system.19 Importantly, zebrafish models of melanoma closely resemble human cancer, both in terms of histopathological features and molecular signatures.20
These models have confirmed the role of relevant oncogenes in melanoma genesis and progression. Moreover, they have proven to be outstanding tools to test and screen for other genes that promote melanoma onset and might represent new therapeutic targets, and even, in the near future, tumor biomarkers for personalized cancer therapy. An excellent example has been reported by Ceol and colleagues. They have used transgenic zebrafish overexpressing BRAFV600E on a p53 mutant background to test genes in a recurrently amplified region on chromosome 1. The histone methyltransferase SETDB1 has been found to cooperate with BRAFV600E and accelerate melanoma. Its relevance in human malignant melanoma has also been demonstrated, and therefore, SETDB1 has been revealed as a novel oncogene in melanoma.21 RAC and RSK1, whose hyperactivation has been detected in human melanoma, have been shown in distinct studies to contribute to melanoma progression when constitutively activated in mitfa:HRASG12V and mitfa:BRAFV600E;p53−/− transgenic backgrounds, respectively.22,23 Lister et al.24 have used a temperature-sensitive mitfa mutant to show the oncogenic activity of Mitfa transcription factor in BRAFV600E transgenic zebrafish and the regression of BRAFV600E mitfa melanoma after Mitfa activity abrogation, thus presenting Mitfa as a promising therapeutic target. The use of zebrafish to identify novel oncogenes begins to exemplify how this model organism will be utilized to overcome tumor heterogeneity through precision oncology.
The significance of zebrafish melanoma models in translational medicine and precision oncology is not limited to the discovery and characterization of potential therapeutic targets, as various studies have already shown the efficacy of zebrafish in identifying, discovering, and testing drugs for the development of new melanoma treatments. Small molecule inhibitors of MEK and PI3K/mTOR, known players in melanoma, have been validated in vivo as targeted drugs suppressing melanocyte hyperplasia phenotype in HRASG12V transgenic embryos.25 Moreover, a zebrafish screen of FDA-approved compounds led to the discovery of two new potential drugs cooperating with MEK inhibitors to suppress the growth of transformed melanocytes.25 Transgenic mitfa:BRAFV600E;p53−/− zebrafish embryos demonstrate a gene signature enriched for markers of multipotent neural crest cells. A chemical genetic screen was, therefore, performed to identify small molecule suppressors of the neural crest lineage. A positive result was obtained with the inhibitors of dihydroorotate dehydrogenase, whose activity as an anti-melanoma agent was then confirmed in vitro and through mouse xenograft models.26 To fully realize the utility of zebrafish in precision oncology, translating these types of drug identification and validation studies to a patient sample size of one is the ultimate goal, such that treatments can be tailored to the individual patient based on zebrafish surrogates of the individual’s tumor.
Neurological tumors
Neurological tumors have also been modeled via transgenic expression of oncogenes, demonstrating the potential to define relevant actionable alterations driving cancer progression and to successfully test specific drugs targeting those alterations.27,28 The focus of most zebrafish studies on brain tumors is malignant glioma, which accounts for 70% of malignant primary brain tumors, and in particular glioblastoma, the most aggressive primary brain cancer, accounting for 70% of malignant gliomas.29,30 Transgenic models of malignant peripheral nerve sheath tumors have also been described.27,31,32,33,34,35 Jung and colleagues established a transgenic zebrafish that overexpressed dominant active, human AKT1 at the ptf1a domain leading to gliomagenesis. Pharmacological tests identified AKT1/2 inhibitor as a targeted drug capable of effectively suppressing gliomagenesis, inhibiting cellular proliferation, and inducing apoptosis in established gliomas.28 The scope of available brain tumor models offers promise for using zebrafish to tailor specific treatment approaches to individual neurological cancer patients.
Pancreatic cancer
The Gal4/UAS transgenic system, based on the ability of the Gal4 transcriptional activator to drive the expression of multiple transgenes under the regulation of UAS (upstream activator sequence) regulatory elements, is widely used to model KRAS-initiated pancreatic cancer in zebrafish.36 Pancreatic cancer is a deadly genetic disease, with a dismal ~9% five year survival rate.37 The majority of pancreatic cancers are pancreatic ductal adenocarcinomas (PDACs) and over 90% of them carry an activating point mutation in the KRAS gene.36 Genetic models based on the Gal4/UAS system enable assessment of the effects of different KRAS mutations and the ability of other proteins to alter the response to oncogenic KRAS, potentially leading to the identification of new targets for precision oncology therapeutic strategies.38,39 In this regard, the involvement of a variety of core signaling pathways, including TGFβ, Wnt, Notch, and Hedgehog, in pancreatic cancer development has also been investigated using Gal4/UAS system.36,38,40
Leukemia
Leukemia, the ninth most common cancer type is a cancer of blood-forming tissues usually involving dysfunction of white blood cells.41 Leukemia has been modeled mainly through transgenesis. A zebrafish model of T-cell acute lymphoblastic leukemia (T-ALL), the most common type of childhood leukemia,42,43 was created in the early 2000s expressing a mouse c-Myc transgene fused to green fluorescent protein (GFP) under the control of a zebrafish rag2 promoter.44 Visualization of GFP+ leukemic cells has demonstrated leukemia originates in the thymus, disseminates to the gill arches and surrounding retro-orbital soft tissue, and then spreads to skeletal muscle and abdominal organs.44 Feng and colleagues have subsequently improved this model by developing conditional, heat-inducible activation of the c-Myc oncogene resulting in greater penetrance of T-ALL and increased control of disease onset.45 Similarly, another zebrafish model of T-ALL has been created by expressing the truncated human NOTCH1 protein fused to EGFP (ICN1-EGFP) under the control of the zebrafish rag2 promoter.46 While these transgenic zebrafish developed T-ALL by 5 months of age, onset of leukemia was dramatically accelerated when crossed to zebrafish overexpressing anti-apoptotic protein, Bcl2. The oncogenic synergy between NOTCH1 and Bcl2 in this model suggests genetic modifier screens may reveal other genes that interact with NOTCH1 to promote T-ALL.46 All of these rag2 promoter-driven transgenic zebrafish models are amenable to drug and genetic screening to identify individualized treatment strategies for leukemia and lymphoma patients. Indeed, a zebrafish screen for therapeutics that alter the number of hematopoietic stem cells (HSCs) has led to the development of Prohema, a derivative of prostaglandin E2 (PGE2), currently in Phase II clinical trials for use in leukemia and lymphoma patients receiving blood transplantations.47,48
Thyroid cancer
Stable transgenic expression of oncogenic BRAF (BRAFV600E) in thyroid epithelial cells has recently been shown to induce thyroid cancer in adult zebrafish. Combinatorial treatment with BRAF and MEK inhibitors rescue normal follicular architecture, restore thyroid hormone production, and reduce epithelial mesenchymal transition stimulated by BRAFV600E. The model has demonstrated in vivo the genetic requirement for Twist expression downstream of BRAFV600E, as ablation of twist3 by CRISPR-Cas9 suppressed BRAF-mediated oncogenesis.49 The in vivo validation of targeted molecular therapies for the treatment of thyroid cancer demonstrates the applicability of the zebrafish system to precision oncology approaches.
