Abstract
Astrocytes are glycolytically active cells in the central nervous system playing a crucial role in various brain processes from homeostasis to neurotransmission. Astrocytes possess a complex branched morphology, frequently examined by fluorescent microscopy. However, staining and fixation may impact the properties of astrocytes, thereby affecting the accuracy of the experimental data of astrocytes dynamics and morphology. On the other hand, phase contrast microscopy can be used to study astrocytes morphology without affecting them, but the post-processing of the resulting low-contrast images is challenging. The main result of this work is a novel approach for recognition and morphological analysis of unstained astrocytes based on machine-learning recognition of microscopic images. We conducted a series of experiments involving the cultivation of isolated astrocytes from the rat brain cortex followed by microscopy. Using the proposed approach, we tracked the temporal evolution of the average total length of branches, branching, and area per astrocyte in our experiments. We believe that the proposed approach and the obtained experimental data will be of interest and benefit to the scientific communities in cell biology, biophysics, and machine learning.
Similar content being viewed by others
Introduction
Astrocytes that are star-like glial cells widely distributed in the central nervous system (CNS), play a crucial role in various CNS functions, including brain development and plasticity, synaptic transmission, blood flow regulation, metabolic control within the neurovascular unit1. Particularly, glycolytically active astrocytes ensure neuron-astroglial metabolic coupling meeting the metabolic needs of stimulated neuronal cells, control local microcirculation and permeability of the blood–brain barrier2. The role of astrocytes in neuroinflammation and regulation of neurogenesis in also well-known. Thus, it is not surprising that altered astroglial structure and functional activity is always seen in brain injury, stroke, neurodevelopmental disorders and neurodegeneration1,3,4,5,6,7. The morphology of astrocytes reflects their metabolic and developmental status8,9. The activation of astrocytes is usually associated with the rise in the cytosolic free calcium which triggers the activity or numerous signaling cascades. Some of them result in fast and transient or postponed and long-lasting changes in astroglial morphology required for astroglial adaptation to the microenvironment, migration, and interaction with other cells. Therefore, astroglial morphology is different in various brain regions (e.g. protoplasmic vs fibrous astrocytes), in loci of inflammation (resting vs polarized astrocytes), within the so-called astroglial synticium (when connexon-coupled astrocytes may simultaneously respond to extracelllular stimuli), and in a close vicinity to synaptically coupled neurons or microvessels (more branched astrocytes migth produce more contacts with activated neuronal cells or endothelial cells). Thus, detecting, counting, and tracking astrocyte growth dynamics and their morphology are critical for better understanding astroglial contribution to the regulation of brain functions.
The morphology features of astrocytes can be studied experimentally, for example, using microfluidic chips10,11 various in situ12 and in vivo13 methods, as well as using computer modeling (in silico)14,15,16,17,18,19,20,21,22. Despite the fact that time-reducing experimental methods based on Generative adversarial network were developed23, cellular dynamics simulation methods significantly reduce time and costs and could lead to improved experiment quality, enhancing our understanding of processes occurring at the cellular and subcellular levels24. Moreover, it enables the study of phenomena that are inaccessible experimentally. Similar situation occurred, for example, in the field of condensed matter physics when computational modeling methods such as classical or quantum molecular dynamics significantly improved our understanding of processes occurring at the atomic level. Computational modeling methods have already been successfully applied for studying blood-brain barrier permeability21, creating detailed mesh models for visual analysis22, reconstructing and exploring various types of astroglia15, astrocytic signal transduction through cytosolic \(Ca^{2+}\) dynamics16, diffusion-weighted signaling17, the impact of different substances (in this case, glutamate) on astrocytic signal transduction20, interaction between neuronal and astrocytic networks18. Besides, using computer cell modeling25,26, it is possible to study tumor growth27, study the formation of blood vessels during angiogenesis and vasculogenesis28,29,30,31,32,33,34,35. However, any computer model must be validated based on experimental data, and in the case of living systems, this data should arise from various multiomics approaches. The morphology data is one of many types used to verify computer models17,18. Considering the significant variability within living systems (such as cellular noise), this data should be averaged across a substantial representative sample. This imposes significant requirements for the scalability of experiments that verify computer models.
For the experimental study of astroglial morphology various types of microscopy are utilized, including phase contrast36, electronic37, fluorescent38, etc. These methods were used to study the interaction of astrocytes within the astroglial network37,39, the role of astrocytes in cerebral metabolic interactions40, mitochondrial metabolism41, differences in ensembles of astrocytes in the cortex42 and other brain regions43. Microscopy is always used to observe brain cells exposed to the action of neurotransmitters, gliotransmitters, cytokines44 or cytotoxic substances45 either in vitro or in vivo, as well as in affected brain regions in vivo, e.g. in brain injury or neurodegeneration46,47,48,49. The role of astrocytes in the formation and maintenance of synapses was also studied using microscopy50,51,52,53. The most common method of visualizing cells is fluorescence microscopy, that usually requires fixation and staining of cells. This method allows for more accurate visualization of cells and organelles. However, these techniques mainly deal with non-viable cells due to procedure of fixation (excluding the application of protocols with fluorescent probes added to alive cells in culture or in the tissue). As a result, the accuracy of estimation of astroglial structure might be greatly compromised. In the conditions in vitro, the morphology of astrocytes might be additionally affected by their adhesion to the the surface with high stiffness (glass or plastics) and establishment of non-physiological monolayers in 2D cultures, or by their inappropriate positioning within 3D polymers. Therefore, if it is necessary to study the parameters and properties of cells in a long-term experiment and at different points in their life cycle, this method will not be suitable. Another solution is the use of phase contrast microscopy, which does not require cell fixation and staining, but it has a drawback of low-contrast images (compared to fluorescence microscopy), thereby making post-processing challenging. Note that despite the lower resolving power of phase contrast microscopy compared to staining methods, the corresponding data still hold significant value for various research purposes.
