Abstract
The invasive brown widow spider, Latrodectus geometricus (Araneae: Theridiidae), has spread in multiple locations around the world and, along with it, brought associated organisms such as endosymbionts. We investigated endosymbiont diversity and prevalence across putative native and invasive populations of this spider, predicting lower endosymbiont diversity across the invasive range compared to the native range. First, we characterized the microbial community in the putative native (South Africa) and invasive (Israel and the United States) ranges via high throughput 16S sequencing of 103 adult females. All specimens were dominated by reads from only 1–3 amplicon sequence variants (ASV), and most individuals were infected with an apparently uniform strain of Rhabdochlamydia. We also found Rhabdochlamydia in spider eggs, indicating that it is a maternally-inherited endosymbiont. Relatively few other ASV were detected, but included two variant Rhabdochlamydia strains and several Wolbachia, Spiroplasma and Enterobacteriaceae strains. We then diagnostically screened 118 adult female spiders from native and invasive populations specifically for Rhabdochlamydia and Wolbachia. We found Rhabdochlamydia in 86% of individuals and represented in all populations, which suggests that it is a consistent and potentially important associate of L. geometricus. Wolbachia was found at lower overall prevalence (14%) and was represented in all countries, but not all populations. In addition, we found evidence for geographic variation in endosymbiont prevalence: spiders from Israel were more likely to carry Rhabdochlamydia than those from the US and South Africa, and Wolbachia was geographically clustered in both Israel and South Africa. Characterizing endosymbiont prevalence and diversity is a first step in understanding their function inside the host and may shed light on the process of spread and population variability in cosmopolitan invasive species.
Similar content being viewed by others
Introduction
When moving into new habitats, invasive species may bring along microbial associates that can influence the invasion process1,2. Some microbes are vertically inherited endosymbionts that are restricted to the invasive species, but may yet influence interactions between the invasive and native species. Maternally-inherited endosymbionts have been shown to affect traits important to host fitness such as dispersal3, fecundity4, and defenses against natural enemies5, potentially providing an advantage to the invasive species6. Some endosymbionts affect the organism’s reproductive biology, for example, by modifying offspring sex ratio in infected populations, which can affect the speed of invasive spread7. For example, acting as both a mutualist and reproductive manipulator, Rickettsia caused whiteflies to have higher fitness and a higher proportion of daughters, and quickly spread in invasive populations8.
Assessing endosymbiont prevalence across geographically distant populations can provide a key to understanding the role of a symbiont. Widespread prevalence of a facultative endosymbiont suggests that the symbiont plays a functional role in its host, such as providing fitness benefits or manipulating reproduction9,10. The latter is often manifested by sex ratio distortions, although the most common reproductive manipulation is cytoplasmic incompatibility (CI), which causes incompatibilities between infected males and uninfected females but does not alter the sex ratio of the population11. Our understanding of the dynamics and prevalence of facultative endosymbiont infection during invasive spread is limited, especially for non-insect arthropod endosymbionts.
Invasive populations are predicted to exhibit reduced endosymbiont prevalence and diversity compared to native populations. During founding events, often few individuals are initially introduced into the invasive range12, in which case only a subset of endosymbionts found in the native range might be introduced to the new location13. However, in most biological invasions, multiple introductions are common14, and so endosymbiont diversity might be lower initially, and then increase over time as more individuals arrive from various localities15. Comparing endosymbiont diversity across invasive and native populations can provide valuable insights into the gain and loss of microbial communities during the invasion process.
The brown widow spider, Latrodectus geometricus (Theridiidae), is a medically important spider with neurotoxic venom. Latrodectus geometricus has spread recently to multiple locations around the world from the putative native range in southern Africa, most likely via cargo shipments16,17. Evidence suggests that during invasion, establishment and spread, spider traits related to dispersal, fecundity, and body size shifted across populations that were established over different time periods18. In addition to these shifts in ecologically important traits, associations with other organisms, such as parasitoids19 or endosymbionts, may have also changed during the invasion spread.
Endosymbionts of widow spiders (genus Latrodectus) are poorly known. A previous study on L. geometricus identified the endosymbiont Rhabdochlamydia, but only examined a few adult females in a single, inbred lab population in Florida, USA20. The same study did not detect Rhabdochlamydia in two other Latrodectus species. Hence, a further study across field-collected individuals worldwide is necessary to assess the presence of Rhabdochlamydia more broadly across populations of L. geometricus. The family Rhabdochlamydiaceae (Phylum: Chlamydiota) is predicted to be the most diverse chlamydial family21. It includes important vertebrate and human pathogens and is widespread across soil and aquatic ecosystems with many yet unknown hosts22. The genus Rhabdochlamydia has been found in a few distantly-related invertebrate hosts, including a cockroach23, a tick24, a dwarf spider25, and a terrestrial isopod26, although it was not found at a high prevalence within any of these species.
Also previously found in invasive populations of L. geometricus was Wolbachia, as a facultative associate in varying prevalence across populations27. Wolbachia infection is common in arthropods, with 40–60% of species infected28, as well as in other invertebrates including nematodes29. Wolbachia is known to affect the fitness and reproduction of many of its hosts, which could have implications for successful invasive establishment and spread30.
In this study, we compared endosymbiont presence and diversity across populations of the brown widow spider, L. geometricus, from the putative native range in South Africa to populations in the invasive range in the United States and Israel, using both high-throughput sequencing and diagnostic PCR screens. Our objectives were to (1) characterize the dominant endosymbionts in L. geometricus, (2) compare prevalence and diversity across purported native and known invasive ranges, and (3) investigate geographic patterns of endosymbiont infection within countries. We predicted that, due to founder effects, some endosymbionts would be lost and infection rates would be lower in invasive populations in the U.S. and Israel compared to putative native populations in South Africa, and that geographic patterns of endosymbiont loss would reflect the proposed routes of invasive spread of L. geometricus within each country.