Liver cancer
Liver cancer is the second leading cause of cancer-related death. Hepatocellular carcinoma (HCC) accounts for 90–95% of liver cancer cases.50 Several zebrafish models of liver cancer have been developed utilizing different expression systems (reviewed well by Lu et al.50), including extensive contributions by Dr. Gong’s group.51,52,53,54,55 As a recent example, they developed a transgenic system for a liver-specific, mifepristone-inducible expression of oncogenic krasV12 via permanent genomic recombination mediated by the Cre-loxP system,55 which will facilitate the study of liver tumors that originate from a single cell or a small number of precursor cells through clonal expansion. In all, 20–40% of HCC are defined by an activating mutation in the gene encoding β-catenin. Evason and colleagues created a transgenic zebrafish expressing hepatocyte-specific activated β-catenin. They used the model to screen for druggable pathways that mediate β-catenin-induced liver growth and identified two c-Jun N-terminal kinase (JNK) inhibitors and two anti-depressants as potential targeted therapeutics.56 As is true in other tumor types discussed in this section, zebrafish have contributed to drug development to treat liver cancer.
Transplantation cancer models
With the first experiment reported in 2005,57 xenotransplantation of human cells into zebrafish represent a young frontier in zebrafish cancer modeling. However, the field has evolved rapidly, and xenograft zebrafish models utilizing various injection sites, developmental stages, and transplanted specimens (i.e., human cell lines, patient-derived primary cancer cells, patient-derived tumor tissue explants) have been developed58,59 (Table 3). Engraftment of a diverse range of human, murine, and zebrafish tumor cells has been demonstrated. Zebrafish transplantation models offer the possibility to study many hallmarks of cancer and steps of cancer progression, such as self-renewal, tumor-induced angiogenesis, invasion and dissemination, interaction between tumor and host, and drug responses.58,59 Cancer specimen transplantation into embryos is certainly the most commonly used zebrafish developmental stage for undeniable advantages, including the ease of producing and injecting many embryos in a short amount of time. Furthermore, the immature state of the immune system of embryos avoids the requirement of immune suppressing agents or irradiation.58,59 Fluorescently labeled tumor cells have been transplanted at developmental stages varying from the blastula stage to 72 h post fertilization (hpf) in injection sites such as blastodisc, yolk sac, bloodstream, perivitelline space, and orthotopic sites, including the hindbrain ventricle and vitreous cavity.9,58,59,60,61,62 The transplanted cells can be studied for up to 21 days post fertilization (dpf), at which point the zebrafish has developed a fully functional innate and adaptive system.59 While embryonic xenotransplantation offers numerous advantages, a limitation is that many of the tumor types being modeled occur predominantly in adults.
Xenotransplantation in juvenile and adult zebrafish seeks to overcome the limitation of translating embryonic zebrafish models to mature human cancer patients. Transplantation of human cancer cells in 30 dpf zebrafish has been established by Stoletov and colleagues in 2007 by injecting cells into the peritoneal cavity and treating the fish with dexamethasone to prevent rejection.63 The study of cancer cell transplantation in adult fish requires immune suppression by irradiation or dexamethasone pre-conditioning, and the use of transparent transgenic zebrafish allows the rapid identification of the transplanted cells.58,59 Casper fish, a cross between the nacre and roy mutant lines, lacking all types of pigments, are commonly used for this purpose.64 More recently, Tang et al.65 developed an optically clear immunocompromised transgenic mutant zebrafish line for optimized cell transplantation and direct visualization of fluorescently labeled cancer cells in the adult fish. Although juvenile and adult zebrafish transplantation models more closely match the developmental state and age of humans afflicted with cancer, the requirement for immune-deficient zebrafish represents the downside.
Melanoma
Both embryonic and adult zebrafish have been used as transplantation models to study drug efficacy in melanoma. van der Ent and colleagues injected different human uveal melanoma cell lines generated from primary tumors and metastases into the yolk of 48 hpf zebrafish embryos. They have shown that targeted inhibition of known pathways by specific drugs proves effective in counteracting cancer cells migration and proliferation, thus demonstrating the applicability of the zebrafish xenograft model for drug screening and discovery.66 Other xenograft zebrafish models have been used to explore relevant pathways in melanoma, such as Nodal in cellular plasticity and tumorigenicity,67 and TGF-β in cellular resistance to MEK inhibitors.68 Recently, a drug treatment system for the long-term administration of anti-melanoma drugs in adult casper zebrafish transplanted with a zebrafish melanoma cell line has been developed. The adult environment may be relevant to test and identify new promising treatments.69
Neurological tumors
A variety of zebrafish neurological tumor xenograft models have been established, primarily through xenotransplantation of different human malignant glioma cells (reviewed in Vittori et al.70). However, several studies have transplanted cells from mouse models of pediatric brain tumors and vestibular Schwannoma.71,72 A glioblastoma xenograft model has been proposed as a system for high-throughput screening of anti-angiogenic compounds. Transplanted cancer cells have been shown to be capable of inducing angiogenesis, which was enhanced by TGF-β1 and inhibited by targeted drugs, such as JNK, ERK, and PI3K inhibitors.73 Another zebrafish glioma xenograft model has been used to evaluate the in vivo efficacy of a novel small molecule radiation sensitizer identified through an in vitro drug screening in human glioma cells.74 The biological behavior of glioma cancer stem cells (GSCs) has also been explored in a zebrafish xenograft model. GSCs induced angiogenesis, which was inhibited by several anti-angiogenic agents. Moreover, the model revealed the in vivo activity of a synthetic compound, Nordy, previously found to promote GSCs differentiation in vitro. Nordy suppressed angiogenesis, tumor invasion, and proliferation of the zebrafish GSC xenograft.75,76 Taken together, zebrafish glioblastoma and glioma xenograft models have proven to be valuable systems for testing various molecularly targeted therapies.