At present, various approaches for cell image analysis exist, with one of the most commonly used being machine learning. Particularly, the open-source tool ilastik54 is actively used for interactive image classification, segmentation, and analysis. However, although modern machine-learning approaches could handle heterogeneity by combining multiple ML models trained on each single cell sub-track55 and using peer prediction paradigm56, traditional machine learning methods are not suitable for astrocytes detection due to their complex morphology57. It was also demonstrated how to derive deep learning-based features to characterize cell morphodynamics by combining time-lapse microscopy with resilient deep learning software designs58. Additionally, methods were outlined showing that these descriptors remain largely unaffected by common sources of artifacts in bioimaging analysis59.
Using deep learning methods, especially considering advances of deep convolutional neural networks (DCNN) in pattern recognition and image classification problems60,61, would be more appropriate. In last few years, a lot of different DCNN models were created62,63,64,65,66,67, which has been used in different fields, for instance biomedical images analysis66,68,69,70,71, physisc and chemistry tasks, etc.72,73,74,75. Thus, DCNN can successfully process complex morphology because of multiple layers of feature extraction that makes it suitable for detection and segmentation76,77.
In the present paper, we propose a new deep learning-based approach for recognition and morphological analysis of isolated astrocytes. For this purpose, we conducted a series of experiments involving the culture of primary cultures of rat brain cortex astrocytes, followed by the phase contrast microscopy analysis. We labeled the astrocytes in the acquired images and used this data to train the DCNN-based model. Using the proposed approach, we tracked the time evolution of the average total length of branches, branching, and area covered by an astrocyte. The novelty of the proposed approach lies in its ability to recognize morphological data from unstained cell microscopic images. We believe that the proposed approach and the obtained experimental data will be of interest and benefit to the scientific communities in cell biology, biophysics, and machine learning.
Results
Astrocyte detection methods with an accuracy (precision P = 0.86) comparable to that of human experts already exist57,78. However, these methods were created for the detection of immunohistochemically stained astrocytes in high-contrast images, and are not suitable for our experiments. For this reason, the Mask R-CNN neural network79, designed for segmentation80, was used to solve the problem of segmentation of living astrocytes. We used a Mask R-CNN model pre-trained on the COCO dataset with Inception Resnet v281 on 1024x1024 pixel images. We chose Mask-RCNN over the popular Segment Anything Model82 for the following reasons. Firstly, the pre-trained model Mask R-CNN show high performance when trained in segmentation living cells83. Secondly, the authors84 showed that they were able to outperform the segmentation accuracy of the latest SAM model using improvements to the existing MASK-RCNN model. Thirdly, it was concluded85 that SAM’s zero-shot segmentation performance is considerably inferior to that of traditional deep learning-based methods. Thus, the selection of the most accurate segmentation model with the fitting of hyperparameters deserves a separate study.
The Mask-RCNN model can process images up to a resolution of 1024 by 1024 pixels. However, to reduce computational efforts, we used resized images for model training. The final size of the input images is shown in Table 1.
The neural network was trained on images labeled using LabelMe86; examples are shown in Fig. 1. For training, microscopic images were pre-processed. They were divided into fragments of 1024x1024 pixels (160x160 \(\mu\)m), with each fragment containing up to 6 astrocytes. The image size of 1024 \(\times\) 1024 pixels for raw images determined by equipment (video camera). Fragments in which astrocytes were not detected were discarded. As a result, we obtained 221 images of 1024 \(\times\) 1024 pixels, containing in total 332 labeled astrocytes recorded on different days of time evolution. All labeled images were divided into three datasets: 75% were included in the training dataset, 15% in the validation dataset, and 10% in the test dataset. To improve the detection of astrocytes87, the training and validation dataset were artificially expanded using the following methods: (i) X-axis flip, random adjustment, (ii), contrast, (iii) brightness, and (iv) random cropping. The dataset size increased to 1105 images by augmentation.
We utilized the acquired datasets to train three models, and the corresponding information is summarized in Table 1. For training the following parameters were adjusted: total steps, warm steps, image resizing. The main difference among the models lay in the resolution of the resized images used during training: (i) Model 20 correspond to 256 \(\times\) 256 px, (ii) Model 30 correspond to 512 \(\times\) 512 px, and (iii) Model 50 correspond to 700 \(\times\) 700 px. We found that Model 50 achieved the highest segmentation accuracy (dice coefficient) on the test set (see Table 1). This is attributed to the high resolution of the input images used for training. However, the training of this model required the greate number of steps and computational efforts. The model trained on 256 \(\times\) 256 px images without augmentation (Model 20) has fewer false positive (FP) detection results than the model trained on 700 \(\times\) 700 px images with augmentation (Model 50). Meanwhile, the model trained on 512 \(\times\) 512 px images with augmentation (Model 30) also has fewer FP detection results than Model 50. It can be concluded that by increasing the training dataset, one can achieve a reduction in FP detection results for Model 50. Based on the data from Table 1, we concluded that Model 20 has the worst characteristics, and therefore, it was not further considered.
The comparison between the segmentation obtained by models 30 and 50, using images that were not included in either the training or test sets, is depicted in Fig. 2. It can be observed that Model 50 detects a higher number of branches compared to Model 30, as evident in the second and third rows of Fig. 2. Additionally, in the second row of Fig. 2, the branches segmented by Model 30 are longer than those segmented by Model 50, but the masks have slight distortions along them. As previously discussed, this discrepancy is associated with the difference in the resolutions of the input images (512 \(\times\) 512 pixels in Model 30 versus 700 \(\times\) 700 pixels in Model 50) used during training. Note that although Model 50 achieves better segmentation, Model 30 exhibits superior performance in minimizing false positives results. It is important, since FP results contribute significant error to the overall statistics as they detect an area that was not labeled as an astrocyte. The high number of false negatives (FN) across all models (astrocyte was labeled but not detected) can be explained by potential errors in cell annotation. In Fig. 3, the model erroneously segmented the intercellular space instead of an astrocyte.
