Abstract
In contrast to acute diarrhoea, the aetiology of persistent digestive disorders (≥ 14 days) is poorly understood in low-resource settings and conventional diagnostic approaches lack accuracy. In this multi-country study, we compared multiplex real-time PCR for enteric bacterial, parasitic and viral pathogens in stool samples from symptomatic patients and matched asymptomatic controls in Côte d’Ivoire, Mali and Nepal. Among 1826 stool samples, the prevalence of most pathogens was highest in Mali, being up to threefold higher than in Côte d’Ivoire and up to tenfold higher than in Nepal. In all settings, the most prevalent bacteria were EAEC (13.0–39.9%) and Campylobacter spp. (3.9–35.3%). Giardia intestinalis was the predominant intestinal protozoon (2.9–20.5%), and adenovirus 40/41 was the most frequently observed viral pathogen (6.3–25.1%). Significantly different prevalences between symptomatic and asymptomatic individuals were observed for Campylobacter, EIEC and ETEC in the two African sites, and for norovirus in Nepal. Multiple species pathogen infection was common in Côte d’Ivoire and Mali, but rarely found in Nepal. We observed that molecular testing detected multiple enteric pathogens and showed low discriminatory accuracy to distinguish between symptomatic and asymptomatic individuals. Yet, multiplex PCR allowed for direct comparison between different countries and revealed considerable setting-specificity.
Similar content being viewed by others
Introduction
Digestive disorders rank among the leading causes of morbidity and mortality worldwide, with diarrhoea playing the most important role1. Although mortality has declined considerably over the past decades, diarrhoea still ranks as the fourth leading cause of death in children under the age of 5 years2. Globally, an estimated 1.65 million people die from diarrhoea annually, among them 446,000 children before they reach their fifth birthday2. The highest burden of digestive disorders is observed in low- and middle-income countries (LMICs), exacerbated by lack of clean water, poor sanitation and hygiene standards1,3. Compared to acute diarrhoea (defined as passage of three or more loose or liquid stools per day that lasts several hours to 13 days), less attention has been paid to persistent diarrhoea (defined as any diarrhoea lasting at least 14 days) and persistent abdominal pain (defined as self-reported, aetiologically unclear abdominal pain in children and adolescents aged 1–18 years). The few well-designed studies revealed a complex aetiology with considerable local idiosyncrasies4,5,6,7. Indeed, it is impossible to identify the causative agent solely based on the clinical symptomatology, and sensitive laboratory examinations are required to determine the causative agents of persistent digestive disorders6,8. However, the most widely used conventional methods are insensitive and only target a narrow spectrum of pathogens. Molecular diagnostic methods, such as multiplex real-time PCR, are gaining importance6. Multi-centred studies may provide new insights into key pathogens in different geographical settings and might help to select the most useful diagnostic tests9.
In order to investigate the aetiology of persistent digestive disorders in LMICs, the Neglected Infectious diseases DIAGnosis (NIDIAG) research network was set up and run for 5 years with funding from the European Commission10,11. Within NIDIAG, we designed a case–control study and examined stool samples in two African and one Asian site. One NIDIAG study was designed to prospectively investigate the aetiology of persistent digestive disorders. Here, we report on a post-hoc molecular analysis carried out on preserved stool samples stemming from both symptomatic patients and asymptomatic controls. The aims of this investigation were to comparatively describe the aetiological spectrum of persistent digestive disorders in the different settings, and to assess whether a threshold to identify pathogens with clinical significance can be identified.
Methods
Ethics statement
The protocol for this NIDIAG multi-country study was approved by the ethics committees of the University of Antwerp in Belgium, the ‘Ethikkommission beider Basel’ (EKBB) in Basel, Switzerland, the ‘Institut National de Recherche en Santé Publique’ in Bamako, Mali, the Ministry of Health in Abidjan in Côte d’Ivoire and the Nepal Health Research Council in Nepal. This study is registered on ClinicalTrials.gov (identifier: NCT02105714) and a detailed study protocol has been published elsewhere12.