Many brain tumor xenograft models have been obtained by orthotopically implanting cancer cells in embryonic or even larvae and juvenile brain.70 Direct transplantation into zebrafish brain facilitates cancer cell survival and proliferation, which leads to more relevant results. Glioblastoma cells have been injected in the hindbrain ventricle at 48–72 hpf to develop a xenograft assay to discover and prioritize compounds impacting glioblastoma progression. The utility of the assay was demonstrated by the ability of a drug with a known anti-cancer effect in cell culture to inhibit proliferation and invasion in the xenograft model.60 Another study described a platform to study the efficacy of drugs for the treatment of pediatric brain tumors. Mouse ependymoma, glioma, and choroid plexus carcinoma cells were transplanted orthotopically into the brain of zebrafish juveniles. As a proof of principle that these models can be used to assess drug efficacy, ERBB2-driven gliomas were successfully inhibited by treating zebrafish with a cytotoxic chemotherapeutic agent (5-fluorouracil) or a tyrosine kinase inhibitor.71 When feasible, orthotopic xenograft models in zebrafish brain tissue offer the advantage of more faithfully replicating human disease by utilizing the same anatomical tumor microenvironment.
Pancreatic cancer
Xenotransplantation of pancreatic cancer cells in zebrafish has also been proposed for the screening of new anti-cancer compounds. Guo and colleagues established a pancreatic adenocarcinoma xenograft model in zebrafish embryos and adults and found that a known small molecule inhibitor, U0126, targeting the KRAS signaling pathway, represses proliferation and migration of the transplanted cancer cells in zebrafish larvae.77 These results suggest this model could be used to identify new therapies for pancreatic cancer.
Retinoblastoma
Two studies have shown an orthotopic transplantation zebrafish model may represent a powerful tool for the development of specific drugs for the treatment of retinoblastoma, the most common intraocular childhood cancer, which often invades the brain and metastasizes.61,62 Injection of retinoblastoma cells into the vitreous cavity of the zebrafish embryo has permitted quantitative analysis of the tumor cells’ proliferative potential and the anti-cancer effect of systemically administered drugs. This model offers a potential screening platform for retinoblastoma anti-cancer drugs.62
Patient-derived transplantation models
Xenotransplantation of human cancer cells directly derived from individual patients (patient-derived xenograft, PDX) represents a fascinating and forthcoming opportunity for the development of zebrafish-based patient-specific clinical approaches for cancer treatment. Such patient-derived xenografts in zebrafish offer a platform for real-time in vivo evaluation of patient prognosis and drug responses, aimed at identifying the most appropriate individualized therapy (Fig. 1, right). Although only several examples of direct transplantation of patient-derived cancer cells in zebrafish have been reported thus far,6,8 the rapidly increasing number of zebrafish xenograft cancer models suggests that zebrafish xenografts are on the road to a clinical application in precision oncology.
Welker and colleagues standardized a patient-derived orthotopic zebrafish xenograft model of glioblastoma. They transplanted two patient-derived glioblastoma cell lines, serum-grown adherent and neurospheres, into the midbrain region of embryonic zebrafish. In vivo tumor growth and cancer cell proliferation, migration, and differentiation were described, with different characteristics in adherent and neurosphere glioblastoma cell lines. Furthermore, currently used glioblastoma therapeutics decreased xenotransplant tumor burden and significantly rescued survival. These results provide proof of principle for the use of the model as a platform for drug screening.9
A preclinical human cancer xenotransplantation platform has been recently developed in zebrafish to inform therapeutic decisions in T-ALL patients (Fig. 2a).5 The authors previously tested the in vitro sensitivity of three T-ALL cell lines, with specific mutations in PTEN and NOTCH1 genes, to three different inhibitors (targeting mTOR, AKT, and NOTCH1) and demonstrated that the same cell lines were sensitive to the same drugs upon xenotransplantation in the zebrafish embryos. The relevance of the zebrafish xenotransplantation model as a preclinical platform for a personalized therapy was demonstrated by xenotransplanting two primary patient-derived bone marrow samples into zebrafish embryos and treating with the three inhibitors. One patient sample responded drastically to NOTCH1 inhibitor (Fig. 2b, c), suggesting a mutation in the NOTCH pathway, which was subsequently confirmed to be a NOTCH1 mutation prevalent in T-ALL. The ability to assess a patient’s responsiveness to such a targeted treatment in a zebrafish avatar (i.e., likeness, surrogate or embodiment of an individual) within 1 week following biopsy, highlights how the zebrafish xenotransplantation response can direct personalized therapy in real-time.5
Precision oncology approach to leukemia drug screening using zebrafish. a Flow chart demonstrating the timeline used. Patient-derived leukemia cells were xenotransplanted into zebrafish embryos, which were administered various drugs. Leukemia cell number was used to assess drug efficacy in the zebrafish avatar corresponding to an individual leukemia patient. Assessment of drug efficacy is completed within 8 days, leading to a fast, effective, and individualized cancer treatment. hpf: hours post-fertilization, hpi: hours post-injection of cells, hpt: hours post-treatment. b Bright-field and fluorescence images of zebrafish injected with patient-derived leukemia cells. Embryos were treated with vehicle (control), Rapamycin (Rap) or Compound E (CE). Images were taken at 72 hpt. Scale bars are 500 µM. c A baseline number of leukemia cells was determined at 96 hpi. An increase in the number of leukemia cells when compared to the baseline data demonstrates cell proliferation in the zebrafish model. In patient sample one, data demonstrates a significant (p < 0.0001) response to the Notch inhibition (CE). The patient sample was subsequently sequenced and a gain of function mutation in the Notch pathway was found. Patient sample two did not demonstrate significant results, suggesting the mutation was not in the Notch pathway, which was subsequently confirmed through sequencing. Reproduced with permission and adapted from: Bentley, V.L. et al. Haematologica 100, 70–76 (2015)5
Transplantation metastasis models
Most cancer deaths are caused by metastasis, as opposed to primary tumors. Metastases result from the spread of cancer from the primary site to distant organs where new tumors form. Metastatic cancers have acquired the capacity to escape the primary malignant lesion site through intravasation into the bloodstream, migration, extravasation, and colonization of a distant site.78 Metastases are associated with poor prognosis due to the difficulty of treating such a complex and diffuse process. Moreover, unlike most other cancer processes, such as tumor initiation, proliferation, apoptosis, invasion etc., metastasis cannot be well modeled in vitro and the development and utilization of in vivo models of the dynamic sequence of steps from the local invasion to the distant colonization remains challenging. The transparency and ease of genetic manipulation of zebrafish embryos, coupled with the emerging opportunities offered by the xenograft models, represents an exceptional frontier to model and visualize the entire process of metastasis at single-cell resolution.