Methods
Study species
Latrodectus geometricus, the brown widow spider, is a globally invasive species that has established populations in parts of North and South America, the Middle East, Australia, and Asia17. In the United States, L. geometricus was first detected in Miami, Florida in 193631, was confined to southern Florida until the late 1990s, and was subsequently detected in Texas and California in the 2000s32. In Israel, L. geometricus was first detected in the Tel Aviv area in 198033, and in the Negev region after 200034. Throughout the global invasive range, L. geometricus is found in urban and settled habitats, and builds nests on and around buildings, on fences, garden furniture, trash bins, and in playgrounds17.
Study sites
We collected L. geometricus adult females from urban environments across the United States (Edisto Island, South Carolina n = 10; Gainesville, Florida n = 10; Austin, Texas n = 6; Los Angeles, California n = 7), Israel (Haifa n = 7, Tel Aviv n = 10, Be’er Sheva n = 10, Yeruham n = 8, Midreshet Ben-Gurion n = 10, Eilat n = 1), and South Africa (Modimolle n = 10, Pretoria n = 5, Johannesburg n = 5, Kimberley n = 8, Cape Town n = 5, Riebeeck-Kasteel n = 6, George n = 7). Spiders were deprived of food for one week before they were preserved in 100% ethanol. Starved individuals have minimal gut content and are less likely to result in false positives for endosymbionts found in the spider’s prey35. To learn about the potential for vertical transmission, we also collected L. geometricus egg sacs from two sites in South Africa: Kimberley (n = 1) and Riebeeck Kasteel (n = 2), and sampled egg sacs produced in the laboratory from Midreshet Ben-Gurion (n = 3) and Tel Aviv (n = 2), Israel.
Bacterial 16S sequencing
We surface-sterilized each adult female L. geometricus specimen (n = 125) with a series of bleach and ethanol rinses36 before longitudinally dividing the abdomen in half and extracting DNA from one half using DNeasy Blood and Tissue extraction kits (Qiagen, Germantown, MD) according to manufacturer’s instructions. In addition, we extracted DNA from the legs of two specimens, as well as from the eggs of 8 L. geometricus egg sacs to assess endosymbiont presence outside reproductive tissues and the potential for maternal transmission, respectively. Extraction quality for each sample was verified by PCR amplification of a ~ 650 bp segment of the COI gene (forward primer, lco1490: 5′-GGTCAACAAATCATAAAGATATTGG-3′, reverse primer, hco2198: 5′-TAAACTTCAGGGTGACCAAAAAATCA-3′; cycling conditions: one cycle of 94 °C for 3 min, followed by 35 cycles of 95 °C for 30 s, 53 °C for 30 s, 72 °C for 1 min, final extension at 72 °C for 5 min37. If COI failed to amplify, we attempted a second extraction with the other half of the abdomen. If this extraction failed to amplify product as well, we assumed sample preservation had been poor and eliminated the specimen from the dataset entirely (7/125 specimens).
To investigate which endosymbionts were present in these specimens, we profiled the microbiomes using high-throughput sequencing of the bacterial community. We amplified the V4 region of bacterial 16S rRNA for each sample using dual indexed 515F/806R primers38. We visualized the resulting products, and multiplexed 1 µl aliquots from successful amplifications into one of two libraries that were purified with GenCatch PCR Cleanup Kits. Samples that failed to amplify (6/118 samples) were not included in the library. Each library also included specimens from other projects that are not reported here, and received a PhiX spike to increase sequence heterogeneity among the amplified sequences. Libraries were sequenced at the University of Kentucky genomics core facility on an Illumina Miseq instrument using a paired-end strategy and 250 bp reads. Sequences from each run were demultiplexed, trimmed and quality filtered within BaseSpace (Illumina, basespace.illumina.com), then imported into QIIME2 (v2021.11, https://qiime2.org39) using a manifest. We conducted additional quality control using deblur40 implemented in QIIME2 using default parameters and a trim length of 251 bases. Resulting amplicon sequence variants (ASV) were taxonomically classified using a naïve Bayes classifier that was trained on the 515F/806R V4 region of the Greengenes 13_8 99% OTUs reference database41. We filtered out 15 ASV that originated from other specimens in the sequencing run (e.g., obligate endosymbionts of other host taxa, see42 for discussion of index swapping), which collectively constituted only a small minority (0.14%) of the 3.57 × 106 reads associated with the L. geometricus samples. Following filtering, L. geometricus samples with less than 1000 reads were excluded from further analysis (9/112 adult samples, 6/8 egg samples). For the remaining samples, we blasted high prevalence ASV sequences (> 1% of any L. geometricus sample) against the NCBI nt database using the megablast algorithm, to identify bacterial taxa that may not have been included in the reference database. For ASV that appeared at very high prevalence or frequency (> 90% of reads for any specimen, or found in multiple specimens across multiple locations), we amplified a longer segment of 16S using universal primers from specimen(s) dominated by that taxon, to aid in taxonomic placement (Forward primer, 27F: 5′-AGAGTTTGATCMTGGCTCAG-3′, reverse primer 1492R: 5′-GGTTACCTTGTTACGACTT-3′, cycling conditions: one cycle of 95 °C for 2 min, followed by 35 cycles of 92 °C for 30 s, 55 °C for 30 s, 72 °C for 30 s, final extension at 72 °C for 6 min43.
