Abstract
Combining information from the tumor microenvironment (TME) with PAM50 Risk of Recurrence (ROR) score could improve breast cancer prognostication. Caveolin-1 (CAV1) is a marker of an active TME. CAV1 is a membrane protein involved in cell signaling, extracellular matrix organization, and tumor-stroma interactions. We sought to investigate CAV1 gene expression in relation to PAM50 subtypes, ROR score, and their joint prognostic impact. CAV1 expression was compared between PAM50 subtypes and ROR categories in two cohorts (SCAN-B, n = 5326 and METABRIC, n = 1980). CAV1 expression was assessed in relation to clinical outcomes using Cox regression and adjusted for clinicopathological predictors. Effect modifications between CAV1 expression and ROR categories on clinical outcome were investigated using multiplicative and additive two-way interaction analyses. Differential gene expression and gene set enrichment analyses were applied to compare high and low expressing CAV1 tumors. All samples expressed CAV1 with the highest expression in the Normal-like subtype. Gene modules consistent with epithelial-mesenchymal transition (EMT), hypoxia, and stromal activation were associated with high CAV1 expression. CAV1 expression was inversely associated with ROR category. Interactions between CAV1 expression and ROR categories were observed in both cohorts. High expressing CAV1 tumors conferred worse prognosis only within the group classified as ROR high. ROR gave markedly different prognostic information depending on the underlying CAV1 expression. CAV1, a potential mediator between the malignant cells and TME, could be a useful biomarker that enhances and further refines PAM50 ROR risk stratification in patients with ROR high tumors and a potential therapeutic target.
Similar content being viewed by others
Introduction
Breast cancer remains a clinical challenge1. Despite improvements in care, a significant proportion of breast cancer patients relapse2,3. During the past decade, molecular profiling of tumors has been implemented in the clinical setting to improve prognostication and treatment selection3,4,5,6. One example is the PAM50 Risk of Recurrence (ROR) score classification is now clinically used worldwide for a subgroup of breast cancer. The PAM50 ROR score is validated for postmenopausal breast cancer patients with ER+ /HER2− tumors receiving five years of endocrine therapy. The ROR score provides prognostic information and can be used to select patients for adjuvant chemotherapy4,7,8. However, further prognostication and treatment prediction refinement is still needed3. Additional information may be gained by looking beyond the malignant cells of the tumor, which most molecular risk scores are based on, and incorporating information from the tumor microenvironment (TME).
The TME has gained increasing attention for its role in breast cancer development and treatment response9,10. It has become increasingly clear that the tumor depends on its surroundings to be able to grow, survive, and metastasize9,10. Currently, there are few prognostic markers derived from the TME and these markers are mostly related to immune cells, such as tumor infiltrating lymphocytes9,10,11. Stromal cells, in particular cancer-associated fibroblasts (CAFs), constitute a significant part of the TME and play a key role in modulating various processes such as epithelial-mesenchymal transition (EMT), hypoxia, and angiogenesis, all important for the development of metastasis12,13.
Challenges remain in translating findings related to the TME into relevant biomarkers useful in clinical practice, since the TME is highly heterogenous and comprises several distinct cell types9,10,14. Depending on the composition of the TME and its interplay with malignant cells, the TME can either be tumor promoting or suppressing9,10,12. An interesting biomarker of an active TME is Caveolin-1 (CAV1). CAV1 is a master regulator of cell signaling and vesicular transport and is located in cholesterol-rich plasma membrane raft domains, known as caveolae15,16. CAV1 modulates key pathogenic processes involving the TME, including drug internalization, tumor-stroma interactions, hypoxia response, cellular metabolism, inflammation, and EMT15,16,17.
Furthermore, studies have reported that CAV1 protein expression in stromal cells may serve as a prognostic biomarker in breast cancer16,18,19,20,21. However, the prognostic impact of CAV1 is both context and localization dependent as previously reported18,19. It is unclear how the interplay between CAV1 and clinically used molecular risk scores, such as PAM50 ROR, relates to prognosis. Herein, we investigate the role of CAV1 gene expression in relation to PAM50 subtypes, ROR scores, and their joint impact on clinical outcome in two large breast cancer cohorts.
Results
Relationship with clinicopathological variables
For SCAN-B, 5326 of 7743 patients were assessed in the analysis, Fig. 1. For METABRIC the entire dataset of 1980 patients was used. All samples in both SCAN-B and METABRIC expressed CAV1. The distribution of CAV1 across PAM50 subtypes was similar for both cohorts, with the CAV1 expression being highest in Normal-like and followed by Luminal A subtype (both Ps < 0.001), Fig. 2A,B. Likewise, the correlations between CAV1 expression and the eight gene modules were similar in both cohorts, showing strong correlations between CAV1 expression and the Lipid and Stroma modules and ROR category dependent correlation with the Steroid and Immune response modules, Supplementary file 1: Supplementary Fig. 1. Notably, there was an inverse correlation with ROR category in both cohorts (both r < − 0.34 and P < 0.001), Fig. 2C,D, Supplementary file 1: Supplementary Fig. 1. Even after adjusting for other predictors of PAM50 ROR category, the highest expression of CAV1 (T3) was strongly negatively associated with ROR high in both SCAN-B (adjusted OR 0.26 95% CI 0.19–0.35, P < 0.001) and METABRIC (adjusted OR 0.32 95% CI 0.21–0.47, P < 0.001), Supplementary file 1: Supplementary Fig. 2. CAV1 gene expression was also negatively correlated with most genes that are a part of the PAM50 ROR, except for KRT14, KRT15, KRT5, and SFRP1, Supplementary file 1: Supplementary Fig. 2. CAV1 tertiles in relation to clinicopathological factors are presented in Table 1 for SCAN-B and METABRIC.
