Abstract
The Tambaqui is one of the most representative Amazon fish species, being highly exploited in fisheries, aquaculture and as a research model. Nonetheless, data about functional genome are still required to evaluate reproductive and nutrition parameters as well as resistance to pathogens. The of next-generation sequencing has allows assessing the transcriptional processes in non-model species by providing comprehensive gene collections to be used as a database in further genomic applications and increased performance of captive populations. In this study, we relied on RNAseq approach to generate the first transcriptome of the telencephalon from adult males and females of Colossoma macropomum, resulting in a reference dataset for future functional studies. We retrieved 896,238 transcripts, including the identification of 267,785 contigs and 203,790 genes. From this total, 91 transcripts were differentially expressed, being 63 and 28 of them positively regulated for females and males, respectively. The functional annotation resulted in a library of 40 candidate genes for females and 20 for males. The functional enrichment classes comprised reproductive processes (GO:0,048,609; GO:0,003,006; GO:0,044,703; GO:0,032,504; GO:0,019,953) being related to sex differentiation (e.g., SAFB) and immune response (e.g., SLC2A6, AHNAK, NLRC3, NLRP3 and IgC MHC I alpha3), thus indicating that the genes in the neurotranscriptome of Tambaqui participate in sex differentiation and homeostasis of captive specimens. These data are useful to design the selection of genes related to sex determination and animal welfare in raising systems of Tambaqui.
Similar content being viewed by others
Introduction
The Tambaqui is the most produced native fish in Brazilian aquaculture1 because of some favorable traits, such as tolerance to hypoxic waters, resistance to pathogens, easy production of fingerlings and rapid growth in captivity2. Despite this, there is still huge potential to expand its production1. Genomic studies represent potential strategies to improve aquaculture systems since the data generated by advanced sequencing and bioinformatic approaches can be successfully used in selection and breeding programs3,4,5
In fact, the evaluation of genes that play a key role in the metabolism of several fish species, including Colossoma macropomum, has been already carried out. These reports provide a database for advanced research to be tested in improvement of fish culture6,7. For instance, functional genome information based on transcriptome of distinct tissues from Tambaqui is available, representing a promising tool for the development of captive stocks, such as evaluation of expressed genes in specimens exposed to pathogens and distinct climate conditions or genetic pathways related to breeding, nutrition and physical traits8,9,10,11,12,13.
A diversified transcriptome dataset is essential to the full understanding of animal physiology inasmuch as the gene expression will vary according to the analyzed tissue, development stage, sex and environmental conditions of each species14. In particular, brain tissue has been widely studied in fishes since the neurotranscriptome analyses has provided several insights about behavioral patterns. Nonetheless, little is known about the genes expressed in distinct areas of encephalon in Tambaqui. A report by10 analyzed the transcriptome of telencephalon of Tambaqui raised under distinct conditions of physical activity, revealing that specimens exposed to exercises were characterized by an increased number of positively regulated genes related to memory and learning as well as faster growth. However, this transcriptome was obtained from juvenile specimens while data about adult and sexually mature fish remain unpublished.
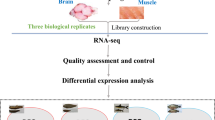
A variety of hormones are regulated by neuroendocrine signals that exhibit complex reaction cascades, like sex steroids, neuroendocrine-immune interactions, and stress response15,16,17,18,19. These mechanisms may play different roles in organisms and may be associated with other unclear functions in Tambaqui. Furthermore, the captive specimens are exposed to stressful environments distinct from those found in wild. Even after replicating natural conditions such as water currents, the natural breeding of Tambaqui has not been accomplished in captivity and positive regulation of genes related to stress stimuli has been reported10. Therefore, taking into account the importance of Tambaqui in aquaculture and the reliability of next generation molecular data in analyses of functional genomics, the present study provided useful information for further research by generating the first functional annotation of differentially expressed genes in the Neurotranscriptoma of adult males and females of C. macropomum. A graphical summary is available in Fig. 1, showing the main steps.
Results
Characterization of the dataset
The seven sequenced libraries of Tambaqui telencephalon comprised from 245,166.54 to 707,867.80 reads, with a mean value of 13,903,652.29 per library. After trimming, these values ranged from 229,145.61 to 681,344.40 reads with a mean GC content of 44% and length between 10 and 149 bp (Table 1).
The compiled libraries generated a total of 107,450,263.00 raw sequences that after trimming were reduced to 97,325,566.00 reads with an overall alignment rate of 97.71%. These values and other measurements of this dataset are shown in Table 3. The assembled reads using de novo method retrieved a transcriptome composed of 896,238 transcripts and 267,785 contigs, including isoforms (Table 2) and N50 values described in Supplementary Material 1.
Functional annotation of transcriptome
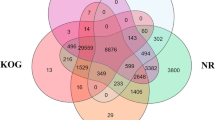
In order to provide an enhanced description of the Tambaqui transcriptome, the 267,785 contigs were compared to the Swiss-Prot and RefSeq datasets. As a result, we retrieved 203,790.00 significant equivalent sequences (76.10%) related to 67 GO terms (Fig. 2A), totaling 533,155 putative genes distributed among the three Gene Ontology (GO) classes. When comparing the ontology terms of the Tambaqui with those of the zebra fish, we obtained 50,947 and 30,568 respectively (Fig. 2B).