Diagnostic PCR
We diagnostically screened all samples (all 118 adult female, 8 egg, and 2 leg samples) for the two bacterial genera previously identified from L. geometricus: Wolbachia (Class Alphaproteobacteria, Order Rickettsiales, Family Anaplasmataceae) and Rhabdochlamydia (Class Chlamydiia, Order Chlamydiales, Family Rhabdochlamydiaceae20,27). For Wolbachia, we followed previously published protocols44, using primers specific to the Wolbachia surface protein (wsp) gene (Forward primer, wspF1: 5′-GTCCAATARSTGATGARGAAAC-3′, reverse primer wspR1: 5′-CYGCACCAAYAGYRCTRTAAA-3′, cycling conditions: one cycle of 94 °C for 2 min, followed by 36 cycles of 94 °C for 30 s, 59 °C for 45 s, 72 °C for 1 min 30 s, final extension at 70 °C for 10 min44. For Rhabdochlamydia, we designed new primers in Primer345 to amplify a ~ 540 bp segment of 16S: Rhabdo_108F 5′-ACACTGCCCAAACTCCTACG-3′ and Rhabdo_647R 5′-TTAGCTWCGACACAGCCAGG-3′. All reactions were run in 10 µl volume; the Rhabdochlamydia reactions included 3 μl purified water, 1 μl 10× Buffer (New England Biolabs), 1.2 μl 10 mM dNTPs, 1.5 μl 25 mM MgCl2, 0.6 μl each of forward and reverse primers at 5 μM, and 0.1 μl of 5 U New England Biolabs Taq Polymerase. PCR reactions received one cycle of 94 °C for 2 min, followed by 25 cycles of 95 °C for 15 s, 56 °C at 15 s, 68 °C for 45 s. We electrophoresed and visualized the products on 1% agarose gels stained with Gel Red (Biotium) alongside known positive and negative (reagents-only) controls. Samples with initial negative diagnoses were retested before being categorized as uninfected. For a subset of the samples with positive evidence of infection, we repeated the PCR at a 20 µl volume and purified the PCR product with either GenCatch PCR Cleanup or Gel Extraction Kits (Epoch Life Sciences, Missouri City, TX) according to manufacturer’s instructions. Products were then submitted for Sanger sequencing (Eurofins, Louisville, KY). Resulting sequences were compared to the NCBI nucleotide database using the megablast algorithm, and specimens returning a 97% or higher match to the expected bacterial genus were scored as positive. For each strain of Wolbachia, we sequenced 5 MLST genes (coxA, fbpA, ftsZ, gatB and hcpA) and the Wolbachia surface protein (wsp) according to Baldo et al.44.
For Rhabdochlamydia, we ran phylogenetic analyses to place the L. geometricus strains, using a set of accessions across Chlamydiia with Oligosphaera ethanolica as an outgroup. For each analysis, multiple alignments were assembled using the MAFFT server (v. 7; https://mafft.cbrc.jp/alignment/server/46) using the Q-INS-I alignment method that takes secondary structure into account. Maximum likelihood phylogenetic analyses were conducted on 1576-character aligned datasets using Garli (v. 2.0147). We applied the most complex model available (GTR + I + G48) as per recommendations of Huelsenbock and Rannala49 for likelihood-based analyses. We conducted a 100-replicate ML search for the tree of highest log-likelihood and a 500-replicate ML bootstrap analysis50 with two search replicates per individual bootstrap replicate. All analyses used the default settings.
We used the same approach to generate a Wolbachia phylogeny. We used a concatenated data set containing 5 MLST genes (coxA, fbpA, ftsZ, gatB and hcpA; total of 2079 characters) with 38 Wolbachia strains pulled from the Wolbachia PubMLST website (https://pubmlst.org/organisms/wolbachia-spp51). Because rooting Wolbachia trees is challenging52, and our objective was only placement of our new strains within established Wolbachia supergroups, we chose to simply root the tree within Supergroup A.
Individual specimens were scored for the presence of Rhabdochlamydia and Wolbachia based on the combination of diagnostic, high-throughput, and Sanger sequencing data. For a sample to be scored positive, a positive diagnostic PCR needed to be corroborated by either high-throughput or Sanger sequencing validation. For a sample to be scored negative, consistent negative diagnostic PCRs needed to be accompanied by positive validation of spider COI and/or other bacterial taxa.
Statistical methods
All analyses were conducted in R version 4.0.253. To compare the prevalence of the dominant strains of Rhabdochlamydia and Wolbachia across South Africa, Israel, and the United States, we used a general linear model (“lme4” package54) with a binomial link function, with Rhabdochlamydia1 or Wolbachia1 presence or absence in an individual as the response variable, and country as the predictor. Maps showing collection localities in South Africa, Israel, and the United States were generated using the R package ggspatial55.
Results
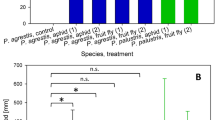
Compared to most microbiomes in arthropods, L. geometricus spiders have a depauperate microbial fauna. Of 103 adult female spiders that produced sufficient read depth (mean ± SE of 33,844 ± 2026 sequences per sample), all were dominated by one to three bacterial strains that accounted for greater than 90% of the reads (Fig. 1). In 64 samples, a single strain accounted for greater than 99% of reads. In most samples, the most prevalent bacterial ASV was Rhabdochlamydia (83/103 samples) although a few samples each were dominated by ASVs corresponding to Wolbachia (6 samples), Enterobacteriaceae (10 samples), Providencia (2 samples), Wohlfahrtimonas (1 sample) and a bacteria that could not be placed by the Greengenes reference database, but which our analyses (see below) place within the Chlamydiales (Chlamydiales1, 3 samples, Fig. 1).