Study area, design and population
The original NIDIAG study was designed as a prospective case–control study pertaining to persistent digestive disorders, which are defined as persistent diarrhoea (≥ 14 days) in individuals aged ≥ 1 year and/or persistent abdominal pain (≥ 14 days) in infants and adolescents aged 1–18 years. Persistent diarrhoea was defined according to World Health Organization (WHO) guidelines, i.e. three or more liquid bowel movements per day lasting ≥ 14 days. Persistent abdominal pain was defined as localised or diffuse abdominal pain ≥ 14 days with possible intermittence. Sample size calculations for the clinical study sites were based on available data on the frequency of persistent digestive disorders in published studies from resource-constrained areas12.
The study in Mali was carried out in the district reference healthcare centre based in Niono, a sub-regional administrative town, approximately 300 km northeast of Mali’s capital Bamako from August 2014 to May 2015. In Côte d’Ivoire, the study site was a regional reference hospital (Hôpital Méthodiste) in Dabou, a medium-sized city approximately 30 km west of the country’s economic capital Abidjan. Participants were enrolled between October 2014 and December 2015. In Nepal, the study was carried out at the B P Koirala Institute of Health Sciences in Dharan, a town approximately 350 km southeast of the country’s capital Kathmandu, and at the Dhankuta District Hospital in the town of Dhankuta, situated approximately 50 km north of Dharan. Sample collection in Nepal was carried out between July 2014 and August 2015.
Recruitment was conducted as previously described13. To each enrolled patient, one control without any gastrointestinal complaints in the 2 months preceding enrolment was matched by age group, sex and geographical location of residence. The reason to include matched asymptomatic controls in a 1:1 ratio to symptomatic patients was to investigate the aetiology of persistent digestive disorders and to provide discriminative data on the distribution of pathogens in order to enhance interpretation of causal associations.
Field and laboratory procedures
A series of conventional bacteriological and parasitological tests was carried out in the different settings, which have been described in detail elsewhere13. For the molecular diagnostic investigation reported here, participants provided stool samples in pre-labelled containers of which 500 mg of solid or 500 µl of fluid sample was transferred into 1 ml Eppendorf tubes. The samples were gently vortexed with 1–2 ml of 96% ethanol and stored in a fridge at 4 °C14. In all study countries, the samples were transferred within a few days from the study site to regional diagnostic centres, where they were stored in freezers at − 20 °C. At the end of the study (after a maximum storage period of up to 14 months), all samples were transferred on dry ice to the Institute of Medical Microbiology and Hygiene in Homburg, Germany, pending molecular diagnostic tests adhering to standard operating procedures. All research activities and microbiological diagnostic tests were performed in accordance with relevant guidelines and regulations.
Molecular post-hoc testing
At the Institute of Medical Microbiology and Hygiene in Homburg, 200 μl of each sample was taken of the 500 μl aliquots for automated nucleic acid extraction with the Promega Maxwell® 16 instrument (Promega Corporation; Fitchburg, United States of America). All samples were purified with the tissue LEV Blood DNA Purification Kit by the same manufacturer, and 1 µl of internal control RNA (for the viral stool panel) or DNA (for the bacterial and viral stool panel) was added. This purification kit was recommended by the manufacturer based on own testing protocols. However, at study inception, we comparatively assessed ten samples using the LEV Blood DNA Purification Kit and the LEV Blood RNA Purification Kit, but did not observe any differences with regard to the subsequent detection of RNA viruses. After extraction, multiplex real-time PCR was carried out on a Stratagene Mx3005P instrument, adhering to the supplier’s instructions (R-Biopharm AG; Darmstadt, Germany).