Most of GO terms were related to Molecular Function (MF) (178,971), encompassing 16 terms, being followed by Biological Process (BP) (178,545 distributed into 31 categories), and Cellular Component (CC) (175,639) corresponding to 20 GO terms (Table 3).
The classes with the highest correspondence within BP were the cellular processes (77.3%), metabolic processes (63.1%) and biological regulation (39.5%). As for the CC class, the cell (77.9%), cell part (77.9%) and organelles (32.4%) were the most representative GO terms. In turn, most of terms within MF were related to binding (68.3%), catalytic activity (49.5%) and transport activity (6.9%) (Fig. 3).
Taking into account the BP class, the following GO terms should be pointed out: regulation of immune system processes (GO:0,002,682), response to stimuli (GO:0,009,719, GO:0,009,607, GO:0,009,628), response to stress (GO:0,006,950), development of the immune system (GO:0,002,520), reproductive processes in multicellular organisms (GO:0,048,609), development of processes involved in breeding (GO:0,003,006), reproductive processes in multicellular organisms (GO:0,044,703), reproduction of multicellular organisms (GO:0,032,504) and sexual reproduction (GO:0,019,953).
Analysis of differential expression
The correlation matrix taking into account the 267,785.00 contigs revealed significant clusters among individuals, separating males and females into two distinct closely related groups (Fig. 4A). When the relative expression levels of each transcript in each sample was compared between male and female clusters, we identified 91 differentially expressed transcripts. The results based on heatmap analyses showed that the transcripts of each sex form separate clades, including 63 upregulated transcripts for females and 28 transcripts related to the males of C. macropomum (Fig. 4B). Supplementary materials 2:
The expression patterns in transcripts were significantly different between males and females. A total of 61 out of the 63 transcripts (96.82%) were expressed in females (reaching expression values of up to 20,107) but not in males. Even though two transcripts were shared between both groups, their expression values were highly divergent, ranging from 30,170 to 269,017 in females and from 0.000 to 9,257 in males (Supplementary materials 3: Table S1).
In turn, 26 out of the 28 upregulated sequences in males (92.85%) were expressed exclusively in this group, with expression values varying from 0.816 to 38,412. Two of these transcripts were also expressed in females but at extremely low levels (0.000 to 2,025) while they were overexpressed in males (values between 3,385 and 27,220) (Supplementary materials 3: Table S1).
Functional annotation of DEGs in males and females of Tambaqui
To obtain a detailed analysis of DEGs, we evaluated the functional annotation of DEGs separately, resulting in 34 GO classes corresponding to Biological Processes (14), Molecular Function (5) and Cellular Component (15) (Fig. 5). The most representative GOs in MF were binding, catalytic activity and transport activity. As for the CC classes, the most relevant GO terms were cell, cell part and organelles. In for BP, cellular process, localization and metabolic processes were the predominant terms. We also highlight the role of putative genes related to reproduction, immune system and response to stimuli.
Considering the abovementioned GO terms, we added the description of 60 novel candidate genes. The females represented the group with the highest number of transcripts corresponding to putative genes, totaling 40 equivalences from the 63 upregulated transcripts. On the other hand, 20 matches were reported among the 28 upregulated transcripts of males (Supplementary materials 4: Table S2). Additional information about the functional annotation based on Blast2Go is available in Supplementary materials 5: Table S3.
The candidate genes identified in the present study play distinct roles in several and important biological pathways. For instance, SLC2A6, AHNAK, NLRC3 and IgC MHC I alpha3 are involved in inborn immunity, response to inflammation and other immune processes regulated by inflammatory stimuli. After a refined manual inspection of the present dataset, we identified upregulated (TRINITY_DN286_c26_g1) sequences in females, characterized by a similarity of 95% with the XM_017693343.2 sequence from Pygocentrus nattereri via NCBI BlastX. This sequence corresponds to the Factor B for Scaffold Attachment (SAFB), a gene that plays a key role as a co-repressor of alpha receptors of estrogen, an important female sex hormone.
Discussion
Transcriptome as source of information
The RNASeq method provides a wide coverage of non-repetitive genes and their isoforms, thus allowing accurate measures of expression profiles in transcriptome and analyses of phenotypic variation among individuals14,20,21,22. In fact, these sequences have been added to public databases worldwide being particularly helpful to describe and predict the role of thousands of proteins that take part in major biological pathways. Therefore, novel data about these genes are essential to relate the retrieved sequences to their putative functions4,23.
In addition, the sequencing method and the experimental design might directly influence the amount and the quality of retrieved reads24. In the present study, a significant number of reads with adequate coverage was generated inasmuch as libraries composed of up to 24,516,654 reads were obtained, as well as an assembled transcriptome characterized by 97% of aligned sequences and a high number of annotated genes (203.790). These results indicate the Illumina platform and the sampling strategy was successful to the goals of this work, thus determining satisfactory levels of functional annotation.