High throughput analysis of bacterial associates in Latrodectus geometricus. Proportional distribution of 16S sequencing reads from L. geometricus adult females collected from South Africa (a), the United States (b), and Israel (c). All bacterial strain types that exceeded 1% of reads in any sample are depicted. All remaining strains are collected within the “other” category. See Supplementary table 1 for taxonomic identification and raw read numbers for all amplicon sequence variants.
Most samples had at least some Rhabdochlamydia representation. Nine samples from several locations in South Africa and the United States had negligible representation (< 0.1% of reads) of Rhabdochlamydia. The number of Rhabdochlamydia reads in the latter samples ranged from 0 (out of 4222 reads) to 359 (out of 37,618 reads), and most fell below the number of Rhabdochlamydia reads seen in blanks (9–81 reads). Two samples were diagnostically positive for Rhabdochlamydia despite low numbers of reads, and were additionally validated by Sanger sequencing of the diagnostic product, thus were counted as Rhabdochlamydia positive in the final dataset. In the remaining seven specimens, the low number of proportional reads and the diagnostic absence supports the genuine absence of Rhabdochlamydia. Of the additional 15 samples that were excluded from high throughput analysis due to poor initial amplification or insufficient read depth, six were validated to have Rhabdochlamydia and nine did not.
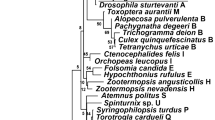
To gain insight into the occurrence of strains of the major endosymbionts found, we used Sanger sequencing data to distinguish among strains of the same symbiont clade. Most detected Rhabdochlamydia strains were identical (GenBank Accession #OP598824). Two variant strains were detected, each in one individual. The variant strain from a Modimolle, South Africa specimen (#OP598825) was 99.8% similar to the dominant strain, differing at only 1/480 bases of 16S. The variant strain from Eilat, Israel (#OP598826) was 98.8% similar, differing at 6/480 bases of 16S. Phylogenetically, all three strains were clustered together within the genus Rhabdochlamydia and family Rhabdochlamydiaceae (Fig. 2).
Phylogenetic placement of Chlamydial bacterial associates of Latrodectus geometricus. Tree of highest log likelihood from 500 maximum likelihood searches of a 35 OTU 16S data set containing 1576 characters conducted with Garli (v. 2.01) using the default settings. Taxa in bold are the new strains from L. geometricus (labeled Rhabdochlamydia1, 2, 3 and Chlamydiales1). Numbers above the nodes are bootstrap values above 50 (500 bootstrap replicates with 2 searches per replicate).
Wolbachia was much less common than Rhabdochlamydia, found in 14% (17/118) of individuals, but represented in spiders collected from all three regions. We were able to sequence all MLST genes and wsp for all three strains of Wolbachia (accession numbers OP612314-OP612330), except gatB in L. geometricus Wolbachia3. The most widespread and characteristic strain of Wolbachia in L. geometricus, Wolbachia1, was present in 13/118 specimens (11%), and phylogenetic analysis placed the strain in Wolbachia Supergroup F (Supplementary Fig. 1). In contrast, L. geometricus Wolbachia2, which was found in four specimens across three localities in South Africa, belongs to a different Wolbachia clade, Supergroup B. A third Wolbachia strain, L. geometricus Wolbachia3, which was found in a single sample that had not been included in high throughput sequencing but was validated with diagnostic PCR and subsequent sequencing, was placed in Supergroup A.
Only 16 other ASV, besides Rhabdochlamydia and Wolbachia, were ever found at > 1% prevalence in any sample, and the majority of these (nine) were each found in single specimens. Enterobacteriaceae1 represented a substantial proportion of reads in 12 individuals across several locations in South Africa and the United States, and was the dominant ASV in eight individuals. When blasted against the NCBI database, a 1359bp segment of 16S from this bacterium (#OP598828) was not closely aligned to any other accessions, bearing greatest resemblance to aphid secondary symbionts (e.g., EU348326 at 96.8%) or Gilliamella, a specialized honeybee gut symbiont (e.g., CP048265 at 95.84%). Enterobacteriaceae1 was absent from Israel, although a different Enterobacteriaceae ASV was detected from two individuals collected from one location in Israel. Two other gammaproteobacteria ASVs, Providencia and Wohlfahrtiimonas, were present in two and one specimens, respectively. One bacterial strain, which was found in four individuals across two locations in the southeast U.S., was not able to be placed against the Greengenes database in the QIIME2 pipeline, but a 498 bp segment of 16S aligns most closely with other Chlamydiales in GenBank (e.g. FJ976094 at 87.2%). Our chlamydial phylogeny (Fig. 2), also supports placement within this order, hence we have designated it Chlamydiales1. Other bacterial ASV were only found at a low percentage of reads across spiders (two Acinetobacter ASV, two Spiroplasma ASV, and one each of Entomoplasmatales, Sporosarcina, Bacillus, Enterococcus, and Lactococcus).
Comparing across the three countries, a higher proportion of spiders collected in Israel were infected with the dominant strain of Rhabdochlamydia, Rhabdochlamydia1, than spiders from South Africa (GLM, z = − 2.128, p = 0.033) or the U.S. (GLM, z = − 2.538, p = 0.011). We found no differences in prevalence of the dominant Wolbachia strain, Wolbachia1, across countries (GLM, US-Israel, z = − 0.689, p = 0.491; US-South Africa, z = − 1.268, p = 0.205; Israel-South Africa, z = − 0.669, p = 0.504). Using diagnostic PCR screening, we found evidence for Rhabdochlamydia in 100% (8/8) of L. geometricus eggs tested from South Africa and Israel. In contrast, only two out of eight egg sacs showed signal of Wolbachia, both from Tel Aviv, consistent with the proportional infection rate in adults from the source populations.
