Abstract
Asynchronously cycling cells pose a challenge to the accurate characterization of phase-specific gene expression. Current strategies, including RNAseq, survey the steady state gene expression across the cell cycle and are inherently limited by their inability to resolve dynamic gene regulatory networks. Single cell RNAseq (scRNAseq) can identify different cell cycle transcriptomes if enough cycling cells are present, however some cells are not amenable to scRNAseq. Therefore, we merged two powerful strategies, the CDT1 and GMNN degrons used in Fluorescent Ubiquitination-based Cell Cycle Indicator (FUCCI) cell cycle sensors and the ribosomal protein epitope tagging used in RiboTrap/Tag technologies to isolate cell cycle phase-specific mRNA for sequencing. The resulting cell cycle dependent, tagged ribosomal proteins (ccTaggedRP) were differentially expressed during the cell cycle, had similar subcellular locations as endogenous ribosomal proteins, incorporated into ribosomes and polysomes, and facilitated the recovery of cell cycle phase-specific RNA for sequencing. ccTaggedRP has broad applications to investigate phase-specific gene expression in complex cell populations.
Similar content being viewed by others
Introduction
Dynamic changes in gene expression influence the ability of cells to maintain identity, adapt to environmental conditions during homeostasis, respond to ligand-mediated signals, and coordinate growth. Organs and tissues are composed of heterogeneous populations of cells that can obfuscate the interpretation of gene expression profiles of specific cell subpopulations. Strategies to profile cell-specific gene expression include the physical isolation of cells of interest by laser capture microdissection or dissociation of single cells followed by cell sorting or antibody-based enrichment (immunopanning)1. Other strategies, including RiboTrap or RiboTag, employ the expression of transgenic constructs encoding epitope-tagged ribosomal proteins (for example, RPL10a or RPL22) to immunoprecipitate mRNAs from complex tissues2,3,4. TU-tagging uses the spatially restricted uracil phosphoribosyltransferase (UPRT) expression with 4-thiouracil (4TU) delivery to label and purify cell type specific-RNA from intact complex tissues5,6,7. Despite the advantage of purifying RNAs from specific cell types within complex tissues, these strategies track steady state gene expression across the cell cycle and are thus inherently limited by their inability to identify changes specific to cell phases.
More recent advances in single cell RNAseq (scRNAseq) have facilitated the molecular characterization of diverse cell types within tissues, organs, and whole animals8,9,10,11,12,13,14,15. There are, however, limitations to scRNAseq. scRNAseq methods require the physical dissociation of cells, a process known to alter the transcriptional landscape during the isolation process. These technologies also tend to survey the steady state gene expression across the cell cycle, limiting the ability to identify gene regulatory networks specific to G1, S, and G2/M in vivo. Moreover, scRNAseq can pose unique problems when attempting to isolate cell cycle-specific transcripts. Firstly, some cell types are not amenable to single cell sorting (i.e., cardiomyocytes) and subsequent single cell RNA sequencing16. Secondly, subpopulations can be too rare to adequately discern from artifact. Thirdly, low sequencing depth can limit the ability to accurately characterize cell subpopulations17,18. Finally, scRNAseq is significantly more expensive than bulk RNA sequencing. Alternative strategies are necessary to address some of these limitations.
Degron-mediated degradation is one mechanism used to regulate the expression of proteins during phases of the cell cycle19,20,21,22,23. FUCCI technologies employ chimeras of fluorescent proteins and degrons to detect cell cycling in vitro and in vivo22,23. In general, FUCCI reporters rely on the expression of SCFskp2 and APCCdh1 to regulate Chromatin Licensing and DNA Replication Factor 1 (CDT1) and Geminin (GMNN) degrons. Second generation bicistronic FUCCI constructs contain internal protein cleavage sites, such as T2A from Thosea asigna virus 2A, to initially produce equimolar amounts of reporter proteins, whose subsequent levels are a function of the cell cycle phase via degron-mediated proteolysis22.
In this study, we merged two powerful strategies, the CDT1 and GMNN degrons of FUCCI and the epitope-tagged ribosomal proteins of RiboTrap/Tag technologies, to create cell cycle-dependent, tagged ribosomes named ccTaggedRP with the goal of discerning cell cycle phase-specific gene expression changes. The ccTaggedRPs were incorporated into ribosomes and polysomes as determined by sucrose gradient centrifugation, had similar subcellular location as endogenous RPL10a by immunofluorescence, and demonstrated predicted changes in protein expression that were dependent on the phase of the cell cycle. Here, we describe our first generation ccTaggedRP and employed the strategy to identify transcripts enriched during G2-M phase from asynchronously cycling HEK293-T cells. The findings also corresponded to transcripts identified by scRNAseq datasets of HEK293-T cells, suggesting that ccTaggedRP is a potential new strategy to detect differentially expressed transcripts during the cell cycle.
Results:
ccTaggedRPs are differentially expressed during the cell cycle and have similar subcellular locations as endogenous ribosomal proteins after transient transfection
To test the hypothesis that chimeras of epitope-tagged ribosomal proteins containing degrons would add new functionality to existing RiboTrap/Tag technology, we created a bicistronic 3xFlagTag-RPL10a-hCdt(30–120)-T2A-3xHA-Tag-RPL10a-hGeminin(1–110) (denoted “ccTaggedRP” hereafter). The encoded ccTaggedRP is translated as a single polypeptide and undergoes cleavage at the T2A site to initially produce 3xFlagTag-RPL10a-hCdt(30–120) and 3xHA-Tag-RPL10a-hGeminin(1–110) in equimolar amounts (Fig. 1A). However, the 3′ hCdt insertion targets the 3xFlagTag-RPL10a-hCdt(30–120) recombinant protein (denoted “Flag-RPL10a-hCdt” hereafter) for degradation during late S through G2 and M phase while the 3’ hGeminin insertion targets the 3xHA-Tag-RPL10a-hGeminin recombinant protein (denoted “HA-RPLA10a-hGem” hereafter) for degradation during late G1 through early S phase. Consequently, the steady-state concentrations of HA-RPL10a-hGem and Flag-RPL10a-hCdt recombinant proteins are dependent on the cell cycle phase. Specifically, Flag-RPL10a-hCdt and HA-RPL10a-hGem recombinant proteins are expected to be enriched in G0/G1 and G2-M, respectively (Fig. 1B). Of note, unlike FUCCI, the ccTaggedRP constructs do not contain fluorescent reporter proteins. The Flag-RPL10a-hCdt and HA-RPL10a-hGem recombinant proteins were expressed at the predicted sizes after transient transfection of HEK293-T cells with recombinant ccTaggedRP plasmid (Fig. 1C). Protein bands of lower molecular weight were observed in the Flag and HA blots, consistent with degradation products mediated by the hCdt(30–120) and hGeminin(1–110) degrons, respectively. Immunofluorescent staining of HEK-293 cells transiently transfected with ccTaggedRP showed similar patterns of Flag and RPL10a locations. Suggesting that the subcellular location of ccTaggedRP was similar to endogenous RPL10a in HEK293-T cells (Fig. 1D).