Regarding bacterial and parasitic infections, the current study employed the following, commercially available multiplex PCR kits from R-Biopharm (https://clinical.r-biopharm.com/): RIDA®GENE Bacterial Stool Panel (for detection of Salmonella spp., Campylobacter spp. and Yersinia enterocolitica), RIDA®GENE EAEC Stool Panel (for enteroaggregative Escherichia coli), RIDA®GENE ETEC/EIEC Stool Panel (for enteroinvasive E. coli (EIEC)/Shigella spp. (which are both characterised by the presence of ipaH) and enterotoxigenic E. coli (ETEC) with the subtypes LT and ST) and the RIDA®GENE Parasitic Stool Panel I (targeting Cryptosporidium spp., Dientamoeba fragilis, Entamoeba histolytica and Giardia intestinalis (synonymous: G. lamblia, G. duodenalis)). In brief, 5 μl of each extracted sample was added to a PCR mix containing 19.9 μl of reaction mix and 0.1 μl of Taq polymerase.
For detection of viral nucleic acids, the RIDA®GENE Viral Stool Panel I for adenovirus 40/41 (target gene: hexon), astrovirus (target gene: CAP), norovirus (target gene: ORF1/ORF2 junction region) and rotavirus (target gene: NSP3) was employed. Five µl of each sample extraction was added to 20 µl of master mix comprising 12.5 µl of reaction mix, 6.9 µl of primer–probe-mix and 0.7 µl of enzyme mix. Negative and positive controls for each stool panel were enclosed with 1 µl of internal control RNA (ICR) and 1 μl of internal control DNA (ICD), respectively, and were part of each PCR run. Samples were analysed using MxPro® QPCR Data Analysis Software.
Infection intensity
We determined infection intensities based on median cycle threshold (Ct) values obtained by real-time PCR analysis per pathogen, that were stratified by cases and controls and classified into four groups: (i) high intensity (Ct ≤ 24.9); (ii) medium (Ct 25.0–29.9); (iii) low (Ct 30.0–34.9); and (iv) very low (Ct ≥ 35.0) infection intensity.
Statistical analysis
Results of multiplex real-time PCR for each study country were tabulated in a Microsoft Excel file, version 14.0 (edition 2010, Microsoft Corporation). Prevalence and co-infections were calculated with Microsoft Excel, while distributional differences were assessed by Pearson’s χ2. A p-value of < 0.05 was considered statistically significant. All stool samples of sufficient quantity to perform multiplex PCR were considered for further analysis.
Results
Study cohort
A total of 1826 stool samples were collected, including 916 subjects with persistent diarrhoea and/or persistent abdominal pain and 910 matched controls. The study cohorts in Mali, Côte d’Ivoire and Nepal comprised 1100 (553 cases and 547 controls), 519 (260 cases and 259 controls) and 207 (103 cases and 104 controls) individuals, respectively. Details on the predominant symptomatology and the age distribution in the study sites are presented in Table 1.
Bacterial, parasitic and viral infections
EAEC was the predominant bacterium in all study sites, with an overall prevalence of 39.9% in Mali, 21.6% in Côte d’Ivoire and 13.0% in Nepal, followed by Campylobacter spp. (Table 2). Campylobacter spp., EIEC/Shigella spp. and ETEC were significantly more frequent in patients than in controls at the two African sites, while this was observed for EAEC only in Côte d’Ivoire. No statistically significant difference between cases and controls was observed for any bacteria in Nepal. Salmonella and Yersinia were only very rarely found in the entire study cohort (≤ 1%).
Among the intestinal protozoa, G. intestinalis and D. fragilis were frequently detected in Mali and Côte d’Ivoire, with G. intestinalis being significantly more prevalent in symptomatic patients. In all study sites, the prevalence of D. fragilis was higher among controls, though a statistical significance was only observed among participants in Mali. In Nepal, the overall prevalence of G. intestinalis and D. fragilis was below 7% and no infections with Cryptosporidium spp. or E. histolytica were observed.
Adenovirus was the most frequent viral infection, with particularly high prevalence in the two African sites. In Côte d’Ivoire, the prevalence of the different viruses was significantly elevated in cases as compared to controls, except for norovirus. In Nepal, only norovirus was significantly more prevalent in symptomatic patients. Rotavirus and astrovirus were rare in all three settings (≤ 5%).