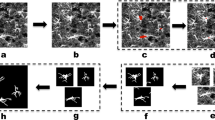
Then, we used Models 30 and 50 for post-processing all available experimental data to assess the morphological characteristics of isolated astrocytes during their time evolution. In particular, we focused on three characteristics: the total length of branches, the number of branchings, and the area of the astrocyte (including its branches). To detect astrocyte branches, we extract their soma based on the local isotropy of the image88,89,90. The directional ratio ranges from low values for vessel-like structures to 1 for drop-shaped structures. This approach has been shown to be very effective in separating soma from branches in fluorescent imaging of neurons91. In contrast to work91, we used masks obtained through a neural network rather than fluorescent images. Example of soma identification is shown in Fig. 4a–c.
Segmentation of the soma in an astrocyte. The (a) is the original microscope image, the (b) is the image colored according to the directional ratio, the (c) is the segmented soma and the (d) is the segmented mask of an astrocyte and its skeletonized branches, different colors correspond to the recognized segments.
After extracting the soma, we separated it from the whole astrocyte body, keeping only the branches. For this purpose, we subtracted the Boolean mask of the soma from the Boolean mask of the segmented astrocyte. The obtained branch masks underwent skeletonization using the scikit-image library function. The border pixels of the astrocyte branches were identified and removed under the condition that their removal did not disrupt the connectivity of the respective object. After skeletonization, the thickness of the astrocyte branch became one pixel (each pixel has two neighboring pixels except begining or end of the branch). Each branch consists of nodes and branch segments coming out of them. Search of segments was developed using a modernized deep-first search graph method: the number of neighboring pixels in skeletonized branch is sequentially calculated and in cases where the number of neighboring pixels exceeds 2 and the previous pixel is not a node, it indicates that this particular pixel serves as a node because more than one segment comes out from it, as shown in Fig. 4d. Note, if a border segment had a length less than 10 pixels (below the typical thickness of branches), we excluded it from consideration. The total length of the branch was calculated by summing the lengths of all segments between the nodes for each astrocyte. Based on this result, we identified nodes, branches and segments of the branches, allowing us to measure the dynamics of cell growth in the experiment.
The dependency of the number of detected astrocytes by Models 30 and 50 in experimental images across different days of astrocyte cultivation are shown in Fig. 5. The maximum number of astrocytes is reached around days 5-6, but by day 10, the count of astrocytes significantly decreases. We attribute this to the fact that after the 6th day, astrocytes form a monolayer with a confluence of approximately 95%, that make them indistinguishable. Examples of a random set of experimental images obtained on different days are shown in Fig. 6.
Histograms of the lengths of all astrocyte branches, astrocyte areas, and the number of nodes measured on the 7th day, during which both models allowed the detection of a sufficient number of astrocytes, are presented in Fig. 7. It can be noted that overall both models lead to similar distributions. Finally, Fig. 8. shows the temporal dependencies of the mean values of the parameters we measured: total length of astrocyte branches L, number of nodes \(N_b\), area S, as well as the corresponding variances D[L], \(D[N_b]\), and D[S]. Orange circles represent data obtained using Model 30, while blue diamonds represent data by Model 50, points measured from samples of fewer than 20 astrocytes are colored gray. Overall, it can be noted that Model 50 estimates longer branches, recognizes a greater number of nodes, and consequently identifies a larger area of the astrocyte (which is considered along with the area of its branches). As noted earlier, this is because Model 50 segments astrocytes with high accuracy. At the same time, both models provide similar values of the investigated parameters dispersion. Important to note that the lengths and number of branchings we measure are notably lower compared to those measured when using cell staining92. This is due to limitations imposed by phase contrast microscopy, which does not allow resolving branches with a diameter of less than \(\lesssim 0.8\) micrometers, consequently restricting the observation of smaller branching networks. The question of the possibility of reconciling data from phase contrast microscopy with fluorescent microscopy of stained cells, as well as the development of an extrapolative model to estimate the total length of branches based on phase contrast microscopy data, deserves further study. The values presented in Fig. 8 are summarized in Table 2 for Model 30 and in Table 3 for Model 50.
The histograms of astrocyte parameters, as measured on the 7th day of experiments. (a) The distribution by lengths, (b) the distribution by areas, and (c) the distribution by the number of nodes. The orange bars represent the results of Model 30, while the blue bars represent the results of Model 50. The touching blue and red bars refer to the same value.
The temporal dependencies of astrocyte parameters are shown: (a) illustrates the average length of branches L, (b) shows the average number of nodes per astrocyte \(N_b\), and (c) presents the average area S of the astrocyte. (d–f) The dependencies of dispersion of the considered characteristics, normalized to the mean value squared, \(D[L]/L^2\), \(D[N_b]/N_b^2\), and \(D[S]/S^2\) respectively. The orange symbols represent the results of Model 30, while blue symbols represent the results of Model 50. Data points calculated from samples of fewer than 20 astrocytes are colored gray.
For in vitro studies, the method should be sensitive enough to various changes in cell morphology induced by the local microenvironment (such as stressful conditions or paracrine and autocrine signaling). To demonstrate the ability of our approach to detect such effects, we run a special series of experiments with Nicotinamide as a well-known modulator of intracellular NAD+ metabolism. Physiology and morphology of astrocytes could be easily affected by altered NAD+ levels (e.g. in aging, ischemia, or neurodegeneration) because glycolytic activity and NAD+-dependent enzymes control their activation, migration and interactions with neurons. Nicotinamide (0.1 mmol) was added to astrocytes on the second day of their culture. Morphological parameters of astrocytes at different time points of nicotinamide exposure are presented in Table 4 for Model 30 and in Table 5 for Model 50. Note that our method allows us to identify noticeable deviations in the mean values and variances of the tested morphological characteristics in the astroglial culture exposed to Nicotinamide.
It is worth noting that the problem of analyzing the evolution of morphological parameters has also been addressed in other studies, for example in93, where the NeuriTES platform was developed to track the dynamic evolution of neurons. This method requires manually labeling the neuron whose evolution is to be tracked and using a measure of transfer entropy to study the relationships between parameters relevant to the evolving system. In contrast, in our work, we proposed to collect statistics of morphological features by days. That is, by imaging a large region, automatically recognizing astrocytes, and extracting morphological parameters of interest.