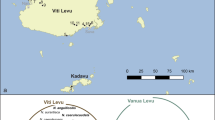
Wolbachia prevalence was too low for formal spatial analysis, but visually appeared to have some level of clustering (Fig. 3). In South Africa, both Wolbachia1 and Wolbachia2 were found in northeastern populations (Johannesburg, Pretoria, and Modimolle) but were not detected elsewhere in the country. Likewise, in Israel, Wolbachia1 was present in central and northern populations (Tel Aviv and Haifa), but was not detected in the southern Negev populations (Beer Sheva, Yeruham, Sede Boqer, Eilat). Among the four U.S. populations, Wolbachia1 was found in spiders collected from Florida and Texas, Wolbachia3 was in South Carolina, but no Wolbachia was detected in spiders from California, the most recently detected invasive population.
Proportion of adult female L. geometricus infected with Rhabdochlamydia1 and/or Wolbachia1 detected through PCR screening across 17 localities in (a) South Africa, (b) Israel, and (c) the United States. Blue represents individuals infected with just Rhabdochlamydia1, purple represents individuals infected with both Rhabdochlamydia1 and Wolbachia1, red represents individuals infected with just Wolbachia1, and white represents individuals infected with neither Wolbachia1 nor Rhabdochlamydia1. Size of pie charts corresponds to the number of individual spiders screened from each site (range = one specimen from Eilat, Israel to 10 specimens from Edisto Island, SC, USA, see Supplementary table 1 for sample sizes and collection localities).
Discussion
Latrodectus geometricus spiders have maintained a characteristic microbiome throughout their global spread. We identified one predominant endosymbiont, Rhabdochlamydia1 in almost all spiders (86%), and represented in all collection locations. We also found a characteristic Supergroup F Wolbachia (Wolbachia1) represented in all countries, albeit in fewer individuals (11% of spiders). We detected both Rhabdochlamydia1 and Wolbachia1 in L. geometricus eggs, indicating that both are vertically transmitted endosymbionts.
The widespread presence of Rhabdochlamydia suggests that it might be important functionally for the host. In other arthropods, endosymbionts found at consistently high frequency across wide geographic ranges have often subsequently been found to have important fitness or reproductive consequences for their hosts56,57. Little is known about the functional role of Rhabdochlamydia in arthropods. It was described from a variety of mostly non-insect arthropods and was generally found at low prevalence in the tested populations23,24,26. In a terrestrial isopod, Rhabdochlamydia had pathogenic effects26. The high prevalence (86%) and vertical transmission of Rhabdochlamydia in L. geometricus argue against a strongly pathogenic role for this bacterial strain within our system. Genomic analysis of Rhabdochlamydia found in other arthropod hosts, an isopod and a tick, found pathways for polyamine synthesis22, which are relevant for virulence and stress responses, suggesting that some strains of this bacteria are potentially beneficial in their host.
We also detected Rhabdochlamydia in L. geometricus legs, consistent with the work of Dunaj et al.20, which indicated that the bacteria is found throughout the body and not just restricted to reproductive tissue. Dunaj et al.20 also found that the bacterial community of L. geometricus was dominated by Rhabdochlamydia, lacking the microbial diversity of the other spider species they examined, and speculated that this result may have been an artifact of laboratory-reared, inbred L. geometricus spiders. Our field collected spiders from locations around the world suggest that their result was not an artifact, but a genuine representation of a characteristic and depauperate bacterial community in L. geometricus. Vertically transmitted bacterial symbionts often dominate the sampled microbiomes of their hosts, overwhelming the signal from more casual bacterial associates11,25,35.
Importantly, maternal transmission of Rhabdochlamydia suggests the possibility of reproductive manipulation of host by symbiont. Reproductive manipulation is extremely common in vertically transmitted symbionts, and the list of bacteria that have been demonstrated to induce such manipulations is rapidly expanding11,58. Rhabdochlamydia has not yet been tested for host reproductive manipulation. The widespread prevalence and vertical transmission of Rhabdochlamydia in L. geometricus would make this system an excellent prospect for such investigations.
Latrodectus geometricus was host to several strains of Wolbachia, a bacterial clade well known for reproductive manipulation. Wolbachia is common in spiders, but most strains belong to Supergroup A or B, as is the case in insects30. In contrast, the dominant Wolbachia strain in L. geometricus belongs to Supergroup F, which has rarely been reported for spiders. Supergroup F has been found sporadically in arthropods, including South African scorpions59, termites60, quill mites61, and nematodes62. Preliminary work on L. geometricus suggested that Wolbachia might induce mild CI in this species63, but the strain of Wolbachia was not characterized, and additional experiments will be necessary to fully validate CI in this system.