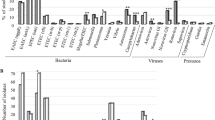
The prevalence of most pathogens was highest in Mali, being 1.5- to 3-fold higher than in Côte d’Ivoire and 3- to 10-fold higher than in Nepal. Considering any bacterial infection, the prevalence in Mali was 67.1% as compared to 45.5% in Côte d’Ivoire and 19.3% in Nepal, with similar country-specific patterns regarding the overall prevalence of parasitic and viral infection (Fig. 1).
Overall prevalence of patients with bacterial, parasitic and viral infections in a multi-centre case–control study on persistent digestive disorders, as defined as persistent diarrhoea (≥ 14 days; individuals aged 1 year and older) and persistent abdominal pain (≥ 14 days; children and adolescents aged 1–18 years) conducted in Mali, Côte d’Ivoire and Nepal.
Infection intensity
The majority of infections was of rather low infection intensity, as identified by Ct values. Except for EIEC/Shigella spp., Salmonella, norovirus, rotavirus and astrovirus, infection intensity of most pathogens was not significantly different between cases and controls in the different study sites. Details are presented in a heat map in Table 3.
Co-infections
Co-infections were rare in Nepal, but common in the two African sites, particularly pertaining to bacterial co-infections among cases (Fig. 2). No clear trend towards distinct differences between the number of overall co-infections and the presence or absence of symptoms was found.
Discussion
In this study, we found that the prevalence of most pathogens investigated, including co-infection rate, was highest in Mali, followed by Côte d’Ivoire. The study setting in Nepal showed the lowest prevalence for individual pathogens and concurrent infections among the three sites. In all settings, the most prevalent bacteria were EAEC and Campylobacter spp., while G. intestinalis was the predominant parasite and adenovirus the predominant viral pathogen. In the two African sites, Campylobacter, EIEC/Shigella spp. and ETEC were significantly more prevalent in symptomatic patients than asymptomatic controls. Co-infections with two or more pathogens were common in the two African sites, but absent in Nepal. Most infection intensities were low.
Even though remarkable progress has been made to reduce diarrhoeal diseases and associated morbidity, diarrhoea remains a leading cause of death in all age groups, with children under the age of 5 years most severely affected2,15, in particular due to resulting growth faltering, undernutrition and immunosuppression16. Additionally, the burden of persistent abdominal pain has not yet been thoroughly addressed in large clinical studies. Indeed, these long-term effects might account for more morbidity than the actual acute health effects, as previously shown for cryptosporidiosis17. Sanitation constitutes one of the major modifiable risk factors for diarrhoeal diseases18 and improved access to water, sanitation and hygiene (WASH) correlates with mortality reductions in most studies19. The link between poverty and infectious diseases has been specifically addressed for parasitic diseases20. At the country level, researchers observed a correlation between the human development index (HDI; a measure that is calculated based on the average healthy life span, level of education and standard of living) and the ‘worm index’ (i.e. the number of individuals requiring preventive chemotherapy with anthelminthic drugs against intestinal helminth infections, schistosomiasis and lymphatic filariasis divided by the total population in this country)21. A strong inverse correlation between HDI and worm index was observed, which might illustrate the long-term negative impact of these infections on the general well-being, productivity and economic development. According to the most recent HDI, Nepal, Côte d’Ivoire and Mali rank at positions 143, 159 and 186, respectively, among 191 countries listed22. Our study corroborates such findings. Indeed, we observed considerably higher prevalences of overall infections, individual pathogens and number of co-infections in countries with a lower HDI (e.g. the prevalence of EAEC was 39.9% in Mali, 21.6% in Côte d’Ivoire and 13.0% in Nepal). Additional research is required to support or refute these associations, and multi-country investigations using the same methodologies across sites are particularly relevant.
Even though the implementation of molecular diagnostics in resource-constrained settings remains challenging, PCR assays are key to identify and attribute diarrhoeal diseases to their microbiological causes, and have important ramifications for the development of clinical diagnosis-treatment algorithms23. From an epidemiological perspective, centralised post-hoc analyses of preserved stool samples with highly standardised molecular tests allow for comparable assessments across different sites, regardless of the respective diagnostic capacities in the field. In contrast, our findings underscore that PCR-based approaches alone without an individual clinical assessment cannot accurately differentiate symptomatic patients from asymptomatic controls.
