Abstract
Despite Plasmodium ovale curtisi (Poc) and wallikeri (Pow) being important human-infecting malaria parasites that are widespread across Africa and Asia, little is known about their genome diversity. Morphologically identical, Poc and Pow are indistinguishable and commonly misidentified. Recent rises in the incidence of Poc/Pow infections have renewed efforts to address fundamental knowledge gaps in their biology, and to develop diagnostic tools to understand their epidemiological dynamics and malaria burden. A major roadblock has been the incompleteness of available reference assemblies (PocGH01, PowCR01; ~ 33.5 Mbp). Here, we applied multiple sequencing platforms and advanced bioinformatics tools to generate new reference genomes, Poc221 (South Sudan; 36.0 Mbp) and Pow222 (Nigeria; 34.3 Mbp), with improved nuclear genome contiguity (> 4.2 Mbp), annotation and completeness (> 99% Plasmodium spp., single copy orthologs). Subsequent sequencing of 6 Poc and 15 Pow isolates from Africa revealed a total of 22,517 and 43,855 high-quality core genome SNPs, respectively. Genome-wide levels of nucleotide diversity were determined to be 2.98 × 10–4 (Poc) and 3.43 × 10–4 (Pow), comparable to estimates for other Plasmodium species. Overall, the new reference genomes provide a robust foundation for dissecting the biology of Poc/Pow, their population structure and evolution, and will contribute to uncovering the recombination barrier separating these species.
Similar content being viewed by others
Introduction
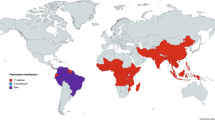
Plasmodium ovale curtisi (Poc) and Plasmodium ovale wallikeri (Pow) are the least-studied human-infecting Plasmodium parasites. Large gaps remain in our understanding of these elusive parasites, from their full geographic distribution to antimalarial susceptibility1,2. Historically, P. ovale was considered a single parasite species (defined by blood-film morphology) associated with benign malaria which rarely presented severe complications including jaundice, anaemia, and fatal pulmonary impairments3,4,5. In 2010, it was demonstrated that ovale malaria was in fact caused by two non-recombining sympatric species subsequently named Poc and Pow2. P. ovale spp. infections mostly occur in Africa (94.5%) followed by Asia (5.3%)6,7,8,9,10. However, the prevalence of P. ovale spp. has been historically underestimated due to asymptomatic infections being unnoticed or undetected, a lack of an accurate rapid diagnostic test (RDT), and microscopy-based misclassification11,12,13,14,15. As such P. ovale spp. infections are commonly reported as mixed infections, alongside P. falciparum or P. vivax, where the presence of another parasite causes an infected individual to become symptomatic and seek medical attention16,17,18. Worryingly, a multi-centre study in Kenya reported an increase in the prevalence of P. ovale spp. co-infections between years 2008 and 201619, but there is a lack of data from other settings. Such reports were prior to the SARS-CoV-2 pandemic, which has subsequently caused a global increase in malaria incidence across all Plasmodium species due to the disruption of intervention efforts20,21.
To date, the genomic characterization of Poc and Pow remains limited, especially when compared to other human infecting Plasmodium species such as P. falciparum and P. vivax22. Addressing this gap in our knowledge is vital, from the development of new treatments and diagnostics to facilitating accurate parasite population surveillance. Most genomic analyses require a complete reference genome, but both currently available assemblies (PocGH01 ~ 33.4 Mbp and PowCR01 ~ 33.5 Mbp; from Ghana) are incomplete, averaging > 2700 and > 7700 unknown nucleotides per megabase, respectively23. To fill this gap, we have sequenced two DNA samples (Poc221 and Pow222), sourced from mono-infections arising in Africa, utilising both Illumina and Oxford Nanopore Technology platforms. The resulting bioinformatic and validation analysis, provides two new reference genomes (Poc221 and Pow222), which are closer to being complete and more robust than the existing P. ovale spp. assemblies, and can be used for high resolution population genomic analysis across large numbers of sequenced samples.
Results
Assembly of the new reference genomes
For the generation of new Poc and Pow reference genomes, two DNA samples (Poc221, Pow222) from travellers returning to the UK from South Sudan and Nigeria respectively, were obtained from the UK Health Security Agency Malaria Reference Laboratory (UKHSA MRL). Both samples were initially sequenced using the Oxford Nanopore Technology (ONT) MinION platform under adaptive sampling conditions, negatively selecting against the human genome followed by P. ovale spp. specific enrichment via selective whole genome amplification (SWGA). The samples were subsequently sequenced using additional ONT MinION runs and the Illumina platform (see Methods). Together this yielded 13.68 Gb and 13.78 Gb of sequence data for samples Poc221 (11.58 Gb Illumina, 2.10 Gb ONT) and Pow222 (10.32 Gb Illumina, 3.46 Gb ONT), respectively. Following quality control (see Methods), 2.59 and 2.40 Gb of WGS data remained for samples Poc221 and Pow222, respectively (Supplementary Table S1). A hybrid assembly approach using Spades software was then implemented to generate the new reference sequences for Poc221 and Pow222 (see Supplementary Figure S1; Methods), which resulted in nuclear (14 chromosomes) and organellar (mitochondrion, apicoplast) genomes.