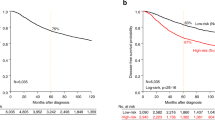
CAV1 expression by PAM50 and ROR category. CAV1 expression (continuous) by PAM50 molecular subtype in SCAN-B (A) and METABRIC (B). CAV1 expression (continuous) by PAM50 ROR category in SCAN-B (C) and METABRIC (D). Kaplan–Meier estimates of PAM50 ROR category among all patients in relation to distant metastasis-free interval in SCAN-B (E) and breast cancer-specific survival in METABRIC (F). Kaplan–Meier estimates of combined ROR category and CAV1 expression (in tertiles) among all patients in relation to distant metastasis-free interval in SCAN-B (G) and breast cancer-specific survival in METABRIC (H). The number of patients is indicated at each time-point.
Survival analysis
In SCAN-B, the median follow-up for the 4158 patients still at risk was 5.45 (IQR 5.07–8.15) years. Follow-up was restricted to ten years in METABRIC and all events after ten years were censored. This was done for two reasons; to make METABRIC more comparable to SCAN-B and because the PAM50 ROR score was developed to predict the risk of distant metastasis within 10 years22. The median follow-up for the 1089 patients still at risk in METABRIC was 10.0 years (IQR 10.0–10.0). The hazards were proportional for the tertiles for all endpoints.
In the univariable survival analyses of the complete cohorts, patients with ROR high had an increased risk of distant metastasis and breast cancer-specific survival compared to ROR low as expected, Fig. 2E,F. Moreover, the addition of CAV1 expression further stratified the distant metastasis-risk and breast cancer-specific survival in the univariable models Fig. 2G,H. The highest expression of CAV1 (T3) was associated with lower risk of recurrence, HR 0.74 (95% CI 0.60–0.92) in SCAN-B and HR 0.66 (95% CI 0.54–0.80) in METABRIC; lower risk of distant metastasis, HR 0.65 (95% CI 0.51–0.84) in SCAN-B and HR 0.61 (95% CI 0.50–0.76) in METABRIC; and lower risk of death, HR 0.68 (95% CI 0.58–0.80) in SCAN-B and HR 0.70 (95% CI 0.59–0.84) in METABRIC, Fig. 3. The highest expression of CAV1 (T3) also conferred a lower risk of breast cancer-related death HR 0.70 (95% CI 0.57–0.87) in METABRIC, Fig. 3.
Univariable survival analyses of CAV1 expression. Kaplan–Meier estimates of CAV1 expression (in tertiles) among all patients in relation to recurrence-free interval in SCAN-B (A) and METABRIC (B), distant metastasis-free interval in SCAN-B (C) and METABRIC (D), overall survival in SCAN-B (E) and METABRIC (F), and breast cancer-specific survival (G). The number of patients is indicated at each time-point.
In the multivariable analyses, the highest expression of CAV1 (T3) in SCAN-B instead conferred an increased risk of recurrence and distant metastasis but not death, Fig. 2. and Supplementary file 1: Supplementary Fig. 3. CAV1 tertiles were therefore adjusted for each variable used in the multivariable model, one at a time to see which variable affected the hazard ratio the most, which was the ROR category. Subsequently, interaction analyses were performed between ROR category and CAV1 tertiles on RFI and DMFI, revealing significant additive interactions and effect modifications of both ROR category on CAV1 and vice versa, Table 2. In SCAN-B, when stratifying by ROR category, the highest expression of CAV1 (T3) conferred increased risk of recurrence adjusted HR 1.57 (95% CI 1.10–2.24) and distant metastasis adjusted HR 1.60 (95% CI 1.08–2.37) only in patients with tumors classified as ROR High but not in ROR Low/Intermediate tumors, Supplementary file 1: Supplementary Fig. 4 and 5.
The distribution of ROR categories in the two cohorts differed with a larger proportion of tumors classified as ROR High in METABRIC than in SCAN-B. Considering the interactions between ROR categories and CAV1 expression in SCAN-B, interaction analyses between ROR categories and CAV1 tertiles for all four endpoints were performed to investigate if potential effect modifications previously seen in SCAN-B was the underlying reason for the discrepant findings on prognosis in METABRIC. (In METABRIC, associations between CAV1 tertiles and either one of the endpoints were not statistically significant in the multivariable analysis, Supplementary file 1: Supplementary Fig. 3.) Subsequently, it became clear that there were significant multiplicate and additive interactions between CAV1 T3 and ROR category on DMFI, OS, and BCSS in METABRIC, Table 2 and Supplementary file 2: Supplementary Table 1, 2, and 3. Similarly, there were effect modifications of both ROR category on CAV1 and vice versa concerning prognosis, Table 2 and Supplementary file 1: Supplementary Table 1, 2, and 3. Interestingly, in both SCAN-B and METABRIC, CAV1 tertiles could identify tumors where the predictive potential of ROR was the highest, Table 2 and Supplementary file 1: Supplementary Table 1, 2, and 3. Similar to DMFI in SCAN-B, when stratifying by ROR category in METABRIC, the highest expression of CAV1 (T3) conferred borderline increased risk of breast cancer-specific death adjusted HR 1.24 (95% CI 0.95–1.62) only in patients with tumors classified as ROR High but not in ROR Low/Intermediate tumors, Table 2, Fig. 2, Supplementary file 1: Supplementary Fig. 6 and 7. The additive interaction was present in both cohorts but was stronger in SCAN-B RERI 1.18 (95% CI 0.10–2.26, P = 0.016) than in METABRIC RERI 0.54 (95% CI 0.20–0.89, P = 0.001) Table 2. For METABRIC, the additive interaction was even stronger when breast cancer-specific survival was used as endpoint RERI 0.77 (95% CI 0.43–1.12, P < 0.001), Fig. 2H. Furthermore, CAV1 tertiles could also delineate in which group the ROR category was prognostic. In the CAV1 T1 tumors, the ROR category did not predict risk of distant metastasis, Supplementary file 1: Supplementary Fig. 8. In CAV1 T2 tumors, the ROR category predicted distant metastasis risk in SCAN-B but not METABRIC Supplementary file 1: Supplementary Fig. 9. For the CAV1 T3 tumors, ROR category was strongly associated with distant metastasis risk in both SCAN-B and METABRIC Supplementary file 1: Supplementary Fig. 10.