The xenotransplantion procedure is well optimized and automated quantitative assays are available to study invasion and metastasis of cancer cells.79,80 Zebrafish xenograft models of human cancer cell invasion, metastasis, and responsiveness to pharmacological or genetic intervention have been correlated to tumorigenicity of analogous human tumor cells in mouse xenograft models.79,81 Furthermore, Tulotta et al.82 demonstrated cross communication between zebrafish and human ligands and receptors, which enables the study of the interactions between human cancer cells and host microenvironment during the metastatic processes.
Outstanding examples of zebrafish metastasis models have recently emerged (Table 3). Stoletov et al.83 used real-time intravital imaging to study the dynamic process of intravascular locomotion and extravasation of fluorescent human cancer cells injected into the pericardium of 48 hpf zebrafish embryos, thus providing new insights into the underlying molecular regulation, which involves β1 integrin, Twist and VEGFA. Au et al.10 elegantly showed the migration dynamics of clusters of circulating tumor cells isolated from the blood of breast cancer and melanoma patients. The zebrafish xenograft metastasis model represents a valuable model to test the metastatic potential of human precision oncology target genes. We recently adapted Stoletov’s model to demonstrate neuropilin-2 promotes extravasation and metastasis of human pancreatic cancer and renal cell carcinoma cells in zebrafish (Fig. 3) by interacting with endothelial α5 integrin. We demonstrate synergy of the zebrafish extravasation model with mammalian metastasis models by also exhibiting the metastatic potential of neuropilin-2 in mice (Fig. 3).84 Thus, zebrafish metastasis models serve as an excellent in vivo platform for validating and supporting murine metastasis data and vice versa. Our data from a separate zebrafish study revealed knockdown of a metastasis suppressor gene, non-metastatic 2 (NME2), promotes extravasation of A549 human lung cancer cells.85 In conjunction with studies of tumor transcriptomes, survival data, and prevalence of lymph node metastases in human lung cancer patients, we demonstrated that NME2 decreases metastatic potential through transcriptional repression of focal adhesion factor vinculin.85 In a study highlighting the mechanism by which T-lymphoblastic lymphoma progresses to the metastatic T-ALL, Feng et al.86 focused on the intravasation process as one of the first steps in metastasis, identifying the role of Sphingosine-1-phosphate receptor 1 (S1P1) and intracellular adhesion molecule 1 (ICAM1) in counteracting cancer cells from entering the vasculature. In a xenograft model of PDAC metastasis, the calcium binding protein S100P was shown to facilitate cancer cell intravasation and extravasation.87 Thus, various stages of cancer metastasis have been accurately modeled in zebrafish transplantation settings.
Human cancer cell xenograft models of extravasation in zebrafish and metastasis in mice. a–h Human 786-O renal cell carcinoma cells overexpressing retroviral control vector (a–d) or neuropilin-2 (NRP-2; e–h) were transiently labeled with cell tracker orange dye, microinjected into the pericardium of 3 dpf Tg(Fli-GFP) zebrafish, and imaged 1 day later. a–d control 786-O cells stay in the ISVs. e, f 786-O cells overexpressing NRP-2 extravasate from the ISVs. i–j 2 × 106 luciferase-labeled 786-O cells suspended in PBS were subcutaneously injected into the right flank of female nude mice. Prior to the tumor growing to 10% of body weight, the subcutaneous tumors were surgically resected. Luciferase imaging was performed on the mice for 4–6 months to monitor metastasis, and the 786-O NRP-2 knockdown group (top) exhibited significantly fewer lung metastases than the control cohort (bottom). k–r Human ASPC-1 pancreatic cancer cells were transduced with control shRNA (k–n) or NRP-2 shRNA (o–r), transiently labeled, microinjected, and imaged as described above. k, l Extravasated control shRNA ASPC-1 cells. m, l Actively extravasating control shRNA ASPC-1 cells. o–r NRP-2 knockdown ASPC-1 cells stay in the ISV. s–t Male SCID mice were orthotopically injected with 2 × 106 GFP-labeled ASPC-1 pancreatic cancer cells suspended in PBS, and after 15 days liver metastases were assessed by xenogen imaging. Reproduced with permission and adapted from: Cao Y. et al. Cancer Res 73, 4579–4590 (2013)84
Zebrafish metastasis models have emerged as a promising system and continued development is likely necessary for applications in personalized medicine. Nevertheless, these models have already demonstrated their usefulness. They facilitate the investigation of new molecules shown to regulate various aspects of the metastasis process and aid in the discovery of targeted drugs that will potentiate our capacity to counteract metastasis in specific pathological conditions. For instance, Ghotra and colleagues have identified SYK as a candidate kinase target for the treatment of advanced prostate cancer. SYK was found to be upregulated in human prostate cancer and associated with malignant progression. They then used a zebrafish xenograft model to show that pharmacologic inhibitors of SYK kinase, currently in phase I–II trials for other indications, prevent metastatic dissemination of prostate cancer cells.88 A zebrafish xenograft model was also used to demonstrate the anti-metastatic activity of the small molecule VPC-18005, targeting the DNA-binding ETS domain of ERG, against prostate cancer cells aberrantly expressing ERG.89 Liekens et al.90 proposed the FDA-approved anti-DNA virus agent cidofovir as an anti-metastatic agent based on its ability to inhibit metastasis of FGF2-driven tumor cells in zebrafish embryos and mice. Amplification of ADP-ribosylation factor 1 (ARF1) is associated with poor outcomes of patients with breast cancer. A zebrafish xenograft model demonstrated that specific inhibition of Arf1 by small molecule LM11 impairs metastatic capability of breast cancer cells. LM11 may, therefore, represent a potential precision oncology treatment for patients with ARF1 amplification.91 Similarly, a zebrafish xenograft model was used to show the anti-metastatic effect of novel compounds designed to antagonize P2 × 7 receptor, previously reported as a key mediator in cancer metastasis. Significant inhibition of the invasion of MDA-MB-231 triple-negative breast cancer cells has been reported, thus identifying new potential drugs for individualized therapy.92 SIRT1 and CXCR4 were identified as novel pro-metastatic players, respectively, in Ewing sarcoma and triple-negative breast cancer, and the anti-metastasis effect of pharmacologically antagonizing these targets was demonstrated in zebrafish.82,93 Continued development of these zebrafish metastasis models, as well as establishment of new zebrafish-based systems, will accelerate their implementation in precision oncology applications.