Large research consortia have recently improved our understanding of the aetiology of acute diarrhoeal infections. In the Enteric Infections and Malnutrition and the Consequences for Child Health study (MAL-ED) in rural areas of South America, Africa and Asia, similar results pertaining to the prevalence of Campylobacter spp., EAEC and G. intestinalis in symptomatic and asymptomatic infants were found24. In the Global Enteric Multicenter Study (GEMS), a case–control study conducted in sub-Saharan Africa and South Asia, Campylobacter spp. was identified as a major cause of moderate-to-severe diarrhoea5,9. In contrast to our study, in which Cryptosporidium spp. was not highly prevalent, cryptosporidiosis was identified as one of the most significant infections in the MAL-ED, GEMS and VIDA studies24,25,26,27. However, a direct comparison between our study and the GEMS and MAL-ED studies is difficult, as the majority (77.9%) of all patients presenting with persistent digestive disorders in our investigation were children and adolescents presenting with persistent abdominal pain rather than diarrhoea as the main symptomatology. Indeed, the actual etiologies of persistent abdominal pain and diarrhoea are poorly recognised and likely to be multifactorial28. Yet, persistent diarrhoea has recently been acknowledged as a growing public health concern, particularly in sub-Saharan Africa, as evidenced by the VIDA study29. The high prevalence of D. fragilis, which was significantly more common among asymptomatic controls, further questions its potential pathogenic relevance30, although this protozoon was also recently associated with persistent abdominal pain in returning travelers31. Recent studies suggested that the severity of symptoms depends on the cumulative number of co-infections rather than on a single causative pathogen. Indeed, several studies did not find convincing evidence of a specific pathogen being associated with persistent diarrhoea in children aged below 6 years in LMICs32,33. It might be hypothesised that a primary infection facilitates a subsequent secondary infection with a possibly lower virulent pathogen, and hence, may increase the risk to develop diarrhoea24,32,34. The high rate of pathogen detection in asymptomatic children might be explained with residual nucleic acid shedding that may persist for months after previous infections24. Quantification of the pathogen load may help to distinguish clinically relevant infections, even though we were unable to define a Ct-based threshold. Longitudinal molecular analyses and parallel bacterial cultures might help to define such clinically useful cut-off values. However, it is important to note that even subclinical PCR findings of pathogens in the stool were recently shown to be associated with considerable long-term negative effects regarding childhood growth and stunting35.
In contrast to acute diarrhoea, few research initiatives have targeted persistent diarrhoea and persistent digestive disorders, which were in the focus of the NIDIAG consortium. Indeed, most data stem from single-centre studies in returning travelers28. In a recent case–control study from Ethiopia, 14% of children with diarrhoea complained about prolonged or persistent diarrhea, and hence, the authors urged for the rapid development of specific guidelines for these conditions36.
Our study has several limitations. First, the molecular analyses were carried out several months after stool collection, and, even though cold chain conditions were meticulously maintained, nucleic acid degradation may have occurred and might have led to lower positivity rates, at least for some of the pathogens investigated. Second, we did not perform bacterial and viral culture to correlate these findings to PCR results. Third, we relied on patients’ self-reports of persistent diarrhoea and persistent abdominal pain. Hence, we cannot ascertain whether the symptomatology had indeed been present for at least 2 weeks in all patients. Fourth, the design of our study does not allow inferring distinct causalities between a specific pathogen and the related clinical symptomology. Fifth, even though our PCR approach covered a wide range of pathogens, other intestinal pathogens (e.g. helminths) are missing. More complex approaches, such as metagenomics or more recently commercialised and more comprehensive multiplex PCR panels, hold promise to provide a more thorough characterisation of the intestinal microbial communities in patients and controls37. Sixth, no comprehensive information on previous clinical and preventive measures in participants such as e.g. previous intake of antimicrobials and rotavirus vaccination in early childhood was readily available.