DGE and GSEA analysis for CAV1 Tertile 3 vs Tertile 1
To elucidate potential biological explanations behind the differential impact of CAV1 according to ROR category, DGE analyses were performed separately in ROR categories for tumors with the highest (T3) versus the lowest (T1) CAV1 expression.
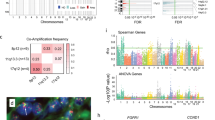
In ROR high tumors, a total of 223 genes were found to be upregulated in high expressing (T3) vs low expressing (T1) CAV1 tumors, and no genes were downregulated, Supplementary file 2: Supplementary Table 4. Notably, several other genes coding for proteins involved in caveolae formation, e.g. CAV2 and CAVIN2 were higher expressed in ROR high/CAV1 high tumors, supporting a potential association with caveolae abundance. In ROR Low/Intermediate tumors, 450 genes were upregulated, and 18 genes were downregulated in high expressing (T3) vs low expressing (T1) CAV1 tumors, Supplementary file 2: Supplementary Table 5. In both ROR categories, genes related to stromal activation, EMT, CAFs and adipogenesis (SOX10, STAC2, FGF2, PTGFR, IGF1, IGF2, GDF10, ADAM33, CD36, PLIN4, PLIN1, MME, PENK, among others) were upregulated, Supplementary file 2: Supplementary Table 4 and 5. Only in ROR low/intermediate tumors were some potential tumor suppressor genes down-regulated (EEF1A2, GRM4, ROBO2 CHGB, CEACAM5, among others), Supplementary file 2: Supplementary Table 5.
Significantly enriched gene sets in high expressing (T3) CAV1 tumors in both ROR categories included EMT, TGF-β signaling, fatty acid metabolism, hypoxia, myogenesis, angiogenesis, xenobiotic metabolism among others, Fig. 4 and Supplementary file 2: Supplementary Tables 6 and 7. In low expressing (T1) CAV1 tumors regardless of ROR category, the MYC targets gene set was enriched, Fig. 4 and Supplementary file 2: Supplementary Tables 6 and 7. The main differences in gene set enrichment between high expressing (T3) CAV1 tumors and low expressing (T1) CAV1 tumors were related to immune response. Among high expressing (T3) CAV1 tumors, interferon-α response and complement hallmarks were enriched only in ROR high, while interferon-γ response hallmark were enriched only in ROR low/intermediate, Fig. 4 and Supplementary file 2: Supplementary Tables 6 and 7. Similar patterns were seen regarding GO terms, Supplementary file 2: Supplementary Tables 8 and 9.
Molecular analyses of CAV1 expression. Volcano plot showing significant up- and downregulated genes (red) in high expressing (T3) in relation to low expressing (T1) CAV1 separately by ROR High category (A) and ROR Low/Intermediate category (B). Gene Set Enrichment Analysis (GO categories) of genes ranked by fold change (log2FC) and p-value < 0.05, up- and downregulated genes (red) in high expressing (T3) in relation to low expressing (T1) CAV1 tumors separately by ROR High category (C) and ROR Low/Intermediate category (D). Categories found enriched in both subgroup analyses are indicated by red text. Dot plot showing activated and suppressed Hallmark Signatures in high expressing (T3) in relation to low expressing (T1) CAV1 tumors separately by ROR High category (E) and ROR Low/Intermediate category (F).
Discussion
Herein, we report that high CAV1 gene expression conferred an especially poor prognosis in patients whose tumors were classified as ROR high. In addition, ROR gave markedly different prognostic information depending on the underlying CAV1 expression, even after taking PAM50 subtype, other clinical predictors, and treatments into account. To our knowledge, this is the first study to examine the CAV1 mRNA gene expression in relation to molecular subtypes and prognosis in large breast cancer cohorts.
Moreover, CAV1 expression was associated with extracellular matrix remodeling, EMT myogenesis, hypoxia, angiogenesis, and stromal activation in both ROR High and Low/Intermediate classified tumors, as corroborated by both GSEA results and their correlations with the stromal gene module. It is known that CAV1 can remodel the extra cellular matrix through activation of stromal cells, elongating and facilitating invasion and metastasis23. Functionally, alterations in caveolae in stromal cells of the TME promote paracrine tumor growth via TGFβ, which activates EMT and myofibroblast differentiation, favoring tumor growth and metastasis24,25. EMT and myogenesis are markers of increased cell motility and loss of adhesion, both required for metastasis26.
Furthermore, CAV1 is linked to angiogenesis, endothelial permeability, and vascular endothelial growth factor (VEGF) response, which is required for tumor survival and the ability to enter the circulation. However, the exact role of CAV1 is unclear27. Hypoxia and angiogenesis are interlinked, and CAV1 is a direct transcriptional target of hypoxia-inducible factors 1α and 2α (HIF1α and 2α) that lead to increased dimerization and phosphorylation of the epidermal growth factor receptor (EGFR) conferring enhancing proliferative, migratory, and invasive capacities of malignant cells28. Further, hypoxia induces metabolic reprogramming in the tumor and CAV1 alterations confer a shift from mitochondrial respiration to tumor promoting aerobic glycolysis through attenuation of MYC expression29, corroborated by our data as seen in the downregulation of MYC response.
So far, the role of CAV1 in the immunomodulatory properties of the TME remains unexplored and further studies are warranted. The types of immune signals enriched in high expressing CAV1 tumors were dependent on ROR category and activation of immune response appeared higher in ROR low/intermediate tumors. The stromal microenvironment has immunomodulatory functions and can inhibit immune cells and decrease their efficacy in targeting and killing malignant cells, through regulation of extravasation and local immune cell replication9,10,11.