Actually25, reported that Illumina MiSeq presented some advantages when compared to other available platforms at that time, particular in relation to the quality parameters. Nonetheless, the Illumina platform demands a high computational effort to analyses of large datasets24. Therefore, the development of additional methods that refine these data by reducing the number of sequences used in gene annotation, such as differential expression approaches, has been important to generate fast discrimination of experimental groups26.
Differential expression: males and females
This is the first transcriptome generated from the telencephalon of adult specimens of Tambaqui. As a portion of the central nervous system, the telencephalon which participates in several mechanisms linked to the emotional state and endocrine system of fishes, including reproduction and animal homeostasis27.
The transcripts expressed in the neurotranscriptome of Tambaqui generated a divergent pattern between males and females, revealing the importance of telencephalon as a major source of information to differential gene expression between sexes28. On average, 98.8% of transcripts were overexpressed exclusively in one group but not in the other, while females were characterized by a larger number of differentially expressed genes or DEGs (63). In contrast29, suggested that upregulated transcripts tend to be more numerous in brain of males, thus differing from our results. Even though these authors analyzed a similar number of libraries when compared to the present work, most values referred to the male brain library (86,112,076 reads) what could lead to a biased amount of DEGs in male tissue. Furthermore, the increased amount of DEGs in females of Tambaqui reported in our study represents an important landmark to investigate genes related to sex determination.
The identification of transcripts related to major reproductive pathways in the neurotranscriptome of Tambaqui indicates that the brain of this species might encompass additional information that can be used to elucidate the mechanisms of sex differentiation in C. macropomum. Therefore, further studies are recommended to seek sex-related genes beyond those related to DEGs, which is accomplishable even for non-model organisms. For example29, successfully annotated 82% and 90% of upregulated transcripts in the brain transcriptome of females and males of tropical gar, respectively.
Several research related to the early differentiation between males and females have been carried out in farm animals, particularly related to increase market gains, since a specific sex might be more advantageous than the other in animal production, as the masculinization of Tilapia females to increase the fish weight30. In the case of Tambaqui, the females present fast growth rates, with weight gains up to 16% higher than those reported in males, making the females more attractive for the production in captivity [31; 32].
In our study, females weighed 1.4 kg, 1.5 kg and 2.6 kg, and were most likely at the same stage of gonadal development, since females of this species, when bred in captivity, present late gonadal development compared to males, starting with approximately 1,200 kg, and do not complete their maturation before 3 kg, which contributes to these showing greater growth31.
The mechanisms involved in the reproductive process and sex determination of fishes are particularly diversified, unlike other groups of vertebrates, such as mammals. These aspects are not phylogenetically conserved and distinct strategies have evolved even in closely related species. Such plasticity of sex differentiation in teleosteans is directly related to sex steroid hormones, possibly regulated by specific genes expressed in brain networks18,28.
The Tambaqui is a rheophilic fish, therefore, environmental stimuli are extremely important for their reproduction. When these fish are cultivated in the northern region of Brazil, males can reach sexual maturity at any time of the year, however, it is believed that the lack of environmental stimuli inhibits the production of progestin, or produce little, which prevents spawning from occurring in captivity31.
Accordingly, GO terms associated with fish reproduction (GO:0,048,609; GO:0,003,006; GO:0,044,703; GO:0,032,504; GO:0,019,953) were presently observed among the metabolic pathways in the neurotranscriptome of Tambaqui. The presence of these transcripts is expected since major hormones are produced and/or activated in the hypothalamus. Some of these genes have already been reported in zebrafish, a model species, like arginine vasotocin receptors33 and gonadotropin-releasing hormone 3 receptor34, but these and some other genes were not differentially expressed in Tambaqui neurotranscriptome.
Most likely, this evidence is explained by the fact that the Tambaqui is a non-model organism in spite of their economic importance and hence the lack of information about this species hindered a more precise functional annotation. Recently13, identified that cyp19a1a and cyp19a1b genes would play a key role in the differentiation of males, but the genes related to similar processes in females are largely unknown. In the present study, high gene expression values were observed for the SAFB in females (11,496; 1,526; 5,939) while it remained unexpressed in males. The SAFB is highly expressed in brain tissues, encoding a co-repressor of estrogen receptors that interact with alpha receptors (ERα). In mice, the overexpression of SAFB in brain was associated with a normal distribution of gonadal steroid hormones at specific development stages, besides playing a putative role in the establishment of genitalia, formation of dimorphic brain structures and reproductive sex behavior35.
Therefore, the presence of SAFB in females of Tambaqui is a strong indicator that this gene is somehow related to the sex differentiation of females. The estrogen receptor (ER) exhibits a close relationship with 17β-estradiol, a hormone used in the feminization process of captive Tambaqui, which is co-expressed along with SAFB and induces significant changes in sex differentiation36. The Tambaqui is a rheophilic fish in which the release of steroid hormones and spawning are related to upstream migration under normal conditions in wild. The presence of overexpressed SAFB co-repressor in captive females may be a key feature that prevents the natural female maturation in culture systems, since maturity has only been achieved in females fed on diets rich in 17β-estradiol36.