ccTaggedRPs are differentially expressed during the cell cycle and have similar subcellular locations as endogenous ribosomal proteins. (A) Schematic of the CAG-3xFlagTag-RPL10a-hCdt(30–120)-T2A-3xHA-Tag-RPL10a-hGeminin(1–110) construct and cleavage at the T2A site. (B) Schematic of the predicted change in 3xFlagTag-RPL10a-hCdt(30–120) and 3xHA-Tag-RPL10a-hGeminin(1–110) during the cell cycle. (C) Immunoblots of asynchronously cycling HEK293-T cells transiently transfected with empty plasmid, ccTaggedRP, or not transfected. Blots were probed with antibodies against Flag Tag, HA Tag, or mRPL10a. Bands corresponding to Flag recombinant protein, HA recombinant protein, and endogenous RPL10a isoforms are denoted by arrows. See Supplemental Fig. 6 for uncut blots. (D) Immunofluorescence of asynchronously cycling HEK293-T cells transiently transfected with ccTaggedRP stained for RPL10a and Flag Tag. Cells expressing ccTaggedRP are denoted by arrows. Cells not expressing ccTaggedRP are denoted by asterisks. DAPI was used to stain for nuclear DNA. DAPI is blue, RPL10a is red, and FLAG is green. Scale bars are included in the lower right corner and correspond to 10 μm.
ccTaggedRPs stably expressed in HEK293-T cells were incorporated into endogenous ribosomes and polysomes
To control for the copy number of recombinant DNA, we created the stable expression of ccTaggedRP through Flp-In insertion of a single copy of ccTaggedRP into HEK293-T cells (Fig. 2A). Immunofluorescent staining of Flag and HA identified distinct cells expressing Flag-RPLa10-hCdt alone, Flag-RPL10a-hCdt and HA RPL10a-hGem, and HA-RPLa10-hGem alone (Fig. 2b). The majority of cells were only positive for Flag-RPLa10-hCdt alone, consistent with G1 phase of the cell cycle.
(A) Immunoblots of asynchronously cycling HEK293-T cells transiently transfected with ccTaggedRP and HEK293-T-FlpIn cells stably expressing ccTaggedRP probed with antibodies to Flag Tag or HA Tag. See Supplemental Fig. 7 for uncut blots. (B) Immunofluorescence of HEK293-T-FlpIn cells expressing ccTaggedRPs showing anti-Flag and anti-HA antibody statins, and DAPI. A merged image is shown. Cells denoted 1, 2, 3 and 4 express Flag-RPL10a-hCdt and HA-RPL10a-hGem, cell denote 5 expressed HA-RPL10a-hGem alone. All other cells express Flag-RPL10a-hCdt alone. Scale bar in the lower right corner corresponds to 20 μm.
To evaluate the incorporation of recombinant ccTaggedRPs into ribosomes, we performed sucrose gradient centrifugation of lysates prepared from HEK-293 T cells stably expressing ccTaggedRPs. Cell lysates were fractionated on a 10% to 50% sucrose gradient while measuring the absorbance at 260 nm to detect ribosomal subunits (40S and 60S), in addition to 80S ribosomes, and translating ribosomes (polysomes) (Fig. 3A). Immunoblots of fractions identified the presence of recombinant Flag-RPL10a-hCdt and HA-RPL10a-hGem in the 60S large ribosomal subunits, 80S ribosomes, and polysomes, with a distribution similar to RPL10a. (Fig. 3B–D). Negative staining electron microscopy of fraction #5 demonstrated that 80S ribosomes were enriched in this fraction as expected (Fig. 3E). The findings support that Flag-RPL10a-hCdt and HA-RPL10a-hGem expressed from recombinant ccTaggedRP were incorporated into ribosomes and polysomes.
Ribosome analysis using sucrose gradient centrifugation of lysates from HEK293T-FlpIN cells stably expressing ccTaggedRPs. (A) Absorbance at 260 nm (A260) was measured during fractionation and showed distinct peaks corresponding to 40S and 60S subunits, 80S ribosomes, and polysomes. (B–D) Western blots corresponding to fractions from the sucrose gradient in panel A, using anti-Flag tag (B), anti-HA tag (C), and anti-RPL10a (denoted eRPL10a) (D) antibodies. (E) Negative stain electron microscopy showed enrichment of 80S ribosomes from fraction #5 of the corresponding A260 peak for 80S ribosomes. Scale bar is 100 nm. See Supplemental Fig. 9 for uncropped blots. Recombinant Flag-RPL10a-hCdt and HA-RPL10a-hGem were detected in fractions corresponding to the 60S ribosomal subunits, 80S ribosomes, and polysomes.
ccTaggedRP expression changes during cell cycle synchronization
The HEK293-T Flp-In cells expressing ccTaggedRP were synchronized via serum starvation, which is known to increase the percentage of cells in G0/G1 as starvation persists. In accordance with expectations, the expression of Flag-RPL10a-hCdt recombinant protein was progressively increased with starvation, suggesting an enrichment in the Flag-RPL10a-hCdt construct in G0/G1 while the expression of HA-RPL10a-hGem recombinant protein progressively decreased, suggesting an enrichment in HA-RPL10a-hGem construct in G2/M. (Supplemental Fig. 1A, B). Specifically, compared to serum-fed HEK293-T Flp-In cells, HA-RPL10a-hGem expression decreased by 82% after 48 h of serum starvation. (Supplemental Fig. 1C).