Our findings and existing literature suggest that CAV1 promotes metastasis and relapse through several critical pathways regardless of genomic risk classification. CAV1 can be considered as an essential protein that regulates paracrine signaling and the interplay between the malignant cells and TME. However, CAV1 expression only yielded additional prognostic information in tumors considered ROR high. A potential explanation for this finding might be that malignant cells that already acquired the intrinsic potential to metastasize still need an active and tumor promoting environment to do so. It might explain why high CAV1 expression in tumors identified patients for whom the ROR score provided most prognostic information. In contrast, the ROR score only gave little prognostic information in patients whose tumors had low CAV1 expression. It has been hypothesized that both an active tumor promoting TME, and oncogenic intrinsic features of malignant cells are needed for the tumor to be able to metastasize9,10, which is in line with our findings.
Our study examined mRNA rather than protein levels. In addition to mRNA expression, protein levels are also affected by translation, post-translational modifications, and regulation of the rate of protein decay30. The global correlation between mRNA and protein is expected to be high30. We have previously reported a correlation of Rs = 0.47 for CAV118. Consequently, the present study results must be interpreted in the context of the biological phenotype related to high CAV1 mRNA expression.
The standard treatment regimens differ between SCAN-B and METABRIC31,32,33,34,35 mainly due to samples being collected during different time periods31,34,35. Therefore, there is a large discrepancy in the type of treatments between the older METABRIC and the contemporary SCAN-B cohort. Differences in treatments could explain why the results regarding prognosis were not fully replicated. CAV1 has been shown to modulate treatment efficacy of chemotherapy (including epirubicin and taxanes) and trastuzumab in breast cancer and other cancers36,37,38,39. These treatments were rarely or not at all used in METABRIC. Our findings in SCAN-B may partly be explained by how CAV1 modulates these treatments since patients with ROR high tumors are more likely to receive chemotherapy and trastuzumab, where treatment efficacy partly depends on CAV1 expression. Unfortunately, a lack of more detailed information on treatments makes it hard to evaluate the role of CAV1 expression in response to specific treatments in our study. Assessment of CAV1 expression in tumor samples from previous randomized clinical trials is warranted to confirm whether CAV1 expression may further refine ROR score prediction.
It should be mentioned that the PAM50 ROR score is used clinically for risk prediction in postmenopausal patients with ER+ /HER2− tumors to identify patients where the recurrence risk is low enough to omit chemotherapy4,7,8. However, studies have shown that in ER negative disease, (ER− /HER2+ and TNBC), both PAM50 subtype and ROR score can predict neoadjuvant treatment response40,41,42,43. Further, in TNBC disease, PAM50 subtype categorization could predict sensitivity to taxanes and capecitabine treatment44. Similarly, PAM50 subtype and ROR score were shown to be prognostic in HER2+ disease45. Therefore, PAM50 ROR could play a role in more than one clinical subgroup of breast cancer, and we believe it is of interest to study the ROR score in a broader context. Our data indicate that CAV1 expression could identify tumors where ROR score was prognostic, potentially broadening the applicability of the PAM50 ROR score, beyond the subgroup of ER+ /HER2− tumors.
Beyond the type of treatment, there could be several other reasons why the results regarding prognosis were not precisely replicated in the METABRIC. First, the METABRIC is a smaller cohort, hence, sample sizes in the tests are smaller and potential survival associations may not be as readily detectable. METABRIC also consists of more advanced tumors and is not population-based46, which is reflected by differences between the cohorts in the distribution of clinicopathological variables, including ROR category. Since nodal status is the key factor for determining ROR category and nodal status was substantially higher in METABRIC than in SCAN-B, this fact may in part explain why the prognostic impact of CAV1 gene expression differed somewhat between the cohorts. The underlying risk of recurrence and death in METABRIC is considerably higher than in SCAN-B, making direct comparisons regarding prognosis difficult46. The derived CAV1 tertile classifications are relative to a population and not based on absolute cut-offs for each tumor. The tertile cut-offs were applied separately for each cohort, meaning that some tumors would be reclassified if a unform cut-off had been applied. One might expect that relatively more tumors in METABRIC would have be classified as low CAV1 expressing due to the inverse association between CAV1 expression and tumor aggressiveness46.
It should be noted that the gene expression data is derived from bulk tumors, which reflects the averaged gene expression across thousands of cells and different cell types47. Therefore, it was not possible to definitively infer which cell types CAV1 was located in47 and, subsequently, the role CAV1 plays in these cell types in the context of breast cancer. Unfortunately, to date, no available assays can apply single-cell resolution for large-scale cohorts such as SCAN-B and METABRIC to evaluate prognostic biomarkers.
Nonetheless, the population-based contemporary SCAN-B cohort offers unique advantages, which lies in the large-scale RNAseq analysis of consecutively enrolled breast cancers32,33,48. To our knowledge, this is the largest cohort of its kind to date. Due to a rigorous population-based approach with consistently high inclusion rates because of seamless integration of patient enrollment and tissue sampling incorporated into routine clinical practice, the study cohort can be considered representative of the general patient demographics in the catchment area32,33,48. Therefore, SCAN-B allows for the evaluation of biomarkers in a contemporary real-world setting. Further, most findings were confirmed, showing stable associations of CAV1 expression with clinicopathological factors and tumor biology, consistent with the literature. In both cohorts, similar additive interactions with ROR regarding clinical outcome were shown as well as the underlying CAV1 expression being able to markedly change the prognostic information yielded by PAM50 ROR.
In conclusion, our findings indicate that high CAV1 gene expression is associated with a particularly poor prognosis in patients with ROR high tumors. As CAV1 can mediate between malignant cells and the TME it may also be a promising therapeutic target. The underlying CAV1 expression markedly modified the prognostic information provided by PAM50 ROR. We have shown in two independent datasets that PAM50 ROR was only prognostic in tumors with high CAV1 expression. Thus, CAV1 expression could be a useful biomarker that may enhance and further refine PAM50 ROR risk stratification for patients with ROR high tumors.
Materials and methods
SCAN-B
The Swedish Cancerome Analysis Network—Breast (SCAN-B: ClinicalTrials.gov ID NCT02306096) is an ongoing population-based study that have enrolled breast cancer patients at seven hospitals in South Sweden and two additional hospitals (Uppsala and Jönköping)32,33. The enrollment of patients is integrated in clinical routine33 and all patients with newly diagnosed or suspected breast cancer are invited to participate. The Swedish National Quality Registry for Breast Cancer is used for collection of clinicopathological data, treatment information, and follow-up32,33,48.