Benchmarking to previous reference genomes
The new reference genomes were benchmarked against the existing available references, PocGH01 (https://plasmodb.org/: v54) and PowCR01 (https://www.ncbi.nlm.nih.gov/: GCA_900090025.2) (Table 1). Gains in core genome contiguity were made across both P. ovale ssp. (> 4.2 Mbp), reflected by > 5- and eightfold improvements in N50, a common metric to assess assembly quality, alongside a 76% and 86% reduction in the number of gaps for both Poc and Pow, respectively. When assessing contiguity improvements on a chromosomal level (Supplementary Figure S2), gains were made for 13 Poc chromosomes, with a maximum increase of at least 670 Kbp in chromosome 8 and minimum increase of at least 70 Kbp in chromosome 9. Similarly, gains were made across 13 Pow chromosomes, with a maximum increase of > 1.1 Mbp for chromosome 10. For both species, most contiguity gains were in sub-telomeric regions (Fig. 1). However, for Pow the most important contiguity gain was on chromosome 10, covering extended core and sub-telomeric regions, which were missing in the PowCR01 assembly. The Poc and Pow nuclear chromosomes have an average homology of 84.1% and 81.0%, respectively, between the new genomes and historic assemblies, which is again reflected in the comparable GC content obtained between new and historic references for each species (Table 1).
Visualizing the new genomic references (A) Poc221 and (B) Pow222. From outer ring to inside: (1) Representation of each nuclear chromosome; (2) Black regions represent chromosomal specific islands of homology identified between the new and historic references. (3). Green regions represent core ortholog genes identified in the new references and Red regions represent members of the PIR multigene family. 4) SNP density.
Annotation enhancements
Poc221 and Pow222 reference genomes were annotated using a combination of Companion and Metaeuk software (see Methods), and then compared to PocGH01 and PowCR01 (Table 2). The new reference genomes had an increased number of protein coding genes (Poc + 769, Pow + 668) and less pseudogenes (Poc -93, Pow -249). Whilst the number of non-coding genes, including ncRNA, snoRNA, snRNA and tRNA remained comparable. To assess the completeness of the references created, ortholog analysis was performed using the 2 new and 2 historic P. ovale spp. reference genomes, along with 13 other Plasmodium species (see Methods; Supplementary Table S2). A total of 6,916 orthogroups were identified across the 17 Plasmodium species references analysed. Of these, 4,268 were identified as being single copy core orthogroups, which were shared across all 13 comparator Plasmodium species. Nearly all these orthogroups were represented in Poc221 (99.9%; 4263/4268) and Pow222 (99.9%; 4262/4268), superior to PocGH01 (99.7%; 4255/4268) and PowCR01 (90.1%; 3847/4268), and represent a gain of + 8 and + 415 single copy core orthologs for Poc and Pow, respectively. This result was subsequently confirmed by BUSCO genome analysis which marked a + 0.3% (Poc) and + 0.4% (Pow) improvement (Table 2). From the missing single copy core orthogroups, 4 were not identified in any P. ovale spp. (Supplementary Table S3), including orthogroup OG0004883, associated with a putative AP2 transcription factor involved in regulating the Plasmodium life cycle. Only one single copy core orthogroup (OG0004738; PF3D7_0416500) was present in Pow but absent in Poc, and vice versa, two were present in Poc but not Pow (OG0004794; PF3D7_0934500, OG0004804; PF3D7_1460700). The single copy orthologs present in the new P. ovale spp. references and the 13 other Plasmodium species were used to investigate the phylogeny of P. ovale spp. The estimated tree topology was in line with previous reports (Supplementary Figure S3), with both Poc and Pow clustering together. The P. ovale spp. clade shares a most recent common ancestor with rodent infecting Plasmodium species, including P. berghei ANKA, P. chabaudi chabaudi, P. vinckei brucechwatti, P. vinckei lentum, P. vinckei vinckei, and P. yoelii yoelii.