To further validate the cell cycle dependence of Flag-RPL10a-hCdt and HA-RPL10a-hGem expression, HEK293-T Flp-In cells expressing ccTaggedRP were synchronized by treatment with Lovastatin, Hydroxyurea or Nocodazole and enrichment in G1, S, and G2/M were confirmed by fluorescent activated cell sorting (FACS) (Fig. 4A–C and Supplemental Fig. 2). Cells treated with Lovastatin were 80.7% G1, 0.5% S, and 18.8% G2/M. Hydroxyurea were 25.1% G1, 57.7% S, and 17.2% G2/M. Cell treated with Nocodazole were 19.5% G1, 5.2% S, and 75.3% G2/M (Fig. 4B). Based on the FACS data and predicted ideal behavior of the degrons with Flag expressed in G1 and S and HA expressed in S and G2/M, the relative changes in Flag and HA expression were estimated (Fig. 4C). Western blotting of cells treated with Lovastatin, Hydroxyurea, or Nocodazole showed reciprocal changes in the expression of Flag-RPL10a-hCdt and HA-RPL10a-hGem (Fig. 4d). Flag-RPL10a-hCdt was increased and decreased in expression in G1 and G2M phases, respectively. HA-RPL10a-hCdt was decreased and increased in expression in G1 and G2M phases, respectively.
The expression of Flag-RPL10a-hCdt and HA-RPL10a-hGem is dependent on the phase of the cell cycle. (A) FACS quantitation of HEK293-T-Flp-In cells stably expressing ccTaggedRPs in G1, S, and G2/M treated with after synchronization with Lovastatin, Hydroxyurea or Nocodazole. The percentage of cells in G1, S, and G2/M under each treatment is shown. B525/50-A EdU AF488 represents EdU intensity and V450/50-A DAPI represents DNA content. These measurements were used to gate cells in G1, S, and G2M. (B) Quantitation of G1, S, and G2/M of cells treated with Lovastatin, Hydroxyurea, or Nocodazole. Values are mean ± SEM. P-values were calculated by ANOVA with Tukey test for multiple comparisons. N = 4 plates per condition. (C) Predicted relative expression of Flag and HA based on the results shown in panel B. (D) Western blots of Flag-RPL10a-hCdt (top), HA-RPL10a-hGem (middle), and GAPDH (bottom) corresponding to the cells treated with Lovastatin, Hydroxyurea, or Nocodazole and analyzed by FACS in panel A. See Supplemental Fig. 10 for uncropped blots.
ccTaggedRP identifies genes expressed during G2-M in asynchronously cycling cells
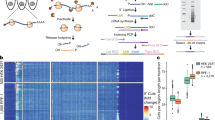
Asynchronous HEK293-T Flp-In cells stably expressing ccTaggedRP were cultured to 60% confluency prior to non-denaturing lysis. Recombinant Flag-RPL10a-hCdt and HA-RPL10a-hGem proteins were then immunoprecipitated (IP) using HA or Flag antibodies, and their associated RNAs were recovered for bulk RNA-sequencing (Fig. 5A). Principal component analysis (PCA) displayed distinct clustering of the HA- and Flag-IP samples (Fig. 5B). This method resolved 2,125 differentially expressed genes with an adjusted p-value ≤ 0.05 between the HA- and Flag-IP RNA samples (Fig. 5C and Supplemental Table 1). Transcripts enriched in the HA-IPs were associated with regulation of the mitotic cell cycle and G2-M transition of the mitotic cell cycle while transcripts enriched in the Flag-IPs were associated with the negative regulation of both the mitotic cell cycle and the G2-M transition of the mitotic cell cycle (Fig. 5D). Additionally, Gene Set Enrichment (GSE) Analysis identified enrichment of G2-M cell cycle and checkpoint pathways in the HA-IPs (Fig. 5E). Furthermore, expression of G2-M genes was significantly increased in the HA-IPs compared to the Flag-IPs (Fig. 5F).
ccTaggedRP identifies genes expressed during G2-M in asynchronously cycling cells. (A) Schematic of the immunoprecipitation of 3xFlagTag-RPL10a-hCdt(30–120) and 3xHA-Tag-RPL10a-hGeminin(1–110) for the purification and sequencing of RNA from asynchronously cycling HEK293-T-Flp-In cells expressing ccTaggedRP. (B) Principal component analysis (PCA) plot of RNA sequencing of mRNAs from immunoprecipitated Flag recombinant protein (red dots) and HA recombinant protein (blue dots). (C) Volcano plot of RNA sequencing of mRNAs from immunoprecipitated Flag recombinant protein and HA recombinant protein. The gray line denotes an adjusted p-value of 0.05. Genes with increased and decreased expression are denoted by red and blue dots, respectively. (D) Waterfall plot of the 1090 genes enriched in the 3xFlagTag-RPL10a-hCdt(30–120) and 1035 genes enriched in the 3xHA-Tag-RPL10a-hGeminin(1–110) immunoprecipitated samples. GO Terms are denoted with adjusted p-values shown. (E) Gene set enrichment (GSE) analysis of RNAseq of mRNAs from immunoprecipitated Flag recombinant protein and HA recombinant protein. Adjusted p-vales are shown. (F) Heat map of the expression of the top 50 G2-M genes from mRNAs from immunoprecipitated Flag recombinant protein and HA recombinant protein.