Stress- and immune-related genes
In fish, the hypothalamic-pituitary-interrenal (HPI) system participates in the mechanisms of response to stress by promoting a cascade of reactions that triggers the release of adrenocorticotropin (ACTH), thus stimulating the production of cortisol to assist in the regulation of the homeostasis of both organs and tissues37. As a matter of fact, the functional annotation in the present study revealed several GO terms in the Biological Processes associated with the responses to stress and stimuli. Captive fishes are particularly susceptible to adverse conditions such as hypoxia, high stock densities and poor water quality during transportation, translocation to fish tanks, changes in diets during growth among others38,39,40.
In addition, fish farms are usually established close to large animal culture systems thus being potentially affected by pollution and contamination from these systems eventually leading to effects on physiological and biochemical parameters of captive fish stocks41. Because of these impacts, studies focused on animal welfare have increased over the last years and thus understanding the genetic mechanisms underlying these pathways is extremely relevant.
The changes of environmental conditions in captivity are stressful to aquatic organisms, thus activating their defense immune responses42. In fact, analyses of neurotranscriptome have already been useful to identify stress mechanisms in fishes43,44,45. The functional annotation in the present study identified the following genes related to fish immune system: NLRP (3 and 12) and IgC MHC I alpha3.
Two domains (3 and 12) of the NLRP gene were annotated in this work. This gene has already been widely reported in mammalian species it has been expressed in fish samples exposed to infectious agents, such as in the Japanese sole after bacterial infection46, and in the mucus of Seriola dumerili parasitized by Neobenedenia girlellae47. Similarly, the upregulated expression of IgC MHC I alpha 3 has also been related to exposure of fishes to pathogens, as reported in the transcriptome of specimens of Salmo salar after viral infection48. Thus, the expression of these genes in the neurotranscriptome of Tambaqui suggests a metabolic response of captive specimens to microorganisms from their environment in order to maintain animal homeostasis.
Even though this research was focused on the characterization of expression profiles, these results provide a baseline for further studies to the development of control markers related to animal welfare. This approach might be useful to direct proper strategies against parasites, contaminants and stressful agents that could jeopardize the animal growth.
The goal of this study was to assemble, predict and discuss the differential expression in transcripts of the neurotranscriptome in adult Tambaqui in order to provide a database that can guide further sex-related gene selection strategies. The neurotranscriptome herein reported encompassed 91 differentially expressed genes related to major ontology classes, such as reproductive process, stress response and immune system. In particular, an upregulated SAFB gene was identified in females, was identified, which putatively plays a key role in the sex differentiation of females, as well as candidate genes related to the immune system, such as the NLRP (3 and 12) and the IgC MHC I alpha3. In future, additional research focusing on detailing the role of these genes can contribute to the development of improved practices in the production of captive Tambaqui such as the formation of feminized stocks and reduced environmental stress in culture systems.
Material and methods
Ethics statement and sampling
The specimens used in this study were collected and transported according to the license granted by Instituto Chico Mendes da Biodiversidade (ICMBio, n. 60,833–1). Prior to the collection of biological material, the animals were anesthetized by immersion in a solution containing a concentration of 200 mg/L of benzocaine and water49 and slaughtered by section of spinal cord and major blood vessels according to the procedures approved by the National Counsel of Animal Experimentation Animal (CONCEA, law 11.794) and by the Committee of Experimentation and Use of Animals (CEUA, n. 7,399,260,721). We also declare that the experimental design and the entire study were carried considering the ARRIVE Guidelines (available at: https://arriveguidelines.org).
The specimens were collected during the rainy season (Amazonian winter). However, all were obtained from a commercial fish farm, where they were in controlled conditions of captivity. Therefore, seasonal variations that occur due to changes in seasons, they would not be affecting the physiology and metabolism of these fish, and consequently the animals' Transcriptome.
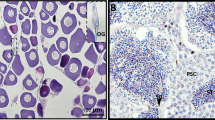
A portion of the telencephalon was selected for the present study. The tissues were obtained from seven adult (four males and three females) specimens of C. macropomum raised under the same management conditions in the fish station “Menino Deus”, municipality of Bonito, PA, northern Brazil (01°18′45.6″ S 047°21′51.3″ W). All specimens were weighed and measured (total length and total height) (Table 4). Besides the removal of telencephalon for RNA isolation, a muscle fragment was obtained from each fish for isolation of genomic DNA. The telencephalon samples were placed individually in 10-mL tubes containing RNA-preservative solution (RNA later® RNA Stabilization Solution, Ambion, Thermo Fisher, USA) and stored at − 20 °C. The muscle samples were stored in 2-mL Eppendorf tubes with 100% ethanol.
Identification of pure and hybrid specimens
Total DNA was isolated from the muscle fragments using the Wizard Genomic DNA purification kit (Promega, Madison, WI, USA). The quality of the DNA was visualized via electrophoresis in 1% agarose gel, stained with blue juice and red gel. The DNA samples were submitted to multiplex PCR for identification of pure or hybrid specimens according to the band profile after amplification of the alpha-Tropomyosin nuclear gene, as described by50. The genotyping confirmed that all samples corresponded to pure specimens of C. macropomum.