Due to improvements in genome contiguity, there were additional ortholog chromosomal assignments for both Poc221 (+ 256) and Pow222 (+ 478) compared to the historic Poc and Pow reference genomes (Supplementary Table S4). Complete mitochondrial sequences were obtained for both Poc221 (5974 bp) and Pow222 (5975 bp), representing almost a 400 bp improvement when compared to the PowCR01 mitochondrial reference (5584 bp). In addition, there were improvements in apicoplast contiguity (3 Kbp added, compared to GenBank entries KX611805 and LT594519), ensuring all 30 core orthogroups were represented (Supplementary Table S5). This result marked an improvement for the Pow reference genome with a gain of 11 apicoplast-associated core orthologs.
Poc and Pow divergence
Expansion of multigene families in P. ovale spp. relative to other Plasmodium species is a major driver behind genome change23,24. The Plasmodium interspersed repeat (PIR) multi-gene family is known to be the largest multigene family in most Plasmodium species25. Additional PIR genes were characterized for both Poc221 (1955 vs. PocGH01 1493) and Pow222 (1606 vs. PowCR01 1338) (Table 2). Of the PIR genes identified, 87.5% (1710) and 95.7% (1429) were clustered into 596 shared orthogroups for Poc221 and PocGH01, respectively. Similarly, 89.6% (1439) and 96.9% (1296) of the PIR genes identified for Pow222 and PowCR01, respectively, are clustered across 586 orthogroups. The results reinforced the greater annotated PIR expansion observed in Poc compared to Pow (PocGH01 vs. PowCR01: + 155; Poc221 vs Pow222: + 349). Most proteins in the expanded surfin-related subtelomeric protein 1 (STP1/SURFIN) multi-gene family in both P. ovale spp. (and P. malariae) contain a Schizont-infected cell agglutination (SICA) domain, with SICAvars known to have an evolutionary link with SURFIN and P. vivax STP1 proteins24. There were similar number of STP1 genes characterized in Pow222 (97) compared to PowCR01 (100), but more in Poc221 (88) compared to PocGH01 (67). When assessing divergence between shared Poc221 and Pow222 genes, single copy orthologs had a high average protein sequence identity (92.7%). When comparing equivalent nuclear chromosomes between Poc221 and Pow222, they average 81.3% genomic homology, including in sub-telomeric regions. No large-scale chromosomal rearrangements were identified between Poc and Pow, with translocated regions mostly found in sub-telomeric or telomeric regions (Fig. 2).
Variant calling
To assess the integrity of the new reference genomes, whole genome sequence (WGS) data was analysed for 12 Poc and 22 Pow samples, sourced publicly (n = 20) and from additional DNA samples from UK travellers (n = 14) (Supplementary Table S6). All samples were mapped to their respective existing and new reference genomes, PocGH01 and Poc221 or PowCR01 and Pow222. There was no significant difference in mean coverage or percentage of reads mapped when comparing the LSHTM sequenced and publicly available samples (all T-tests p > 0.05)26. When comparing the percentage of mapped reads between new and existing reference genomes, a slight increase was observed for Poc221 compared to PocGH01 (+ 0.03%: ~ 600 reads) but a significant gain was observed when comparing Pow222 to PowCR01 (+ 0.8%, 204,300 reads, P < 3 × 10–6). Using a set of 4834 common biallelic SNPs across the 34 isolates, a principal component analysis confirmed the expected two distinct clusters representing the Poc and Pow species (Supplementary Figure S4).
From the 6 Poc and 15 Pow isolates with high quality genome-wide data, a total of 349,408 and 421,404 genome-wide SNPs were identified, averaging 128,320 and 103,771 SNPs per sample, when utilizing the Poc221 and Pow222 references, respectively. The distribution of nucleotide diversity across both genomes revealed the expected high diversity peaks in sub-telomeric / telomeric regions (Fig. 1), and lower levels in the inferred core genome (Supplementary Table S7). After removing hyper-variable regions, leaving the core genome, 89.3% and 89.0% of the full nuclear genome remained for Poc221 and Pow222, from which a total of 22,517 and 43,855 SNPs were identified from the Poc and Pow samples, averaging 6,887 and 7,473 SNPs per sample, respectively. Subsequently the nucleotide diversity was estimated (Poc 2.98 × 10–4; Pow 3.43 × 10–4), in line with other Plasmodium studies27,28,29. Using the SNP data, neighbour-joining trees were constructed for both species, leading to some clustering of Pow isolates sourced from Cameroon and Senegal (Fig. 3). Species-specific SNPs in the mitochondrion genomes were confirmed across all isolates, further validating the new reference genomes, and supporting the established paradigm that there is no recombination between Poc and Pow (Supplementary Table S8).
Neighbour joining tree of the 15 Pow and 6 Poc isolates from West Africa (red), Central Africa (green), East Africa (blue), when aligned to Pow222 and Poc221 respectively. Corresponding BioSample IDs can be found in Supplementary Table S6.