Sample collection followed established SCAN-B procedures and protocols32,33. In brief, the remaining fresh collected tumor samples from surgical specimens were preserved in RNAlater (Qiagen, Hilden, Germany). Core needle biopsies were taken before neoadjuvant treatment and preserved in RNAlater. Gene expression profiling of the tumors was performed by massive parallel paired-end sequencing of mRNA (RNA-seq) using a custom SCAN-B workflow32,48. Details on library preparation, quality control, the analysis pipeline, and software used are described elsewhere32,33,48.
All clinicopathological data and gene expression data for SCAN-B patients used here were downloaded from the Supplementary Information and Data from Staaf et al.48. Expression levels were expressed in fragments per kilobase of exon per million mapped reads (FPKM) in an expression matrix48. To all FPKM data an offset of + 0.1 was added, and then the data was log2 transformed.
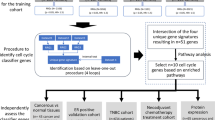
Patients were enrolled between September 1, 2010, and May 31, 2018, and followed until November 202148. From the beginning 7743 patients were included with a total of 8350 gene expression profiles (GEXs), as previously described48. After exclusion of GEXs from noninvasive cancer or lymph nodes, a total of 7142 patients remained with 7650 GEXs in the current study. In case multiple gene expression profiles from a single tumor passed quality control, the profile with the highest RNA concentration measured by NanoDrop spectrophotometry was chosen, as previously described48.This procedure left one GEX per patient for analysis. Further, patients with bilateral cancer or no available follow-up for distant metastasis were excluded, Fig. 1. After exclusions, GEX profiles from a total of 5326 patients were available for analysis. Information on PAM50 subtype and ROR category was obtained from Staaf et al.48 who assigned these categories using single sample predictors48.
Metabric
Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) is collection of clinically annotated primary fresh-frozen breast cancer specimens from five tumor banks in the UK and Canada34. The patients were diagnosed with non-metastatic breast cancer between 1977 and 200534. Manual curation and basic quality control of the clinicopathological data including treatment information was performed31. None of the HER2+ patients received trastuzumab. For a subset of 1980 patients, known as the METABRIC molecular dataset, gene-expression data from microarrays is available31,34,35. Details on sample handling, gene expression profiling and workflow are described elsewhere34,35. The METABRIC molecular dataset was downloaded from https://www.cbioportal.org/study/summary?id=brca_metabric and corresponding clinical data from Rueda et al.31. The genefu package49 was used to assign PAM50 subtype using nearest centroid correlation22 and calculate the PAM50 ROR score based on centroid correlations, tumor size and proliferation score according to the ROR equation with nodal status dependent cut-offs to assign categories, as described7,50,51.
In both SCAN-B and METABRIC, eight gene expression modules representing different biological functions in breast cancer were calculated as previously described52.
Statistical analysis
Differences in CAV1 mRNA expression depending on PAM50 subtype were evaluated using Kruskal–Wallis test. Correlations between CAV1 expression, ROR category and the eight gene modules52 were assessed using Pearson’s correlation (r). Pearson’s correlation was also used to assess correlations between CAV1 mRNA expression and mRNA expression of the PAM50 genes. Logistic regression was used to test whether CAV1 mRNA expression was independently associated with ROR category after adjusting for potential confounders (age at diagnosis, axillary lymph node status (pN1/2/3), tumor size (pT2/3/4), Grade (III vs I or II), ER+, PR+, HER2+, PAM50 subtype (Luminal A as reference), and (neo)adjuvant treatments.
Endpoints used for survival analysis were recurrence-free interval (RFI), distant metastasis-free interval (DMFI), and overall survival (OS) for both SCAN-B and METABRIC, as previously defined31,34,48. Breast cancer-specific survival (BCSS) was used as additional endpoint for METABRIC31,34.
For the survival analysis, log2 transformed CAV1 mRNA expression were categorized into tertiles, tertile 1 (T1), tertile 2 (T2), and tertile 3 (T3) to allow for non-linear effects. The lowest expression of CAV1 (T1) was used as reference. For survival analyses, the R packages ‘survival’ and ‘survminer’ were used.
Univariable survival analyses were performed using the Kaplan–Meier method and the Log-rank test. Cox proportional hazards models were used to obtain crude and adjusted Hazard ratios (HRs) with 95% confidence intervals (CI). The multivariable models were adjusted for age (binned in 5-year intervals for SCAN-B or continuous for METABRIC), tumor characteristics; axillary lymph node status (pN1/2/3), tumor size (pT2/3/4), Grade (III vs I or II), ER+, PR+, HER2+, PAM50 subtype (Luminal A as reference), PAM50 ROR category (High vs Low/Intermediate); and (neo)adjuvant treatments (endocrine treatment and chemotherapy for both SCAN-B and METABRIC and trastuzmab for SCAN-B only).
Schoenfeld’s residuals were used to test and graphically examine the proportional hazard assumption for the CAV1 tertiles in the adjusted model. To investigate effect modifications between the CAV1 tertiles and PAM50 ROR category, two-way interaction analyses on multiplicative and additive scales were performed in the multivariable model using the ‘interactionR’ package53.
Differential gene expression (DGE) analysis was conducted in SCAN-B using the ‘Limma-Voom’ package54 to find differentially expressed genes (DEGs) between the highest tertile (T3) and the lowest tertile (T1) of CAV1 expression. The criteria used to define DEGs is a false discovery rate (FDR) of ≤ 0.05 and log2 fold change (log2FC) ≥ 1.5 for up-regulated genes and log2FC ≤ − 1.5 for down-regulated genes. To correct for batch effects, batch was included in the Limma models. Gene set enrichment analysis (GSEA) was performed in ‘clusterprofiler’55 to find the statistically significant, concordant gene sets that differed between the highest tertile (T3) and the lowest tertile (T1) of CAV1 expression. Gene sets were grouped according to Gene Ontology (GO) and Hallmark Signature annotations56,57.