Conclusions
We provide insights into the role of bacterial, protozoal and viral pathogens, some of which are frequently missed by conventional methods, based on detailed molecular examination of stool samples obtained from patients with persistent digestive disorders and asymptomatic controls in Côte d’Ivoire, Mali and Nepal. We observed high setting-specificity, but Campylobacter spp., G. intestinalis and norovirus were significantly more prevalent among symptomatic patients in most study areas. Our data indicate a correlation between the average pathogen load in the individual sites and the corresponding country HDI, which might be governed by poor WASH. Further studies are warranted to assess the long-term effects of intestinal infections beyond diarrhoeal disease manifestations.
Data availability
The datasets are available from the corresponding author on reasonable request.
References
GBD Collaborators. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: A systematic analysis for the Global Burden of Disease Study 2015. Lancet Infect. Dis. 17, 909–948. https://doi.org/10.1016/S1473-3099(17)30276-1 (2017).
Troeger, C. et al. Estimates of the global, regional, and national morbidity, mortality, and aetiologies of diarrhoea in 195 countries: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Infect. Dis. 18, 1211–1228. https://doi.org/10.1016/S1473-3099(18)30362-1 (2018).
Prüss-Ustün, A. et al. Burden of disease from inadequate water, sanitation and hygiene in low- and middle-income settings: A retrospective analysis of data from 145 countries. Trop. Med. Int. Health 19, 894–905. https://doi.org/10.1111/tmi.12329 (2014).
Pawlowski, S. W., Warren, C. A. & Guerrant, R. Diagnosis and treatment of acute or persistent diarrhea. Gastroenterology 136, 1874–1886. https://doi.org/10.1053/j.gastro.2009.02.072 (2009).
Kotloff, K. L. et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. Lancet 382, 209–222. https://doi.org/10.1016/S0140-6736(13)60844-2 (2013).
Becker, S. L. et al. Combined stool-based multiplex PCR and microscopy for enhanced pathogen detection in patients with persistent diarrhoea and asymptomatic controls from Côte d’Ivoire. Clin. Microbiol. Infect. 21, 591. https://doi.org/10.1016/j.cmi.2015.02.016 (2015).
Carvajal-Vélez, L. et al. Diarrhea management in children under five in sub-Saharan Africa: Does the source of care matter? A countdown analysis. BMC Public Health 16, 830. https://doi.org/10.1186/s12889-016-3475-1 (2016).
Li, L. L. et al. Aetiology of diarrhoeal disease and evaluation of viral-bacterial coinfection in children under 5 years old in China: A matched case-control study. Clin. Microbiol. Infect. 22, 381. https://doi.org/10.1016/j.cmi.2015.12.018 (2016).
Kotloff, K. L. et al. The Global Enteric Multicenter Study (GEMS) of diarrheal disease in infants and young children in developing countries: Epidemiologic and clinical methods of the case/control study. Clin. Infect. Dis. 55, S232–S245. https://doi.org/10.1093/cid/cis753 (2012).
Becker, S. L. et al. Persistent digestive disorders in the tropics: Causative infectious pathogens and reference diagnostic tests. BMC Infect. Dis. 13, 37. https://doi.org/10.1186/1471-2334-13-37 (2013).
Boelaert, M., Consortium Network. Clinical research on neglected tropical diseases: Challenges and solutions. PLoS Negl. Trop. Dis. 10, e0004853. https://doi.org/10.1371/journal.pntd.0004853 (2016).
Polman, K. et al. Diagnosis of neglected tropical diseases among patients with persistent digestive disorders (diarrhoea and/or abdominal pain ≥14 days): A multi-country, prospective, non-experimental case–control study. BMC Infect. Dis. 15, 338. https://doi.org/10.1186/s12879-015-1074-x (2015).
Becker, S. L. et al. Experiences and lessons from a multicountry NIDIAG study on persistent digestive disorders in the tropics. PLoS Negl. Trop. Dis. 10, e0004818. https://doi.org/10.1371/journal.pntd.0004818 (2016).