Discussion
Regarded as the most recent speciation event within Plasmodium malaria parasites, the divergence of the Poc and Pow genomes is a natural experiment that illuminates the evolutionary forces shaping the adaptive radiation of the genus. To provide insights, we have utilised a combination of paired-end short-read genome sequences with long-read outputs to produce new references genomes for the sibling species Poc and Pow. These were derived from high-quality material from two clinical isolates harbouring only a single species each, with DNA undergoing further selective whole-genome amplification. Compared to previously available references, which were substantially assembled from co-infecting P. ovale spp DNA detected in existing genome sequence data from P. falciparum-infected blood samples23, we improved overall completeness adding > 400 single copy ortholog gene sequences to the reference annotation of the Pow genome. In addition, for both Poc and Pow, we extended sub-telomeric multi-gene annotations and obtained full organellar genomes for the mitochondrion and apicoplast.
Our robust, two-phase approach provided important new information in three areas. Firstly, more complete nuclear and organellar genome sequence were captured for these species than historic assemblies (PocGH01, PowCR01), with enhanced contiguity. Secondly, telomeric and sub-telomeric hypervariable regions harbouring multigene families such as PIR, believed to play a key role across all life cycle stages of the parasite25,30, are comprehensively represented for both species. Thirdly, the long read data identify for the first time a number of chromosomal translocations that are likely to reflect the evolutionary divergence of Poc and Pow (Fig. 2). This, and the notably greater expansion of the PIR multi-gene family in the genome of Poc compared to Pow, warrant further investigation and may help us understand the speciation of these two parasites. For example, one open question is whether the varying expansion supports the multi-jump speciation hypothesis, whereby the most common ancestor of Poc and Pow, was introduced to early homonids via 2 independent host transitions with enough evolutionary time in between to facilitate allopatric speciation by preventing recombination2. Further long-read sequencing across many Poc and Pow isolates, including from non-African sources, could help test whether the observed inter-chromosomal translocations are fixed and potentially contribute to the puzzling lack of recombination between Poc and Pow, despite their sympatry and the numerous documented co-infections in a single host31. Taken together, our data irrefutably support elevation of these two parasite taxa to full species status. This confirms that the Poc and Pow nomenclature of Sutherland et al.2, used throughout this report, is incorrect and needs to be changed, as pointed out recently by others4,5.
Certain caveats need to be considered in interpreting the data. Clinical ovale malaria infections are invariably of low density, and obtaining samples with sufficient parasite material to generate high quality genome sequences at high coverage is extremely challenging. Our access to additional material to use in our nucleotide diversity analysis was an advantage, but the sample size is very small (only 12 and 22 samples for Poc and Pow, respectively) and not all produced genome data of the required quality. Further, our Poc reference is from East Africa, but ten of our 12 comparators were West African. Conversely, our Pow reference is from West Africa, whereas only two isolates of East African origin were available to contribute to the diversity analysis for that species. Therefore, the true pattern of genetic diversity in both ovale species can only be determined with broader samples encompassing all of Africa, and parasites from Asia and Oceania. In lieu of such studies, our work provides the first insights into genomic diversity of Poc and Pow.
With the global objective of malaria eradication, ensuring control and treatment strategies are effective across all human-infecting Plasmodium species is essential. The new genome data provided here will improve selection of conserved drug targets for the development of new antimalarials and the development of rapid diagnostic tools which accurately identify these neglected species. These advances are essential to ensure that control activities focusing on P. falciparum and P. vivax can be modified to also address the burden of morbidity due to ovale malaria.
Methods
P. ovale spp. samples and DNA processing
Sixteen P. ovale spp. DNA samples were extracted from blood samples from returning travellers to the UK, who were diagnosed with malaria between 2019 and 2020, confirmed by the UKHSA MRL at LSHTM. Samples were initially designated as P. ovale spp. infections by nested PCR and qPCR according to standard practice. The UK National Research Ethics Service (Ref: 18/LO/0738) and LSHTM Research Ethics Committee (Ref: 14710) provided approval for the whole study under “Drug susceptibility and genetic diversity of imported malaria parasites from UK travellers”, and all methods were performed in accordance with relevant guidelines and regulations, and informed consent was obtained from all UK study participants.