Isolation of RNA, construction of libraries and next-generation sequencing
The total RNA was isolated from 30 mg of from telencephalon tissue macerated in liquid nitrogen using the RNeasy Mini kit protocol (© QIAGEN). The quality and quantification (ng/µl) of RNA samples was evaluated in a Nanodrop 1000 (Thermo Scientific) and integrity in a 2100 Bioanalyzer Instrument (Table 4).
The RNA molecules were then fragmented and transcribed into cDNA. Individual libraries were constructed per cDNA sample, starting with 2000 ng of total RNA, using the SureSelect Strand Specific RNA Library Prep ILM Kit (Agilent Technologies) according to the manufacturer's protocol. In this process, unique barcodes for each sample and the adapters compatible with the sequencing platform were added, thus allowing the samples to be grouped and then individually identified (Table 4). Then, the libraries were transferred to the flow cell of the Illumina HiSeq 2500 platform, using the TruSeq SBS v3-HS kit. The procedures from RNA isolation to sequencing that resulted in raw sequence reads, paired-end: 2 × 100, of three female and four male replicates was carried out in the Genetic Center of the National Institute of Cancer José de Alencar Gomes da Silva (INCA) in Rio de Janeiro. The raw sequence reads were submitted to the Sequence Read Archive (SRA) database of the National Center for Biotechnology Information (NCBI) (accession code: PRJNA1016488).
Pre-processing and de novo assembly
The raw sequences generated in Illumina platform were clustered and converted into FastQ files according to their index and barcode tags. The quality parameters of these sequences were analyzed individually in the software FastQC v 0.1851. Afterwards, the adapters, short or low-quality reads were discarded using the software Trimmomatic v. 0.4052, according to the trimming criteria reported by53 for non-model organisms.
After removing the low-quality reads, the contigs were assembled using de novo approach in default parameters of the software Trinity 2.13.254,55. Since one of the major goals of this study was to analyze the differentially expressed genes (DEG) between males and females of Tambaqui, the de novo methodology is highly recommended56 since it allows identifying the most relevant DEG when compared to Ab initio approaches, even though a reference genome is already available for C. macropomum.
Analysis of differential expression and functional annotation
Based on the assembled telencephalon transcriptome, we performed a comparative analysis by separating the biological replicates into two groups (males and females). The quantification of gene expression was carried out mapping back the assembled transcriptome to the trimmed RNA seq data through software Salmon 1.8.057. The generated matrix of gene expressing obtained from the quantification step was used for the differential expression analysis using EdgeR v. 2.26.058 implemented in R package v. 4.1.0 (2019), thus resulting in the identification of differentially expressed transcripts. For expression value normalization in this study, we employed the Trimmed Mean of M-values (TMM) method, which was also utilized for hierarchical cluster analysis. The selection of TMM was based on its ability to address variations in library size and composition, ensuring a more accurate representation of gene expression levels55,59.
Two strategies were used for the functional annotation of transcripts: (1) annotation of total transcriptome, and (2) annotation of DEGs. Therefore, the files were compared to distinct sequence datasets to provide the most complete coverage of functional annotation, starting from those available in UniProtKB/Swiss-Prot and UniProtKB/TrEMBL (Release 2022_11) and complemented by non-redundant databases from NCBI, Refseq protein, Uniref, GenPept, NCBI ntm and Refseq RNA (all accessed on November, 18, 2022), via Blast2GO60, considering E-values of 1e-3, and a minimum coverage of 90%. For sequences identified across multiple databases, we prioritized the best hit based on the identification criteria described earlier.
As for the DEGs, we carried out a manual inspection of hits. The unrecorded transcripts were manually annotated after searches in GeneCards (https://www.genecards.org) dataset to check the identified gene followed by search of similar gene sequences in AmiGO Gene Ontology61, using the database available in the model species Danio rerio (zebrafish) as reference. The official gene symbols were confirmed by the HUGO Gene Nomenclature Committee (HGNC) and NCBI gene database. The gene classes identified in Gene Ontology were then inserted in the annotation file.
At last, we related each transcript from both total transcriptome and DEGs to the each one of the three GO classes: BP, MF and CC (Gene Ontology Consortium). The resulting graphs were plotted using the online tool WEGO 2.0 available in https://wego.genomics.cn62. Using the same software, we also built a graph relating the annotation and the GO terms retrieved in Tambaqui to those reported in zebrafish to provide a comparative scenario between both fish species.
Data availability
Accession codes: PRJNA1016488. Submitted data will be made available after publication. Reviewers can access it via the following link: https://dataview.ncbi.nlm.nih.gov/object/PRJNA1016488?reviewer=ac8607v0k50tqfdl4i4vlgvadt.
References
Peixe, Br. 2023. Anuário 2023 Peixe Br da Piscicultura. A força do peixe brasileiro. Associação Brasileira de Piscicultura. https://www.peixebr.com.br/anuario/ (2023).
Morais, I. D. S. & O’Sullivan, F. D. A. Biologia, habitat e cultivo do tambaqui Colossoma macropomum (CUVIER, 1816). Sci. Amazon. 6(1), 81–93 (2017).