Patient-derived transplantation models of metastasis
Zebrafish are amenable to modeling the metastatic potential of cancer cells derived from patient specimens and transplanted in the embryo. Initial progress was demonstrated by Marques et al.6 by transplanting direct explants from gastrointestinal human tumors into zebrafish embryos and larvae, into the yolk sac and organotopically in the liver, respectively. They showed the zebrafish model permits rapid analysis (micrometastasis formed within 24 h after transplantation) of primary human tumor specimen metastasis. Weiss and colleagues similarly transplanted pancreatic carcinoma cells and resected specimens of human pancreatic carcinoma into zebrafish embryos. The model was used to demonstrate the anti-metastatic in vivo activity of retinoid acid receptor antagonists, following the identification of the retinoid acid target miR-10A as a key mediator of metastasis in pancreatic cancer.8 The possibility of a near future application of the zebrafish model in precision oncology emerges from the study of Mercatali et al.7 They injected zebrafish embryos with a primary culture of bone metastasis derived from a 67 years old patient with breast cancer and compared its metastatic potential with that of established cancer cell lines. Importantly, primary cell behavior reflected the clinical course of the patient’s medical history, underscoring the noteworthy benefits that such an approach might signify for the evaluation of the patient prognosis and the identification of the most appropriate individualized therapy.
Genome editing
The advent of genome editing presents another promising strategy by which zebrafish will likely be utilized to tailor cancer therapies to individual patients. Whole-genome sequencing is commonly used to determine the molecular and genetic signature of a specific patient’s tumor. Upon identification of actionable mutations and alterations, transcription activator-like effector nucleases (TALENs) or clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated systems (Cas) genome editing tools could be used to mimic key oncogenic mutations/alternations in a zebrafish avatar of an individual patient’s disease. Ekker and colleagues have used TALENs to achieve precise locus-specific DNA breaks in somatic and germline tissue of zebrafish as well as in vivo targeted knock-ins through homology directed repair.94 Similarly, the CRISPR/Cas9 system has been shown to be a simple, quick, and scalable technique for in vivo editing of zebrafish genes.95 Thus, the genome editing toolbox offers strategies for recreating oncogenic mutations and perhaps even mimicking chromosomal rearrangements or fusions through knock-in technology. Genome editing in zebrafish can be achieved in a matter of months, which makes it possible to create a genome edited zebrafish that faithfully replicates key drivers of an individual cancer patient’s tumor, in sufficient time to use the zebrafish to identify the ideal individualized treatment strategy. A similar precision medicine approach using genome edited zebrafish has been proposed to treat cardiovascular disease.96 Taken together, genome editing in zebrafish offers an efficient strategy for modeling the key molecular characteristics driving tumor progression in a specific patient and screening drugs to identify the ideal treatment approach in real-time.
Challenges and shortcomings
The majority of zebrafish xenotransplantation studies require the injection of cancer cells into embryos. While this methodology has advantages as described (i.e., high-throughput injections, immunosuppression is not required, translucent embryos are amenable to in vivo fluorescent imaging, etc.), shortcomings of this approach must also be considered. The modeled tumors typically initiate during in adulthood naturally, and the embryonic environment presents undeniable differences that may impact cancer biology. For instance, embryonic development relies on the activation of specific programs and signaling pathways that are not active in adult organs under physiological conditions, but can become aberrantly activated during pathological stress, such as tumor initiation, progression, and metastasis. Various studies have utilized xenografts in adult zebrafish69,77,86 to overcome these concerns, but additional time and optimization is necessary for adult xenograft models to become as widely used and standardized as embryonic xenograft platforms.
Another limitation of modeling human cancer in zebrafish embryos is that such studies ignore the contribution of the immune system on cancer cell behavior, as embryos have not yet developed a functional immune system.58,59 This is also a drawback of human cancer cell xenotransplantation using adult zebrafish, which requires previous immunosuppression or the use of genetically immunocompromised zebrafish lines.65,69,77,86 A few attempts to develop immunocompetent xenograft systems have been reported. Cancers arising in syngeneic donors can be directly transplanted into a sibling recipient without the need of irradiation or dexamethasone pre-conditioning. However, only tumors made in clonal zebrafish lines can be transplanted into related recipients, and syngeneic fish are difficult to rear and not widely available.59,97 The impact of limitations that arise from using immunocompromised zebrafish models can be lessened by validating results in immunocompetent genetic cancer models established in adult zebrafish or other model organisms.
Long-term study of transplanted tumor cell behavior and drug response is particularly challenging. Embryo xenograft models are used in short-term assays, as long-term approaches compromise host viability. Xenotransplantation in adult fish after irradiation also prevents long-term studies because the immune system recovers within 20 days of irradiation. Dexamethasone-mediated chemical ablation of the immune system is a solution effective for solid tumor transplantation but not for leukemia. Nevertheless, mutant zebrafish lines harboring mutations that eliminate the immune response are emerging and represent a promising evolution toward the goal of long-term cancer biology and precision oncology studies in adult zebrafish.59,97
A recent report described a novel protocol for the long-term orthotopic transplantation of zebrafish brain tumor tissue into immunocompetent recipients. The method is based on the injection of zebrafish brain tumor cells into the fourth ventricle of a 48 hpf embryo. This allows tumors to grow in immunocompetent animals over the life of the zebrafish, enabling long-term monitoring of tumor cell behavior and drug response, including re-transplantation of tumors over many generations for potential studies on tumor evolution or drug relapse mechanisms.97
Lastly, a minor but intrinsic limitation of zebrafish xenografts of human tumor cells is the contrasting temperatures of the host (human cancer cells) and receipt (zebrafish). Zebrafish are typically reared at 28–29 °C, while human cells thrive at 37 °C. The compromise usually adopted is to raise zebrafish embryos at 33–35 °C following xenotransplantation of human cancer cells. Importantly, a possible impact of such temperature difference on the physiology of the fish and the biology of the tumor cells cannot be ruled out.
Conclusions and future directions
Given the value of zebrafish as a disease model, its power as a tool in precision oncology is becoming increasingly appreciated and applications are dawning. This advancement has been fueled by studies utilizing zebrafish cancer models to define molecular players of carcinogenesis, identify actionable alterations, and develop targeted therapies. Results to date provide a basis for individualized medicine research directions as well as platforms, workflows, and systems for applications in precision oncology (Table 1).