To enrich Plasmodium DNA, a P. ovale spp. selective whole genome amplification (SWGA) primer set was designed, utilizing a software tool (https://github.com/eclarke/swga)32, to preferentially amplify Poc(PocGH01) and Pow (PowCR01) over the human genome (GRCh38). The top outputs were identified and overlapping primers combined to form a final set of 7 primers: CGAAAAA*A*C, CGAAAT*T*G, TCGTAAA*A*A, CGTAAT*A*A, TTTACGT*A*T, ATTTTCG*A*T, and TATCGT*T*A, where an asterisk (*) represents the presence of a phosphorothioate bond which minimises primer degradation by the 3’ exonuclease activity of Phi29. When combined, the SWGA primer set has a total of 28.12 and 28.43 binding sites per 100kbp of PocGH01 and PowCR01, respectively (Supplementary Table S9). There is a total of 2 bindings sites per 100kbp for the GRCh38 human reference genome, indicating a > 14-fold preference for the Plasmodium target. Samples were subject to SWGA following previously published protocols33. All SWGA reactions were carried out in a UV Cabinet for PCR Operations (UV-B-AR, Grant-Bio) to eliminate potential contamination. A maximum of 80 ng of gDNA (minimum of 5 ng) was added to a total 50 µl reaction alongside 5 µl of 10 × Phi29 DNA Polymerase Reaction Buffer (New England BioLabs), 0.5 µl of Purified 100 × BSA (New England BioLabs), 0.5 µl of 250 µM Primer mix, 5 µl 10 mM dNTP (Roche), 30 units Phi29 DNA Polymerase (New England BioLabs) and Nuclease-Free Water (Ambion, The RNA Company) to reach a final reaction volume of 50 µl. The reaction was carried out on a thermocycler with the following step-down program: 5 min at 35 °C, 10 min at 34 °C, 15 min at 33 °C, 20 min at 32 °C, 25 min 31 °C, 16 h at 30 °C and 10 min at 65 °C. After SWGA, samples were purified using a 1:1 ratio of AMPure XP beads (Beckman-Coulter), following manufacturer’s instructions and quantified via Qubit assay.
Library preparation and whole genome sequencing (WGS)
Short read sequencing (paired end 150 bp reads) of the DNA samples (n = 16; NCBI accession: PRJNA1015456) was performed on an Illumina NovaSeq 6000 platform by The Applied Genome Centre, LSHTM. For the two isolates (Poc221; SAMN37357391; Pow222 SAMN37357402), selected to create the new reference genomes, long-read sequencing data was obtained in two rounds using ONT MinION made available by The Applied Genome Centre, LSHTM. The two isolates were first prepared for sequencing using the ONT LSK-109 and EXP-NBD104 barcoding kit as per manufacturer's instruction. To select for fragments of greater mass during the library preparation procedure, LSB buffer was used during magnetic bead clean-up and 120 ng of library was loaded onto the R10 flow cell for sequencing with adaptive sampling rejecting reads associated with the human genome (GRCh38). Following SWGA enrichment and T7 endonuclease (NEB-M0302S) treatment, as per manufacturer’s protocol (WAL_9070_v109_revQ_14Aug2019), samples were again prepared following same methodology and sequenced using a R10 flow cell. All resulting fast5 files were base-called using Bonito (ONT) (models: dna_r10.3 and dna_r10.4) and reads generated subsequently trimmed and demultiplexed using Porechop software (v0.2.4). The ONT reads generated had mean read lengths (quality score) of 1.15 Kbp (Q18.2) and 1.17 Kbp (Q18.8) for Poc221 and Pow222 isolates, respectively. All sample specific long-read data obtained was subsequently combined for de novo assembly. In addition, WGS for 20 publicly available P. ovale spp. samples were incorporated in this study (ENA project accession: PRJEB51041).
WGS data quality control
All short read data was filtered via Trimmomatic software (v0.39; parameters: LEADING:3 TRAILING:3 SLIDINGWINDOW:4:20 MINLEN:36)34. Long read data was self-mapped via minimap2 (v2.17-r941; parameters: -x ava-ont -g 500)35, and scrubbed via yacrd (v0.6.2; parameters: default)36. Subsequently all sequence data was filtered via Kraken2 (v2.0.7-beta; parameters: default)37 and KrakenTools (v1.2; parameters; –exclude –taxid 2 2157 10239 9443 –include-children)38, to remove all bacteria, archaea, virus, and primate classified reads. Subsequently all filtered data was mapped to the human reference genome (GRCh38), utilizing minimap2 (v2.17-r941; parameters: -ax map-ont) for long read data and bwa mem (0.7.17-r1188; parameters: default). Following mapping, the samtools software suite was used to extract all reads which were not mapped in a proper pair (v;1.16.1 parameters: view -F 2)39,40. Genotypes at biallelic SNPs common to Poc and Pow were called using GATK, and a principal component analysis was performed to confirm the separation of each species.