scRNAseq of HEK293-T cells identifies populations of G1-S and G2-M
To further validate ccTaggedRP, scRNAseq data for HEK293-T cells were sourced from 10× Genomics for comparison of transcripts expressed during G1/S and G2/M. Cells within the dataset were assigned to either “G1/S” or “G2/M” identities based on gene expression (Fig. 6A and Supplemental Figs. 3 and 4, and Supplemental Table 2). G1/S cells exhibited increased expression of MCM5, a protein important in DNA synthesis and upregulated in G1/S while G2/M cells exhibited increased expression of CCNB1, a protein important in cell cycle division and upregulated in G2/M (Fig. 6B, C). In this dataset, G1/S cells and G2/M cells accounted for 62.5% and 37.5% of the total number of cells, respectively (Fig. 6D). Differential expression analysis between the G1/S and G2/M cells revealed 530 differentially expressed genes with an adjusted p-value ≤ 0.05 (Fig. 6E). Hypergeometric testing of the differentially expressed genes indicated an enrichment in DNA replication processes and mitotic cell cycle processes for cells in G1/S and G2/M respectively (Fig. 6F, G). Furthermore, G2/M cells displayed increased expression of G2-M genes compared to G1/S cells (Fig. 6H).
scRNAseq of HEK293-T cells identifies populations of G1-S and G2-M. (A) UMAP of cells assigned to G1/S and G2/M identity in HEK293-T cell culture. (B,C) UMAP of MCM5 and CCNB1 expression. (D) Percentage of total cells assigned to G1/S and G2/M identities. (E) Volcano plot of differential gene expression between G1/S and G2/M clusters. (F,G) Gene Ontology results via hypergeometric testing of genes enriched in the G1/S cluster and G2/M cluster. (H) Heat map of the average expression for the top 40 G2/M genes between the G1/S and G2/M clusters.
Comparison of genes expressed in G2-M during asynchronous cycling from ccTaggedRP and scRNAseq
To investigate the differences between the two methodologies, functionalization of shared and method-specific differentially expressed genes was performed. Comparison of differentially expressed genes from the ccTaggedRP method and scRNAseq found 2019 genes specific to the ccTaggedRP method, 423 genes specific to the scRNAseq method, and 107 genes shared between the two (Supplemental Fig. 5A). Furthermore, hypergeometric testing of the shared differentially expressed genes indicated enrichment of genes associated with mitotic cell cycle processes (Supplemental Fig. 5B).
Discussion
We sought to develop an extension of the RiboTrap/RiboTag technologies that would facilitate the characterization of transcribed genes during distinct cell cycle phases of asynchronously cycling cells. RiboTrap and RiboTag use an EGFP-tagged RPL10a ribosomal protein subunit or HA-tagged RPL22 ribosomal protein subunit, respectively, for translating ribosome affinity purification (TRAP) of mRNA populations and characterization of gene expression2,3,4,24,25,26,27. Both methods have provided new insights regarding the gene expression of specific cell types in complex tissues. For example, RiboTrap has been used to investigate the differential gene expression of excitatory neurons and interneuron subtypes28, interstitial/pericytes, myeloid cells, nephron cells, and endothelial cells29 while Ribotag has been used to characterize gene expression in neurons3, microglia30, oocyte maturation31, and cardiomyocyte and endothelial cells4 in diverse disease states. Notably, the gene expression is obtained from bulk RNA sequencing of pooled mRNAs, meaning that the total pool of expressed transcripts dilutes the cell cycle phase-specific transcripts and modifies interpretation. To overcome this limitation, we describe the first-generation ccTaggedRP technology based on merging aspects of FUCCI and RiboTag methods. FUCCI employs degrons that target proteins for degradation during the G1/S or G2/M phases of the cell cycle22,23. By creating ccTaggedRP, which contains different epitope-tagged RPL10a proteins with either hCdt1 or hGem1 degrons, the mRNAs expressed during cell cycle phases can be enriched by TRAP. The initial polypeptide is bicistronic and cleaved into distinct tagged RPL10a chimeric proteins whose subsequent abundance is dictated by the phase of the cell cycle via hCdt1- and hGem-mediated degradation. Additionally, the targeted insertion of a single copy of ccTaggedRP in HEK293-T-Flp-In cells avoided the effects of multiple copy insertion or insertion into a transcriptionally silenced region of the genome, which could adversely affect the expression of ccTaggedRP.
3xFlagTag-RPL10a-hCdt(30–120) and 3xHA-Tag-RPL10a-hGeminin were differentially expressed during G1/S and G2/M based on synchronization of different phases of the cycle using inhibitors, had similar subcellular locations as endogenous ribosomal proteins, were incorporated into the 60S ribosomal subunit, 80S ribosomes, and polysomes, and facilitated TRAP of mRNAs expressed during G1/S and G2/M. Bulk RNA sequencing of mRNAs associated with the immunoprecipitation of 3xHA-Tag-RPL10a-hGeminin identified transcripts enriched in the regulation of the mitotic cell cycle and G2-M transition of the mitotic cell cycle. Conversely, the immunoprecipitation of 3xFlagTag-RPL10a-hCdt(30–120) identified transcripts enriched in the negative regulation of the mitotic cell cycle and the G2-M transition of the mitotic cell cycle. Collectively, our data suggests the predominance of the HA recombinant protein and Flag recombinant protein in the G2/M and G1/S phases of the cell cycle, respectively.
Next, we compared the results of TRAP using ccTaggedRP with scRNAseq results of HEK293-T cells obtained by 10× Genomics. Cells from the scRNAseq were assigned to either G1/S or G2/M phases based on their expression profiles. The differential expression between the cell cycle clusters was then compared to those obtained by ccTaggedRP. A comparison of differentially expressed genes from the ccTaggedRP method and scRNAseq found 2019 genes specific to the ccTaggedRP method, 423 genes specific to the scRNAseq method, and 107 genes shared between the two methods. Evaulation of the shared differential gene expression confirmed similar gene expression using the two methods; however, the ccTaggedRP approach resolved nearly four-fold more differentially expressed genes between conditions. Moreover, the transcript enrichment from the ccTaggedRP method was similar if not greater than that of the scRNAseq method. Additionally, the significantly lower cost of bulk RNAseq and its consequent opportunity for multiple biological replicates further bolster the potential of ccTaggedRP as an alternative to scRNAseq for resolving cell cycle-specific gene expression.