Although symbiont communities were largely similar across our sampled regions, we did find some subtle differences between the likely native and invasive ranges. Rhabdochlamydia was found at highest prevalence in Israel compared to populations in the U.S. and South Africa. Multiple strains of Rhabdochlamydia, Wolbachia, and the Enterobacteriaceae were found in South Africa, the putative native population. The dominant strain of Enterobacteriaceae was found in South Africa and the U.S., but absent in Israel, the newest invasive region that we sampled. From a previous study, Wolbachia prevalence in L. geometricus in the U.S. was highest near the initial site of introduction in Florida27. In comparison, we found lower Wolbachia prevalence in other locations in the southeastern and central U.S, and absence in spiders from California, the most recently established population. Similarly, in Israel, Wolbachia was absent in recently established populations in southern Israel. These patterns are consistent with the loss of endosymbionts during the invasion process, but more localities, specimens, and more knowledge of the invasion route is needed. Climatic differences such as hotter, dryer conditions in the Negev Desert in southern Israel could also contribute to reduction of Wolbachia64, although deeper sampling effort would be needed to assess whether Wolbachia is entirely absent from these locations.
Further work will test the functional role and fitness effects of endosymbiont presence in L. geometricus, as well as compare patterns of host-endosymbiont diversity during invasive spread. Invasive L. geometricus are highly dispersive18, and are less susceptible to parasitism by parasitoids compared to native widow species in the invasive range19. It would be valuable to test whether these advantages and others during invasion are related to interactions with endosymbionts. In particular, the dominance and high prevalence of Rhabdochlamydia across global populations of L. geometricus suggests an important role of this endosymbiont. Characterizing potentially important and widespread endosymbionts is a step towards understanding their relevance to ecological interactions and responses to rapid environmental changes.
Data availability
The datasets generated and/or analyzed during the current study are available via NCBI SRA, Bioproject PRJNA1068539: https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1068539.
References
Chalkowski, K., Lepczyk, C. A. & Zohdy, S. Parasite ecology of invasive species: Conceptual framework and new hypotheses. Trends Parasitol. 34, 655–663. https://doi.org/10.1016/j.pt.2018.05.008 (2018).
Sepúlveda, D. A., Zepeda-Paulo, F., Ramírez, C. C., Lavandero, B. & Figueroa, C. C. Diversity, frequency, and geographic distribution of facultative bacterial endosymbionts in introduced aphid pests. Insect Sci. 24(3), 511–521. https://doi.org/10.1111/1744-7917.12313 (2017).
Leonardo, T. E. & Mondor, E. B. Symbiont modifies host life-history traits that affect gene flow. Proc. R. Soc. B Biol. Sci. 273, 1079–1084. https://doi.org/10.1098/rspb.2005.3408 (2006).
Vorburger, C. & Gouskov, A. Only helpful when required: A longevity cost of harbouring defensive symbionts. J. Evol. Biol. 24, 1611–1617. https://doi.org/10.1111/j.1420-9101.2011.02292.x (2011).
Oliver, K. M. & Martinez, A. J. How resident microbes modulate ecologically-important traits of insects. Curr. Opin. Insect Sci. 4, 1–7. https://doi.org/10.1016/j.cois.2014.08.001 (2014).
Jaenike, J. Population genetics of beneficial heritable symbionts. Trends Ecol. Evol. 27, 226–232. https://doi.org/10.1016/j.tree.2011.10.005 (2012).
Rey, O. et al. Distribution of endosymbiotic reproductive manipulators reflects invasion process and not reproductive system polymorphism in the little fire ant Wasmannia auropunctata. PLoS One 8, e58467. https://doi.org/10.1371/journal.pone.0058467 (2013).
Himler, A. G. et al. Rapid spread of a bacterial symbiont in an invasive whitefly is driven by fitness benefits and female bias. Science 332, 254–256. https://doi.org/10.1126/science.1199410 (2011).
Duron, O., Hurst, G. D. D., Hornett, E. A., Josling, J. A. & Engelstädter, J. High incidence of the maternally inherited bacterium Cardinium in spiders. Mol. Ecol. 17, 1427–1437. https://doi.org/10.1111/j.1365-294X.2008.03689.x (2008).
Łukasik, P. et al. Unrelated facultative endosymbionts protect aphids against a fungal pathogen. Ecol. Lett. 16, 214–218. https://doi.org/10.1111/ele.12031 (2013).
Rosenwald, L. C., Sitvarin, M. I. & White, J. A. Endosymbiotic Rickettsiella causes cytoplasmic incompatibility in a spider host. Proc. Biol. Sci. 287, 20201107. https://doi.org/10.1098/rspb.2020.1107 (2020).
Dlugosch, K. M. & Parker, I. M. Founding events in species invasions: Genetic variation, adaptive evolution, and the role of multiple introductions. Mol. Ecol. 17, 431–449. https://doi.org/10.1111/j.1365-294X.2007.03538.x (2008).
Shoemaker, D. D., Ross, K. G., Keller, L., Vargo, E. L. & Werren, J. H. Wolbachia infections in native and introduced populations of fire ants (Solenopsis spp.). Insect Mol. Biol. 9, 661–673. https://doi.org/10.1046/j.1365-2583.2000.00233.x (2000).
Bertelsmeier, C. & Keller, L. Bridgehead effects and role of adaptive evolution in invasive populations. Trends Ecol. Evol. 33, 527–534. https://doi.org/10.1016/j.tree.2018.04.014 (2018).
Desneux, N. et al. Intraspecific variation in facultative symbiont infection among native and exotic pest populations: Potential implications for biological control. Biol. Control 116, 27–35. https://doi.org/10.1016/j.biocontrol.2017.06.007 (2018).
Garb, J. E., González, A. & Gillespie, R. G. The black widow spider genus Latrodectus (Araneae: Theridiidae): Phylogeny, biogeography, and invasion history. Mol. Phylogenet. Evol. 31, 1127–1142. https://doi.org/10.1016/j.ympev.2003.10.012 (2004).