Morozova, O. & Marra, M. A. Applications of next-generation sequencing technologies in functional genomics. Genomics 92(5), 255–264. https://doi.org/10.1016/j.ygeno.2008.07.001 (2008).
Houston, R. D. et al. Harnessing genomics to fast-track genetic improvement in aquaculture. Nat. Rev. Genet. 21(7), 389–409. https://doi.org/10.1038/s41576-020-0227-y (2020).
Guan, W. Z. & Qiu, G. F. Transcriptome analysis of the growth performance of hybrid mandarin fish after food conversion. PloS One 15(10), e0240308. https://doi.org/10.1371/journal.pone.0240308 (2020).
Machado, A. M., Ferraz, R., de Ribeiro, A. R., Ozório, R. & Castro, L. F. C. From the Amazon: A comprehensive liver transcriptome dataset of the teleost fish tambaqui Colossoma macropomum. Data Br. 23, 103751. https://doi.org/10.1016/j.dib.2019.103751 (2019).
Varela, E. S. et al. A high-density linkage map and sex-linked markers for the Amazon Tambaqui Colossoma macropomum. BMC Genom. 22(1), 1–10. https://doi.org/10.1186/s12864-021-08037-8 (2021).
Prado-Lima, M. & Val, A. L. Transcriptomic characterization of tambaqui (Colossoma macropomum, Cuvier, 1818) exposed to three climate change scenarios. PLoS One. 11(3), e0152366. https://doi.org/10.1371/journal.pone.0152366 (2016).
Ariede, R. B. et al. Microsatellites associated with growth performance and analysis of resistance to Aeromonas hydrophila in tambaqui Colossoma macropomum. Front. Genet. 9(3), 1–8. https://doi.org/10.3389/fgene.2018.00003 (2018).
Pereira, P. D. et al. Environmental enrichment improved learning and memory, increased telencephalic cell proliferation, and induced differential gene expression in Colossoma macropomum. Front. Pharmacol. 11(840), 1–14. https://doi.org/10.3389/fphar.2020.00840 (2020).
Ferraz, R. B. et al. The fatty acid elongation genes elovl4a and elovl4b are present and functional in the genome of tambaqui (Colossoma macropomum). Comp. Biochem. Physiol. B Biochem. Mol. Biol. 245, 110447. https://doi.org/10.1016/j.cbpb.2020.110447 (2020).
da Costa, J. C., de Souza, S. S. & Val, A. L. Impact of high temperature, CO2 and parasitic infection on inflammation, immunodepression and programmed cell death in Colossoma macropomum at the transcriptional level. Microb. Pathog. 172, 105804. https://doi.org/10.1016/j.micpath.2022.105804 (2022).
Paixão, R. V. et al. Phylogenomic and expression analysis of Colossoma macropomum cyp19a1a and cyp19a1b and their non-classical role in tambaqui sex differentiation. Gene 843, 146795. https://doi.org/10.1093/nar/gkv1189 (2022).
Wickramasinghe, S., Cánovas, A., Rincón, G. & Medrano, J. F. RNA-sequencing: a tool to explore new frontiers in animal genetics. Livest. Sci. 166, 206–216. https://doi.org/10.1016/j.livsci.2014.06.015 (2014).
Holloway, A. C. & Leatherland, J. F. Neuroendocrine regulation of growth hormone secretion in teleost fishes with emphasis on the involvement of gonadal sex steroids. Rev. Fish Biol. Fish. 8, 409–429. https://doi.org/10.1023/A:1008824723747 (1998).
Verburg-Van Kemenade, B. L., Stolte, E. H., Metz, J. R. & Chadzinska, M. Neuroendocrine–immune interactions in teleost fish. Fish. Physiol. 28, 313–364. https://doi.org/10.1016/S1546-5098(09)28007-1 (2009).
Cerdá-Reverter, J. M. & Canosa, L. F. Neuroendocrine systems of the fish brain. Fish. Physiol. 28, 3–74. https://doi.org/10.1016/S1546-5098(09)28001-0 (2009).
Godwin, J. Neuroendocrinology of sexual plasticity in teleost fishes. Front. Neuroendocrinol. 31(2), 203–216. https://doi.org/10.1016/j.yfrne.2010.02.002 (2010).
Nardocci, G. et al. Neuroendocrine mechanisms for immune system regulation during stress in fish. Fish Shellfish Immunol. 40(2), 531–538. https://doi.org/10.1016/j.fsi.2014.08.001 (2014).
Bushmanova, E., Antipov, D., Lapidus, A. & Prjibelski, A. D. RnaSPAdes: a de novo transcriptome assembler and its application to RNA-Seq data. GigaScience 8(9), 1–13. https://doi.org/10.1093/gigascience/giz100 (2019).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 10(1), 57–63. https://doi.org/10.1038/nrg2484 (2009).
García-Nieto, P. E., Wang, B. & Fraser, H. B. Transcriptome diversity is a systematic source of variation in RNA-sequencing data. PLoS Comput. Biol. 18(3), e1009939. https://doi.org/10.1371/journal.pcbi.1009939 (2022).