Conclusion
We performed the experiments with primary cultures of rat brain cortical astrocytes visualized with phase contrast microscopy at different time points of their growth in vitro. We labeled the data and trained three models based on deep convolutional neural networks that enable the recognition of astrocytes in low-contrast images (obtained without using cell staining) and the analysis of their morphology. Using the obtained models, we measured how morphological characteristics such as total branch length, branching count, and overall size of an astrocyte are changed along their growth in the culture. The results of our study show that the application of a neural network to process experimental data with the study of astrocyte growth dynamics can significantly improve the speed of analysis. The novelty of the proposed approach lies in its ability to recognize morphological data from unstained cell microscopic images. Our approach allows us to automate the data processing, making it more efficient. We have demonstrated that training of the neural network with the data obtained in samples that have been thoroughly analyzed by qualified neuroscientists allows achieving good results in predicting the outcomes of astroglia branching. Our results significantly extend the potential use of neural networks in neurobiology and represent an important contribution to the development of experimental data processing methods in this field.
In the conclusion, it is possible to formulate a series of problems and directions for further studies, in which the obtained results and methods can serve as a basis. Firstly, by augmenting the training dataset with data containing a priori known other cell types, the proposed approach can be generalized for the recognition of various cell types in phase-contrast microscopy data. Furthermore, having sufficient datasets for cell morphology analysis based on phase contrast and fluorescence methods, it may be possible to establish accurate correspondences between them, potentially speeding up and reducing the cost of a range of cell morphology studies by eliminating the need for staining. Furthermore, when properly generalized and integrated with a system for automatic periodic scanning of plates with cell culture, our method can enable the development of a system capable of tracking and analyzing the morphology of each individual cell during its cultivation. Finally, the analysis of three-dimensional cellular structures is of great interest, which requires the development of separate methods and algorithms. One possible approach to developing such methods is the synthesis of computer-generated 3D structures based on a series of two-dimensional images, which is actively used in solving similar tasks in the field of soft matter physics. It is worth noting that the proposed method, as well as possible extensions to 3D structures, tracking individual cells, and identifying their types, will be useful for the development of digital twins of individual cells and organoids. For example, these digital twins could be based on cellular Potts models and implemented in packages such as CompuCell3D or other agent-based approaches. In particular, such models require significant statistics of the morphological parameters of cells under different conditions for parameterizing the models or their final verification. In turn, digital twins will allow us to better understand the mechanisms of various processes occurring in cell systems, as well as reduce the number of experiments. All of the above indicates the potential benefit of the proposed method for the scientific communities in biology, biophysics, medicine, neuroscience, and bioimaging.
Materials and methods
Isolation and cultivation of astrocytes from the rat brain cortex
A dissociated culture of astrocytes was obtained from newborn Wistar rats using the method of enzymatic-mechanical dissociation according to a previously described method94,95. The isolated structures were rinsed with calcium and magnesium-free phosphate-buffered saline (PBS, pH 7.4, Gibco Life Technologies, USA), minced with a scalpel, and incubated for 15 min at 37 \(^{\circ }\)C in a 0.05\(\%\) trypsin and 0.02\(\%\) EDTA solution (Gibco Life Technologies, USA). After two washes with phosphate buffer, the structures were mechanically dissociated in culture medium using stepwise pipetting. The culture medium consisted of 90\(\%\) minimum essential medium (MEM Gibco, UK), 10\(\%\) fetal bovine serum (FBS), 2 mM glutamax (Gibco, UK), 10 mM HEPES buffer (Sigma, USA), pH 7.2–7.4. The cell suspension was centrifuged for 3 minutes at 1000 rpm, the pellet was resuspended in the culture medium, and cells were seeded into two T-25 culture flasks. Cultures were maintained in a CO\(_2\) incubator (RWD Life Science, China) at 37 \(^{\circ }\)C, 5\(\%\) CO\(_2\), and 98\(\%\) relative humidity. On the second day of cultivation, a complete change of the culture medium was performed to a medium containing 10% FBS. Upon monolayer formation, astrocytic cultures were transferred to 96-well plastic plates (Servicebio, China) with 100 \(\upmu\)l of a suspension containing \(2\times 10^6\) cells per ml. This two actions also serves as a step in purifying the astrocyte culture. Cell counting and viability assessment were performed using the Countess automated cell counter (Invitrogen), which recorded total cell count, live and dead cells, survival percentage, and cell size distribution. All experimental protocols were approved by the Ethical Committee of the “Research Center of Neurology”.
Immunofluorescence assay
The isolated primary culture cells were transferred onto cover slides and incubated in a 10\(\%\) FBS culture medium upon monolayer formation. The media was discarded, and cells were washed in PBS. The cells were further fixed with fresh 5\(\%\) paraformaldehyde (PFA) for 15 min, samples were treated with 0.1\(\%\) TritonX-100 (Calbiochem Biochemicals, USA) and blocked with 5\(\%\) bovine serum albumin (BSA, Sigma, Germany). The primary antibodies were diluted in the IHCDiluent (Leica), placed on cover slides, incubated at 4 \(^\circ\)C for 6 h. The selected antibodies were rabbit anti-GFAP antibody (1:250, Abcam, ab68428), rabbit anti-NeuN antibody (1:200, ABclonal, A19086), rabbit anti- IBA1 antibody (1:200, Affinity Biosciences, DF6442), rabbit anti-PDGFRB antibody (1:300, Elabscience, E-AB-32531). After five washes with PBS, the cells were incubated with Goat Anti-Rabit IgG(H+L) Alexa Fluor 594-cojugated (1:250, Elabscience, E-AB1060) or Alexa Fluor 594-conjugated Donkey Anti-Rabit IgG(H+L) (1:200, Fine Test, FNSA-0063), FITC Goat Anti-Rabbit IgG (H+L) (1:200, ABclonal, AS011) or FITC-linked Guinea pig Anti-Rabbit IgG Polyclonal antibody (1:1000, Cloud-Clone, SAA544Rb58) secondary antibodies for 2 h at room temperature. Then cells were washed with PBS twice and stained with Fluoroshield and DAPI (Sigma). The slides proceeded without primary antibodies were used as a negative control. Images were captured using EVOS M7000 system (ThermoFisher Scientific).