Sadir, M. & Marske, K. A. Urban environments aid invasion of brown widows (Theridiidae: Latrodectus geometricus) in North America, constraining regions of overlap and mitigating potential impact on native widows. Front. Ecol. Evol. https://doi.org/10.3389/fevo.2021.757902 (2021).
Mowery, M. A., Lubin, Y. & Segoli, M. Invasive brown widow spiders disperse aerially under a broad range of environmental conditions. Ethology 128, 564–571. https://doi.org/10.1111/eth.13314 (2022).
Mowery, M. A., Arabesky, V., Lubin, Y. & Segoli, M. Differential parasitism of native and invasive widow spider egg sacs. Behav. Ecol. 33, 565–572. https://doi.org/10.1093/beheco/arac017 (2022).
Dunaj, S. J., Bettencourt, B. R., Garb, J. E. & Brucker, R. M. Spider phylosymbiosis: Divergence of widow spider species and their tissues’ microbiomes. BMC Evol. Biol. 20, 104. https://doi.org/10.1186/s12862-020-01664-x (2020).
Halter, T. et al. Ecology and evolution of chlamydial symbionts of arthropods. ISME Commun. 2, 1–11. https://doi.org/10.1038/s43705-022-00124-5 (2022).
Halter, T. et al. One to host them all: Genomics of the diverse bacterial endosymbionts of the spider Oedothorax gibbosus. Microb. Genom. 9(2), 943. https://doi.org/10.1099/mgen.0.000943 (2023).
Corsaro, D. et al. ‘Candidatus Rhabdochlamydia crassificans’, an intracellular bacterial pathogen of the cockroach Blatta orientalis (Insecta: Blattodea). Syst. Appl. Microbiol. 30, 221–228. https://doi.org/10.1016/j.syapm.2006.06.001 (2007).
Pillonel, T. et al. Sequencing the obligate intracellular Rhabdochlamydia helvetica within its tick host Ixodes ricinus to investigate their symbiotic relationship. Genome Biol. Evol. 11, 1334–1344. https://doi.org/10.1093/gbe/evz072 (2019).
Vanthournout, B. & Hendrickx, F. Endosymbiont dominated bacterial communities in a dwarf spider. PLoS One 10, e0117297. https://doi.org/10.1371/journal.pone.0117297 (2015).
Kostanjšek, R., Štrus, J., Drobne, D. & Avguštin, G. ‘Candidatus Rhabdochlamydia porcellionis’, an intracellular bacterium from the hepatopancreas of the terrestrial isopod Porcellio scaber (Crustacea: Isopoda). Int. J. Syst. Evol. Microbiol. 54, 543–549. https://doi.org/10.1099/ijs.0.02802-0 (2004).
Arrington, B. D. The prevalence and effect of Wolbachia infection on the brown widow spider (Latrodectus geometricus) (Georgia Southern University, 2014).
Zug, R. & Hammerstein, P. Still a host of hosts for Wolbachia: Analysis of recent data suggests that 40% of terrestrial arthropod species are infected. PLoS One 7, e38544. https://doi.org/10.1371/journal.pone.0038544 (2012).
Fenn, K. et al. Phylogenetic relationships of the Wolbachia of nematodes and arthropods. PLoS Pathog. 2, e94. https://doi.org/10.1371/journal.ppat.0020094 (2006).
Kaur, R. et al. Living in the endosymbiotic world of Wolbachia: A centennial review. Cell Host Microbe 29, 879–893. https://doi.org/10.1016/j.chom.2021.03.006 (2021).
Pearson, J. F. W. Latrodectus geometricus Koch in southern Florida. Science 83, 522–523 (1936).
Vincent, L. S., Vetter, R. S., Wrenn, W. J., Kempf, J. K. & Berrian, J. E. The brown widow spider Latrodectus geometricus C. L. Koch, 1841, in southern California. Pan-Pac. Entomol. 84, 344–349. https://doi.org/10.3956/2008-07.1 (2009).
Levy, G. & Amitai, P. Revision of the widow-spider genus Latrodectus (Araneae: Theridiidae) in Israel. Zool. J. Linn. Soc. 77, 39–63. https://doi.org/10.1111/j.1096-3642.1983.tb01720.x (1983).
Mowery, M. A., Lubin, Y., Harari, A., Mason, A. C. & Andrade, M. C. B. Dispersal and life history of brown widow spiders in dated invasive populations on two continents. Anim. Behav. https://doi.org/10.1016/j.anbehav.2022.02.006 (2022).
White, J. A. et al. Endosymbiotic bacteria are prevalent and diverse in agricultural spiders. Microb. Ecol. 79, 472–481. https://doi.org/10.1007/s00248-019-01411-w (2020).
Curry, M. M., Paliulis, L. V., Welch, K. D., Harwood, J. D. & White, J. A. Multiple endosymbiont infections and reproductive manipulations in a linyphiid spider population. Heredity 115, 146–152. https://doi.org/10.1038/hdy.2015.2 (2015).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120. https://doi.org/10.1128/AEM.01043-13 (2013).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. https://doi.org/10.1038/nmeth.f.303 (2010).
Bokulich, N. A. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59. https://doi.org/10.1038/nmeth.2276 (2013).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072. https://doi.org/10.1128/AEM.03006-05 (2006).
Larsson, A., Stanley, G. M., Sinha, R., Weissman, I. & Sandberg, R. Computational correction of index switching in multiplexed sequencing libraries. Nat. Methods https://doi.org/10.1038/nmeth.4666 (2018).