Jensen, P. Behavior genetics and the domestication of animals. Annu. Rev. Anim. Biosci. 2(1), 85–104. https://doi.org/10.1146/annurev-animal-022513-114135 (2014).
Hong, M. et al. RNA sequencing: New technologies and applications in cancer research. J. Hematol. Oncol. 13(1), 1–16. https://doi.org/10.1186/s13045-020-01005-x (2020).
Quail, M. A. et al. A tale of three next generation sequencing platforms: Comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genom. 13(341), 1–13. https://doi.org/10.1186/1471-2164-13-341 (2012).
Robles, J. A. et al. Efficient experimental design and analysis strategies for the detection of differential expression using RNA-Sequencing. BMC Genom. 13(484), 1–14. https://doi.org/10.1186/1471-2164-13-484 (2012).
Sado, R. Y., Souza, F. C., Behr, E. R., Mocha, P. R. E. & Baldisserotto, B. Anatomy of Teleosts and elasmobranchs. In Biology and Physiology of Freshwater Neotropical Fish (eds Baldisserotto, B. et al.) 21–47 (Elsevier, 2020). https://doi.org/10.1016/B978-0-12-815872-2.00002-6.
Wong, R. Y., McLeod, M. M. & Godwin, J. Limited sex-biased neural gene expression patterns across strains in Zebrafish (Danio rerio). BMC Genom. 15(905), 1–9. https://doi.org/10.1186/1471-2164-15-905 (2014).
Cribbin, K. M., Quackenbush, C. R., Taylor, K., Arias-Rodriguez, L. & Kelley, J. L. Sex-specific differences in transcriptome profiles of brain and muscle tissue of the tropical gar. BMC Genom. 18(283), 1–9. https://doi.org/10.1186/s12864-017-3652-3 (2017).
Habibah, A. N. et al. Growth and gonadal development of female Nile tilapia (Oreochromis niloticus) exposed to sex reversing thermal treatment. Aquaculture 531, 735865. https://doi.org/10.1016/j.aquaculture.2020.735865 (2021).
Almeida, F. D. et al. Early puberty of farmed tambaqui (Colossoma macropomum): Possible influence of male sexual maturation on harvest weight. Aquaculture 452, 224–232. https://doi.org/10.1016/j.aquaculture.2015.10.031 (2016).
Lobo, I. K. C. et al. Transcriptome of tambaqui Colossoma macropomum during gonad differentiation: Different molecular signals leading to sex identity. Genomics 112(3), 2478–2488. https://doi.org/10.1016/j.ygeno.2020.01.022 (2020).
Iwasaki, K., Taguchi, M., Bonkowsky, J. L. & Kuwada, J. Y. Expression of arginine vasotocin receptors in the developing zebrafish CNS. Gene Expr. Patterns. 13(8), 335–342. https://doi.org/10.1016/j.gep.2013.06.005 (2013).
Marvel, M., Levavi-Sivan, B., Wong, T. T., Zmora, N. & Zohar, Y. Gnrh2 maintains reproduction in fasting zebrafish through dynamic neuronal projection changes and regulation of gonadotropin synthesis, oogenesis, and reproductive behaviors. Sci. Rep. 11(1), 1–16. https://doi.org/10.1038/s41598-021-86018-3 (2021).
Hashimoto, T. et al. Scaffold attachment factor B: Distribution and interaction with Erα in the rat brain. Histochem. Cell. Biol. 153, 323–338. https://doi.org/10.1007/s00418-020-01853-1 (2020).
Reis, V. R. & Almeida, F. L. Effect of 17β-oestradiol on the sex ratio of tambaqui Colossoma macropomum. Aquac. Res. 50(1), 154–161. https://doi.org/10.1111/are.13878 (2019).
Konstantin, A. D. et al. Understanding neurobehavioral effects of acute and chronic stress in zebrafish. Stress 24(1), 1–18. https://doi.org/10.1080/10253890.2020.1724948 (2021).
Odhiambo, E., Angienda, P. O., Okoth, P. & Onyango, D. Stocking density induced stress on plasma cortisol and whole blood glucose concentration in Nile tilapia fish (Oreochromis niloticus) of lake Victoria, Kenya. Int. J. Zool. 2020, 1–8. https://doi.org/10.1155/2020/9395268 (2020).
Abdel-Tawwab, M., Monier, M. N., Hoseinifar, S. H. & Faggio, C. Fish response to hypoxia stress: Growth, physiological, and immunological biomarkers. Fish Physiol. Biochem. 45, 997–1013. https://doi.org/10.1007/s10695-019-00614-9 (2019).
Segner, H. et al. Welfare of fishes in aquaculture. FAO Fish. Aquac. Circ. 2019, 1–18 (2019).
Ahmed, I., Reshi, Q. M. & Fazio, F. The influence of the endogenous and exogenous factors on hematological parameters in different fish species: A review. Aquac. Int. 28, 869–899. https://doi.org/10.1007/s10499-019-00501-3 (2020).