All statistical analyses were conducted in R version 4.2.2. P-values < 0.05 was considered statistically significant. All P-values were two-tailed. This study followed the Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK) criteria58.
Ethics approval and consent to participate
Ethical approvals for the cohorts studied (SCAN-B and METABRIC) were obtained in relation to the primary projects and publications31,32,33,34,35,48. The SCAN-B study was approved by the Lund University ethics committee32,33,48. The METABRIC study was approved by the ethics committees at the University of Cambridge and the British Columbia Cancer Research Centre 31,34,35. All participants signed written informed consent. No separate approval was obtained for this specific study since it is based on previously published data. The study was conducted in accordance with the ethical principles of the Declaration of Helsinki.
Data availability
RNA-sequencing-based gene expression data for the SCAN-B cohort is a publicly accessible dataset from Staaf et al.48 available at Mendeley Data. Microarray-based gene-expression data for METABRIC is publicly available from Curtis et al.34 and Pereira et al.35 at cBioPortal and clinical data is a publicly accessible dataset from Rueda et al.31.
Abbreviations
- BCSS:
-
Breast cancer-specific survival
- CAF:
-
Cancer-associated fibroblast
- CAV1:
-
Caveolin-1
- DEGs:
-
Differentially expressed genes
- DGE:
-
Differential gene expression
- DMFI:
-
Distant metastasis-free interval
- EGFR:
-
Epidermal growth factor receptor
- ER:
-
Estrogen receptor
- EMT:
-
Epithelial-mesenchymal transition
- FC:
-
Fold change
- FDR:
-
False discovery rate
- FPKM:
-
Fragments per kilobase of exon per million mapped reads
- GEX:
-
Gene expression profile
- GO:
-
Gene ontology
- GSEA:
-
Gene set enrichment analysis
- HER2:
-
Human epidermal growth factor receptor 2
- HIF1α:
-
Hypoxia-inducible factor 1 α
- HIF2α:
-
Hypoxia-inducible factor 2 α
- HR:
-
Hazard ratio
- IHC:
-
Immunohistochemistry
- OS:
-
Overall survival
- METABRIC:
-
Molecular taxonomy of breast cancer international Consortium
- PR:
-
Progesterone receptor
- RFI:
-
Recurrence-free interval
- ROR:
-
Risk of recurrence
- RNA-seq:
-
Massive parallel paired-end sequencing of mRNA
- SCAN-B: :
-
Swedish cancerome atlas network-Breast
- TGFβ:
-
Transforming growth factor-beta
- TME:
-
Tumor microenvironment
- TNBC:
-
Triple-negative breast cancer
- VEGF:
-
Vascular endothelial growth factor
References
Harbeck, N. et al. Breast cancer. Nat. Rev. Dis. Primers 5, 66. https://doi.org/10.1038/s41572-019-0111-2 (2019).
An overview of the randomised trials. Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival. Lancet 365, 1687–1717. https://doi.org/10.1016/s0140-6736(05)66544-0 (2005).
Kwa, M., Makris, A. & Esteva, F. J. Clinical utility of gene-expression signatures in early stage breast cancer. Nat. Rev. Clin. Oncol. 14, 595–610. https://doi.org/10.1038/nrclinonc.2017.74 (2017).
Harris, L. N. et al. Use of biomarkers to guide decisions on adjuvant systemic therapy for women with early-stage invasive breast cancer: American society of clinical oncology clinical practice guideline. J. Clin. Oncol. 34, 1134–1150. https://doi.org/10.1200/JCO.2015.65.2289 (2016).
Sparano, J. A. et al. Clinical and genomic risk to guide the use of adjuvant therapy for breast cancer. N. Engl. J. Med. 380, 2395–2405. https://doi.org/10.1056/NEJMoa1904819 (2019).
Ohnstad, H. O. et al. Prognostic value of PAM50 and risk of recurrence score in patients with early-stage breast cancer with long-term follow-up. Breast Cancer Res. 19, 120. https://doi.org/10.1186/s13058-017-0911-9 (2017).
Gnant, M. et al. Predicting distant recurrence in receptor-positive breast cancer patients with limited clinicopathological risk: using the PAM50 Risk of Recurrence score in 1478 postmenopausal patients of the ABCSG-8 trial treated with adjuvant endocrine therapy alone. Ann. Oncol. 25, 339–345. https://doi.org/10.1093/annonc/mdt494 (2014).
Lænkholm, A. V. et al. PAM50 risk of recurrence score predicts 10-year distant recurrence in a comprehensive danish cohort of postmenopausal women allocated to 5 years of endocrine therapy for hormone receptor-positive early breast cancer. J. Clin. Oncol. 36, 735–740. https://doi.org/10.1200/jco.2017.74.6586 (2018).
de Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: From cancer initiation to metastatic outgrowth. Cancer Cell. 41, 374–403. https://doi.org/10.1016/j.ccell.2023.02.016 (2023).
Bejarano, L., Jordāo, M. J. C. & Joyce, J. A. Therapeutic targeting of the tumor microenvironment. Cancer Discov. 11, 933–959. https://doi.org/10.1158/2159-8290.Cd-20-1808 (2021).
Dieci, M. et al. Prognostic and predictive value of tumor-infiltrating lymphocytes in two phase III randomized adjuvant breast cancer trials. Ann. Oncol. 26, 1698–1704 (2015).
Caligiuri, G. & Tuveson, D. A. Activated fibroblasts in cancer: Perspectives and challenges. Cancer Cell. 41, 434–449. https://doi.org/10.1016/j.ccell.2023.02.015 (2023).
Kanzaki, R. & Pietras, K. Heterogeneity of cancer-associated fibroblasts: Opportunities for precision medicine. Cancer Sci. 111, 2708–2717. https://doi.org/10.1111/cas.14537 (2020).
Wagner, J. et al. A single-cell atlas of the tumor and immune ecosystem of human breast cancer. Cell 177, 1330-1345.e1318. https://doi.org/10.1016/j.cell.2019.03.005 (2019).