There are, however, limitations to the ccTaggedRP technology. First, the exchange rates of RPLs into and out of ribosomes are likely different for each RPL and could affect the efficiency of the ccTaggedRP technology (Supplemental Fig. 6). For example, RPLs with slower exchange rates may not degrade efficiently during cell cycle phases and influence the interpretation of TRAP data. Therefore, future ccTaggedRPs that use different RPLs will require careful characterization, given the dynamic properties of RPLs. This problem may be further exacerbated for rapidly cycling tissue, in which there is insufficient time for degradation of the constructs between phases. FUCCI constructs encode fluorescently labeled degrons and may be more amenable to degradation compared to Flag-RPL10a-hCdt and HA-RPL10a-hGem that are incorporated into ribosomes, potentially slowing their degradation. However, HA-RPL10a-hGem demonstrated more pronounced expression changes in response to the cell cycle. As presented in this study, we were able to resolve cell cycle-specific gene expression in a highly replicative embryonic kidney epithelial cell line, suggesting that the kinetics of the construct are sufficiently fast to support analyses in most contexts. Second, the heterogeneity of ribosomes could influence the interpretation of TRAP data32. Third, we acknowledge the presence of lower molecular weight HA and FLAG recombinant proteins, which are likely products of protein degradation. This is to be expected as each of the recombinant proteins have C-terminal degrons. Importantly, despite the presence of these degradation products, cell cycle-specific gene expression was still able to be discerned. Finally, we tested the technology in asynchronous cells in culture, a scenario that provided adequate signal detection. Translating the ccTaggedRP technology to a transgenic mouse for in vivo experiments may be challenging.
Despite these limitations, the ccTaggedRP method is promising and has great potential to identify cell cycle phase-specific gene expression in complex tissues. Moreover, it can be combined with other technologies to create even more powerful approaches. The ccTaggedRP construct could be inserted downstream of tissue-specific promoters to resolve cell cycle-specific gene expression in specific tissues. This may be particularly useful due to the high degree of contamination observed in other purification methods. Furthermore, in tissues with low replicative capacities, this methodology affords the ability to pool multiple biological samples to characterize these sparse populations. Alternatively, the methods could be adapted to create chimeras of epitope-tagged Argonaut proteins containing hCdt1 and hGem to survey the expression of microRNAs expressed during G1/S and G2/M in asynchronous cycling cells in complex tissues.
Methods details
The Flp-In HEK293-T cells stably expressing ccTaggedRP cell line and plasmid are available upon reasonable request to the corresponding author. The CAG-ccTaggedRP plasmid will be deposited into Addgene.
Creation of the ccTaggedRP construct
A DNA construct encoding 3× Flag-Tag, human Cdt1 (amino acid 30 to 120), T2A, 3× HA-Tag, and human Geminin (hGem (amino acids 1 to 110)) and containing 5P BglII and 3P NheI restriction sites was synthesized by IDT DNA, Inc. The hCdt1 and hGem DNA sequences were similar to the sequences used in Fucci2aR22. CMV promoter and multicloning sites were removed from pSF-CAG-Kan (plasmid OGS505, Sigma, Inc) using BglII and NheI and replaced with the sequence containing 3× Flag Tag, hCdt1(30–120), T2A, 3× HA Tag, and hGem(1–110). The DNA sequence corresponding to cDNA of mouse RPL10a was synthesized by IDT DNA, Inc, and modified to add 5P and 3P AvrII or 5P and 3P MluI restriction sites. The add 5P and 3P AvrII or 5P and 3P MluI RPL10a cassettes were subcloned into 3× Flag Tag, hCdt1(30–120), T2A, 3× HA Tag, and hGem(10–110) to create in frame 3× Flag Tag-RPL10a-Cdt1(30–120)-T2A-3× HA Tag-RPL10a-hGem(10–110) in the pSF plasmid. Then, a CAG promoter was subcloned into the 3× Flag Tag-RPL10a-hCdt1(30–120)-T2A-3× HA Tag-RPL10a-hGem(10–110) plasmid to create the CAG-ccTaggedRP construct. We made the Flp-In HEK293-T cells stably expressing ccTaggedRP using the ccTaggedRP DNA construct that subcloned into a plasmid modified to contain a CAG promoter and 3P FRT and Hygromycin sequences corresponding to pcDNA5/FRT (ThermoFisher, Inc.). The Flp-In HEK293-T cells were purchased from Invitrogen, Inc and the Flp-In HEK293-T cells stably expressing ccTaggedRP were made according to the manufacture’s protocol. The DNA sequences of all constructs were validated by Sanger sequencing.
Polysome profiling
Polysome profiling of HEK293T cells was conducted as described previously33. A 100 mg cell pellet was lysed with 500 µL of lysis buffer (20 mM HEPES–KOH pH 7.7, 100 mM KCl, 5 mM MgCl2, 0.5% IGEPAL, 1 mM DTT, 0.5× protease inhibitor cocktail (Promega), 4 units/mL RNase inhibitor (Promega), 100 µg/mL cycloheximide). The lysate was passed through a 23G needle four times and centrifuged at 11,200×g at 4 °C, for 8 min. The clarified lysate was loaded on top of a linear sucrose gradient (10–50% sucrose, 20 mM HEPES–KOH pH 7.7, 100 mM KCl, 10 mM MgCl2, 1 mM DTT). The sample was centrifuged at 230,000×g at 4 °C, for 2.5 h using an SW40 rotor. After centrifugation, a BioComp gradient profiler was used to fractionate the sucrose gradient at 0.2 mL/min into 800 µL fractions. The absorbance at 260 nm was measured during fractionation.