Hongoh, Y., Ohkuma, M. & Kudo, T. Molecular analysis of bacterial microbiota in the gut of the termite Reticulitermes speratus (Isoptera; Rhinotermitidae). FEMS Microbiol. Ecol. 44, 231–242. https://doi.org/10.1016/S0168-6496(03)00026-6 (2003).
Baldo, L. et al. Multilocus sequence typing system for the endosymbiont Wolbachia pipientis. Appl. Environ. Microbiol. 72, 7098–7110. https://doi.org/10.1007/s00284-007-9009-4 (2006).
Untergasser, A. et al. Primer3—New capabilities and interfaces. Nucleic Acids Res. 40, e115. https://doi.org/10.1093/nar/gks596 (2012).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166. https://doi.org/10.1093/bib/bbx108 (2019).
Zwickl, D. J. Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion (The University of Texas at Austin, 2006).
Rodríguez, F., Oliver, J. L., Marín, A. & Medina, J. R. The general stochastic model of nucleotide substitution. J. Theor. Biol. 142, 485–501. https://doi.org/10.1016/S0022-5193(05)80104-3 (1990).
Huelsenbeck, J. P. & Rannala, B. Frequentist properties of Bayesian posterior probabilities of phylogenetic trees under simple and complex substitution models. Syst. Biol. 53, 904–913. https://doi.org/10.1080/10635150490522629 (2004).
Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39, 783–791. https://doi.org/10.2307/2408678 (1985).
Jolley, K. A., Bray, J. E. & Maiden, M. C. J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 3, 124. https://doi.org/10.12688/wellcomeopenres.14826.1 (2018).
Rodrigues, J. et al. Wolbachia springs eternal: Symbiosis in Collembola is associated with host ecology. R. Soc. Open Sci. 10, 230288. https://doi.org/10.1098/rsos.230288 (2023).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2023).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48. https://doi.org/10.18637/jss.v067.i01 (2015).
Dunnington, D., Thorne, B. & Hernangómez, D. ggspatial: Spatial Data Framework for ggplot2 (2023). https://github.com/paleolimbot/ggspatial.
Arif, S. et al. Evidence for multiple colonisations and Wolbachia infections shaping the genetic structure of the widespread butterfly Polyommatus icarus in the British Isles. Mol. Ecol. 30, 5196–5213. https://doi.org/10.1111/mec.16126 (2021).
Cornwell, B. H. & Hernández, L. Genetic structure in the endosymbiont Breviolum ‘muscatinei’ is correlated with geographical location, environment and host species. Proc. R. Soc. B Biol. Sci. 288, 20202896. https://doi.org/10.1098/rspb.2020.2896 (2021).
Pollmann, M. et al. Highly transmissible cytoplasmic incompatibility by the extracellular insect symbiont Spiroplasma. iScience 25, 104335. https://doi.org/10.1016/j.isci.2022.104335 (2022).
Baldo, L., Prendini, L., Corthals, A. & Werren, J. H. Wolbachia are present in southern African scorpions and cluster with Supergroup F. Curr. Microbiol. 55, 367–373. https://doi.org/10.1007/s00284-007-9009-4 (2007).
Lo, N. & Evans, T. A. Phylogenetic diversity of the intracellular symbiont Wolbachia in termites. Mol. Phylogenet. Evol. 44, 461–466. https://doi.org/10.1016/j.ympev.2006.10.028 (2007).
Glowska, E., Dragun-Damian, A., Dabert, M. & Gerth, M. New Wolbachia supergroups detected in quill mites (Acari: Syringophilidae). Infect. Genet. Evol. 30, 140–146. https://doi.org/10.1016/j.meegid.2014.12.019 (2015).
Casiraghi, M. et al. Phylogeny of Wolbachia pipientis based on gltA, groEL and ftsZ gene sequences: Clustering of arthropod and nematode symbionts in the F supergroup, and evidence for further diversity in the Wolbachia tree. Microbiology 151, 4015–4022. https://doi.org/10.1099/mic.0.28313-0 (2005).
Knight, E. Characterizing the complex relationship between the brown widow spider and Its bacterial endosymbiont, Wolbachia. Electronic Theses and Dissertations. https://digitalcommons.georgiasouthern.edu/etd/1825 (2018).
Charlesworth, J., Weinert, L. A., Araujo, E. V. & Welch, J. J. Wolbachia, Cardinium and climate: An analysis of global data. Biol. Lett. 15, 20190273. https://doi.org/10.1098/rsbl.2019.0273 (2019).
Acknowledgements
We thank Ofir Altstein, Ishai Hoffmann, Cayley Buckner, Madison Heisey, Catherine Scott, Nishant Singh, Alyssa Fuller, Astri Leroy, Annari van der Merwe, Fiona Hellmann, Joh Henschel, Theresa Henschel, and Colin Ralston for assistance collecting spiders. This work was supported by a Zuckerman STEM Leadership Postdoctoral Fellowship to MAM. This material is based upon work supported by the National Science Foundation under Grant No. 1953223 and the National Institute of Food and Agriculture, U.S. Department of Agriculture (Hatch No. 1020740).
Author information
Authors and Affiliations
Contributions
MAM, YL, MS, and JAW conceived of and designed the study. MAM, YL, MS, TK, and RL collected specimens. MAM, LCR, and JAW collected the data, and MAM, JAW, and EC analyzed the data. MAM wrote the initial draft of the manuscript. All authors provided editorial comments and made contributions to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mowery, M.A., Rosenwald, L.C., Chapman, E. et al. Endosymbiont diversity across native and invasive brown widow spider populations. Sci Rep 14, 8556 (2024). https://doi.org/10.1038/s41598-024-58723-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-58723-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.