As the number of zebrafish cancer models being developed continues to grow with a greater focus on translation studies, we anticipate zebrafish models to increasingly contribute to personalized medicine. Indeed, studies have already presented zebrafish models of human cancer with specific genetic alterations and have demonstrated models’ abilities to test and identify targeted drugs that inhibit tumor growth and metastatic potential by acting on the specific molecules and pathways responsible for the tumorigenic phenotype. One can easily visualize this approach extending to wide potentialities for defining individualized treatments for oncology patients. Given a specific tumor, driven by different mutations/alterations in different patients, which, therefore, present different responses to cancer treatment, we might imagine a tumor-specific panel of zebrafish mutants/transgenics reproducing the array of the known oncogenic alterations found in patients harboring that tumor type. A future effort to consolidate and standardize the existing models, enriching the panel with models of other significant alterations found in patients, could lead to a complete in vivo platform used to identify the best available drug for each molecular signature. Concurrently, the panel could be used to screen drugs libraries for the identification of new active compounds. Ultimately, such a strategy will define a personalized treatment plan for each patient based on the molecular, genetic, and clinical characteristics of their cancer.
A more extensive use of patient-derived xenografts in zebrafish will also represent in the near future a powerful complementary approach to develop individualized treatments and to define the most appropriate therapeutic strategy for specific alterations found in patients. First insights into the possibility of a direct, real-time application of zebrafish xenograft models in the clinic5 suggest a precision oncology future in which primary specimens from patients diagnosed with cancer could be xenotransplanted in zebrafish embryos to test the responses of the patient cancer cells to various available drugs. The output of the test, obtainable in days, will dictate the most effective treatment for an individual cancer patient.
Taken together, the existing and developing array of zebrafish models within the collective toolbox of the zebrafish and precision oncology research community create a promising future where zebrafish may emerge alongside current clinical applications, such genomic technologies and molecular diagnostics, to improve our ability to precisely tailor individualized cancer therapies and positively impact the clinical outcome of each patient afflicted with cancer.
Data availability
Data sharing is not applicable to this article as no data sets were generated.
References
Schmidt, K. T., Chau, C. H., Price, D. K. & Figg, W. D. Precision oncology medicine: the clinical relevance of patient specific biomarkers used to optimize cancer treatment. J. Clin. Pharmacol. 56, 1–37 (2016).
Millner, L. M. & Strotman, L. N. The future of precision medicine in oncology. Clin. Lab. Med. 36, 557–573 (2016).
White, R., Rose, K. & Zon, L. Zebrafish cancer: the state of the art and the path forward. Nat. Rev. Cancer 13, 624–636 (2013).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Bentley, V. L. et al. Focused chemical genomics using zebrafish xenotransplantation as a pre-clinical therapeutic platform for T-cell acute lymphoblastic leukemia. Haematologica 100, 70–76 (2015).
Marques, I. J. et al. Metastatic behaviour of primary human tumours in a zebrafish xenotransplantation model. BMC Cancer 9, 128 (2009).
Mercatali, L. et al. Development of a patient-derived xenograft (PDX) of breast cancer bone metastasis in a Zebrafish model. Int. J. Mol. Sci. 17, 1375 (2016).
Weiss, F. U. et al. Retinoic acid receptor antagonists inhibit miR-10a expression and block metastatic behavior of pancreatic cancer. Gastroenterology 137, 2136–2145 (2009).
Welker, A. M. et al. Standardized orthotopic xenografts in zebrafish reveal glioma cell line specific characteristics and tumor cell heterogeneity. Dis. Model. Mech. 9, 199–210 (2015).
Au, S. H. et al. Clusters of circulating tumor cells traverse capillary-sized vessels. Proc. Natl Acad. Sci. 113, 4947–4952 (2016).
Miller, A. J. & Mihm, M. C. Melanoma. N. Engl. J. Med. 355, 51–65 (2006).
Siegel, R., Ma, J., Zou, Z. & Jemal, A. Cancer statistics, 2014. Ca. Cancer J. Clin. 64, 9–29 (2014).
Roh, M. R., Eliades, P., Gupta, S. & Tsao, H. Genetics of melanocytic nevi. Pigment Cell Melanoma Res. 28, 661–672 (2015).
Patton, E. E. et al. BRAF mutations are sufficient to promote nevi formation and cooperate with p53 in the genesis of melanoma. Curr. Biol. 15, 249–254 (2005).
Kaufman, C. K. et al. A zebrafish melanoma model reveals emergence of neural crest identity during melanoma initiation. Science 351, aad2197 (2016).
Dovey, M., White, R. M. & Zon, L. I. Oncogenic NRAS cooperates with p53 loss to generate melanoma in zebrafish. Zebrafish 6, 397–404 (2009).
Michailidou, C. et al. Dissecting the roles of Raf- and PI3K-signalling pathways in melanoma formation and progression in a zebrafish model. Dis. Model. Mech. 2, 399–411 (2009).
Santoriello, C. et al. Kita driven expression of oncogenic HRAS leads to early onset and highly penetrant melanoma in zebrafish. PLoS ONE 5, e15170 (2010).
Mouti, M. A., Dee, C., Coupland, S. E. & Hurlstone, A. F. L. Minimal contribution of ERK1 / 2-MAPK signalling towards the maintenance of oncogenic GNAQ Q209P -driven uveal melanomas in zebrafish. Oncotarget 7, 39654–39670 (2016).
Van Der Weyden, L. et al. Cross-species models of human melanoma. J. Pathol. 238, 152–165 (2016).
Ceol, C. J. et al. The SETDB1 histone methyltransferase is recurrently amplified in and accelerates melanoma. Nature 471, 513–517 (2011).
Dalton, L. E. et al. Constitutive RAC activation is not sufficient to initiate melanocyte neoplasia but accelerates malignant progression. J. Invest. Dermatol. 133, 1572–1581 (2013).
Salhi, A. et al. RSK1 activation promotes invasion in nodular melanoma. Am. J. Pathol. 185, 704–716 (2015).
Lister, J. A. et al. A conditional zebrafish MITF mutation reveals MITF levels are critical for melanoma promotion vs. regression in vivo. J. Invest. Dermatol. 134, 133–140 (2013).
Fernandez, L. et al. Reprofiling using a zebrafish melanoma model reveals drugs cooperating with targeted therapeutics. Oncotarget 7, 40348–40361 (2016).
White, R. M. et al. DHODH modulates transcriptional elongation in the neural crest and melanoma. Nature 471, 518–522 (2011).
Ju, B. et al. Oncogenic KRAS promotes malignant brain tumors in zebrafish. Mol. Cancer 14, 1–11 (2015).
Jung, I. H. et al. Glioma is formed by active Akt1 alone and promoted by active Rac1 in transgenic zebrafish. Neuro. Oncol. 15, 290–304 (2013).
Louis, D. N. Molecular pathology of malignant gliomas. Annu. Rev. Pathol. Mech. Dis. 1, 97–117 (2006).