Negative stain EM
For negative stain electron microscopy, a 10 µL sample corresponding to the 80S ribosome peak (fraction 5) was applied directly after fractionation to a glow discharged 400-mesh grid (EMS, CF400-Cu-50) and incubated for 5 min at room temperature. The grid was stained with 2% uranyl acetate twice for 1 min each. Excess fluid was blotted away with Whatman filter paper after each incubation period (Cytiva, 1001-090). Images were collected on an FEI F20 electron microscope operated at 120 kV and equipped with a TVIPS TemCam XF416 camera. Images were taken at a magnification of 62,000×, and a pixel size of 0.17 nm.
Western blotting
For Western Blotting of sucrose gradient fractions, TCA (100% w/v) was added to each fraction at a final concentration of 10% and the samples were incubated for 30 min on ice. The samples were centrifuged at 13,000×g at 4 °C for 15 min. Pellets were washed with ice cold acetone and centrifuged at 13,000 xg for 10 min (2×). The pellets were dried, then resuspended in 100 µL of 1× SDS-PAGE sample buffer (50 mM Tris–HCl pH 6.8, 2% SDS, 6% glycerol, 0.004% bromophenol blue, 1% β-mercaptoethanol). Proteins were resolved on a NuPAGE 4–12% Bis–Tris minigel (Invitrogen, NP0323BOX) and transferred onto a 0.2 µm PVDF membrane (Invitrogen LC2002). The membrane was blocked in 3% BSA in 1× PBST (0.1% Tween-20) and probed with FLAG (Sigma, F1804), HA (Invitrogen, 26183), and RPL10a (Invitrogen, MA5-44710) antibodies diluted in 1% BSA in 1× PBST. Secondary anti-mouse (Invitrogen, A21058) and anti-rabbit (LI-COR, 926-32213) antibodies were used for detection with the LI-COR Odyssey imager.
For Western blotting of cell lysates, HEK293-T cells expressing ccTaggedRP were cultured in DMEM (CAT#:11965-092, ThermoFisher) supplemented with 10% fetal bovine serum (CAT#: 10438026, ThermoFisher) and 500 µg/mL Hygromycin B (CAT#: 10687010, ThermoFisher) until 80% confluency. Cells were washed once with Dulbecco's phosphate-buffered saline (DPBS) (CAT#: 14040133, ThermoFisher) and lysed in RIPA buffer (CAT#: 89900, ThermoFisher) supplemented with protease inhibitors (Roche, 04693159001). Protein concentration was quantified with the Pierce BCA kit (CAT#: 23225, ThermoFisher) and normalized. 4× loading buffer was added to samples. 25 µg of total protein was run on SDS gels (CAT#: 4561034, Bio-rad) and transferred to PVDF membranes with 45 µm pores (CAT#: 84-895, Prometheus). The membranes were blocked in 5% non-fat milk (Great Value) in Tris-buffered saline with 0.1% Tween 20 (TBST). Membranes were then incubated overnight with primary antibodies against FLAG (1:1000, CAT#: F1804, Invitrogen), HA (1:1000, CAT#: 26183, Invitrogen), or GAPDH (1:1000, CAT#: ab9485, Abcam) in 5% milk solution. Membranes were washed three times with TBST and subsequently incubated in TBST with HRP-conjugated secondary antibodies (CAT#: G-21234, Invitrogen). Membranes were washed twice with TBST and once with TBS. Chemiluminescent reaction was catalyzed by addition of 1:1 1 Ambersham ECL Detection Kit. Membranes were imaged on a Bio-Rad ChemiDoc Touch Imaging System (Bio-Rad, Inc.). Densitometry was quantified using Bio-Rad Image Lab 6.1 (Rio-Rad, Inc.).
Serum starvation of cultured cells
HEK293-T cells expressing ccTaggedRP were cultured in DMEM (CAT#:11965-092, ThermoFisher) supplemented with 10% fetal bovine serum (CAT#: 10438026, ThermoFisher) and 500 µg/mL Hygromycin B (CAT#: 10687010, ThermoFisher) until 80% confluency. Starved samples were then cultured in DMEM supplemented with only fetal bovine serum for the indicated amount of time.
Pharmacologic synchronization of cultured cells
HEK293-T cells expressing ccTaggedRP were cultured in Dulbecco's Modified Eagle Medium (DMEM) (CAT#:11965-092, ThermoFisher) supplemented with 10% fetal bovine serum (CAT#: 10438026, ThermoFisher) and 500 µg/mL Hygromycin B (CAT#: 10687010, ThermoFisher) until 70% confluency. Cells were then treated with vehicle, 100 μM Lovastatin (CAT#: 400046, Sigma) for 48 h, 20 µg/ml hydroxyurea (CAT#: 400046, Sigma) for 24 h, or 5 µg/ml nocodazole (M1404, Sigma) for 48 h.
Flow cytometry
Cells were incubated with 10 µM EdU for 1.5 h prior to harvest. Following harvest, cells were incubated with fixable LIVE/DEAD stain (CAT#: L34992, ThermoFischer) for 20 min in the dark and then washed with PBS. Following staining, cells were fixed with 4% paraformaldehyde (PFA) for 20 min and then were washed with 1% BSA. Cells were permeabilized and underwent the Click-it reaction according to manufacturer's instructions (CAT#: C10633, ThermoFischer). Cells were then washed with permeabilization buffer and stained with FxCycle Violet (CAT#: F10347, ThermoFischer) for 1 h in the dark. Data was obtained with a BD LSR Fortessa and analyzed with OMIQ (Dotmatics). Gating strategy is illustrated in Supplemental Fig. 2.