Identification of culture cells
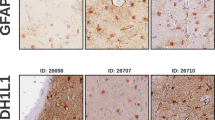
Immunophenotyping of neuronal, astroglial and microglial cells was performed to confirm the purity of the primary culture. As shown in Fig. 9a, the majority of isolated cells was positively stained with the astrocyte marker GFAP. Negative results have been obtained in immunostaining with NeuN antibodies that recognize mature neurons, thereby confirming the absence of neuronal cells in the primary culture. The cells in the primary culture were positively stained with a microglial marker Iba1 (Fig. 9b), but not more than 10\(\%\) of Iba1-immunopositive cells have been found. Also, cells in the primary culture were positively stained with a pericyte marker PDGFRB, the number of PDGFRB-positive cells was no more than 8\(\%\) (Fig. 9c). Thus, we confirm that astrocytes are the predominant type of cells within the primary culture which was further used for the application of machine learning algorithms to recognize astrocytes in phase-contrast images.
Identification of culture cells: (a) astrocytes in the primary culture: the cell bodies and processes are stained with GFAP antibody (red), the nuclei are stained with DAPI (blue), (b) microglia in the primary culture: cell bodies and processes are stained with Iba1 antibody (red), the nuclei are stained with DAPI (blue), (c) pericytes in the primary culture: the cell bodies and processes are stained with PDGFRB antibody (green), the nuclei are stained with DAPI (blue).
Phase-contrast microscopy
To study morphology of astrocytes in their native state, we applied the phase contrast microscopy which does not require any staining and fixation. Live observations were conducted using phase-contrast microscopy with the “EVOS M7000” imaging system (Thermo Fisher Scientific, USA). Experimental work was carried out to visualize rat astrocyte cells during their growth. An EVOS M7000 phase contrast microscope with an AMC1000 stage incubator was used for observation. A 96-well plate with cells was installed inside a built-in incubator with motorized stages. We conducted a series of experiments over a duration of 18 days. Microphotography of cells was carried out daily and included photographic recording of a 4x4 frame field in each well of the plate at a magnification of 40x starting from the second day of cultivation. Note that the study area was selected randomly, without preliminary analysis for the presence of astrocytes. Since the microscope has motorized stage movers, it was possible to track the position of the frame according to coordinates in the XY plane throughout the entire well area. This approach allowed the same area of each well to be recorded daily. Thus, as a result of the experiments, photographs of astrocytes were obtained at different moments of their growth. The experimental work involved capturing photographic records of astrocytes through microscopy of 6 wells within the plate. In total, over 1100 astrocytes were recorded at various growth stages. The results of the experiment are low-contrast images of astrocytes, which are subsequently the object of analysis for neural networks. In Fig. 10. we show images acquired during the same experiment (on the same day and on an identical setup) to check for possible bias between experiments under nominally identical conditions.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Cabezas, R. et al. Growth factors and astrocytes metabolism: Possible roles for platelet derived growth factor. Med. Chem. 12, 204–210. https://doi.org/10.2174/1573406411666151019120444 (2016).
Nag, S. & Walker, J. The Blood–Brain and Other Neural Barriers: Reviews and Protocols (Humana Press, 2011).
Siracusa, R., Fusco, R. & Cuzzocrea, S. Astrocytes: Role and functions in brain pathologies. Front. Pharmacol.https://doi.org/10.3389/fphar.2019.01114 (2019).
Freeman, M. R. Specification and morphogenesis of astrocytes. Science 330, 774–778. https://doi.org/10.1126/science.1190928 (2010).
Guillamon-Vivancos, T., Gomez-Pinedo, U. & Matias-Guiu, J. Astrocytes in neurodegenerative diseases (I): Function and molecular description. Neurologia (English Edition) 30, 119–129. https://doi.org/10.1016/j.nrleng.2014.12.005 (2015).
Zhou, Y. et al. Dual roles of astrocytes in plasticity and reconstruction after traumatic brain injury. Cell Commun. Signal.https://doi.org/10.1186/s12964-020-00549-2 (2020).
Sloan, S. A. & Barres, B. A. Mechanisms of astrocyte development and their contributions to neurodevelopmental disorders. Curr. Opin. Neurobiol. 27, 75–81. https://doi.org/10.1016/j.conb.2014.03.005 (2014).
Zhou, B., Zuo, Y.-X. & Jiang, R.-T. Astrocyte morphology: Diversity, plasticity, and role in neurological diseases. CNS Neurosci. Ther. 25, 665–673. https://doi.org/10.1111/cns.13123 (2019).
Wang, S.S.-H. et al. Functional trade-offs in white matter axonal scaling. J. Neurosci. 28, 4047–4056. https://doi.org/10.1523/jneurosci.5559-05.2008 (2008).
Middelkamp, H. H. T. et al. Cell type-specific changes in transcriptomic profiles of endothelial cells, IPSC-derived neurons and astrocytes cultured on microfluidic chips. Sci. Rep.https://doi.org/10.1038/s41598-021-81933-x (2021).
Herland, A. et al. Distinct contributions of astrocytes and pericytes to neuroinflammation identified in a 3D human blood–brain barrier on a chip. PLOS ONE 11, e0150360. https://doi.org/10.1371/journal.pone.0150360 (2016).
Oudart, M. et al. Astrodot: A new method for studying the spatial distribution of mRNA in astrocytes. J. Cell Sci.https://doi.org/10.1242/jcs.239756 (2020).
Yu, X., Nagai, J. & Khakh, B. S. Improved tools to study astrocytes. Nat. Rev. Neurosci. 21, 121–138. https://doi.org/10.1038/s41583-020-0264-8 (2020).
Oschmann, F., Berry, H., Obermayer, K. & Lenk, K. From in silico astrocyte cell models to neuron-astrocyte network models: A review. Brain Res. Bull. 136, 76–84. https://doi.org/10.1016/j.brainresbull.2017.01.027 (2018).
Savtchenko, L. P. et al. Disentangling astroglial physiology with a realistic cell model in silico. Nat. Commun.https://doi.org/10.1038/s41467-018-05896-w (2018).