Patani, N., Martin, L. A., Reis-Filho, J. S. & Dowsett, M. The role of caveolin-1 in human breast cancer. Breast Cancer Res. Treat. 131, 1–15. https://doi.org/10.1007/s10549-011-1751-4 (2012).
Simón, L., Campos, A., Leyton, L. & Quest, A. F. G. Caveolin-1 function at the plasma membrane and in intracellular compartments in cancer. Cancer Metastasis Rev. 39, 435–453. https://doi.org/10.1007/s10555-020-09890-x (2020).
Ketteler, J. & Klein, D. Caveolin-1, cancer and therapy resistance. Int. J. Cancer 143, 2092–2104. https://doi.org/10.1002/ijc.31369 (2018).
Godina, C. et al. Interplay between Caveolin-1 and body and tumor size affects clinical outcomes in breast cancer. Transl. Oncol. https://doi.org/10.1016/j.tranon.2022.101464 (2022).
Scatena, C. et al. New insights in the expression of stromal caveolin 1 in breast cancer spread to axillary lymph nodes. Sci. Rep. 11, 2755. https://doi.org/10.1038/s41598-021-82405-y (2021).
Sloan, E. K. et al. Stromal cell expression of caveolin-1 predicts outcome in breast cancer. Am. J. Pathol. 174, 2035–2043. https://doi.org/10.2353/ajpath.2009.080924 (2009).
Witkiewicz, A. K. et al. An absence of stromal caveolin-1 expression predicts early tumor recurrence and poor clinical outcome in human breast cancers. Am. J. Pathol. 174, 2023–2034. https://doi.org/10.2353/ajpath.2009.080873 (2009).
Parker, J. S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 27, 1160–1167. https://doi.org/10.1200/jco.2008.18.1370 (2009).
Goetz, J. G. et al. Biomechanical remodeling of the microenvironment by stromal caveolin-1 favors tumor invasion and metastasis. Cell 146, 148–163. https://doi.org/10.1016/j.cell.2011.05.040 (2011).
Martinez-Outschoorn, U. E., Sotgia, F. & Lisanti, M. P. Caveolae and signalling in cancer. Nat. Rev. Cancer 15, 225–237. https://doi.org/10.1038/nrc3915 (2015).
Bailey, K. M. & Liu, J. Caveolin-1 up-regulation during epithelial to mesenchymal transition is mediated by focal adhesion kinase. J. Biol. Chem. 283, 13714–13724. https://doi.org/10.1074/jbc.M709329200 (2008).
Mittal, V. Epithelial mesenchymal transition in tumor metastasis. Annu. Rev. Pathol. 13, 395–412. https://doi.org/10.1146/annurev-pathol-020117-043854 (2018).
Bernatchez, P. Endothelial caveolin and its scaffolding domain in cancer. Cancer Metastasis Rev. 39, 471–483. https://doi.org/10.1007/s10555-020-09895-6 (2020).
Wang, Y. et al. Hypoxia promotes ligand-independent EGF receptor signaling via hypoxia-inducible factor-mediated upregulation of Caveolin-1. Proc. Natl. Acad. Sci. USA 109, 4892–4897. https://doi.org/10.1073/pnas.1112129109 (2012).
Wang, S. et al. Caveolin-1 inhibits breast cancer stem cells via c-Myc-mediated metabolic reprogramming. Cell. Death Dis. 11, 450. https://doi.org/10.1038/s41419-020-2667-x (2020).
Larsson, O., Tian, B. & Sonenberg, N. Toward a genome-wide landscape of translational control. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a012302 (2013).
Rueda, O. M. et al. Dynamics of breast-cancer relapse reveal late-recurring ER-positive genomic subgroups. Nature 567, 399–404. https://doi.org/10.1038/s41586-019-1007-8 (2019).
Saal, L. H. et al. The Sweden Cancerome Analysis Network - Breast (SCAN-B) Initiative: A large-scale multicenter infrastructure towards implementation of breast cancer genomic analyses in the clinical routine. Genom. Med. 7, 20. https://doi.org/10.1186/s13073-015-0131-9 (2015).
Rydén, L. et al. Minimizing inequality in access to precision medicine in breast cancer by real-time population-based molecular analysis in the SCAN-B initiative. Br J Surg 105, e158–e168. https://doi.org/10.1002/bjs.10741 (2018).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352. https://doi.org/10.1038/nature10983 (2012).
Pereira, B. et al. The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat. Commun. 7, 11479. https://doi.org/10.1038/ncomms11479 (2016).
Wang, Z. et al. Caveolin-1 mediates chemoresistance in breast cancer stem cells via β-catenin/ABCG2 signaling pathway. Carcinogenesis 35, 2346–2356. https://doi.org/10.1093/carcin/bgu155 (2014).
Zhao, Y. et al. Caveolin-1 expression predicts efficacy of weekly nab-paclitaxel plus gemcitabine for metastatic breast cancer in the phase II clinical trial. BMC Cancer 18, 1019. https://doi.org/10.1186/s12885-018-4936-y (2018).
Pereira, P. M. R. et al. Caveolin-1 mediates cellular distribution of HER2 and affects trastuzumab binding and therapeutic efficacy. Nat. Commun. 9, 5137. https://doi.org/10.1038/s41467-018-07608-w (2018).
Indira Chandran, V. et al. Hypoxia attenuates trastuzumab uptake and Trastuzumab-Emtansine (T-DM1) cytotoxicity through redistribution of phosphorylated Caveolin-1. Mol. Cancer Res. 18, 644–656. https://doi.org/10.1158/1541-7786.Mcr-19-0856 (2020).
Dieci, M. V. et al. Integrated evaluation of PAM50 subtypes and immune modulation of pCR in HER2-positive breast cancer patients treated with chemotherapy and HER2-targeted agents in the CherLOB trial. Ann. Oncol. 27, 1867–1873. https://doi.org/10.1093/annonc/mdw262 (2016).