Immunofluorescence of cultured cells
HEK293-T cells expressing ccTaggedRP were cultured in DMEM (CAT#:11965-092, ThermoFisher) supplemented with 10% fetal bovine serum (CAT#: 10438026, ThermoFisher) and 500 µg/mL Hygromycin B (CAT#: 10687010, ThermoFisher) until 80% confluency. Medium was removed, and cells were fixed in 4% PFA. Cells were washed and incubated overnight in 1× PBS. Slides were then incubated at 90 °C in 1× unmasking solution (CAT#: H-3300, Vector Labs). Slides were cooled to room temperature and then washed three times with 1× PBS. Slides were blocked in 1× PBS containing fish skin gelatin oil (FSG) and Donkey Serum (CAT#: S30-100ML, Millipore Sigma) for 1 h at room temperature. Cell slides were incubated in primary antibodies against FLAG (CAT#: PAB29056, Abnova) and RPL10a (CAT#: PA5-62845, Invitrogen) in 1× PBS contacting FSG overnight at 4 °C. Slides were washed once with 1× PBS containing FSG and 0.1% Tween-20 and once with 1× PBS. Cells were then incubated with secondary antibodies against donkey anti-chicken (CAT#: 703-585-155, Jackson ImmunoResearch) and donkey anti-rabbit IgG (CAT#: 711-475-152, Jackson ImmunoResearch) in 1× PBS containing FSG and 0.1% Tween-20. Slides were washed once with 1× PBS containing FSG and 0.1% Tween-20 and once with 1× PBS. Cover slips were then mounted with VECTASHIELD® Antifade Mounting Medium with DAPI (H-1200-10, Vector Laboratories).
Cell culture, lysis, immunoprecipitations, and RNA recovery
HEK293-T cells expressing ccTaggedRP were cultured in Dulbecco's Modified Eagle Medium (DMEM) (CAT#:11965-092, ThermoFisher) supplemented with 10% fetal bovine serum (CAT#: 10438026, ThermoFisher) and 500 µg/mL Hygromycin B (CAT#: 10687010, ThermoFisher) until 70% confluency. Cells were washed with DPBS (CAT#: 14040133, ThermoFisher) and lysed in Pierce™ IP Lysis Buffer (CAT#: 87787, ThermoFischer) supplemented with 1× protease inhibitor (CAT#: 87785, ThermoFischer) and RNAse Inhibitor (CAT#: AM2682, Invitrogen). Lysed samples were incubated overnight with 5 µg of IP antibodies against FLAG (CAT#: F1804, Invitrogen) or HA (CAT#: 26183, Invitrogen). 0.25 mg of Pierce Protein A/G Magnetic Beads (CAT#: 88802, ThermoFischer) were added, and the resulting mixture was incubated overnight with rocking at 4 °C. Magnetic bead-target complexes were collected with a magnetic stand, and the remaining solution was aspirated. Samples were washed twice with IP lysis buffer and resuspended in RW1 buffer. RNA was then isolated with the RNeasy mini kit (CAT#: 74104, Qiagen) according to the manufacturer's instructions. Isolated mRNA samples were quantified and submitted to Genewiz for further processing and sequencing.
RNA sequencing analyses
cDNA library preparation with adapter ligation, and sequencing using an Illumina® HiSeq® system were performed by Genewiz, Inc. Reads underwent quality control, trimming, alignment, and quantification according to Genewiz standard workflows. Sequence reads were trimmed to remove possible adapter sequences and nucleotides with poor quality using Trimmomatic v.0.36. The trimmed reads were mapped to the Homo sapiens GRCh38 reference genome available on ENSEMBL using the STAR aligner v.2.5.2b. The STAR aligner is a splice aligner that detects splice junctions and incorporates them to help align the entire read sequences. BAM files were generated as a result of this step. Below are the statistics of mapping the reads to the reference genome. Genes with less than ten reads in at least three samples (the minimum sample size for a group) were excluded for subsequent analysis. Differential expression analysis was performed using R and the DEseq2 package. Functionalization was performed using ClusterProfiler package in R and GSEA software.
scRNAseq analysis
A scRNAseq dataset of cultured HEK293-T cells was sourced from 10× Genomics (10× Genomics). Sequencing yielded 737,280 cells with an average of 66.58 reads per cell and 19.45 genes per cell. Poor quality cells were removed, using the following inclusion criteria: cells with greater than 2000 genes, greater than 7500 reads, and less than 7.5% mitochondrial DNA. Cells were assigned a “G2/M” or “G1/S” identity using the CellCycleScoring algorithm in Seurat and previously published data sets (supplemental Fig. 7)34,35. Cells unable to be assigned confidently to either identity were discarded before downstream analysis. After filtering, 2447 cells remained with an average of 15,651.61 reads per cell and 3485.21 genes per cell. Data was normalized and scaled with the Seurat package in R. UMAP dimensionality reduction was performed with 11 principal components and a resolution of 0.5. Differential expression analysis based on the cell cycle cluster was performed using the Seurat package in R. Functionalization was performed using clusterProfiler package in R.
Quantification and statistical analysis
Shapiro Wilk test was performed on continuous variables to evaluate the data distribution. Normally-distributed data and non-normally distributed data were then analyzed for statistical significance by Student t test and Mann–Whitney U test, respectively. Statistical significance for RNAseq and scRNAseq data was determined through Benjamini–Hochberg procedure and reported as the adjusted p-value. All data is presented as mean ± standard deviation.
Inclusion and diversity
We support inclusive, diverse, and equitable conduct of research.
Data availability
All data reported in this paper will be shared upon request. Please contact the corresponding author, M.J.W., for access to data. The RNAseq data files have been deposited into Gene Expression Omnibus (GEO), Accession ID: GSE225385 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE225385). This paper does not report original code. Any additional information required to reanalyze the data deposited in this paper is available from the lead contact upon request.
References
Barres, B. A., Silverstein, B. E., Corey, D. P. & Chun, L. L. Immunological, morphological, and electrophysiological variation among retinal ganglion cells purified by panning. Neuron 1(9), 791–803 (1988).
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135(4), 738–748 (2008).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl. Acad. Sci. USA 106(33), 13939–13944 (2009).
Zhou, P. et al. Interrogating translational efficiency and lineage-specific transcriptomes using ribosome affinity purification. Proc. Natl. Acad. Sci. USA 110(38), 15395–15400 (2013).