Verisokin, A. Y., Verveyko, D. V., Postnov, D. E. & Brazhe, A. R. Modeling of astrocyte networks: Toward realistic topology and dynamics. Front. Cell. Neurosci.https://doi.org/10.3389/fncel.2021.645068 (2021).
Nguyen, K.-V., Hernandez-Garzon, E. & Valette, J. Efficient GPU-based Monte-Carlo simulation of diffusion in real astrocytes reconstructed from confocal microscopy. J. Magnet. Reson. 296, 188–199. https://doi.org/10.1016/j.jmr.2018.09.013 (2018).
Lenk, K. et al. A computational model of interactions between neuronal and astrocytic networks: The role of astrocytes in the stability of the neuronal firing rate. Front. Comput. Neurosci.https://doi.org/10.3389/fncom.2019.00092 (2020).
Hines, M. L. & Carnevale, N. T. The neuron simulation environment. Neural Comput. 9, 1179–1209. https://doi.org/10.1162/neco.1997.9.6.1179 (1997).
Allam, S. L. et al. A computational model to investigate astrocytic glutamate uptake influence on synaptic transmission and neuronal spiking. Front. Comput. Neurosci.https://doi.org/10.3389/fncom.2012.00070 (2012).
Radan, M., Djikic, T., Obradovic, D. & Nikolic, K. Application of in vitro pampa technique and in silico computational methods for blood-brain barrier permeability prediction of novel cns drug candidates. Eur. J. Pharmaceut. Sci. 168, 106056. https://doi.org/10.1016/j.ejps.2021.106056 (2022).
Abdellah, M. et al. Metaball skinning of synthetic astroglial morphologies into realistic mesh models for in silico simulations and visual analytics. Bioinformatics 37, i426–i433. https://doi.org/10.1093/bioinformatics/btab280 (2021).
Comes, M. C. et al. Accelerating the experimental responses on cell behaviors: A long-term prediction of cell trajectories using social generative adversarial network. Sci. Rep.https://doi.org/10.1038/s41598-020-72605-3 (2020).
Goncalves Seabra, A. C., Silva, A. F. D., Stieglitz, T. & Amado-Rey, A. B. In silico blood pressure models comparison. IEEE Sens. J. 22, 23486–23493. https://doi.org/10.1109/jsen.2022.3215597 (2022).
Ghaffarizadeh, A., Heiland, R., Friedman, S. H., Mumenthaler, S. M. & Macklin, P. Physicell: An open source physics-based cell simulator for 3-D multicellular systems. PLOS Comput. Biol. 14, e1005991. https://doi.org/10.1371/journal.pcbi.1005991 (2018).
Swat, M. H. et al. Multi-Scale Modeling of Tissues Using CompuCell3D. 325–366 (Elsevier, 2012).
Roy, M. & Finley, S. D. Metabolic reprogramming dynamics in tumor spheroids: Insights from a multicellular, multiscale model. PLOS Comput. Biol. 15, e1007053. https://doi.org/10.1371/journal.pcbi.1007053 (2019).
Risau, W. Mechanisms of angiogenesis. Nature 386, 671–674. https://doi.org/10.1038/386671a0 (1997).
Ferozepurwalla, Z., Merzah, J., Thielemans, L. & Birdsey, G. Molecular and Cellular Mechanisms of Angiogenesis. 219–226 (Springer, 2019).
Villa, C., Chaplain, M. A. J., Gerisch, A. & Lorenzi, T. Mechanical models of pattern and form in biological tissues: The role of stress-strain constitutive equations. Bull. Math. Biol.https://doi.org/10.1007/s11538-021-00912-5 (2021).
Tosin, A., Ambrosi, D. & Preziosi, L. Mechanics and chemotaxis in the morphogenesis of vascular networks. Bull. Math. Biol. 68, 1819–1836. https://doi.org/10.1007/s11538-006-9071-2 (2006).
Manoussaki, D. A mechanochemical model of angiogenesis and vasculogenesis. ESAIM Math. Model. Numer. Anal. 37, 581–599 https://doi.org/10.1051/m2an:2003046 (2003).
Nakazawa, T., Tasaki, S., Nakai, K. & Suzuki, T. Multicellular model of angiogenesis. AIMS Bioeng. 9, 44–60. https://doi.org/10.3934/bioeng.2022004 (2022).
Wang, Y. et al. On-chip-angiogenesis based on a high-throughput biomimetic three-dimensional cell spheroid culture system. Analyst 148, 3870–3875. https://doi.org/10.1039/d3an00817g (2023).
Mada, J. & Tokihiro, T. Pattern formation of vascular network in a mathematical model of angiogenesis. Jpn. J. Indus. Appl. Math. 39, 351–384. https://doi.org/10.1007/s13160-021-00493-9 (2021).
Wyss-Coray, T. et al. Adult mouse astrocytes degrade amyloid-b in vitro and in situ. Nat. Med. 9, 453–457. https://doi.org/10.1038/nm838 (2003).
Ogata, K. & Kosaka, T. Structural and quantitative analysis of astrocytes in the mouse hippocampus. Neuroscience 113, 221–233. https://doi.org/10.1016/s0306-4522(02)00041-6 (2002).
Zhuo, L. et al. Live astrocytes visualized by green fluorescent protein in transgenic mice. Dev. Biol. 187, 36–42. https://doi.org/10.1006/dbio.1997.8601 (1997).
Aten, S. et al. Ultrastructural view of astrocyte arborization, astrocyte-astrocyte and astrocyte-synapse contacts, intracellular vesicle-like structures, and mitochondrial network. Prog. Neurobiol. 213, 102264. https://doi.org/10.1016/j.pneurobio.2022.102264 (2022).
Kacem, K., Lacombe, P., Seylaz, J. & Bonvento, G. Structural organization of the perivascular astrocyte endfeet and their relationship with the endothelial glucose transporter: A confocal microscopy study. Glia 23, 1–10. https://doi.org/10.1002/(sici)1098-1136(199805)23:1<1::aid-glia1>3.0.co;2-b (1998).