Urbinat, E. C., Zanuzzo, F. S. & Biller-Takahashi, J. D. Stress and immune system in fish. In Biology and Physiology of Freshwater Neotropical Fish (eds Baldisserotto, B. et al.) 93–114 (Elsevier, 2020). https://doi.org/10.1016/B978-0-12-815872-2.00002-6.
Liu, L., Zhang, R., Wang, X., Zhu, H. & Tian, Z. Transcriptome analysis reveals molecular mechanisms responsive to acute cold stress in the tropical stenothermal fish tiger barb (Puntius tetrazona). BMC Genom. 21(1), 1–14. https://doi.org/10.1186/s12864-020-07139-z (2020).
Dai, Y. F. et al. RNA-seq transcriptome analysis of the liver and brain of the black carp (Mylopharyngodon piceus) during fasting. Mar. Biotechnol. 23, 389–401. https://doi.org/10.1007/s10126-021-10032-9 (2021).
Shang, F. et al. Transcriptome analysis identifies key metabolic changes in the brain of Takifugu rubripes in response to chronic hypoxia. Genes 13(8), 1347. https://doi.org/10.3390/genes13081347 (2022).
Chen, H. et al. Characterization of the Japanese flounder NLRP3 inflammasome in restricting Edwardsiella piscicida colonization in vivo. Fish Shellfish Immunol. 103, 169–180. https://doi.org/10.1016/j.fsi.2020.04.063 (2020).
Fernández-Montero, A. et al. Proteomic profile and protease activity in the skin mucus of greater amberjack (Seriola dumerili) infected with the ectoparasite Neobenedenia girellae—an immunological approach. Fish Shellfish Immunol. 110, 100–115. https://doi.org/10.1016/j.fsi.2021.01.001 (2021).
Samsing, F. et al. Transcriptome response of Atlantic salmon (Salmo salar) to a new piscine orthomyxovirus. Pathogens 9(10), 807. https://doi.org/10.3390/pathogens9100807 (2020).
Botelho, H. A., Costa, A. C., de Freitas, R. T. F. & Fernandes, É. M. Bromatological analysis of filet pacu (Piaractus mesopotamicus), pirapitinga (Piaractus brachypomum) and tambaqui (Colossoma macropomum)). Rev. Ciên. Vet. Saúde Públ. 5(2), 158–165 (2017).
Gomes, F. et al. Innovative molecular approach to the identification of Colossoma macropomum and its hybrids. An. Acad. Bras. Cienc. 84, 517–526. https://doi.org/10.1590/S0001-37652012005000025 (2012).
Andrews S. A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30(15), 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
MacManes, M. D. On the optimal trimming of high-throughput mRNA sequence data. Front. Genet. 5, 13. https://doi.org/10.3389/fgene.2014.00013 (2014).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29(7), 644–652. https://doi.org/10.1038/nbt.1883 (2011).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the trinity platform for reference generation and analysis. Nat. Protoc. 8(8), 1494–1512. https://doi.org/10.1038/nprot.2013.084 (2013).
Wang, S. & Gribskov, M. Comprehensive evaluation of de novo transcriptome assembly programs and their effects on differential gene expression analysis. Bioinformatics 33(3), 327–333. https://doi.org/10.1093/bioinformatics/btw625 (2017).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14(4), 417–419. https://doi.org/10.1038/nmeth.4197 (2017).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1), 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome biol. 11(3), 1–9. https://doi.org/10.1186/gb-2010-11-3-r25 (2010).
Conesa, A. et al. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18), 3674–3676. https://doi.org/10.1093/bioinformatics/bti610 (2005).
Carbon, S. et al. AmiGO: Online access to ontology and annotation data. Bioinformatics 25(2), 288–289. https://doi.org/10.1093/bioinformatics/btn615 (2009).
Ye, J. et al. WEGO 2.0: A web tool for analyzing and plotting GO annotations, 2018 update. Nucl. Acids Res. 46(W1), W71–W75. https://doi.org/10.1093/nar/gky400 (2018).
Acknowledgements
Our thanks to “Instituto Nacional de Câncer José de Alencar Gomes da Silva” (INCA) for generating the next-generation sequencing data and to Prof. Horacio Schneider (in memory) for the valuable contribution.
Funding
The financial support to this study was granted by CNPq (process n. 439113/2018-0) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—CAPES (PhD scholarship, grant n. 001). PAPQ-Propesp/Federal University of Pará (UFPA).
Author information
Authors and Affiliations
Contributions
J.M. Conceptualization, Validation, Writing—Original Draft, Writing—Review & Editing. I.V. Methodology, Investigation. C.F. Visualization. P.S. Visualization. I.L. Visualization. C.F. Methodology, Investigation. P.P. Formal analysis, Data Curation. L.R. Formal analysis, Data Curation. C.G-D. Visualization. M. M. Visualization. I.S. Visualization, Resources. M.V. Visualization, Resources. G.E–G. Supervision, Writing—Original Draft, Writing—Review & Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Miranda, J., Veneza, I., Ferreira, C. et al. First neurotranscriptome of adults Tambaquis (Colossoma macropomum) with characterization and differential expression between males and females. Sci Rep 14, 3130 (2024). https://doi.org/10.1038/s41598-024-53734-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-53734-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.