Cleary, M. D., Meiering, C. D., Jan, E., Guymon, R. & Boothroyd, J. C. Biosynthetic labeling of RNA with uracil phosphoribosyltransferase allows cell-specific microarray analysis of mRNA synthesis and decay. Nat. Biotechnol. 23(2), 232–237 (2005).
Gay, L., Karfilis, K. V., Miller, M. R., Doe, C. Q. & Stankunas, K. Applying thiouracil tagging to mouse transcriptome analysis. Nat. Protoc. 9(2), 410–420 (2014).
Gay, L. et al. Mouse TU tagging: A chemical/genetic intersectional method for purifying cell type-specific nascent RNA. Genes Dev. 27(1), 98–115 (2013).
Briggs, J. A. et al. The dynamics of gene expression in vertebrate embryogenesis at single-cell resolution. Science 360(6392), 12 (2018).
Fincher, C. T., Wurtzel, O., de Hoog, T., Kravarik, K. M. & Reddien, P. W. Cell type transcriptome atlas for the planarian Schmidtea mediterranea. Science 360(6391), 1736 (2018).
Lescroart, F. et al. Defining the earliest step of cardiovascular lineage segregation by single-cell RNA-seq. Science 359(6380), 1177–1181 (2018).
Li, G. et al. Transcriptomic profiling maps anatomically patterned subpopulations among single embryonic cardiac cells. Dev. Cell 39(4), 491–507 (2016).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360(6390), 758–763 (2018).
Plass, M. et al. Cell type atlas and lineage tree of a whole complex animal by single-cell transcriptomics. Science. 360(6391), 1723 (2018).
Wagner, D. E. et al. Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360(6392), 981–987 (2018).
Wang, I. E., Wagner, D. E. & Reddien, P. W. Clonal analysis of planarian stem cells by subtotal irradiation and single-cell transplantation. Methods Mol. Biol. 1774, 479–495 (2018).
Litvinukova, M. et al. Cells of the adult human heart. Nature 588(7838), 466–472 (2020).
Haque, A., Engel, J., Teichmann, S. A. & Lonnberg, T. A practical guide to single-cell RNA-sequencing for biomedical research and clinical applications. Genome Med. 9(1), 75 (2017).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: A tutorial. Mol. Syst. Biol. 15(6), e8746 (2019).
Davey, N. E. & Morgan, D. O. Building a regulatory network with short linear sequence motifs: Lessons from the degrons of the anaphase-promoting complex. Mol. Cell. 64(1), 12–23 (2016).
Holt, L. J. Regulatory modules: Coupling protein stability to phopshoregulation during cell division. FEBS Lett. 586(17), 2773–2777 (2012).
Meszaros, B., Kumar, M., Gibson, T. J., Uyar, B. & Dosztanyi, Z. Degrons in cancer. Sci Signal. 10(470), 9982 (2017).
Mort, R. L. et al. Fucci2a: A bicistronic cell cycle reporter that allows Cre mediated tissue specific expression in mice. Cell Cycle 13(17), 2681–2696 (2014).
Sakaue-Sawano, A. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132(3), 487–498 (2008).
Chasse, H., Boulben, S., Costache, V., Cormier, P. & Morales, J. Analysis of translation using polysome profiling. Nucleic Acids Res. 45(3), e15 (2017).
Doyle, J. P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135(4), 749–762 (2008).
Heiman, M., Kulicke, R., Fenster, R. J., Greengard, P. & Heintz, N. Cell type-specific mRNA purification by translating ribosome affinity purification (TRAP). Nat. Protoc. 9(6), 1282–1291 (2014).
Knight, Z. A. et al. Molecular profiling of activated neurons by phosphorylated ribosome capture. Cell 151(5), 1126–1137 (2012).
Huntley, M. A. et al. Genome-wide analysis of differential gene expression and splicing in excitatory neurons and interneuron subtypes. J. Neurosci. 40(5), 958–973 (2020).
Miyazaki, T. et al. Cell-specific image-guided transcriptomics identifies complex injuries caused by ischemic acute kidney injury in mice. Commun. Biol. 2, 326 (2019).
Haimon, Z. et al. Re-evaluating microglia expression profiles using RiboTag and cell isolation strategies. Nat. Immunol. 19(6), 636–644 (2018).
Martins, J. P. S. & Conti, M. Profiling maternal mRNA translation during oocyte development. Methods Mol. Biol. 1818, 43–50 (2018).
Genuth, N. R. & Barna, M. The discovery of ribosome heterogeneity and its implications for gene regulation and organismal life. Mol. Cell. 71(3), 364–374 (2018).
Karamysheva, Z. N. et al. Polysome profiling in leishmania, human cells and mouse testis. J. Vis. Exp. 134, 57600 (2018).
Fischer, M., Grossmann, P., Padi, M. & DeCaprio, J. A. Integration of TP53, DREAM, MMB-FOXM1 and RB-E2F target gene analyses identifies cell cycle gene regulatory networks. Nucleic Acids Res. 44(13), 6070–6086 (2016).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352(6282), 189–196 (2016).
Acknowledgements
Negative stain EM images were taken at the University of Virginia Molecular Electron Microscopy Core facility (RRID:SCR_019031), which is supported in part by the School of Medicine and built with NIH grant G20-RR31199. We thank the University of Virginia Center for Excellence in Genetics (M.J.W.). This work was supported by National Institutes of Health grants (R01HL158718) to M.J.W. and the Medical Scientist Training Program, University of Virginia, Charlottesville, VA to J.C, and by the Searle Scholars Program and departmental startup funds to A.J.
Author information
Authors and Affiliations
Contributions
J.C., T.A.L., L.A.B., K.W.S., A.Y., E.M., A.J., J.V. and M.J.W. designed the experiments and performed research. J.C., E.M, A.J. T.A.L, and M.J.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cochran, J.D., Leathers, T.A., Maldosevic, E. et al. Cell cycle specific, differentially tagged ribosomal proteins to measure phase specific transcriptomes from asynchronously cycling cells. Sci Rep 14, 1623 (2024). https://doi.org/10.1038/s41598-024-52085-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-52085-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.