Jackson, J. G. & Robinson, M. B. Regulation of mitochondrial dynamics in astrocytes: Mechanisms, consequences, and unknowns. Glia 66, 1213–1234. https://doi.org/10.1002/glia.23252 (2017).
Lanjakornsiripan, D. et al. Layer-specific morphological and molecular differences in neocortical astrocytes and their dependence on neuronal layers. Nat. Commun.https://doi.org/10.1038/s41467-018-03940-3 (2018).
Chai, H. et al. Neural circuit-specialized astrocytes: Transcriptomic, proteomic, morphological, and functional evidence. Neuron 95, 531-549.e9. https://doi.org/10.1016/j.neuron.2017.06.029 (2017).
Beretta, C. et al. Extracellular vesicles from amyloid-b exposed cell cultures induce severe dysfunction in cortical neurons. Sci. Rep.https://doi.org/10.1038/s41598-020-72355-2 (2020).
Zhang, C. et al. Effects of dimethyl sulfoxide on the morphology and viability of primary cultured neurons and astrocytes. Brain Res. Bull. 128, 34–39. https://doi.org/10.1016/j.brainresbull.2016.11.004 (2017).
Wakida, N. M. et al. Phagocytic response of astrocytes to damaged neighboring cells. PLOS ONE 13, e0196153. https://doi.org/10.1371/journal.pone.0196153 (2018).
Wakida, N. M., Cruz, G. M. S., Pouladian, P., Berns, M. W. & Preece, D. Fluid shear stress enhances the phagocytic response of astrocytes. Front. Bioeng. Biotechnol.https://doi.org/10.3389/fbioe.2020.596577 (2020).
Teh, D. B. L. et al. Transcriptome analysis reveals neuroprotective aspects of human reactive astrocytes induced by interleukin 1b. Sci. Rep.https://doi.org/10.1038/s41598-017-13174-w (2017).
Allahyari, R. V., Clark, K. L., Shepard, K. A. & Garcia, A. D. R. Sonic hedgehog signaling is negatively regulated in reactive astrocytes after forebrain stab injury. Sci. Rep.https://doi.org/10.1038/s41598-018-37555-x (2019).
Farhy-Tselnicker, I. & Allen, N. J. Astrocytes, neurons, synapses: A tripartite view on cortical circuit development. Neural Dev.https://doi.org/10.1186/s13064-018-0104-y (2018).
Vezzoli, E. et al. Ultrastructural evidence for a role of astrocytes and glycogen-derived lactate in learning-dependent synaptic stabilization. Cereb. Cortex 30, 2114–2127. https://doi.org/10.1093/cercor/bhz226 (2019).
Gavrilov, N. et al. Astrocytic coverage of dendritic spines, dendritic shafts, and axonal boutons in hippocampal neuropil. Front. Cell. Neurosci.https://doi.org/10.3389/fncel.2018.00248 (2018).
Arizono, M. et al. Structural basis of astrocytic Ca2+ signals at tripartite synapses. Nat. Commun.https://doi.org/10.1038/s41467-020-15648-4 (2020).
Berg, S. et al. Ilastik: Interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232. https://doi.org/10.1038/s41592-019-0582-9 (2019).
Comes, M. C. et al. A camera sensors-based system to study drug effects on in vitro motility: The case of pc-3 prostate cancer cells. Sensors 20, 1531. https://doi.org/10.3390/s20051531 (2020).
D’Orazio, M. et al. Deciphering cancer cell behavior from motility and shape features: Peer prediction and dynamic selection to support cancer diagnosis and therapy. Front. Oncol.https://doi.org/10.3389/fonc.2020.580698 (2020).
Suleymanova, I. et al. A deep convolutional neural network approach for astrocyte detection. Sci. Rep.https://doi.org/10.1038/s41598-018-31284-x (2018).
D’Orazio, M. et al. Machine learning phenomics (MLP) combining deep learning with time-lapse-microscopy for monitoring colorectal adenocarcinoma cells gene expression and drug-response. Sci. Rep.https://doi.org/10.1038/s41598-022-12364-5 (2022).
Mencattini, A. et al. Deep-manager: A versatile tool for optimal feature selection in live-cell imaging analysis. Commun. Biol.https://doi.org/10.1038/s42003-023-04585-9 (2023).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444. https://doi.org/10.1038/nature14539 (2015).
Guo, Y. et al. Deep learning for visual understanding: A review. Neurocomputing 187, 27–48. https://doi.org/10.1016/j.neucom.2015.09.116 (2016).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. Adv. Neural Inform. Process. Syst. 25, 133 (2012).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. https://doi.org/10.48550/ARXIV.1409.1556 (2014).
Zeiler, M. D. & Fergus, R. Visualizing and understanding convolutional networks. In Computer Vision—ECCV 2014: 13th European Conference, Zurich, Switzerland, September 6–12, 2014, Proceedings, Part I 13. 818–833 (Springer, 2014).
Szegedy, C. et al. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 1–9 (2015).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, October 5–9, 2015, Proceedings, Part III 18. 234–241 (Springer, 2015).
Xia, X. & Kulis, B. W-net: A deep model for fully unsupervised image segmentation. arXiv preprint arXiv:1711.08506 (2017).
Ciresan, D., Giusti, A., Gambardella, L. & Schmidhuber, J. Deep neural networks segment neuronal membranes in electron microscopy images. Adv. Neural Inform. Process. Syst. 25, 14 (2012).
Jung, C. et al. W-net: A CNN-based architecture for white blood cells image classification. arXiv preprint arXiv:1910.01091 (2019).
Shanthi, T. & Sabeenian, R. Modified alexnet architecture for classification of diabetic retinopathy images. Comput. Electr. Eng. 76, 56–64. https://doi.org/10.1016/j.compeleceng.2019.03.004 (2019).
Kaur, T. & Gandhi, T. K. Automated brain image classification based on VGG-16 and transfer learning. In 2019 International Conference on Information Technology (ICIT). https://doi.org/10.1109/icit48102.2019.00023 (IEEE, 2019).
Lu, T., Han, B. & Yu, F. Detection and classification of marine mammal sounds using alexnet with transfer learning. Ecol. Inform. 62, 101277. https://doi.org/10.1016/j.ecoinf.2021.101277 (2021).