Rousseau, A., Mokhtari, K. & Duyckaerts, C. The 2007 WHO classification of tumors of the central nervous system—what has changed? Curr. Opin. Neurol. 21, 720–727 (2008).
Amsterdam, A. et al. Many ribosomal protein genes are cancer genes in zebrafish. PLoS Biol. 2, E139 (2004).
Astone, M. et al. A GFP-tagged gross deletion on chromosome 1 causes malignant peripheral nerve sheath tumors and carcinomas in zebrafish. PLoS ONE 10, e0145178 (2015).
Berghmans, S. et al. Tp53 mutant zebrafish develop malignant peripheral nerve sheath tumors. Proc. Natl Acad. Sci. USA 102, 407–412 (2005).
Feitsma, H., Kuiper, R. V., Korving, J., Nijman, I. J. & Cuppen, E. Zebrafish with mutations in mismatch repair genes develop neurofibromas and other tumors. Cancer Res. 68, 5059–5066 (2008).
Lai, K. et al. Many ribosomal protein mutations are associated with growth impairment and tumor predisposition in zebrafish. Dev. Dyn. 238, 76–85 (2009).
Liu, S. & Leach, S. D. Screening pancreatic oncogenes in zebrafish using the Gal4/UAS system. Methods Cell Biol. 105, 367–381 (2011).
Howlader, N. et al. SEER Cancer Statistics Review, 1975–2014 (National Cancer Institute, Bethesda, MD) https://seer.cancer.gov/csr/1975_2014/ (2017).
Park, J. T. et al. Differential in vivo tumorigenicity of diverse KRAS mutations in vertebrate pancreas: A comprehensive survey. Oncogene 34, 2801–2806 (2015).
Provost, E. et al. The tumor suppressor rpl36 restrains KRAS(G12V)-induced pancreatic cancer. Zebrafish 11, 551–559 (2014).
Schiavone, M. et al. Zebrafish reporter lines reveal in vivo signaling pathway activities involved in pancreatic cancer. Dis. Model. Mech. 7, 883–894 (2014).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2017. Ca. Cancer J. Clin. 67, 7–30 (2017).
Ridges, S. et al. Zebrafish screen identifies novel compound with selective toxicity against leukemia. Blood 119, 5621–5631 (2012).
Santoriello, C. & Zon, L. I. Hooked! modeling human disease in zebrafish. J. Clin. Investig. 122, 2337–2343 (2012).
Langenau, D. M. et al. Myc-Induced T Cell Leukemia in Transgenic Zebrafish. Science 299, 887–890 (2003).
Feng, H. et al. Heat-shock induction of T-cell lymphoma/leukaemia in conditional Cre/lox-regulated transgenic zebrafish. Br. J. Haematol. 138, 169–175 (2007).
Chen, J. et al. NOTCH1-induced T-cell leukemia in transgenic zebrafish. Leukemia 21, 462–471 (2007).
North, T. E. et al. Prostaglandin E2 regulates vertebrate haematopoietic stem cell homeostasis. Nature 447, 1007–1011 (2007).
Cutler, C. et al. Prostaglandin-modulated Umbilical cord blood hematopoietic stem cell transplantation. Blood 122, 3074–3081 (2013).
Anelli, V. et al. Oncogenic BRAF disrupts thyroid morphogenesis and function via twist expression. Elife 6, 1–19 (2017).
Lu, J.-W. et al. Zebrafish as a disease model for studying human hepatocellular carcinoma. World J. Gastroenterol. 21, 12042–12058 (2015).
Lam, S. H. et al. Conservation of gene expression signatures between zebrafish and human liver tumors and tumor progression. Nat. Biotechnol. 24, 73–75 (2006).
Nguyen, A. T. et al. A high level of liver-specific expression of oncogenic KrasV12 drives robust liver tumorigenesis in transgenic zebrafish. Dis. Model. Mech. 4, 801–813 (2011).
Nguyen, A. T. et al. An inducible krasV12 transgenic zebrafish model for liver tumorigenesis and chemical drug screening. Dis. Model. Mech. 5, 63–72 (2012).
Yan, C., Yang, Q. & Gong, Z. Tumor-associated neutrophils and macrophages promote gender disparity in hepatocellular carcinoma in zebrafish. Cancer Res. 77, 1395–1407 (2017).
Nguyen, A., Koh, V., Spitsbergen, J. & Gong, Z. Development of a conditional liver tumor model by mifepristone-inducible Cre recombination to control oncogenic kras V12 expression in transgenic zebrafish. Sci. Rep. 6, 19559 (2016).
Evason, K. J. et al. Identification of chemical inhibitors of β-catenin-driven liver tumorigenesis in zebrafish. PLoS Genet. 11, e1005305 (2015).
Lee, L. M. J., Seftor, E. A., Bonde, G., Cornell, R. A. & Hendrix, M. J. C. The fate of human malignant melanoma cells transplanted into zebrafish embryos: Assessment of migration and cell division in the absence of tumor formation. Dev. Dyn. 233, 1560–1570 (2005).
Zhang, B., Xuan, C., Ji, Y., Zhang, W. & Wang, D. Zebrafish xenotransplantation as a tool for in vivo cancer study. Fam. Cancer 14, 487–493 (2015).
Moore, J. C. & Langenau, D. M. Allograft cancer cell transplantation in zebrafish. Adv. Exp. Med. Biol. 916, 265–287 (2016).
Wehmas, L. C., Tanguay, R. L., Punnoose, A. & Greenwood, J. A. Developing a Novel Embryo–Larval Zebrafish Xenograft Assay to Prioritize Human Glioblastoma Therapeutics. Zebrafish 13, 317–329 (2016).
Chen, X. et al. Invasiveness and metastasis of retinoblastoma in an orthotopic zebrafish tumor model. Sci. Rep. 5, 10351 (2015).
Jo, D. H. et al. Orthotopic transplantation of retinoblastoma cells into vitreous cavity of zebrafish for screening of anticancer drugs. Mol. Cancer 12, 71 (2013).
Stoletov, K., Montel, V., Lester, R. D., Gonias, S. L. & Klemke, R. High-resolution imaging of the dynamic tumor cell vascular interface in transparent zebrafish. Proc. Natl Acad. Sci. 104, 17406–17411 (2007).
White, R. M. et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2, 183–189 (2008).
Tang, Q. et al. Imaging tumour cell heterogeneity following cell transplantation into optically clear immune-deficient zebrafish. Nat. Commun. 7, 10358 (2016).
van der Ent, W. et al. Modeling of human uveal melanoma in zebrafish xenograft embryos. Invest. Ophthalmol. Vis. Sci. 55, 6612–6622 (2014).