Hybrid genome assembly and annotation
Each stage of the hybrid assembled pipeline used to create the two new references is summarised (Supplementary Figure S1). Initial assemblies were created using Spades software (v3.13.0) under hybrid settings41. Contigs was subsequently extended via NTLink (v1.3.8)42 and SSPACE (v2.0)43 software followed by Pilon (v1.24)44 and sniffles (v2.0.7) based polishing45. The contigs were then scaffolded via RagTag (v2.1.0)46 using the PocGH01 reference as a guide. Gaps in the assembled scaffolds were subsequently closed via Abyss (v2.0.2)47, TGS-GapCloser (v1.1.1)48 and GapFiller (v1.11)49 software tools. Mitochondrial and Apicoplast associated scaffolds were circularized via Circlator (v1.5.5) software50, followed by another round of Pilon-based polishing. Each assembly then went through an initial round of annotation via Companion51 under default parameters with the prior PocGH01 annotation as a guide. Core orthologs were identified OrthoFinder (v2.5.4)52, relative to other Plasmodium species. The assemblies were subsequently manually filtered to remove duplicate and misassembled genes based on annotation and coverage metrics. Following manual filtering, scaffolds were split into contigs based on gapped positions and all stages from RagTag (v2.1.0) (43) onwards were rerun. The final assemblies were annotated using both Companion and MetaEUK software (v5.34c21f2)53, again using the existing PocGH01 annotation as a guide under default parameters and subsequently assessed via BUSCO v5.3.0, using plasmodium_odb database (2020-08-05). Reference genome summaries were made using the python package pyCircos (v.1.0.2) and custom python scripts which can be found at (https://github.com/MatthewHiggins2017/HigginsMPovaleReferences).
Ortholog identification
OrthoFinder software (v2.5.4), was utilised to identify orthogroups between the 4 P. ovale references (Poc221, Pow222, PocGH01 (PlasmoDB: v54), PowCR01 (NCBI: GCA_900090025.2)) and 13 other Plasmodium species; P. berghei ANKA, P. chabaudi chabaudi, P. cynomolgi M, P. falciparum 3D7, P. gallinaceum 8A, P. knowlesi H, P. malariae UG01, P. reichenowi CDC, P. vinckei brucechwatti DA, P. vinckei lentum DE, P. vinckei vinckei CY, P. vivax P01, and P. yoelii yoelii 17X. PIR and STP1 multigene family members were extracted directly from the general feature format (GFF) file for each P. ovale spp. reference.
Phylogenetic analysis
Single copy core orthologroups represented in Poc221 and Pow222 were subsequently extracted and aligned using Mafft software54 under default settings. Each alignment was then processed using the GBlocks software55 under default settings to remove gapped and uninformative positions. All alignments were subsequently combined to form a sequence covering 1,850,442 amino acids. This combined sequence was used to construct multiple maximum likelihood phylogenetic trees via bootstrapping with RAXML-ng software56 utilizing LG substitution model a with gamma distribution. A conserved tree structure for was identified and subsequently visualized via ITOL57.
Genomic Islands of homology
Genomic islands of homology between the new and existing references were identified using progressive Mauve (v2.4.0) software58. The program was run with default “seed families” and default values for all other parameters. The same parameters were also used in identified islands of homology between the new Poc221 and Pow222 references.
Reference validation mapping and variant calling
All Illumina raw sequencing data was filtered using trimmomatic software (v0.39; parameters: LEADING:3 TRAILING:3 SLIDINGWINDOW:4:20 MINLEN:36). Filtered reads were then mapped to the respective new and existing reference genome (Poc221 and PocGH01 or Pow222 and PowCR01) using BWA-MEM alignment (v0.7.17) software. Mapping and coverage statistics were extracted using BWA STATs (v0.7.17) and Samtools (v1.16.1) respectively. SNPs and insertions/deletions (indels) were found using GATK’s HaplotypeCaller (v4.1.4.1)59, and subsequently filtering was performed using the GATK VariantFiltration parameters (filter "QD < 2.0" "QUAL < 30.0" "SOR > 3.0" "FS > 60.0" "MQ < 40.0" -"MQRankSum < -12.5" "ReadPosRankSum < -8.0″). SNP density was calculated using vcftools (v0.1.15)60. Statistical tests to compare mapping metrics were performed using the Scipy python package61. Species-specific SNPs in the mitochondrial cytB gene were extracted, and based on using > 90% of sites in a sample, the isolates were correctly classified into Poc and Pow.
Data availability
All raw sequence data is available from NCBI (project accession number PRJNA1015456). All sample accession codes are lists in Supplementary Table S6. The new reference genomes and all associated files can be found at https://github.com/MatthewHiggins2017/HigginsMPovaleReferences/.
References
Fuehrer, H.-P., Campino, S. & Sutherland, C. J. The primate malaria parasites Plasmodium malariae, Plasmodium brasilianum and Plasmodium ovale spp.: Genomic insights into distribution, dispersal and host transitions. Malar. J. 21, 138 (2022).