Gluz, O. et al. Efficacy of deescalated chemotherapy according to PAM50 subtypes, immune and proliferation genes in triple-negative early breast cancer: Primary translational analysis of the WSG-ADAPT-TN trial. Int. J. cancer 146, 262–271. https://doi.org/10.1002/ijc.32488 (2020).
Filho, O. M. et al. Association of immunophenotype with pathologic complete response to Neoadjuvant chemotherapy for triple-negative breast cancer: A secondary analysis of the brightness phase 3 randomized clinical trial. JAMA Oncol. 7, 603–608. https://doi.org/10.1001/jamaoncol.2020.7310 (2021).
Carey, L. A. et al. Molecular heterogeneity and response to Neoadjuvant human epidermal growth factor receptor 2 targeting in Calgb 40601, a randomized phase iii trial of paclitaxel plus Trastuzumab with or without Lapatinib. J. Clin. Oncol. 34, 542–549. https://doi.org/10.1200/jco.2015.62.1268 (2016).
Asleh, K. et al. Triple-Negative PAM50 Non-Basal Breast Cancer Subtype Predicts Benefit from Extended Adjuvant Capecitabine. Clin. cancer Res. 29, 389–400. https://doi.org/10.1158/1078-0432.Ccr-22-2191 (2023).
Prat, A. et al. Research-based PAM50 subtype predictor identifies higher responses and improved survival outcomes in HER2-positive breast cancer in the NOAH study. Clin. Cancer Res. 20, 511–521. https://doi.org/10.1158/1078-0432.Ccr-13-0239 (2014).
Xie, Y. et al. Breast cancer gene expression datasets do not reflect the disease at the population level. NPJ Breast Cancer 6, 39. https://doi.org/10.1038/s41523-020-00180-x (2020).
Li, X. & Wang, C. Y. From bulk, single-cell to spatial RNA sequencing. Int. J. Oral. Sci. 13, 36. https://doi.org/10.1038/s41368-021-00146-0 (2021).
Staaf, J. et al. RNA sequencing-based single sample predictors of molecular subtype and risk of recurrence for clinical assessment of early-stage breast cancer. NPJ Breast Cancer 8, 94. https://doi.org/10.1038/s41523-022-00465-3 (2022).
Gendoo, D. M. et al. Genefu: an R/Bioconductor package for computation of gene expression-based signatures in breast cancer. Bioinformatics 32, 1097–1099. https://doi.org/10.1093/bioinformatics/btv693 (2016).
Filipits, M. et al. The PAM50 risk-of-recurrence score predicts risk for late distant recurrence after endocrine therapy in postmenopausal women with endocrine-responsive early breast cancer. Clin. Cancer Res. 20, 1298–1305. https://doi.org/10.1158/1078-0432.Ccr-13-1845 (2014).
Wallden, B. et al. Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med. Genom. 8, 54. https://doi.org/10.1186/s12920-015-0129-6 (2015).
Fredlund, E. et al. The gene expression landscape of breast cancer is shaped by tumor protein p53 status and epithelial-mesenchymal transition. Breast Cancer Res. 14, R113. https://doi.org/10.1186/bcr3236 (2012).
Alli, B. Y. InteractionR: An R package for full reporting of effect modification and interaction. Softw. Impacts https://doi.org/10.1016/j.simpa.2021.100147 (2021).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv007 (2015).
Wu, T. et al. Clusterprofiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141. https://doi.org/10.1016/j.xinn.2021.100141 (2021).
Ashburner, M. et al. Gene ontology: Tool for the unification of biology. Gene. Ontol. Consortium. Nat. Genet. 25, 25–29. https://doi.org/10.1038/75556 (2000).
Liberzon, A. et al. The molecular signatures database (MSIGDB) hallmark gene set collection. Cell. Syst. 1, 417–425. https://doi.org/10.1016/j.cels.2015.12.004 (2015).
McShane, L. M. et al. Reporting recommendations for tumor marker prognostic studies (REMARK). J. Natl. Cancer Inst. 97, 1180–1184. https://doi.org/10.1093/jnci/dji237 (2005).
Acknowledgements
The authors would like to acknowledge the patients who provided consent to participate in the SCAN-B study, the central SCAN-B laboratory at the Division of Oncology, Lund University for sample processing and RNA sequencing, the Swedish National Quality Register for Breast Cancer for clinical and histopathological data, Regional Cancer Center South, and the South Swedish Breast Cancer Group.
Funding
Open access funding provided by Lund University. The Swedish Cancer Society (CAN 20 0763 and CAN 23 2952), the Faculty of Medicine at Lund University, the Mrs Berta Kamprad Foundation, the South Swedish Health Care Region (Region Skåne ALF 40620), and the Skåne University Hospital fund. AB and KI holds a young researcher award from ALF (Region Skåne). The funders had no role in study design and conduct of the study, data collection and analysis, data interpretation, or manuscript preparation and decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
C Godina: Conceptualization, Data curation, Formal analysis, Methodology, Software, Visualization, Writing—original draft, Writing—review & editing. M Belting: Conceptualization, Writing—review & editing. J Vallon-Christersson: Resources, Data curation, Writing – review & editing. K Isaksson: Funding acquisition, Writing—review & editing. A Bosch: Conceptualization, Supervision, Writing—review & editing. H Jernström: Conceptualization, Funding acquisition, Formal analysis, Methodology, Project administration, Resources, Supervision, Writing—original draft, Writing—review & editing. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Ana Bosch has received institutional honoraria from Pfizer, Roche and Lilly for consultation and lectures. She has participated in Advisory Board meetings for Pfizer and Novartis. Co-founder and chair of the board for SACRA therapeutics. Karolin Isaksson has received speaker honorarium from Pierre Fabre. The other authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Godina, C., Belting, M., Vallon-Christersson, J. et al. Caveolin-1 gene expression provides additional prognostic information combined with PAM50 risk of recurrence (ROR) score in breast cancer. Sci Rep 14, 6675 (2024). https://doi.org/10.1038/s41598-024-57365-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-57365-8
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.