Topczewska, J. M. et al. Embryonic and tumorigenic pathways converge via Nodal signaling: role in melanoma aggressiveness. Nat. Med. 12, 925–932 (2006).
Smith, M. P. et al. Effect of SMURF2 targeting on susceptibility to MEK inhibitors in melanoma. J. Natl. Cancer Inst. 105, 33–46 (2013).
Dang, M., Henderson, R. E., Garraway, L. A. & Zon, L. I. Long-term drug administration in the adult zebrafish using oral gavage for cancer preclinical studies. Dis. Model. Mech. 9, 811–820 (2016).
Vittori, M., Motaln, H. & Turnsek, T. L. The study of glioma by xenotransplantation in zebrafish early life stages. J. Histochem. Cytochem. 63, 749–761 (2015).
Eden, C. J. et al. Orthotopic models of pediatric brain tumors in zebrafish. Oncogene 34, 1736–1742 (2015).
Lee, H. -J. et al. Development of a vestibular schwannoma xenograft zebrafish model for in vivo antitumor drug screening. Laryngoscope 126, E409–E415 (2016).
Yang, X. J. et al. TGF-β1 enhances tumor-induced angiogenesis via JNK pathway and macrophage infiltration in an improved zebrafish embryo/xenograft glioma model. Int. Immunopharmacol. 15, 191–198 (2013).
Lally, B. E. et al. Identification and biological evaluation of a novel and potent small molecule radiation sensitizer via an unbiased screen of a chemical library. Cancer Res. 67, 8791–8799 (2007).
Yang, X. J. et al. A novel zebrafish xenotransplantation model for study of glioma stem cell invasion. PLoS ONE 8, 1–9 (2013).
Yang, X. et al. A synthetic dl-nordihydroguaiaretic acid (nordy), inhibits angiogenesis, invasion and proliferation of glioma stem cells within a zebrafish xenotransplantation model. PLoS ONE 9, e85759 (2014).
Guo, M. et al. U0126 inhibits pancreatic cancer progression via the KRAS signaling pathway in a zebrafish xenotransplantation model. Oncol. Rep. 34, 699–706 (2015).
Chambers, A. F., Groom, A. C. & MacDonald, I. C. Dissemination and growth of cancer cells in metastatic sites. Nat. Rev. Cancer 2, 563–572 (2002).
Ghotra, V. P. S. et al. Automated whole animal bio-imaging assay for human cancer dissemination. PLoS ONE 7, e31281 (2012).
Annila, T. et al. ZebIAT, an image analysis tool for registering zebrafish embryos and quantifying cancer metastasis. BMC Bioinforma. 14(Suppl 10), S5 (2013).
Drabsch, Y., He, S., Zhang, L., Snaar-jagalska, B. E. & Dijke, P. Transforming growth factor- β signalling controls human breast cancer metastasis in a zebrafish xenograft model. Breat Cancer Res. 15, R106 (2013).
Tulotta, C. et al. Inhibition of signaling between human CXCR4 and zebrafish ligands by the small molecule IT1t impairs the formation of triple-negative breast cancer early metastases in a zebrafish xenograft model. Dis. Model. Mech. 9, 141–153 (2016).
Stoletov, K. et al. Visualizing extravasation dynamics of metastatic tumor cells. J. Cell Sci. 123, 2332–2341 (2010).
Cao, Y. et al. Neuropilin-2 promotes extravasation and metastasis by interacting with endothelial α5 integrin. Cancer Res. 73, 4579–4590 (2013).
Thakur, R. K. et al. Non-metastatic 2 (NME2)-mediated suppression of lung cancer metastasis involves transcriptional regulation of key cell adhesion factor vinculin. Nucl. Acids Res. 42, 11589–11600 (2014).
Feng, H. et al. T-lymphoblastic lymphoma cells express high levels of BCL2, S1P1, and ICAM1, leading to a blockade of tumor cell intravasation. Cancer Cell 18, 353–366 (2010).
Barry, S. et al. S100P is a metastasis-associated gene that facilitates transendothelial migration of pancreatic cancer cells. Clin. Exp. Metastas. 30, 251–264 (2013).
Ghotra, V. P. S. et al. SYK is a candidate kinase target for the treatment of advanced prostate cancer. Cancer Res. 75, 230–240 (2015).
Butler, M. S. et al. Discovery and characterization of small molecules targeting the DNA-binding ETS domain of ERG in prostate cancer. Oncotarget 8, 42438–42454 (2017).
Liekens, S. et al. The broad-spectrum anti-DNA virus agent cidofovir inhibits lung metastasis of virus-independent, FGF2-driven tumors. Oncotarget 6, 4633–4648 (2014).
Xie, X., Tang, S., Cai, Y., Pi, W. & Deng, L. Suppression of breast cancer metastasis through the inactivation of ADP-ribosylation factor 1. Oncotarget 7, 58111–58120 (2016).
Park, J. -H. et al. Potent suppressive effects of 1-piperidinylimidazole based novel P2X7 receptor antagonists on cancer cell migration and invasion. J. Med. Chem. 59, 7410–7430 (2016).
Ban, J. et al. Suppression of deacetylase SIRT1 mediates tumor-suppressive NOTCH response and offers a novel treatment option in metastatic Ewing sarcoma. Cancer Res. 74, 6578–6588 (2014).
Bedell, V. M. et al. In vivo genome editing using a high-efficiency TALEN system. Nature 491, 114–118 (2012).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Campbell, J. M., Hartjes, K. A., Nelson, T. J., Xu, X. & Ekker, S. C. New and TALENted genome engineering toolbox. Circ. Res. 113, 571–587 (2013).
Casey, M. J. et al. Transplantation of zebrafish pediatric brain tumors into immune-competent hosts for long-term study of tumor cell behavior and drug response. J. Vis. Exp. 123, 1–10 (2017).
Phelps, R. A. et al. A two-step model for colon adenoma initiation and progression caused by APC loss. Cell 137, 623–634 (2009).
Acknowledgements
L.H. is supported by NIH/NCI R00-CA187035 and institutional Paint the Town Pink pilot project funding.
Author information
Authors and Affiliations
Contributions
M.A., E.D., S.A., and L.H. participated in literature review, idea generation, and constructive discussion. M.A., E.D., and L.H. wrote the manuscript and produced figures. M.A. and L.H. revised the manuscript. L.H. conceived the concept and supervised efforts. All authors approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Astone, M., Dankert, E.N., Alam, S.K. et al. Fishing for cures: The alLURE of using zebrafish to develop precision oncology therapies. npj Precision Onc 1, 39 (2017). https://doi.org/10.1038/s41698-017-0043-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-017-0043-9