Sutherland, C. J. et al. Two nonrecombining sympatric forms of the human malaria parasite Plasmodium ovale occur globally. J. Infect. Dis. 201, 1544–1550 (2010).
Kotepui, M., Kotepui, K. U., Milanez, G. D. & Masangkay, F. R. Severity and mortality of severe Plasmodium ovale infection: A systematic review and meta-analysis. PLoS One 15, e0235014 (2020).
Snounou, G., Sharp, P. M. & Culleton, R. The two parasite species formerly known as Plasmodium ovale. Trends Parasitol. 40, 21–27 (2024).
Šlapeta, J., Sutherland, C. J. & Fuehrer, H.-P. Calling them names: Variants of Plasmodium ovale. Trends Parasitol. https://doi.org/10.1016/j.pt.2023.12.010 (2023).
Mahittikorn, A., Masangkay, F. R., Kotepui, K. U., Milanez, G. D. J. & Kotepui, M. Comparison of Plasmodium ovale curtisi and Plasmodium ovale wallikeri infections by a meta-analysis approach. Sci. Rep. 11, 6409 (2021).
Fançony, C. et al. Various pfcrt and pfmdr1 genotypes of Plasmodium falciparum cocirculate with P. malariae, P. ovale spp., and P. vivax in northern Angola. Antimicrob. Agents Chemother. 56, 5271–5277 (2012).
Doderer-Lang, C. et al. The ears of the African elephant: Unexpected high seroprevalence of Plasmodium ovale and Plasmodium malariae in healthy populations in Western Africa. Malar. J. 13, 240 (2014).
Fuehrer, H.-P. et al. Indigenous Plasmodium ovale malaria in Bangladesh. Am. J. Trop. Med. Hyg. 83, 75–78 (2010).
Nguyen, H. T. T. et al. Case report: Diagnostic challenges in the detection of a mixed plasmodium vivax/ovale infection in a non-endemic setting. Am. J. Trop. Med. Hyg. 103, 1085–1087 (2020).
Chavatte, J.-M., Tan, S. B. H., Snounou, G. & Lin, R. T. P. V. Molecular characterization of misidentified Plasmodium ovale imported cases in Singapore. Malar. J. 14, 454 (2015).
Mitchell, C. L. et al. Under the radar: Epidemiology of plasmodium ovale in the democratic Republic of the Congo. J. Infect. Dis. 223, 1005–1014 (2021).
Tanizaki, R. et al. Performance of rapid diagnostic tests for plasmodium ovale malaria in Japanese travellers. Trop. Med. Health 42, 149–153 (2014).
Leonard, C. M. et al. Missed Plasmodium ovale infections among symptomatic persons in Angola, Mozambique, and Ethiopia. Open Forum Infect Dis 9, ofac261 (2022).
Talman, A. M. et al. Evaluation of the intra- and inter-specific genetic variability of Plasmodium lactate dehydrogenase. Malar. J. 6, 140 (2007).
Senn, H., Alattas, N., Boggild, A. K. & Morris, S. K. Mixed-species Plasmodium falciparum and Plasmodium ovale malaria in a paediatric returned traveller. Malar. J. 13, 78 (2014).
Gurevitch, J. & Laufer, A. A mixed infection with Plasmodium vivax and P. ovale revealed after splenectomy. Ann. Trop. Med. Parasitol. 46, 238–239 (1952).
Kim, G. et al. Mixed infection with plasmodium falciparum and plasmodium ovale in a returned traveller: The First Case in Korea. J. Korean Med. Sci. 34, e23 (2019).
Akala, H. M. et al. Plasmodium interspecies interactions during a period of increasing prevalence of Plasmodium ovale in symptomatic individuals seeking treatment: An observational study. Lancet Microbe 2, e141–e150 (2021).
Lone, S. A. & Ahmad, A. COVID-19 pandemic: An African perspective. Emerg. Microbes Infect. 9, 1300–1308 (2020).
Sicuri, E., Ramponi, F., Lopes-Rafegas, I. & Saúte, F. A broader perspective on the economics of malaria prevention and the potential impact of SARS-CoV-2. Nat. Commun. 13, 2676 (2022).
MalariaGEN, et al. An open dataset of Plasmodium falciparum genome variation in 7000 worldwide samples. Wellcome Open Res 6, 42 (2021).
Rutledge, G. G. et al. Plasmodium malariae and P. ovale genomes provide insights into malaria parasite evolution. Nature 542, 101–104 (2017).
Ansari, H. R. et al. Genome-scale comparison of expanded gene families in Plasmodium ovale wallikeri and Plasmodium ovale curtisi with Plasmodium malariae and with other Plasmodium species. Int. J. Parasitol. 46, 685–696 (2016).