Marotz, C. et al. Evaluation of the effect of storage methods on fecal, saliva, and skin microbiome composition. mSystems 6, 20. https://doi.org/10.1128/mSystems.01329-20 (2021).
Troeger, C. et al. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: A systematic analysis for the Global Burden of Disease Study 2015. Lancet Infect. Dis. 17, 909–948. https://doi.org/10.1016/S1473-3099(17)30276-1 (2017).
Troeger, C. et al. Global disability-adjusted life-year estimates of long-term health burden and undernutrition attributable to diarrhoeal diseases in children younger than 5 years. Lancet Glob. Health 6, e255–e269. https://doi.org/10.1016/S2214-109X(18)30045-7 (2018).
Khalil, I. A. et al. Morbidity, mortality, and long-term consequences associated with diarrhoea from Cryptosporidium infection in children younger than 5 years: A meta-analyses study. Lancet Glob. Health 6, e758–e768. https://doi.org/10.1016/S2214-109X(18)30283-3 (2018).
Troeger, C. E. et al. Quantifying risks and interventions that have affected the burden of diarrhoea among children younger than 5 years: An analysis of the Global Burden of Disease Study 2017. Lancet Infect. Dis. 20, 37–59. https://doi.org/10.1016/S1473-3099(19)30401-3 (2020).
Reiner, R. C. Jr. et al. Mapping geographical inequalities in childhood diarrhoeal morbidity and mortality in low-income and middle-income countries, 2000–17: Analysis for the Global Burden of Disease Study 2017. Lancet 395, 1779–1801. https://doi.org/10.1016/S0140-6736(20)30114-8 (2020).
King, C. H. Parasites and poverty: The case of schistosomiasis. Acta Trop. 113, 95–104. https://doi.org/10.1016/j.actatropica.2009.11.012 (2010).
Kang, S., Damania, A., Majid, M. F. & Hotez, P. J. Extending the global worm index and its links to human development and child education. PLoS Negl. Trop. Dis. 12, e0006322. https://doi.org/10.1371/journal.pntd.0006322 (2018).
United Nations. Human Development Report 2019. http://hdr.undp.org/en/content/table-1-human-development-index-and-its-components-1 (2019).
Platts-Mills, J. A. et al. Use of quantitative molecular diagnostic methods to assess the aetiology, burden, and clinical characteristics of diarrhoea in children in low-resource settings: A reanalysis of the MAL-ED cohort study. Lancet Glob. Health 6, e1309–e1318. https://doi.org/10.1016/S2214-109X(18)30349-8 (2018).
Platts-Mills, J. A. et al. Pathogen-specific burdens of community diarrhoea in developing countries: A multisite birth cohort study (MAL-ED). Lancet Glob. Health 3, e564–e575. https://doi.org/10.1016/S2214-109X(15)00151-5 (2015).
Fotedar, R. et al. Laboratory diagnostic techniques for Entamoeba species. Clin. Microbiol. Rev. 20, 511–532. https://doi.org/10.1128/CMR.00004-07 (2007).
Pop, M. et al. Diarrhea in young children from low-income countries leads to large-scale alterations in intestinal microbiota composition. Genome Biol. 15, R76. https://doi.org/10.1186/gb-2014-15-6-r76 (2014).
Hossain, M. J. et al. Clinical and epidemiologic features of Cryptosporidium-associated diarrheal disease among young children living in sub-Saharan Africa: The vaccine impact on diarrhea in Africa (VIDA) study. Clin. Infect. Dis. 76, S97–S105. https://doi.org/10.1093/cid/ciad044 (2023).
DuPont, H. L. Persistent diarrhea: A clinical review. JAMA 315, 2712–2723. https://doi.org/10.1001/jama.2016.7833 (2016).
Buchwald, A. G. et al. Etiology, presentation, and risk factors for diarrheal syndromes in 3 sub-Saharan African countries after the introduction of rotavirus vaccines from the vaccine impact on diarrhea in Africa (VIDA) study. Clin. Infect. Dis. 76, S12–S22. https://doi.org/10.1093/cid/ciad022 (2023).