Mashrur, F. R., Dutta Roy, A. & Saha, D. K. Automatic identification of arrhythmia from ECG using Alexnet convolutional neural network. In 2019 4th International Conference on Electrical Information and Communication Technology (EICT). https://doi.org/10.1109/eict48899.2019.9068806 (IEEE, 2019).
Lu, X., Duan, X., Mao, X., Li, Y. & Zhang, X. Feature extraction and fusion using deep convolutional neural networks for face detection. Math. Probl. Eng. 1–9, 2017. https://doi.org/10.1155/2017/1376726 (2017).
Yu, B., Yin, H. & Zhu, Z. St-unet: A spatio-temporal u-network for graph-structured time series modeling. arXiv preprint arXiv:1903.05631 (2019).
Jiang, Y., Yang, M., Wang, S., Li, X. & Sun, Y. Emerging role of deep learning-based artificial intelligence in tumor pathology. Cancer Commun. 40, 154–166. https://doi.org/10.1002/cac2.12012 (2020).
Chen, T. & Chefd’hotel, C. Deep Learning Based Automatic Immune Cell Detection for Immunohistochemistry Images. 17–24 (Springer, 2014).
Labate, D. & Kayasandik, C. Advances in quantitative analysis of astrocytes using machine learning. Neural Regener. Res. 18, 313. https://doi.org/10.4103/1673-5374.346474 (2023).
He, K., Gkioxari, G., Dollár, P. & Girshick, R. Mask r-CNN. In Proceedings of the IEEE International Conference on Computer Vision. 2961–2969 (2017).
Tsai, H.-F., Gajda, J., Sloan, T. F., Rares, A. & Shen, A. Q. Usiigaci: Instance-aware cell tracking in stain-free phase contrast microscopy enabled by machine learning. SoftwareX 9, 230–237 (2019).
Girshick, R., Radosavovic, I., Gkioxari, G., Dollár, P. & He, K. Detectron. https://github.com/facebookresearch/detectron (2018).
Kirillov, A. et al. Segment anything. In Proceedings of the IEEE/CVF International Conference on Computer Vision. 4015–4026 (2023).
Edlund, C. et al. Livecell—A large-scale dataset for label-free live cell segmentation. Nat. Methods 18, 1038–1045 (2021).
Zhang, W. et al. Tobacco leaf segmentation based on improved mask RCNN algorithm and SAM model. IEEE Access (2023).
He, S., Bao, R., Li, J., Grant, P. E. & Ou, Y. Accuracy of segment-anything model (SAM) in medical image segmentation tasks. arXiv preprint arXiv:2304.09324 (2023).
Russell, B. C., Torralba, A., Murphy, K. P. & Freeman, W. T. Labelme: A database and web-based tool for image annotation. Int. J. Comput. Vis. 77, 157–173 (2008).
Shijie, J., Ping, W., Peiyi, J. & Siping, H. Research on data augmentation for image classification based on convolution neural networks. In 2017 Chinese Automation Congress (CAC). 4165–4170 (IEEE, 2017).
Kayasandik, C. B. & Labate, D. Improved detection of soma location and morphology in fluorescence microscopy images of neurons. J. Neurosci. Methods 274, 61–70 (2016).
Labate, D., Laezza, F., Negi, P., Ozcan, B. & Papadakis, M. Efficient processing of fluorescence images using directional multiscale representations. Math. Model. Nat. Phenomena 9, 177–193 (2014).
Ozcan, B., Labate, D., Jiménez, D. & Papadakis, M. Directional and non-directional representations for the characterization of neuronal morphology. In Wavelets and Sparsity XV. Vol. 8858. 12–22 (SPIE, 2013).
Ozcan, B., Negi, P., Laezza, F., Papadakis, M. & Labate, D. Automated detection of soma location and morphology in neuronal network cultures. PloS one 10, e0121886 (2015).
Hayashi, M. K., Sato, K. & Sekino, Y. Neurons induce tiled astrocytes with branches that avoid each other. Int. J. Mol. Sci. 23, 4161. https://doi.org/10.3390/ijms23084161 (2022).
Mencattini, A. et al. Neurites. monitoring neurite changes through transfer entropy and semantic segmentation in bright-field time-lapse microscopy. Patterns 2 (2021).
Weikert, S. et al. Rapid Ca2+-dependent no-production from central nervous system cells in culture measured by no-nitrite/ozone chemoluminescence. Brain Res. 748, 1–11. https://doi.org/10.1016/s0006-8993(96)01241-3 (1997).
Stel’mashuk, E. V. et al. Vliianie induktora neirovospaleniia na komponenty neirovaskuliarnoi edinitsy golovnogo mozga in vitro. Rossiiskii Fiziol. Zh. IM Sechenova 108, 686–696 (2022).
Acknowledgements
The post-processing and analysis of experiments was supported by the Russian Science Foundation, Grant No. 22-72-10128.
Author information
Authors and Affiliations
Contributions
E.V.Y, N.P.K. and N.A.K. developed experimental work; I.V.S. developed an approach for recognition and morphological analysis; N.A.K., S.V.N., A.A.S., D.E.N., E.V.Y. conducted the experiments; N.A.K., S.V.N. obtained and cultivated cells; I.V.S., A.A.S., D.E.N., I.R.D., K.D.G., I.N.S., S.V.N. created the dataset for training a neural network; I.V.S., K.D.G., N.P.K. post-processed experimental data; N.P.K., E.V.Y., A.D.N. I.V.S., wrote the manuscript; N.P.K., E.V.Y., I.V.S., I.N.S., I.R.D. prepared the figures; N.P.K., E.V.Y., I.V.S., N.A.K., A.B.S., S.O.Y. discussed the results; N.P.K. and E.V.Y. conceived, developed, and directed the study. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yakovlev, E.V., Simkin, I.V., Shirokova, A.A. et al. Machine learning approach for recognition and morphological analysis of isolated astrocytes in phase contrast microscopy. Sci Rep 14, 9846 (2024). https://doi.org/10.1038/s41598-024-59773-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-59773-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.