Little, T. S. et al. Analysis of pir gene expression across the Plasmodium life cycle. Malar. J. 20, 445 (2021).
Joste, V., Guillochon, E., Clain, J., Coppée, R. & Houzé, S. Development and optimization of a selective whole-genome amplification to study plasmodium ovale Spp. Microbiol Spectr 10, e0072622 (2022).
Ye, R. et al. Genome-wide analysis of genetic diversity in plasmodium falciparum isolates from China-Myanmar border. Front. Genet. 10, 1065 (2019).
de Oliveira, T. C. et al. Genome-wide diversity and differentiation in New World populations of the human malaria parasite Plasmodium vivax. PLoS Negl. Trop. Dis. 11, e0005824 (2017).
Ibrahim, A. et al. Population-based genomic study of Plasmodium vivax malaria in seven Brazilian states and across South America. Lancet Reg Health Am 18, 100420 (2023).
Brashear, A. M. et al. A glance of the blood stage transcriptome of a Southeast Asian Plasmodium ovale isolate. PLoS Negl. Trop. Dis. 13, e0007850 (2019).
Sutherland, C. J. Persistent parasitism: The adaptive biology of malariae and ovale malaria. Trends Parasitol. 32, 808–819 (2016).
Clarke, E. L. et al. swga: A primer design toolkit for selective whole genome amplification. Bioinformatics 33, 2071–2077 (2017).
Ibrahim, A. et al. Selective whole genome amplification of Plasmodium malariae DNA from clinical samples reveals insights into population structure. Sci. Rep. 10, 10832 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Marijon, P., Chikhi, R. & Varré, J.-S. yacrd and fpa: Upstream tools for long-read genome assembly. Bioinformatics 36, 3894–3896 (2020).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Lu, J. et al. Metagenome analysis using the Kraken software suite. Nat. Protoc. 17, 2815–2839 (2022).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv [q-bio.GN] (2013).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Coombe, L., Warren, R. L., Wong, J., Nikolic, V. & Birol, I. ntLink: A toolkit for De Novo genome assembly scaffolding and mapping using long reads. Curr Protoc 3, e733 (2023).
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
Walker, B. J. et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963 (2014).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 23, 258 (2022).
Paulino, D. et al. Sealer: A scalable gap-closing application for finishing draft genomes. BMC Bioinform. 16, 230 (2015).
Xu, M. et al. TGS-GapCloser: A fast and accurate gap closer for large genomes with low coverage of error-prone long reads. Gigascience 9 (2020).
Nadalin, F., Vezzi, F. & Policriti, A. GapFiller: A de novo assembly approach to fill the gap within paired reads. BMC Bioinform. 13 Suppl 14, S8 (2012).
Hunt, M. et al. Circlator: Automated circularization of genome assemblies using long sequencing reads. Genome Biol. 16, 294 (2015).
Steinbiss, S. et al. Companion: A web server for annotation and analysis of parasite genomes. Nucleic Acids Res. 44, W29-34 (2016).
Emms, D. M. & Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Levy Karin, E., Mirdita, M. & Söding, J. MetaEuk-sensitive, high-throughput gene discovery, and annotation for large-scale eukaryotic metagenomics. Microbiome 8, 48 (2020).
Katoh, K., Misawa, K., Kuma, K.-I. & Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: A fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Darling, A. C. E., Mau, B., Blattner, F. R. & Perna, N. T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 14, 1394–1403 (2004).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. bioRxiv 201178 (2018) https://doi.org/10.1101/201178.
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Virtanen, P. et al. SciPy 1.0: Fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Acknowledgements
We wish to thank staff at the UKHSA Malaria Reference Laboratory. TGC and SC are funded by UKRI grants (ref. BB/X018156/1, MR/X005895/1, and EP/Y018842/1). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. DN and CJS are supported by the UKHSA contract for malaria reference service provision by the MRL at LSHTM.
Author information
Authors and Affiliations
Contributions
MH, CJS, TGC and SC conceived and designed the study. DN and CJS contributed parasite DNA for sequencing. MH and SC coordinated the sequencing of samples. EM, DW, and JEP contributed bioinformatic tools. MH performed the bioinformatic and statistical analysis, under the supervision of SC and TGC. MH wrote the first draft of the manuscript, and the final version included edits from all authors. The final manuscript was read and approved by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Higgins, M., Manko, E., Ward, D. et al. New reference genomes to distinguish the sympatric malaria parasites, Plasmodium ovale curtisi and Plasmodium ovale wallikeri. Sci Rep 14, 3843 (2024). https://doi.org/10.1038/s41598-024-54382-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54382-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.