Incani, R. N. et al. Diagnosis of intestinal parasites in a rural community of Venezuela: Advantages and disadvantages of using microscopy or RT-PCR. Acta Trop. 167, 64–70. https://doi.org/10.1016/j.actatropica.2016.12.014 (2017).
Gefen-Halevi, S. et al. Persistent abdominal symptoms in returning travellers: Clinical and molecular findings. J. Travel Med. 29, 11. https://doi.org/10.1093/jtm/taac011 (2022).
Bardhan, P. K. et al. Small bowel and fecal microbiology in children suffering from persistent diarrhea in Bangladesh. J. Pediatr. Gastroenterol. Nutr. 26, 9–15 (1998).
Abba, K., Sinfield, R., Hart, C. A. & Garner, P. Pathogens associated with persistent diarrhoea in children in low and middle income countries: Systematic review. BMC Infect. Dis. 9, 88. https://doi.org/10.1186/1471-2334-9-88 (2009).
Sarker, S. A., Ahmed, T. & Brüssow, H. Persistent diarrhea: A persistent infection with enteropathogens or a gut commensal dysbiosis? Environ. Microbiol. 19, 3789–3801. https://doi.org/10.1111/1462-2920.13873 (2017).
Rogawski, E. T. et al. Use of quantitative molecular diagnostic methods to investigate the effect of enteropathogen infections on linear growth in children in low-resource settings: Longitudinal analysis of results from the MAL-ED cohort study. Lancet Glob. Health 6, e1319–e1328. https://doi.org/10.1016/S2214-109X(18)30351-6 (2018).
Zangenberg, M. et al. Prolonged and persistent diarrhoea is not restricted to children with acute malnutrition: An observational study in Ethiopia. Trop. Med. Int. Health 24, 1088–1097. https://doi.org/10.1111/tmi.13291 (2019).
Schneeberger, P. H. H. et al. Qualitative microbiome profiling along a wastewater system in Kampala, Uganda. Sci. Rep. 9, 17334. https://doi.org/10.1038/s41598-019-53569-5 (2019).
Acknowledgements
The authors are grateful to all the participants from Niono (Mali), Dabou (Côte d’Ivoire), Dharan and Dhankuta (Nepal) for their active participation in the current study. Thanks are addressed to the laboratory technicians at the District Reference Healthcare Centre in Niono, the ‘Institut National de Recherche en Santé Publique’ (INRSP) in Bamako, the ‘Hôpital Méthodiste’ in Dabou, the B P Koirala Institute of Health Sciences in Dharan and the Dhankuta District Hospital.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work is part of the collaborative NIDIAG European research network, which is supported by the European Union’s Seventh Framework Program for research, technological development and demonstration under Grant Agreement No. 260260.
Author information
Authors and Affiliations
Contributions
MB, FC, KP, JU and SLB conceived the study. HKMF, MS, RS, JKC, EKN, JAY, BK, KK, NRB, SR and SLB enrolled the participants and conducted the field work, including on-site diagnostics. JKJ, FB and JV conducted the multiplex PCR. JKJ, RS, JAY and NRB performed the statistical analysis. LvM, EB and FC assisted in clinical tasks related to the study. JKJ drafted the manuscript. All authors read, commented on and approved the final manuscript. Informed consent to participate in this study was obtained from all subjects and/or their legal guardians.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests. The reagents for the Stratagene Mx3005P multiplex real-time PCR and the specific fluorescent-based detection system (MxPro QPCR Software) were provided free of charge by the company R-Biopharm (Darmstadt, Germany). The funders and R-Biopharm had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jasuja, J.K., Bub, F., Veit, J. et al. Multiplex PCR for bacterial, viral and protozoal pathogens in persistent diarrhoea or persistent abdominal pain in Côte d’Ivoire, Mali and Nepal. Sci Rep 14, 10926 (2024). https://doi.org/10.1038/s41598-024-60491-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60491-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.